Abstract
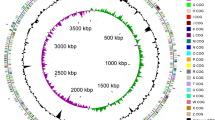
Leptospirillum ferriphilum has been identified as the dominant, moderately thermophilic, bioleaching microorganism in bioleaching processes. It is an acidic and chemolithoautrophic bacterium that gains electrons from ferrous iron oxidation for energy production and cell growth. Genetic information about this microorganism has been limited until now, which has hindered its further exploration. In this study, the complete genome of L. ferripilum ML-04 is sequenced and annotated. The bacterium has a single circular chromosome of 2,406,157 bp containing 2,471 coding sequences (CDS), 2 rRNA operons, 48 tRNA genes, a large number of mobile genetic elements and 2 genomic islands. In silico analysis shows L. ferriphilum ML-04 fixes carbon through a reductive citric acid (rTCA) cycle, and obtains nitrogen through ammonium assimilation. The genes related to “cell envelope biogenesis, outer membrane” (6.9%) and “DNA replication, recombination and repair” (5.6%) are abundant, and a large number of genes related to heavy metal detoxification, oxidative and acidic stress defense, and signal transduction pathways were detected. The genomic plasticity, plentiful cell envelope components, inorganic element metabolic abilities and stress response mechanisms found the base for this organism’s survival in the bioleaching niche.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Acuña, J., J. Rojas, A.M. Amaro, H. Toledo, and C.A. Jerez. 1992. Chemotaxis of Leptospirillum ferrooxidans and other acidophilic chemolithotrophs: comparison with the Escherichia coli chemosensory system. FEMS Microbiol. Lett. 75, 37–42.
Baker-Austin, C. and M. Dopson. 2007. Life in acid: pH homeostasis in acidophiles. Trends Microbiol. 15, 165–171.
Barreto, M., E. Jedlicki, and D.S. Holmes. 2005. Identification of a gene cluster for the formation of extracellular polysaccharide precursors in the chemolithoautotroph Acidithiobacillus ferrooxidans. Appl. Environ. Microbiol. 71, 2902–2909.
Benson, G. 1999. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580.
Blaudeck, N., P. Kreutzenbeck, M. Muller, G.A. Sprenger, and R. Freudl. 2005. Isolation and characterization of bifunctional Escherichia coli TatA mutant proteins that allow efficient Tat-dependent protein translocation in the absence of TatB. J. Biol. Chem. 280, 3426–3432.
Cabreiro, F., C.R. Picot, B. Friguet, and I. Petropoulos. 2006. Methionine sulfoxide reductases: relevance to aging and protection against oxidative stress. Ann. NY Acad. Sci. 1067, 37–44.
Castanie-cornet, M., T.A. Penfound, D. Smith, J.F. Elliott, and J.W. Foster. 1999. Control of acid resistance in Escherichia coli. J. Bacteriol. 181, 3525–3535.
Coram, N.J. and D.E. Rawlings. 2002. Molecular relationship between two groups of the genus Leptospirillum and the finding that Leptosphillum ferriphilum sp. nov. dominates South African commercial biooxidation tanks that operate at 40°C. Appl. Environ. Microbiol. 68, 838–845.
Dahl, C., A. Schulte, Y. Stockdreher, C. Hong, F. Grimm, J. Sander, R. Kim, S. Kim, and D.H. Shin. 2008. Structural and molecular genetic insight into a widespread sulfur oxidation pathway. J. Math. Biol. 384, 1287–1300.
Delepelaire, P. 2004. Type I secretion in gram-negative bacteria. Biochim. Biophys. Acta. 1694, 149–161.
Dixon, R. 1998. The oxygen-responsive NIFL-NIFA complex: a novel two-component regulatory system controlling nitrogenase synthesis in Γ-Proteobacteria. Arch. Microbiol. 169, 371–380.
Ducluzeau, A., S. Ouchane, and W. Nitschke. 2008. The cbb3 oxidases are an ancient innovation of the domain bacteria. Mol. Biol. Evol. 25,1158–1166.
Edwards, K.J., P.L. Bond, and J.F. Banfield. 2000. Characteristics of attachment and growth of Thiobacillus caldus on sulphide minerals: a chemotactic response to sulphur minerals? Environ. Microbiol. 2, 324–332.
Eichhorn, E., J.R. van der Ploeg, and T. Leisinger. 2000. Deletion analysis of the Escherichia coli taurine and alkanesulfonate transport systems. J. Bacteriol. 182, 2687–2695.
