Abstract
In 2006, Shinya Yamanaka first reported that in vitro reprogramming of somatic cells toward pluripotency was achieved by simple induction of specific transcription factors. Induced pluripotent stem cell (iPSC) technology has since revolutionized the ways in which we explore the mechanisms of human diseases and develop therapeutics. Here, I describe the recent advances in human iPSC-based disease modeling and drug discovery and discuss the current challenges. Additionally, I outline potential future applications of human iPSCs in classifying patients based on their response to drugs in clinical trials and elucidating optimal patient-specific therapeutic strategies, which will contribute to reduced attrition rates and the development of precision medicine.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
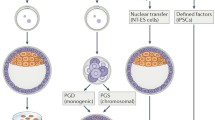
In 1962, it was reported that transfer of the nucleus from a frog somatic cell to an enucleated egg cell generated a fully functional tadpole that was genetically identical to the donor frog (Gurdon et al. 1958; Gurdon 1962). This finding radically changed the prevalent theory in the early 20th century that cellular differentiation was a unidirectional and irreversible process. It was believed that fully differentiated somatic cells were unable to return to dedifferentiated and pluripotent stem-cell states. Since this discovery, much progress has been made in understanding the reprogramming process. In the late 20th century, several researchers described the cloning of mammals, such as sheep and mice, from their adult somatic cells using somatic cell nuclear transfer (Wilmut et al. 1997; Wakayama et al. 1998). In 2006, Takahashi and Yamanaka reported that introduction of four transcription factors (POU5F1, SOX2, KLF4, and MYC) reprogrammed differentiated mouse somatic cells into embryonic stem cell (ESC)-like pluripotent cells called induced pluripotent stem cells (iPSCs) (Takahashi and Yamanaka 2006). A year later, they generated human iPSCs from human fibroblasts using the same combination of four transcription factors described in the 2006 paper (Takahashi et al. 2007). Simultaneously, Thomson and colleagues also reported the generation of human iPSCs through a somewhat different combination of transcription factors (POU5F1, SOX2, NANOG, and LIN28) than that used by Yamanaka and colleagues (Yu et al. 2007). This iPSC technology overcame serious limitations imposed by the generation of patient-specific ESCs and those of previous reprogramming methods. Because iPSCs are generated directly from human adult somatic cells, they are free from the ethical issues associated with ESC-generation procedures and also from the risk of immune rejection following transplantation to recover or replace damaged tissues. Furthermore, iPSC technology is remarkably efficient and simple as compared with previous reprogramming methods such as somatic cell nuclear transfer and cell fusion.
Since its discovery, great efforts have been made to improve human iPSC technology. Methods were developed for the delivery of reprogramming transcription factors into the cell using adenoviruses, Sendai viruses, transposons, plasmids, RNAs, and recombinant proteins in order to generate integration-free iPSCs (Okita et al. 2008; Stadtfeld et al. 2008; Fusaki et al. 2009; Kaji et al. 2009; Kim et al. 2009; Yu et al. 2009; Yusa et al. 2009; Woltjen et al. 2009; Jia et al. 2010; Nishimura et al. 2011; Hou et al. 2013). Specifically, Sendai viruses, episomal plasmids, and synthetic mRNAs are now widely used for integration-free delivery of reprogramming transcription factors. More recently, small molecules were shown to be suitable substitutes for some reprogramming factors (Hou et al. 2013). These advancements enabled the use of iPSC technology in regenerative medicine for the development of patient-specific cell therapy. Additionally, iPSC technology has been used to construct cell-based model systems for studying human diseases and produce screening platforms for the development and validation of therapeutic compounds. Here, I provide an overview of the use of human iPSCs for disease modeling and drug discovery and describe how human iPSC-based disease models have been utilized to assess the efficacy and toxicity of known drugs and potential drug candidates. I also discuss the challenges of human iPSC-based drug discovery and the efforts to overcome these limitations.
Disease modeling using human iPSCs
Modeling of human diseases is crucial for understanding the molecular mechanisms of pathogenesis and developing strategies for the prevention and treatment of diseases. Because primary patient cells are difficult to isolate in most cases and, when possible, are available in very small quantities, researchers have traditionally relied upon patient-derived immortalized cell lines for in vitro assays and animal models for in vivo experiments when studying disease etiologies and developing therapeutic interventions. However, these disease models often do not accurately reproduce human pathophysiology. Although patient-derived immortalized cell lines have been widely used due to their relatively low cost, unlimited supply, and lack of ethical concerns associated with the use of human tissue, cell lines often lose the phenotypic and functional features present in primary human cells over an extended culture period. Therefore, caution is necessary when using immortalized cell lines, and key data should be confirmed in additional experiments using primary human cells.
Animal models are invaluable tools for modeling human diseases by enabling investigation of in vivo pathogenesis. Specifically, mice have long served as animal models for human biology and disease; however, mice and humans have considerable genetic, anatomical, and physiological differences. For example, mice share only 80% gene homology with humans (Church et al. 2009). This genetic divergence between mice and humans may explain many of the differences that distinguish human and mouse biology. On anatomical and physiological levels, mice and humans also exhibit differences in many organs. For example, the heart size and the resting cardiac rate of mice and humans are substantially different (Hamlin and Altschuld 2011). These differences can preclude the recapitulation of human disease phenotypes in animals and may explain why the human response to drug candidates in clinical trials is often difficult to predict from preclinical data. For these reasons, there is a compelling need to develop human models for investigating human pathogenesis and developing new therapeutics.
