Abstract
Clomiphene citrate, a selective estrogen receptor modulator, is metabolized into its 4-hydroxylated active metabolites, primarily by CYP2D6. In this study, we investigated the effects of the most common CYP2D6 variant allele in Asians, CYP2D6*10, on the pharmacokinetics of clomiphene and its two active metabolites (4-OH-CLO and 4-OH-DE-CLO) in healthy Korean subjects. A single 50-mg oral dose of clomiphene citrate was given to 22 Korean subjects divided into three genotype groups according to CYP2D6 genotypes, CYP2D6*wt/*wt (n = 8; *wt = *1 or *2), CYP2D6*wt/*10 (n = 8) and CYP2D6*10/*10 (n = 6). Concentrations of clomiphene and its metabolites were determined using a validated HPLC–MS/MS analytical method in plasma samples collected up to 168 h after the drug intake. There was a significant difference only in the Cmax of clomiphene between three CYP2D6 genotype groups (p < 0.05). Paradoxically, the elimination half-life (t1/2) and AUC of both active metabolites were all significantly increased in the CYP2D6*10 homozygous carriers, compared with other genotype groups (all p < 0.001). The AUCinf of corrected clomiphene active moiety in CYP2D6*10/*10 subjects was 2.95- and 2.05-fold higher than that of CYP2D6*wt/*wt and *wt/*10 genotype groups, respectively (both p < 0.001). Along with the partial impacts on the biotransformation of clomiphene and its metabolites by CYP2D6 genetic polymorphism, further studies on the effects of other CYP enzymes in a multiple-dosing condition can provide more definite evidence for the inter-individual variabilities in clomiphene pharmacokinetics and/or drug response.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Clomiphene citrate (2-[4-[2-chloro-1,2-diphenylethenyl]phenoxy]-N,N-diethyl ethaneaminedihydrogen citrate) is a selective estrogen receptor modulator, which enhances the release of gonadotrophin-releasing hormone by inhibiting negative feedback of estradiol on the hypothalamus. As a result, releasing of follicle stimulating hormone and luteinizing hormone leads to growth of follicle and ovulation. Clomiphene is used as a first-line therapy to treat female infertility with absent or irregular ovulation, especially in those with polycystic ovary syndrome (PCOS) (Dickey 1996).
Clomiphene citrate is orally administered once daily for 5 days from day 2, 3, 4 or 5 of spontaneous or induced bleeding. After starting with the lowest dose, the daily dose of clomiphene citrate can be increased in increments of 50 mg/day per cycle until an ovulatory cycle is achieved (Homburg 2005). The recommended daily dose in the first cycle is 50 mg, but previous reports suggests that only 46% of patients will respond to this dose with ovulation. Furthermore, 8–30% of clomiphene-treated women do not respond and fail to ovulate even at the highest tolerated dose (Rostami-Hodjegan et al. 2004; Homburg 2005). Despite over 45 years of clinical experience with clomiphene citrate, these interindividual variability of clomiphene drug response is not yet clearly demonstrated.
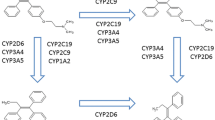
Clomiphene citrate is given as an unequal mixture of two geometric isomers, (E)- and (Z)-clomiphene, in the ratio of 67:33. It is readily absorbed orally, undergoes enterohepatic recirculation, and is excreted principally in the feces. Clomiphene is extensively metabolized in the liver and the intestinal mucosa, via hydroxylation, hydrogenation, N-deethylation, N-oxide formation, methoxylation and combinations of them, and the diverse cytochrome P450 (CYP) enzymes, including CYP3A4, CYP2D6, CYP3A5, CYP2B6, CYP2C9, and CYP2C19, are involved in these metabolic processes (Ghobadi et al. 2008; Mürdter et al. 2012; Mazzarino et al. 2013). These CYP enzymes are genetically polymorphic and numerous variant alleles exist; over 34 alleles for CYP3A4, 113 alleles for CYP2D6, 11 alleles for CYP3A5, 38 alleles for CYP2B6, 60 alleles for CYP2C9, and 45 alleles for CYP2C19 have been observed (PharmVar 2017). Pharmacokinetics and pharmacodynamics of many substrate drugs were affected in various degrees by these genetic polymorphisms (Ma et al. 2012; Hirota et al. 2013; Preissner et al. 2013; Chaudhry et al. 2014; Probst-Schendzielorz et al. 2015; Byeon et al. 2015; Lee et al. 2016; Kim et al. 2017). Among them, CYP3A4 and CYP2D6 were reported as main metabolizing enzymes of clomiphene (Mazzarino et al. 2013), and CYP2D6-mediated 4-hydroxylated active metabolites, 4-hydroxy-clomiphene (4-OH-CLO) and 4-hydroxy-N-desethylclomiphene (4-OH-DE-CLO), are considered to have at least 100-fold higher inhibitory potency at the estrogen receptor than their parent drug (Mürdter et al. 2012; Mazzarino et al. 2013).