Goltsman, D.S.A., V.J. Denef, S.W. Singer, N.C. VerBerkmoes, M. Lefsrud, R.S. Mueller, G.J. Dick, and et al. 2009. Community genomic and proteomic analysis of chemoautotrophic, iron-oxidizing “Leptospirillum rubarum” (Group II) and “Leptospirillum ferrodiazotrophum” (Group III) in acid mine drainage biofilms. Appl. Environ. Microbiol. 75, 4599–4615.
Hanson, T.E. and F.R. Tabita. 2001. A ribulose-1,5-bisphosphate carboxylase/ oxygenase (RubisCO)-like protein from Chlorobium tep idum that is involved with sulfur metabolism and the response to oxidative stress. Proc. Natl. Acad. Sci. USA 98, 4397–4402.
Hutchins, A.M., J.F. Holden, and M.W.W. Adams. 2001. Phosphoenolpyruvate synthetase from the hyperthermophilic archaeon Pyrococcus furiosus. J. Bacteriol. 183, 709–715.
Jeans, C., S.W. Singer, C.S. Chan, N.C. VerBerkmoes, M. Shah, R.L. Hettich, J.F. Banfield, and M.P. Thelen. 2008. Cytochrome 572 is a conspicuous membrane protein with iron oxidation activity purified directly from a natural acidophilic microbial community. ISME J. 2, 542–550.
Jiao, Y. and D.K. Newman. 2007. The pio operon is essential for phototrophic Fe(II) oxidation in Rhodopseudomonas palustris TIE-1. J. Bacteriol. 189, 1765–1773.
Kawasaki, S., J. Ishikura, D. Chiba, T. Nishino, and Y. Niimura. 2004. Purification and characterization of an H2O-forming NADH oxidase from Clostridium aminovalericum: existence of an oxygen-detoxifying enzyme in an obligate anaerobic bacteria. Arch. Microbiol. 181, 324–330.
Kerscher, L., S. Nowitzki, D. Oesterhelt. 1982. Thermoacidophilic archaebacteria contain bacterial-type ferredoxins acting as electron acceptors of 2-oxoacid:ferredoxin oxidoreductases. Eur. J. Biochem. 128, 223–230.
Kurtz, D.M., Jr. 2006. Avoiding high-valent iron intermediates: Superoxide reductase and rubrerythrin. J. Inorg. Biochem. 100, 679–693.
Leigh, J.A. and J.A. Dodsworth. 2007. Nitrogen regulation in bacteria and archaea. Annu. Rev. Microbiol. 61, 349–377.
Leverrier, P., J.P. Vissers, A. Rouault, P. Boyaval, and G. Jan. 2004. Mass spectrometry proteomic analysis of stress adaptation reveals both common and distinct response pathways in Propionibacterium freudenreichii. Arch. Microbiol. 181, 215–230.
Levican, G., J.A. Ugalde, N. Ehrenfeld, A. Maass, and P. Parada. 2008. Comparative genomic analysis of carbon and nitrogen assimilation mechanisms in three indigenous bioleaching bacteria: predictions and validations. BMC Genomics 9, 581.
Li, B., J. Lin, S. Mi, and J. Lin. 2010. Arsenic resistance operon structure in Leptospirillum ferriphilum and proteomic response to arsenic stress. Bioresour. Technol. 101, 9811–9814.
Macnab, R.M. 2004. Type III flagellar protein export and flagellar assembly. Biochim. Biophys. Acta. 1694, 207–217.
Mai, X. and M.W. Adams. 1996. Characterization of a fourth type of 2-keto acid-oxidizing enzyme from a hyperthermophilic archaeon: 2-ketoglutarate ferredoxin oxidoreductase from Thermococcus litoralis. J. Bacteriol. 178, 5890–5896.
Maklashina, E., D.A. Berthold, and G. Cecchini. 1998. Anaerobic expression of Escherichia coli succinate dehydrogenase: functional replacement of fumarate reductase in the respiratory chain during anaerobic growth. J. Bacteriol. 180, 5989–5996.
Michels, M. and E.P. Bakker. 1985. Generation of a large, protonophore-sensitive proton motive force and pH difference in the acidophilic bacteria Thermoplasma acidophilum and Bacillus acidocaldarius. J. Bacteriol. 161, 231–237.
Moat, A.G., J.W. Foster, and M.P. Spector. 2002. Microbial Physiology, fourth ed. Wiley-Liss, New York, NY, USA.