Human iPSCs can be a robust alternative to transformed cell lines and animals as models for investigating human disorders. Human iPSCs are intrinsically able to self-renew indefinitely and differentiate into any cell type, thereby making it possible to obtain a sufficient number of various human cell types. Moreover, a great advantage of iPSCs is that they are derived from individual patients with known disease phenotypes, even when there are multiple unknown contributing genetic mutations. This enables the study of genotype-phenotype relationships in genetically complex or sporadic diseases, models of which are extremely challenging to generate using current techniques. Although nuclease-based genome-editing technology has improved significantly in recent years and is utilized for modeling monogenic disorders using human pluripotent stem cells, it remains difficult to generate models for genetically complex diseases that require genetic alterations at multiple loci (Kim and Kim 2014). Additionally, recent studies reported that many risk alleles that distinguish affected patients from unaffected subjects are located in noncoding regions that exhibit low sequence homology between humans and animals (Avior et al. 2016). This presents a high risk of failure in attempting to recapitulate human disease phenotypes, even after introducing the same human genetic variants in animal models.
Since the initial discovery of iPSCs, many researchers have attempted to generate human disease models using patient-derived iPSCs and reported that the resultant differentiated cells exhibited disease-relevant phenotypes. One of the first human iPSC-based disease models generated was for spinal muscular atrophy caused by a loss-of-function mutation in the SMN1 gene. Motor neurons differentiated from patient-derived iPSCs degenerated much faster than those from normal control iPSCs (Ebert et al. 2009). In a study of Rett syndrome, a severe neurodevelopmental disorder, neurons differentiated from patient-derived iPSCs showed defects in structure and electrophysiological function when compared to neurons derived from either ESCs or normal iPSCs (Marchetto et al. 2010). Cardiomyocytes differentiated from iPSCs derived from patients with type 2 long QT syndrome (LQTS) exhibited the prolonged action-potential duration due to significant reductions of cardiac potassium current and arrhythmogenicity, which are typical phenotypes associated with LQTS (Itzhaki et al. 2011).
The vast majority of iPSC-based human disease models have been generated for monogenic diseases. However, several recent studies described that iPSCs generated from patients with genetically complex, sporadic, or even infectious diseases can be used as disease models. Israel et al. reported that in human iPSCs generated from two patients with sporadic Alzheimer’s diseases (AD), only one of the two patient-derived iPSC lines showed phenotypes resembling those of familial AD patients (Israel et al. 2012). This result indicates that mechanisms underlying sporadic AD may be relevant to the pathogenesis of familial AD, and it also implied the presence of unknown genetic mutations that affect the phenotypic heterogeneity in sporadic AD pathogenesis. Parkinson’s disease (PD) is another sporadic neurodegenerative disease where multiple factors, such as genetic mutations and environmental toxins, contribute to the onset and progression of the disease. Two recent studies described iPSCs generated from patients carrying the G2019S mutation in the LRRK2 gene. This mutation causes sporadic PD with age-dependent variations (Nguyen et al. 2011; Sanchez-Danes et al. 2012). Dopaminergic neurons differentiated from PD patient-derived iPSCs exhibited enhanced sensitivity to stress agents and increased expression of α-synuclein proteins as compared with their levels in control cells. These studies showed for the first time that an LRRK2 mutation causes sporadic human PD. It is also possible to model the pathogenic process of infectious diseases using human iPSCs. In a recent study, human iPSCs-derived cardiomyocytes were generated, infected with a coxsackievirus B3 strain that causes viral myocarditis, and used for the investigation of disease mechanisms and the screening of novel antiviral therapeutics (Sharma et al. 2014). These findings clearly suggested that human iPSC-based disease models were successful in recapitulating human pathogenesis and could be used for other purposes such as drug screening and personalized treatment.
Drug discovery using human iPSCs
Despite significant biotechnological developments and increase in the understanding of human diseases and the genome, the attrition rate remains high during drug development. An overall estimate suggests that the attrition rate of drug candidates is approximately 96%, which has become a cause of concern to the pharmaceutical industry (Paul et al. 2010). Analysis of US Food and Drug Administration (FDA) data on new drug application (NDA) and biologics license application (BLA) from 2003 to 2011 demonstrated that some of the primary reasons for suspension in phase 3 clinical trials and NDA/BLA filings were safety concerns and a lack of efficacy (Hay et al. 2014). To reduce the attrition rate in the clinical-development stages, researchers have focused on the development of more predictive and reliable models for early screening of drug-candidate efficacy and toxicity. In this regard, human iPSC-based models have received increased attention regarding their potential to provide more clinically relevant data based on their accurate reflection of in vivo conditions. Here, I provide several examples of the use of human iPSCs for testing drug-candidate efficacy and toxicity.