There are two previous studies which have shown inconsistent results for the effect of CYP2D6 genetic polymorphisms on the pharmacokinetics of clomiphene and/or its metabolites. In the former study performed by Mürdter et al. (2012), the maximum plasma concentration (Cmax) of (E)-4-OH-CLO and (E)-4-OH-DE-CLO in healthy subjects homozygous for CYP2D6 null alleles (CYP2D6*4 or *5) were 8- and 12-fold lower than those in CYP2D6 EM subjects, respectively. Meanwhile, Ji et al. (2016) have recently reported that no significant differences were observed for the plasma concentrations of clomiphene metabolites between CYP2D6 EM and IM phenotype patients.
CYP2D6 is usually divided into four phenotypic subgroups by the number of functional CYP2D6 alleles; extensive metabolizers (EMs), intermediate metabolizers (IMs), poor metabolizers (PMs), and ultrarapid metabolizers (UMs). In Korean population, two functional alleles (CYP2D6*1 and *2) and one variant allele account for 93.9% of the allele frequency. Thus, in the present study, we investigated the association between CYP2D6*10 allele and the pharmacokinetics of clomiphene and its two active metabolites (4-OH-CLO and 4-OH-DE-CLO) in healthy Korean subjects with CYP2D6*wt/*wt, *wt/*10, and *10/*10 genotypes.
Materials and methods
Subjects
Twenty-two healthy Korean subjects with CYP2D6*wt/*wt (n = 8, *wt = *1 or *2), *wt/*10 (n = 8) and *10/*10 (n = 6) genotypes were enrolled in this pharmacokinetic study (mean age 22.9 ± 2.0 years, mean height 172.2 ± 8.0 cm, mean weight 63.6 ± 9.5 kg, and mean BMI 21.4 ± 2.0 kg/m2, respectively). Genomic DNA was isolated from peripheral blood leukocytes using a Wizard® Genomic DNA purification Kit (Promega, Madison, WI, USA) according to the manufacturer’s instructions. The genotyping for CYP2D6*2 and *10 alleles were performed by polymerase chain reaction-restriction fragment length polymorphism (PCR–RFLP) analyses, and CYP2D6*5 and *X × N alleles by long-PCR analyses, as previously described (Lundqvist et al. 1999; Naveen et al. 2006). All subjects were healthy according to their medical histories, physical examinations, vital signs (blood pressure, heart rate and body temperature) and routine laboratory tests (blood chemistry, hematology, and urine analysis). The subjects were asked to refrain from taking any medications, consuming caffeine, grapefruit products, alcoholic beverages, and smoking for at least one week before and throughout the study period.
Study protocol
All study procedures were performed according to the guidelines of the Declaration of Helsinki and was approved by the Institutional Ethics Committee of the Metro Hospital, Anyang, Republic of Korea. Verbal and written informed consents were obtained from all participants. Each subject administered a single oral dose of 50 mg clomiphene citrate (Clomifene Citrate Tablet, Youngpoong Pharm., Incheon, Korea) with 240 ml of water after an overnight fast. Blood samples were obtained before and at 1, 2, 4, 6, 8, 10, 12, 24, 48, 96 and 168 h after drug administration. The plasma samples for the determination of clomiphene and metabolites concentrations were immediately separated from the whole blood and were stored at − 70 °C until analysis.