Morgan, R.W., M.F. Christman, F.S. Jacobson, G. Storz, and B.N. Ames. 1986. Hydrogen peroxide-inducible proteins in Salmonella typhimurium overlap with heat shock and other stress proteins. Proc. Natl. Acad. Sci. USA 83, 8059–8063.
Muro-Pastor, M.I., J.C. Reyes, and F.J. Florencio. 2005. Ammonium assimilation in cyanobacteria. Photosynth. Res. 83, 135–150.
Preiss, J. 1984. Bacterial glycogen synthesis and its regulation. Annu. Rev. Microbiol. 38, 419–458.
Quatrini, R., C. Appia-Ayme, Y. Denis, E. Jedlicki, D.S. Holmes, and V. Bonnefoy. 2009. Extending the models for iron and sulfur oxidation in the extreme Acidophile Acidithiobacillus ferrooxidans. BMC Genomics 10, 394.
Rawlings, D.E., H. Tributsch, and G.S. Hansford. 1999. Reasons why ‘Leptospirillum’-like species rather than Thiobacillus ferrooxidans are the dominant iron-oxidizing bacteria in many commercial processes for the biooxidation of pyrite and related ores. Microbiology 145, 5–13.
Seibold, G.M., M. Wurst, and B.J. Eikmanns. 2009. Roles of maltodextrin and glycogen phosphorylases in maltose utilization and glycogen metabolism in Corynebacterium glutamicum. Microbiology 155, 347–358.
Simon, J., R. Gross, O. Einsle, P.M.H. Kroneck, A. Kroger, and O. Klimmek. 2000. A NapC/NirT-type cytochrome c (NrfH) is the mediator between the quinone pool and the cytochrome c nitrite reductase of Wolinella succinogenes. Mol. Microbiol. 35, 686–696.
Singer, S.W., C.S. Chan, A. Zemla, N.C. VerBerkmoes, M. Hwang, R.L. Hettich, J.F. Banfield, and M.P. Thelen. 2008. Characterization of cytochrome 579, an unusual cytochrome isolated from an iron-oxidizing microbial community. Appl. Environ. Microbiol. 74, 4454–4462.
Szurmant, H. and G.W. Ordal. 2004. Diversity in chemotaxis mechanisms among the bacteria and archaea. Microbiol. Mol. Biol. Rev. 68, 301–319.
Tuffin, I.M., S.B. Hector, S.M. Deane, and D.E. Rawlings. 2006. Resistance determinants of a highly arsenic-resistant strain of Leptospirillum ferriphilum isolated from a commercial biooxidation tank. Appl. Environ. Microbiol. 72, 2247–2253.
Valdes, J., F. Veloso, E. Jedlicki, and D. Holmes. 2003. Metabolic reconstruction of sulfur assimilation in the extremophile Acidithiobacillus ferrooxidans based on genome analysis. BMC Genomics 4, 51.
van der Ploeg, J.R., E. Eichhorn, and T. Leisinger. 2001. Sulfonate-sulfur metabolism and its regulation in Escherichia coli. Arch. Microbiol. 176, 1–8.
Walkup, L.K.B. and T. Kogoma. 1989. Escherichia coli proteins inducible by oxidative stress mediated by the superoxide radical. J. Bacteriol. 171, 1476–1484.
Wong, H.C., A.L. Fear, R.D. Calhoon, G.H. Eichinger, R. Mayer, D. Amikam, M. Benziman, and et al. 1990. Genetic organization of the cellulose synthase operon in Acetobacter xylinum. Proc. Natl. Acad. Sci. USA 87, 8130–8134.
Yun, N.R., M. Yamamoto, H. Arai, M. Ishii, and Y. Igarashi. 2002. A novel five-subunit-type 2-oxoglutalate:ferredoxin oxidoreductases from Hydrogenobacter thermophilus TK-6. Biochim. Biophys. Res. Commun. 292, 280–286.
Zhang, Q., T. Iwaaki, T. Wakagi, and T. Oshima. 1996. 2-Oxoacid:ferredoxin oxidoreductase from the thermoacidophilic archaeon, Sulfolobus sp. strain 7. J. Biochem. 120, 587–599.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplemental material for this article may be found at http://www.springer.com/content/120956
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Mi, S., Song, J., Lin, J. et al. Complete genome of Leptospirillum ferriphilum ML-04 provides insight into its physiology and environmental adaptation. J Microbiol. 49, 890–901 (2011). https://doi.org/10.1007/s12275-011-1099-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-011-1099-9