Drug screening using iPSCs
Cells differentiated from patient-derived iPSCs are increasingly employed as screening platforms for the development and validation of therapeutic compounds (Table 1). In a human iPSC-based model for familial dysautonomia (FD), a peripheral neuropathy caused by a mutation in the IKBKAP gene, the plant hormone kinetin was validated as a novel drug candidate capable of reversing aberrant IKBKAP splicing and increasing neuronal differentiation and migration (Lee et al. 2009). Similarly, in human iPSC models for amyotrophic lateral sclerosis (ALS) associated with mutations in the SOD1, C9ORF72, FUS1, or TDP43 genes, treatment with a Kv7 channel activator and a histone acetyltransferase inhibitor improved motor-neuron survival and rescued abnormal ALS phenotypes of motor neurons (Egawa et al. 2012; Wainger et al. 2014). Several FDA-approved drugs, including digoxin, methotrimeprazine, and fluphenazine, also exerted substantial neuroprotective effects in motor neurons differentiated from ALS patient-derived iPSCs (Burkhardt et al. 2013; Barmada et al. 2014). Studies of AD, which is the most complex and common neurodegenerative disease, demonstrated that human iPSCs are valuable tools for the identification and validation of new drug candidates. In neurons differentiated from iPSCs derived from patients carrying mutations in the PSEN1 or PSEN2 genes, small-molecule compounds inhibiting or modulating γ-secretase substantially reduced the production of β-amyloid peptides generated by secretase-mediated cleavage of amyloid precursor protein. For cardiovascular diseases, a human iPSC-based drug-screening study was first performed with iPSCs derived from patients with LQTS which is caused by mutations in the KCNQ1 and KCNH2 genes. Cardiomyocytes differentiated from LQTS patient-derived iPSCs were used to evaluate the potency of existing as well as novel pharmacological compounds. Treatment with β-adrenergic receptor inhibitors, such as nifedipine and pinacidil, proved effective at ameliorating disease phenotypes (Itzhaki et al. 2011). As an allosteric modulator of human ERG, the novel small-molecule LUF7346, was also identified as capable of rescuing genetic and drug-induced LQTS phenotypes (Sala et al. 2016). In a study of hepatic disorders, iPSCs were derived from patients with Wilson’s disease caused by mutations in the ATP7B gene (Zhang et al. 2011). In hepatocytes differentiated from patient-derived iPSCs, curcumin was identified as a possible therapeutic compound capable of partial restoration of mutant ATP7B localization and reversal of functional defects
Although many studies using human iPSCs evaluated the efficacy of small sets of drug candidates, the following three studies are particularly notable examples of using human iPSCs for large-scale high-throughput screening (HTS). Burkhardt et al. screened 1757 bioactive compounds on motor neurons differentiated from ALS patient-derived iPSCs (Burkhardt et al. 2013). Several FDA-approved glycosides, including digoxin, lanatoside C, and proscillaridin A, efficiently reduced the formation of transactive response DNA-binding protein-43 aggregates associated with ALS pathogenesis. In the second example of human iPSC-based HTS, neural crest cells differentiated from FD patient-derived iPSCs were employed to screen approximately 7000 compounds (Lee et al. 2012). Among the hits, SKF-86466 was identified as the molecule inducing the transcription of IKBKAP, which is responsible for FD. Treatment with SKF-86466 also rescued the disease-specific loss of autonomic neuron marker expression. In the third study, HTS was performed using hepatocytes differentiated from iPSCs derived from a patient with α−1 antitrypsin deficiency. A screening of more than 3000 clinically-approved compounds in the Johns Hopkins drug library identified five clinical drugs that ameliorated the disease phenotype and could be rapidly tested in clinical trials as novel therapeutics (Choi et al. 2013). These studies demonstrated the usefulness of cells differentiated from patient-derived iPSCs in accurately reflecting drug response in human patients when compared with the use of immortalized cell lines and animal models. Furthermore, these results suggested that patient-derived iPSCs enable the simple evaluation of previously approved drugs on different disease models, thereby promoting the discovery of potential new indications.
Toxicity assessment using iPSCs
Many approved drugs have been subsequently withdrawn from the market due to safety issues. From 1980 to 2009, 118 drugs were withdrawn from the market, with approximately 22% of the withdrawn drugs discontinued based on their toxicities (Qureshi et al. 2011). These post-marketing failures of approved drugs are attributable at least in part to conventional drug-safety assays using animal models. This has resulted in an increased focus on determining the ability of human iPSCs to predict adverse drug response in patients having different genetic backgrounds.
Cardiotoxicity is among the major reasons for drug withdrawals. Based on improved protocols for the differentiation of pluripotent stem cells into cardiomyocytes, several attempts were undertaken to use cardiomyocytes differentiated from human iPSCs for assessment of a drug’s cardiotoxicity. A recent study reported that human iPSC-derived cardiomyocytes can serve as a sensitive and robust platform for testing drug-induced arrhythmias (Navarrete et al. 2013). In another study, human iPSCs were generated from patients with various hereditary cardiac disorders including hereditary LQTS, familial hypertrophic cardiomyopathy, and familial dilated cardiomyopathy (Liang et al. 2013). Cardiomyocytes that were differentiated from these iPSCs, represented disease phenotypes and were used to assess susceptibility to several known cardiotoxic drugs. Additionally, cardiomyocytes from these patient-specific iPSCs exhibited greater sensitivity to drug-induced cardiac toxicity as compared with controls, demonstrating their value in predicting different susceptibility to drugs in patients with different genetic backgrounds. These findings also suggest that they be suitably included in current protocols for preclinical drug-metabolism and toxicity screening.
Drug-induced hepatic toxicity is another leading cause of drug withdrawal. Takayama et al. reported that human iPSC-derived hepatocytes have the potential to predict inter-individual differences in drug-metabolism and drug response (Takayama et al. 2014). Since genetic polymorphisms in cytochrome P450 are responsible for the differences in every individual’s ability to metabolize drug molecules, diverse human iPSCs were generated from individuals having different single-nucleotide polymorphisms in the CYP2D6 gene. When compared to parental primary human hepatocytes, those derived from human iPSCs retained donor-specific cytochrome P450-activity levels and drug responsiveness. These findings suggested that human iPSC-derived hepatocytes would be a powerful tool not only for identification of patient populations at high-risk for hepatic toxicity, but also for stratification of patients based on drug responsiveness.