Determination of plasma clomiphene and its active metabolites concentrations
Plasma concentrations of clomiphene and its two metabolites, 4-OH-CLO and 4-OH-DE-CLO, were determined using a validated high-performance liquid chromatography with tandem mass spectrometry (HPLC–MS/MS) analytical method. Briefly, 200 μl of plasma was spiked with 10 μl of an internal standard (tamoxifen, 3 μg/ml) and then extracted with 2 ml of ethyl acetate. The organic phase was evaporated at 50 °C under a constant flow of nitrogen gas. The residue was reconstituted with 300 μl of mobile phase, and a 10-μl aliquot was injected into the analytical column (Luna, C18, 3 μm, 2.0 mm × 100 mm, Phenomenex, Torrance, CA, USA). The mobile phase consisted of 20:80 ammonium acetate buffer (10 mM, pH 6.15)/acetonitrile at a flow rate of 200 μl/min. The column oven temperature was 30 °C. The mass spectrometer was operated in electrospray ionization (ESI) negative ion mode and ions were detected in multiple-reaction monitoring (MRM) mode. Mass transition pairs of m/z were 406.2 → 100.0, 422.2 → 100.0, 394.2 → 72.0 and 372.1 → 72.0 for clomiphene, 4-OH-CLO, 4-OH-DE-CLO and internal standard (tamoxifen), respectively. The calibration curve of each analyte was linear over a range from 1 to 20 ng/ml for clomiphene, and 0.01 to 0.5 ng/ml for 4-OH-CLO and 4-OH-DE-CLO, respectively. The mean accuracies of clomiphene, 4-OH-CLO and 4-OH-DE-CLO were all between 98.5 and 108.4%. The coefficients of variation (intra-day and inter-day precision) for clomiphene, 4-OH-CLO and 4-OH-DE-CLO were all below 12%.
Pharmacokinetic analysis
The pharmacokinetic parameters of clomiphene and its metabolites were estimated using non-compartmental methods with the BA calc 2007 analysis program (KFDA, Seoul, Korea). Actual blood sampling times were used, while observed values were used for Cmax. The area under the plasma concentration–time curve (AUC) was calculated using the linear-log trapezoidal rule. The elimination rate constant (ke) was estimated from the least-squares regression slope of the terminal plasma concentration. The AUC from 0 to infinity (AUCinf) was calculated as AUCinf = AUC + Ct/ke, where Ct is the most recently measured plasma concentration. The elimination half-life (t1/2) was calculated as ln 2/ke, and the apparent oral clearance (CL/F) of clomiphene was calculated as CL/F = dose/AUCinf. Based on the reported biological potency of each compound (Mürdter et al. 2012), the amount of clomiphene active moiety was corrected and calculated by the sum of the AUCinf of clomiphene, and the AUCinf multiplied by 100 of 4-OH-CLO and 4-OH-DE-CLO.
Statistical analysis
Pharmacokinetic data from the study results were expressed as mean ± standard deviation (SD). Differences in pharmacokinetic parameters in three different CYP2D6 genotype groups were assessed using a one-way analysis of variance (ANOVA) with the Bonferroni t test (as the multiple post hoc comparisons), or Kruskal–Wallis one-way ANOVA on ranks with the Mann–Whitney rank sum test after normality and equal variance tests. All data were analyzed using the statistical program Prism® Windows 6.0 (GraphPad Software, La Jolla, CA, USA). The p values of less than 0.05 were considered statistically significant.
Results
There were no significant differences in the demographic characteristics between the three CYP2D6 genotype groups (data not shown), and none of the subjects showed any adverse reactions associated with clomiphene administration throughout the study period.