Challenges of human iPSC-based disease modeling and drug discovery
Human iPSCs versus ESCs
Human iPSCs are highly similar to human ESCs in terms of marker expression, self-renewal capacity, and differentiation potential. However, more refined genome-wide genetic and epigenetic analyses indicated several differences between these cells, including the persistence of epigenetic memory in human iPSCs, different DNA-methylation signatures, and different extents of genetic aberrations (Robinton and Daley 2012). Since the use of human iPSCs for disease modeling and drug discovery is based on the assumption that human ESCs can be replaced with human iPSCs, it is important to investigate whether subtle differences between them might affect experimental results. The first comparison of human ESC and iPSC disease models was conducted for fragile X syndrome (FXS), which is a common cause of inherited intellectual disability in boys. The mutation leading to FXS is a trinucleotide CGG expansion at the 5′-untranslated region of the FMR1 gene accompanied by epigenetic modification at the promoter region and subsequent silencing of transcription. Studies of human ESCs from FXS-affected embryos showed that FMR1 was expressed in mutated undifferentiated ESCs, but was transcriptionally silenced during differentiation (Eiges et al. 2007; Turetsky et al. 2008; Telias et al. 2013). These findings demonstrated that the FMR1 gene is inactivated at early embryonic developmental stages and that the trinucleotide CGG expansion is necessary, but not sufficient for gene silencing. However, a study of human iPSCs derived from FXS patients reported contrasting results (Urbach et al. 2010). In the undifferentiated iPSC model, the FMR1 gene remained methylated and silenced, which suggests that human iPSCs exhibit epigenetic memory derived from patient somatic cells. This result indicated that human iPSCs are unable to recapitulate certain diseases. It is sometimes worthwhile to generate both ESC and iPSC models for the same disease and investigate similarities and differences between them.
Modeling late-onset disease models
Many neurodegenerative diseases, such as AD and PD, are late-onset diseases, and their disease phenotypes may not manifest themselves over short periods of in vitro culture. Therefore, several attempts were made to artificially age cells differentiated from human iPSCs to enable modeling of late-onset diseases. In a recent study, progerin, which is a truncated form of the lamin A protein involved in premature aging during development of Hutchinson-Gilford progeria syndrome, was ectopically expressed to promote cellular aging processes in dopaminergic neurons differentiated from PD patient-derived iPSCs (Liu et al. 2011). Progerin expression induced PD-related features, such as dendrite degeneration, loss of tyrosine hydroxylase expression, and morphological features, such as enlarged mitochondria or Lewy body precursor inclusions. Additionally, stressors, such as oxidative stress, growth factor deficiency, and excitotoxicity, have also been used to promote the cellular aging process in cells differentiated from human iPSCs (Koch et al. 2011; Nguyen et al. 2011; An et al. 2012).
Selection and generation of appropriate controls
Another critical issue in human iPSC-based disease modeling and drug discovery is the selection and generation of appropriate non-disease controls. Human iPSC lines are variable in their differentiation propensities and phenotypic output due to the incomplete silencing of reprogramming factors, clonal variation, and different genetic backgrounds; it is sometimes difficult to detect disease-specific phenotypes in cells derived from human iPSCs. Inoue et al. suggested that deductive and/or inductive controls are required to set to solve this problem (Inoue and Yamanaka 2011; Inoue et al. 2014). As deductive controls, isogenic cells are generated by inserting or deleting disease-linked gene variants in iPSCs, which are then compared with the controls. For example, disease-linked gene variants in patient-derived iPSCs can be replaced with the wild-type gene, which is called “gene-corrected” deductive control. Conversely, disease-linked gene variants can be inserted into endogenous wild-type gene loci in non-disease iPSCs, which is called “gene-edited” controls (Merkle and Eggan 2013). Comparison of human iPSCs with their isogenic controls helps to determine phenotypic manifestation of candidate disease-linked genes, but not identify other genetic variants contributing to a disease phenotype. On the other hands, inductive controls include iPSCs derived from healthy individuals or from other patients. Although inductive controls are less complicated to generate as compared with deductive controls, generation of multiple different iPSC lines is required to offset the genetic variability present in different individuals.
Future perspective
Despite the challenges to implementing human iPSCs in drug discovery, these cells have great potential to improve the translatability of preclinical information leading to the clinic. As shown in Fig. 1, human iPSCs can be used in several stages of the drug-discovery process. Currently, human iPSCs are used mainly during the earliest stages, where cells are employed as models for understanding human pathogenesis or drug-screening platforms for assessing efficacy and toxicity. As the numbers of genetically diverse human iPSCs increase and efficient and robust differentiation protocols become available, it is possible that the use of human iPSCs will extend to screening patients who are most likely to have therapeutic effects and least likely to experience drug toxicity. This concept, referred to as an “in vitro clinical trial,” involving human iPSCs has enabled identification of patient subsets likely to response to the drug being studied (Table 2). Such a stratification of patients based on drug responsiveness will increase the success rate of clinical trials.