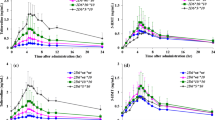
The mean plasma concentration–time profiles of clomiphene, 4-OH-CLO, 4-OH-DE-CLO, and clomiphene active moiety in 22 healthy Korean volunteers with different CYP2D6 genotypes (CYP2D6*wt/*wt, *wt/*10 and *10/*10) after a single oral dose administration of 50 mg clomiphene are illustrated in Fig. 1. There was only a significant difference in the Cmax of clomiphene parent drug between three CYP2D6 genotype groups (p < 0.05). The Cmax of clomiphene in CYP2D6*10/*10 group (29.53 ± 4.06 nM) was 1.45-fold higher than that in CYP2D6*wt/*wt group (20.37 ± 5.64 nM, p < 0.05). Other clomiphene pharmacokinetic parameters, including t1/2, CL/F and AUC, showed similar values and the differences did not reach statistical significance between each genotype group (Table 1).
Plasma concentration–time profiles of clomiphene (a), 4-hydroxyclomiphene (b), and 4-hydroxy-N-desethylclomiphene (c) in CYP2D6*wt/*wt (n = 8, circles), CYP2D6*wt/*10 (n = 8, squares) and CYP2D6*10/*10 (n = 6, triangles) groups after a single oral dose of clomiphene citrate 50 mg. Each value is given as mean ± SD
Pharmacokinetic parameters for 4-OH-CLO and 4-OH-DE-CLO showed more distinct and paradoxical differences than parent drug, according to the CYP2D6 genotype (Table 1). The t1/2 and AUC values for both metabolites were all significantly different between three CYP2D6 genotype groups (p < 0.001). The AUCinf of 4-OH-CLO in CYP2D6*10/*10 group (29.85 ± 4.35 nM h) were 3.48- and 2.05-fold higher than that in CYP2D6*wt/*wt and *wt/*10 group, respectively (8.57 ± 2.68 nM h and 14.54 ± 5.10 nM h; both p < 0.001). Also, 2.69-fold prolonged 4-OH-CLO t1/2 (34.52 ± 5.06 h) were observed in CYP2D6*10/*10 genotype, compared to that in CYP2D6*wt/*wt group (13.30 ± 3.96 h, p < 0.001). In the case of 4-OH-DE-CLO, the t1/2 and AUCinf values in CYP2D6*10/*10 group (37.73 ± 7.14 h and 75.82 ± 17.78 nM h) also showed significant differences compared to other two genotype groups, respectively (15.34 ± 3.99 h and 18.37 ± 5.23 nM h for CYP2D6*wt/*wt group, both p < 0.001; 22.01 ± 4.36 h and 30.79 ± 9.28 nM h for CYP2D6*wt/*10 group; p < 0.01 and < 0.001). The corrected amount of clomiphene active moiety in CYP2D6*10/*10 subjects (12027.68 ± 2421.32 nM h) was 2.95- and 2.05-fold higher than that of CYP2D6*wt/*wt and *wt/*10 genotype groups, respectively (4076.79 ± 817.18 and 5862.19 ± 1383.43 nM h; both p < 0.001, Fig. 2).
Discussion
It is known that the triphenylethylene-derived molecular structure of clomiphene is very similar to that of tamoxifen, a widely used selective estrogen receptor modulator (Mürdter et al. 2012). Tamoxifen requires metabolic activation by CYP3A4/5 and CYP2D6 to elicit its pharmacological activity, and two active metabolites, 4-hydroxytamoxifen and 4-hydroxy-N-desmethyl tamoxifen (endoxifen) are the major therapeutic moieties of tamoxifen (Desta et al. 2004; Lim et al. 2005). Lim et al. (2007) reported that association between CYP2D6 genetic polymorphism and tamoxifen pharmacokinetics and clinical outcomes in patients with breast cancer. In patients carrying the CYP2D6*10/*10 genotype, the steady-state plasma concentrations of both 4-hydroxytamoxifen and endoxifen were 1.6- and 2.3-fold lower than those in CYP2D6*wt/*wt or CYP2D6*wt/*10 genotype group, respectively. Therefore, it is inferred that CYP2D6 can also play a pivotal role in producing potent active metabolites of clomiphene, and genetic polymorphisms in CYP2D6 can lead to the alterations in pharmacokinetics and/or clinical outcomes of clomiphene.
Mürdter et al. (2012) has previously reported that clomiphene metabolites, which have similar structure to the active metabolite of tamoxifen, are considered to be more potent than clomiphene parent drug in vitro, and CYP2D6 genetic polymorphism can affect the pharmacokinetics of clomiphene and its metabolites, based on the data conducted in Caucasians. However, the sample size in their research was too small (total of six subjects, including only two CYP2D6 PMs) to evaluate the clear effects of CYP2D6 genetic polymorphisms on the pharmacokinetics and/or clinical outcomes of clomiphene, and the influences of non-functional CYP2D6 alleles might be exaggerated. In this study, we evaluated the effects of CYP2D6*10 allele on the pharmacokinetics of clomiphene and its active metabolites, with at least six subjects per each genotype group. Consequently, the presence of CYP2D6*10 allele caused no significant changes in most pharmacokinetic parameters of clomiphene, except for Cmax. Moreover, the subjects homozygous for CYP2D6*10 exhibited higher plasma concentrations for two active metabolites, 4-OH-CLO and 4-OH-DE-CLO, than CYP2D6*wt/*wt and/or *wt/*10 genotype groups, which are inconsistent with the previous study results (Mürdter et al. 2012; Ji et al. 2016). A similar tendency was also observed with the comparison of clomiphene active moiety.
Clomiphene has a complex metabolic pathway, and the active metabolites of clomiphene, as well as the parent drug, is metabolized by various CYP isozymes including CYP2D6 (Mürdter et al. 2012). Therefore, we suggest that the genetic polymorphisms of CYP2D6 and alterations in enzymatic activity also influenced the pharmacokinetic profiles of clomiphene metabolites. Increased plasma concentrations of clomiphene active metabolites may lead to the enhancement in the clinical drug response of clomiphene. This is supported by one recent study performed by Ji et al. (2016), which demonstrated that the all patients with CYP2D6 IM genotype in their study were classified as clomiphene responders, while clomiphene non-responders were composed only of CYP2D6 EM patients. Meanwhile, the plasma concentrations of 4-OH-DE-CLO, which is known to be the final metabolite of clomiphene, also significantly increased in subjects with CYP2D6*10/*10 genotype. The reason for pharmacokinetic differences between the three genotype groups is unclear, but the possibility of currently unknown metabolic pathways after conversion to 4-OH-DE-CLO might be considered.
It is known that clomiphene citrate is well tolerated at recommended dosages. Nevertheless, increased amounts of clomiphene active moiety may lead to the development of several adverse events associated with the use of clomiphene. The ovarian hyperstimulation syndrome, one of the most common adverse reactions, is characterized by rapid progression and may lead to a serious medical disorder. The clinical signs of this syndrome in severe cases include gross ovarian enlargement, gastrointestinal symptoms, ascites, dyspnea, oliguria, and pleural effusion. The lowest effective dose is generally recommended when necessary to minimize the occasional abnormal ovarian enlargement and related symptoms due to clomiphene administration, and clomiphene therapy should be discontinued if enlargement of the ovary occurs. Patients should also be advised regarding the risk for visual symptoms, such as blurring, spots or flashes, during the use of clomiphene. These symptoms increase in incidence, and may be irreversible with increased dosage or duration of therapy. Although the etiology is not yet understood, patients with any visual problems should discontinue treatment and get a complete ophthalmological evaluation promptly (CLOMID label information 2012).