Contribution of human iPSCs to drug discovery process. In the drug discovery process, human induced pluripotent stem cells (iPSCs) have been employed as cellular disease models for identifying new disease targets and drug screening platforms for assessing efficacy and toxicity. The use of human iPSCs is expected to be expanded to pre-select drug responders for clinical trials (patient stratification) and for the development of an optimal therapeutic strategy for each patient (precision medicine)
With the availability of next-generation sequencing and bioinformatics, patient-derived iPSCs can also be used to obtain information regarding interactions between genotype, phenotype, and drug response. This pharmacogenomic information may help to identify specific genetic markers in drug responders, and consequently, could lead to a new type of diagnosis and stratification. For example, patients with complex genetic disorders, such as AD, react differently to medications (Freund-Levi et al. 2006; Quinn et al. 2010; Yaffe 2010). In such cases, panels of patient iPSC-derived neurons that represent the genetic variation of AD patients can be established to classify patients for the appropriate treatment and develop the optimal therapeutic strategy for each individual patient in a process known as precision medicine.
With growing recognition of the potential of human iPSCs for disease modeling and drug discovery, several large-scale initiatives (Human Induced Pluripotent Stem Cell Initiative, StemBANCC, California Institute of Regenerative Medicine, New York Stem Cell Foundation, etc.) have been created to establish human iPSC repositories with comprehensive clinical and genetic information and to make them available as worldwide research resources (Soares et al. 2014). It may not take long to see examples of successful application of human iPSCs in biomedical science.
References
An MC, Zhang N, Scott G, Montoro D, Wittkop T, Mooney S, Melov S, Ellerby LM (2012) Genetic correction of Huntington’s disease phenotypes in induced pluripotent stem cells. Cell Stem Cell 11:253–263
Avior Y, Sagi I, Benvenisty N (2016) Pluripotent stem cells in disease modelling and drug discovery. Nat Rev Mol Cell Biol 17:170–182
Barmada SJ, Serio A, Arjun A, Bilican B, Daub A, Ando DM, Tsvetkov A, Pleiss M, Li X, Peisach D, Shaw C, Chandran S, Finkbeiner S (2014) Autophagy induction enhances TDP43 turnover and survival in neuronal ALS models. Nat Chem Biol 10:677–685
Brennand KJ, Simone A, Jou J, Gelboin-Burkhart C, Tran N, Sangar S, Li Y, Mu Y, Chen G, Yu D, Mccarthy S, Sebat J, Gage FH (2011) Modelling schizophrenia using human induced pluripotent stem cells. Nature 473:221–225
Burkhardt MF, Martinez FJ, Wright S, Ramos C, Volfson D, Mason M, Garnes J, Dang V, Lievers J, Shoukat-Mumtaz U, Martinez R, Gai H, Blake R, Vaisberg E, Grskovic M, Johnson C, Irion S, Bright J, Cooper B, Nguyen L, Griswold-Prenner I, Javaherian A (2013) A cellular model for sporadic ALS using patient-derived induced pluripotent stem cells. Mol Cell Neurosci 56:355–364
Chang CY, Chen SM, Lu HE, Lai SM, Lai PS, Shen PW, Chen PY, Shen CI, Harn HJ, Lin SZ, Hwang SM, Su HL (2015) N-butylidenephthalide attenuates Alzheimer’s disease-like cytopathy in down syndrome induced pluripotent stem cell-derived neurons. Sci Rep 5:8744
Charbord J, Poydenot P, Bonnefond C, Feyeux M, Casagrande F, Brinon B, Francelle L, Auregan G, Guillermier M, Cailleret M, Viegas P, Nicoleau C, Martinat C, Brouillet E, Cattaneo E, Peschanski M, Lechuga M, Perrier AL (2013) High throughput screening for inhibitors of REST in neural derivatives of human embryonic stem cells reveals a chemical compound that promotes expression of neuronal genes. Stem Cells 31:1816–1828
Chen C, Jiang P, Xue H, Peterson SE, Tran HT, Mccann AE, Parast MM, Li S, Pleasure DE, Laurent LC, Loring JF, Liu Y, Deng W (2014) Role of astroglia in Down’s syndrome revealed by patient-derived human-induced pluripotent stem cells. Nat Commun 5:4430
Choi SM, Kim Y, Shim JS, Park JT, Wang RH, Leach SD, Liu JO, Deng C, Ye Z, Jang YY (2013) Efficient drug screening and gene correction for treating liver disease using patient-specific stem cells. Hepatology 57:2458–2468
Chung CY, Khurana V, Auluck PK, Tardiff DF, Mazzulli JR, Soldner F, Baru V, Lou Y, Freyzon Y, Cho S, Mungenast AE, Muffat J, Mitalipova M, Pluth MD, Jui NT, Schule B, Lippard SJ, Tsai LH, Krainc D, Buchwald SL, Jaenisch R, Lindquist S (2013) Identification and rescue of alpha-synuclein toxicity in Parkinson patient-derived neurons. Science 342:983–987
Church DM, Goodstadt L, Hillier LW, Zody MC, Goldstein S, She X, Bult CJ, Agarwala R, Cherry JL, Dicuccio M, Hlavina W, Kapustin Y, Meric P, Maglott D, Birtle Z, Marques AC, Graves T, Zhou S, Teague B, Potamousis K, Churas C, Place M, Herschleb J, Runnheim R, Forrest D, Amos-Landgraf J, Schwartz DC, Cheng Z, Lindblad-Toh K, Eichler EE, Ponting CP (2009) Lineage-specific biology revealed by a finished genome assembly of the mouse. PLoS Biol 7:e1000112
Cooper O, Seo H, Andrabi S, Guardia-Laguarta C, Graziotto J, Sundberg M, Mclean JR, Carrillo-Reid L, Xie Z, Osborn T, Hargus G, Deleidi M, Lawson T, Bogetofte H, Perez-Torres E, Clark L, Moskowitz C, Mazzulli J, Chen L, Volpicelli-Daley L, Romero N, Jiang H, Uitti RJ, Huang Z, Opala G, Scarffe LA, Dawson VL, Klein C, Feng J, Ross OA, Trojanowski JQ, Lee VM, Marder K, Surmeier DJ, Wszolek ZK, Przedborski S, Krainc D, Dawson TM, Isacson O (2012) Pharmacological rescue of mitochondrial deficits in iPSC-derived neural cells from patients with familial Parkinson’s disease. Sci Transl Med 4:141RA90
Ding J, Chen X, Gao Z, Dai X, Li L, Xie C, Jiang H, Zhang L, Zhong D (2013) Metabolism and pharmacokinetics of novel selective vascular endothelial growth factor receptor-2 inhibitor apatinib in humans. Drug Metab Dispos 41:1195–1210
Ebert AD, Yu J, Rose FF Jr, Mattis VB, Lorson CL, Thomson JA, Svendsen CN (2009) Induced pluripotent stem cells from a spinal muscular atrophy patient. Nature 457:277–280
Egawa N, Kitaoka S, Tsukita K, Naitoh M, Takahashi K, Yamamoto T, Adachi F, Kondo T, Okita K, Asaka I, Aoi T, Watanabe A, Yamada Y, Morizane A, Takahashi J, Ayaki T, Ito H, Yoshikawa K, Yamawaki S, Suzuki S, Watanabe D, Hioki H, Kaneko T, Makioka K, Okamoto K, Takuma H, Tamaoka A, Hasegawa K, Nonaka T, Hasegawa M, Kawata A, Yoshida M, Nakahata T, Takahashi R, Marchetto MC, Gage FH, Yamanaka S, Inoue H (2012) Drug screening for ALS using patient-specific induced pluripotent stem cells. Sci Transl Med 4:145RA104
Eiges R, Urbach A, Malcov M, Frumkin T, Schwartz T, Amit A, Yaron Y, Eden A, Yanuka O, Benvenisty N, Ben-Yosef D (2007) Developmental study of fragile X syndrome using human embryonic stem cells derived from preimplantation genetically diagnosed embryos. Cell Stem Cell 1:568–577
Freund-Levi Y, Eriksdotter-Jonhagen M, Cederholm T, Basun H, Faxen-Irving G, Garlind A, Vedin I, Vessby B, Wahlund LO, Palmblad J (2006) Omega-3 fatty acid treatment in 174 patients with mild to moderate Alzheimer disease: OmegAD study: a randomized double-blind trial. Arch Neurol 63:1402–1408
Fu M, Zhang J, Lin Y, Zhu X, Ehrengruber MU, Chen YE (2002) Early growth response factor-1 is a critical transcriptional mediator of peroxisome proliferator-activated receptor-gamma 1 gene expression in human aortic smooth muscle cells. J Biol Chem 277:26808–26814
Fusaki N, Ban H, Nishiyama A, Saeki K, Hasegawa M (2009) Efficient induction of transgene-free human pluripotent stem cells using a vector based on Sendai virus, an RNA virus that does not integrate into the host genome. Proc Jpn Acad Ser B 85:348–362
Garbes L, Heesen L, Holker I, Bauer T, Schreml J, Zimmermann K, Thoenes M, Walter M, Dimos J, Peitz M, Brustle O, Heller R, Wirth B (2013) VPA response in SMA is suppressed by the fatty acid translocase CD36. Hum Mol Genet 22:398–407
Germain ND, Chen PF, Plocik AM, Glatt-Deeley H, Brown J, Fink JJ, Bolduc KA, Robinson TM, Levine ES, Reiter LT, Graveley BR, Lalande M, Chamberlain SJ (2014) Gene expression analysis of human induced pluripotent stem cell-derived neurons carrying copy number variants of chromosome 15q11-q13.1. Mol Autism 5:44
Griesi-Oliveira K, Acab A, Gupta AR, Sunaga DY, Chailangkarn T, Nicol X, Nunez Y, Walker MF, Murdoch JD, Sanders SJ, Fernandez TV, Ji W, Lifton RP, Vadasz E, Dietrich A, Pradhan D, Song H, Ming GL, Gu X, Haddad G, Marchetto MC, Spitzer N, Passos-Bueno MR, State MW, Muotri AR (2015) Modeling non-syndromic autism and the impact of TRPC6 disruption in human neurons. Mol Psychiatry 20:1350–1365
Guo X, Disatnik MH, Monbureau M, Shamloo M, Mochly-Rosen D, Qi X (2013) Inhibition of mitochondrial fragmentation diminishes Huntington’s disease-associated neurodegeneration. J Clin Invest 123:5371–5388
Gurdon JB (1962) The developmental capacity of nuclei taken from intestinal epithelium cells of feeding tadpoles. J Embryol Exp Morphol 10:622–640