There are several limitations in our study. First, we did not determine the concentrations of each geometric isomer for clomiphene and/or active metabolites. As mentioned above, clomiphene is given as a mixture of (E)- and (Z)-isomer, and the major metabolic route and pharmacological activity of each isomer is quite different. It is reported that 4-hydroxylated metabolites of clomiphene is observed only from (E)-clomiphene, while the primary metabolites of (Z)-clomiphene are formed via N-deethylation and, to a lesser extent, N-oxidation. The most potent inhibitory activities against the estrogen receptor were observed in (E)-isomers (Mürdter et al. 2012). Nevertheless, the calculated Cmax and AUC of (Z)-clomiphene were 3- and 20-fold higher than those of (E)-clomiphene, respectively, after a single oral dose of 50 mg clomiphene citrate in 9 PCOS patients (Ghobadi et al. 2009). Therefore, extremely low concentrations of 4-OH-CLO and 4-OH-DE-CLO compared with clomiphene in this study might be because the (Z)-clomiphene accounted for the majority of quantified parent drug contents. Second, our study is performed only with healthy volunteers, in a single oral dose condition. In anovulatory patients, the Cmax values for both clomiphene isomers, and AUC value for (Z)-clomiphene showed significant differences when compared to those in healthy subjects (Ghobadi et al. 2009). Moreover, clomiphene therapy regimen generally consists of repeated cycles of increasing dose from 50 to 150 mg per day until ovulation (Mürdter et al. 2012). Thus, a multiple-dose study with patients will be able to suggest more obvious relationship between CYP2D6 genetic polymorphisms and clomiphene pharmacokinetics and/or drug response. A recent study reported that there were no significant differences in clomiphene metabolite concentrations between CYP2D6 EM and IM phenotype patients, after consecutive dose of clomiphene citrate 100 mg/day (Ji et al. 2016). However, they could not collect the blood samples of all subjects at the same time due to their different expected day of ovulation response after clomiphene therapy, affecting the interpretation of the pharmacokinetics in different study groups.
In conclusion, the presence of CYP2D6*10 allele can influence the biotransformation of clomiphene, leading to the alteration in the pharmacokinetics of parent drug and its active metabolites. Considering the involvement of diverse CYP isozymes in the metabolism of clomiphene and its dosage regimen in clinical practice, further studies on the effects of genetic polymorphisms in other CYP enzyme-expressing genes, along with CYP2D6, in a multiple-dosing condition can provide more definite evidence for the inter-individual variabilities in clomiphene pharmacokinetics and/or drug response.
References
Byeon JY, Kim YH, Na HS, Jang JH, Kim SH, Lee YJ, Bae JW, Kim IS, Jang CG, Chung MW, Lee SY (2015) Effects of the CYP2D6*10 allele on the pharmacokinetics of atomoxetine and its metabolites. Arch Pharm Res 38:2083–2091
Chaudhry SR, Muhammad S, Eidens M, Klemm M, Khan D, Efferth T, Weisshaar MP (2014) Pharmacogenetic prediction of individual variability in drug response based on CYP2D6, CYP2C9 and CYP2C19 genetic polymorphisms. Curr Drug Metab 15(7):711–718
CLOMID label information (2012) Sanofi-Aventis U.S. LLC, Bridgewater, NJ, USA. http://www.accessdata.fda.gov/drugsatfda_docs/label/2012/016131s026lbl.pdf. Accessed 27 Oct 2017
Desta Z, Ward BA, Soukhova NV, Flockhart DA (2004) Comprehensive evaluation of tamoxifen sequential biotransformation by the human cytochrome P450 system in vitro: prominent roles for CYP3A and CYP2D6. J Pharmacol Exp Ther 310:1062–1075
Dickey RP (1996) Development, pharmacology and clinical experience with clomiphene citrate. Hum Reprod Update 2:483–506
Ghobadi C, Gregory A, Crewe HK, Rostami-Hodjegan A, Lennard MS (2008) CYP2D6 is primarily responsible for the metabolism of clomiphene. Drug Metab Pharmacokinet 23:101–105
Ghobadi C, Mirhosseini N, Shiran MR, Moghadamnia A, Lennard MS, Ledger WL, Rostami-Hodjegan A (2009) Single-dose pharmacokinetic study of clomiphene citrate isomers in anovular patients with polycystic ovary disease. J Clin Pharmacol 49:147–154
Hirota T, Eguchi S, Ieiri I (2013) Impact of genetic polymorphisms in CYP2C9 and CYP2C19 on the pharmacokinetics of clinically used drugs. Drug Metab Pharmacokinet 28(1):28–37
Homburg R (2005) Clomiphene citrate—end of an era? A mini-review. Hum Reprod 20:2043–2051
Ji M, Kim KR, Lee W, Choe W, Chun S, Min WK (2016) Genetic polymorphism of CYP2D6 and clomiphene concentrations in infertile patients with ovulatory dysfunction treated with clomiphene citrate. J Korean Med Sci 31:310–314
Kim SH, Kim DH, Byeon JY, Kim YH, Kim DH, Lim HJ, Lee CM, Whang SS, Choi CI, Bae JW, Lee YJ, Jang CG, Lee SY (2017) Effects of CYP2C9 genetic polymorphisms on the pharmacokinetics of celecoxib and its carboxylic acid metabolite. Arch Pharm Res 40(3):382–390
Lee HJ, Kim YH, Kim SH, Lee CM, Yang AY, Jang CG, Lee SY, Bae JW, Choi CI (2016) Effects of CYP2C9 genetic polymorphisms on the pharmacokinetics of zafirlukast. Arch Pharm Res 39(7):1013–1019
Lim YC, Desta Z, Flockhart DA, Skaar TC (2005) Endoxifen (4-hydroxy-N-desmethyl-tamoxifen) has anti-estrogenic effects in breast cancer cells with potency similar to 4-hydroxy-tamoxifen. Cancer Chemother Pharmacol 55:471–478
Lim HS, Ju Lee H, Seok Lee K, Sook Lee E, Jang IJ, Ro J (2007) Clinical implications of CYP2D6 genotypes predictive of tamoxifen pharmacokinetics in metastatic breast cancer. J Clin Oncol 25:3837–3845
Lundqvist E, Johansson I, Ingelman-Sundberg M (1999) Genetic mechanisms for duplication and multiduplication of the human CYP2D6 gene and methods for detection of duplicated CYP2D6 genes. Gene 226:327–338
Ma JD, Lee KC, Kuo GM (2012) Clinical application of pharmacogenomics. J Pharm Pract 25(4):417–427
Mazzarino M, Biava M, de la Torre X, Fiacco I, Botrè F (2013) Characterization of the biotransformation pathways of clomiphene, tamoxifen and toremifene as assessed by LC-MS/(MS) following in vitro and excretion studies. Anal Bioanal Chem 405:5467–5487
Mürdter TE, Kerb R, Turpeinen M, Schroth W, Ganchev B, Böhmer GM, Igel S, Schaeffeler E, Zanger U, Brauch H, Schwab M (2012) Genetic polymorphism of cytochrome P450 2D6 determines oestrogen receptor activity of the major infertility drug clomiphene via its active metabolites. Hum Mol Genet 21:1145–1154
Naveen AT, Adithan C, Soya SS, Gerard N, Krishnamoorthy R (2006) CYP2D6 genetic polymorphism in South Indian populations. Biol Pharm Bull 29:1655–1658
PharmVar (2017) CYP allele nomenclature. http://www.pharmvar.org/genes. Accessed 30 Dec 2017
Preissner SC, Hoffmann MF, Preissner R, Dunkel M, Gewiess A, Preissner S (2013) Polymorphic cytochrome P450 enzymes (CYPs) and their role in personalized therapy. PLoS ONE 8(12):e82562
Probst-Schendzielorz K, Viviani R, Stingl JC (2015) Effect of Cytochrome P450 polymorphism on the action and metabolism of selective serotonin reuptake inhibitors. Expert Opin Drug Metab Toxicol 11(8):1219–1232
Rostami-Hodjegan A, Lennard MS, Tucker GT, Ledger WL (2004) Monitoring plasma concentrations to individualize treatment with clomiphene citrate. Fertil Steril 81:1187–1193
Acknowledgements
This research was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science, ICT, and Future Planning (NRF-2016R1A2B4007381).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no potential conflict of interest.
Rights and permissions
About this article
Cite this article
Kim, MJ., Byeon, JY., Kim, YH. et al. Effect of the CYP2D6*10 allele on the pharmacokinetics of clomiphene and its active metabolites. Arch. Pharm. Res. 41, 347–353 (2018). https://doi.org/10.1007/s12272-018-1005-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12272-018-1005-7