Gurdon JB, Elsdale TR, Fischberg M (1958) Sexually mature individuals of Xenopus laevis from the transplantation of single somatic nuclei. Nature 182:64–65
Hamlin RL, Altschuld RA (2011) Extrapolation from mouse to man. Circ Cardiovasc Imaging 4:2–4
Hay M, Thomas DW, Craighead JL, Economides C, Rosenthal J (2014) Clinical development success rates for investigational drugs. Nat Biotechnol 32:40–51
Hibaoui Y, Grad I, Letourneau A, Sailani MR, Dahoun S, Santoni FA, Gimelli S, Guipponi M, Pelte MF, Bena F, Antonarakis SE, Feki A (2014) Modelling and rescuing neurodevelopmental defect of Down syndrome using induced pluripotent stem cells from monozygotic twins discordant for trisomy 21. EMBO Mol Med 6:259–277
Hou P, Li Y, Zhang X, Liu C, Guan J, Li H, Zhao T, Ye J, Yang W, Liu K, Ge J, Xu J, Zhang Q, Zhao Y, Deng H (2013) Pluripotent stem cells induced from mouse somatic cells by small-molecule compounds. Science 341:651–654
Inoue H, Yamanaka S (2011) The use of induced pluripotent stem cells in drug development. Clin Pharmacol Ther 89:655–661
Inoue H, Nagata N, Kurokawa H, Yamanaka S (2014) iPS cells: a game changer for future medicine. EMBO J 33:409–417
Israel MA, Yuan SH, Bardy C, Reyna SM, Mu Y, Herrera C, Hefferan MP, Van Gorp S, Nazor KL, Boscolo FS, Carson CT, Laurent LC, Marsala M, Gage FH, Remes AM, Koo EH, Goldstein LS (2012) Probing sporadic and familial Alzheimer’s disease using induced pluripotent stem cells. Nature 482:216–220
Itzhaki I, Maizels L, Huber I, Zwi-Dantsis L, Caspi O, Winterstern A, Feldman O, Gepstein A, Arbel G, Hammerman H, Boulos M, Gepstein L (2011) Modelling the long QT syndrome with induced pluripotent stem cells. Nature 471:225–229
Jia F, Wilson KD, Sun N, Gupta DM, Huang M, Li Z, Panetta NJ, Chen ZY, Robbins RC, Kay MA, Longaker MT, Wu JC (2010) A nonviral minicircle vector for deriving human iPS cells. Nat Method 7:197–199
Jin ZB, Okamoto S, Osakada F, Homma K, Assawachananont J, Hirami Y, Iwata T, Takahashi M (2011) Modeling retinal degeneration using patient-specific induced pluripotent stem cells. PLoS ONE 6:e17084
Jung HJ, Park K, Kim JJ, Lee JH, Han KO, Han DK (2008) Effect of RGD-immobilized dual-pore poly(L-lactic acid) scaffolds on chondrocyte proliferation and extracellular matrix production. Artif Organs 32:981–989
Kaji K, Norrby K, Paca A, Mileikovsky M, Mohseni P, Woltjen K (2009) Virus-free induction of pluripotency and subsequent excision of reprogramming factors. Nature 458:771–775
Kim H, Kim JS (2014) A guide to genome engineering with programmable nucleases. Nat Rev Genet 15:321–334
Kim KT, Choi HH, Steinmetz MO, Maco B, Kammerer RA, Ahn SY, Kim HZ, Lee GM, Koh GY (2005) Oligomerization and multimerization are critical for angiopoietin-1 to bind and phosphorylate Tie2. J Biol Chem 280:20126–20131
Kim D, Kim CH, Moon JI, Chung YG, Chang MY, Han BS, Ko S, Yang E, Cha KY, Lanza R, Kim KS (2009) Generation of human induced pluripotent stem cells by direct delivery of reprogramming proteins. Cell Stem Cell 4:472–476
Kim KL, Song SH, Choi KS, Suh W (2013a) Cooperation of endothelial and smooth muscle cells derived from human induced pluripotent stem cells enhances neovascularization in dermal wounds. Tissue Eng Part A 19:2478–2485
Kim KL, Yang JH, Song SH, Kim JY, Jang SY, Kim JM, Kim JA, Sung KI, Kim YW, Suh YL, Suh W, Kim DK (2013b) Positive correlation between the dysregulation of transforming growth factor-beta1 and aneurysmal pathological changes in patients with Marfan syndrome. Circ J 77:952–958
Koch P, Breuer P, Peitz M, Jungverdorben J, Kesavan J, Poppe D, Doerr J, Ladewig J, Mertens J, Tuting T, Hoffmann P, Klockgether T, Evert BO, Wullner U, Brustle O (2011) Excitation-induced ataxin-3 aggregation in neurons from patients with Machado-Joseph disease. Nature 480:543–546
Kondo T, Asai M, Tsukita K, Kutoku Y, Ohsawa Y, Sunada Y, Imamura K, Egawa N, Yahata N, Okita K, Takahashi K, Asaka I, Aoi T, Watanabe A, Watanabe K, Kadoya C, Nakano R, Watanabe D, Maruyama K, Hori O, Hibino S, Choshi T, Nakahata T, Hioki H, Kaneko T, Naitoh M, Yoshikawa K, Yamawaki S, Suzuki S, Hata R, Ueno S, Seki T, Kobayashi K, Toda T, Murakami K, Irie K, Klein WL, Mori H, Asada T, Takahashi R, Iwata N, Yamanaka S, Inoue H (2013) Modeling Alzheimer’s disease with iPSCs reveals stress phenotypes associated with intracellular Abeta and differential drug responsiveness. Cell Stem Cell 12:487–496
Kreuger J, Nilsson I, Kerjaschki D, Petrova T, Alitalo K, Claesson-Welsh L (2006) Early lymph vessel development from embryonic stem cells. Arterioscler Thromb Vasc Biol 26:1073–1078
Lee G, Papapetrou EP, Kim H, Chambers SM, Tomishima MJ, Fasano CA, Ganat YM, Menon J, Shimizu F, Viale A, Tabar V, Sadelain M, Studer L (2009) Modelling pathogenesis and treatment of familial dysautonomia using patient-specific iPSCs. Nature 461:402–406
Lee G, Ramirez CN, Kim H, Zeltner N, Liu B, Radu C, Bhinder B, Kim YJ, Choi IY, Mukherjee-Clavin B, Djaballah H, Studer L (2012) Large-scale screening using familial dysautonomia induced pluripotent stem cells identifies compounds that rescue IKBKAP expression. Nat Biotechnol 30:1244–1248
Lee H, Lee JK, Park MH, Hong YR, Marti HH, Kim H, Okada Y, Otsu M, Seo EJ, Park JH, Bae JH, Okino N, He X, Schuchman EH, Bae JS, Jin HK (2014) Pathological roles of the VEGF/SphK pathway in Niemann-Pick type C neurons. Nat Commun 5:5514
Liang P, Lan F, Lee AS, Gong T, Sanchez-Freire V, Wang Y, Diecke S, Sallam K, Knowles JW, Wang PJ, Nguyen PK, Bers DM, Robbins RC, Wu JC (2013) Drug screening using a library of human induced pluripotent stem cell-derived cardiomyocytes reveals disease-specific patterns of cardiotoxicity. Circulation 127:1677–1691
Liu GH, Barkho BZ, Ruiz S, Diep D, Qu J, Yang SL, Panopoulos AD, Suzuki K, Kurian L, Walsh C, Thompson J, Boue S, Fung HL, Sancho-Martinez I, Zhang K, Yates J 3rd, Izpisua Belmonte JC (2011) Recapitulation of premature ageing with iPSCs from Hutchinson-Gilford progeria syndrome. Nature 472:221–225
Liu Q, Waltz S, Woodruff G, Ouyang J, Israel MA, Herrera C, Sarsoza F, Tanzi RE, Koo EH, Ringman JM, Goldstein LS, Wagner SL, Yuan SH (2014) Effect of potent gamma-secretase modulator in human neurons derived from multiple presenilin 1-induced pluripotent stem cell mutant carriers. JAMA Neurol 71:1481–1489
Lu S, Kanekura K, Hara T, Mahadevan J, Spears LD, Oslowski CM, Martinez R, Yamazaki-Inoue M, Toyoda M, Neilson A, Blanner P, Brown CM, Semenkovich CF, Marshall BA, Hershey T, Umezawa A, Greer PA, Urano F (2014a) A calcium-dependent protease as a potential therapeutic target for Wolfram syndrome. Proc Natl Acad Sci USA 111:E5292–E5301
Lu XH, Mattis VB, Wang N, Al-Ramahi I, Van Den Berg N, Fratantoni SA, Waldvogel H, Greiner E, Osmand A, Elzein K, Xiao J, Dijkstra S, De Pril R, Vinters HV, Faull R, Signer E, Kwak S, Marugan JJ, Botas J, Fischer DF, Svendsen CN, Munoz-Sanjuan I, Yang XW (2014b) Targeting ATM ameliorates mutant Huntingtin toxicity in cell and animal models of Huntington’s disease. Sci Transl Med. 6:268RA178
Maetzel D, Sarkar S, Wang H, Abi-Mosleh L, Xu P, Cheng AW, Gao Q, Mitalipova M, Jaenisch R (2014) Genetic and chemical correction of cholesterol accumulation and impaired autophagy in hepatic and neural cells derived from Niemann-Pick Type C patient-specific iPS cells. Stem Cell Rep 2:866–880
Marchetto MC, Carromeu C, Acab A, Yu D, Yeo GW, Mu Y, Chen G, Gage FH, Muotri AR (2010) A model for neural development and treatment of Rett syndrome using human induced pluripotent stem cells. Cell 143:527–539
Merkle FT, Eggan K (2013) Modeling human disease with pluripotent stem cells: from genome association to function. Cell Stem Cell 12:656–668
Navarrete EG, Liang P, Lan F, Sanchez-Freire V, Simmons C, Gong T, Sharma A, Burridge PW, Patlolla B, Lee AS, Wu H, Beygui RE, Wu SM, Robbins RC, Bers DM, Wu JC (2013) Screening drug-induced arrhythmia [corrected] using human induced pluripotent stem cell-derived cardiomyocytes and low-impedance microelectrode arrays. Circulation 128:S3–S13
Ng SY, Soh BS, Rodriguez-Muela N, Hendrickson DG, Price F, Rinn JL, Rubin LL (2015) Genome-wide RNA-Seq of human motor neurons implicates selective ER stress activation in spinal muscular atrophy. Cell Stem Cell 17:569–584
Nguyen HN, Byers B, Cord B, Shcheglovitov A, Byrne J, Gujar P, Kee K, Schule B, Dolmetsch RE, Langston W, Palmer TD, Pera RR (2011) LRRK2 mutant iPSC-derived DA neurons demonstrate increased susceptibility to oxidative stress. Cell Stem Cell 8:267–280
Nishimura K, Sano M, Ohtaka M, Furuta B, Umemura Y, Nakajima Y, Ikehara Y, Kobayashi T, Segawa H, Takayasu S, Sato H, Motomura K, Uchida E, Kanayasu-Toyoda T, Asashima M, Nakauchi H, Yamaguchi T, Nakanishi M (2011) Development of defective and persistent Sendai virus vector: a unique gene delivery/expression system ideal for cell reprogramming. J Biol Chem 286:4760–4771
Okita K, Nakagawa M, Hyenjong H, Ichisaka T, Yamanaka S (2008) Generation of mouse induced pluripotent stem cells without viral vectors. Science 322:949–953
Paul SM, Mytelka DS, Dunwiddie CT, Persinger CC, Munos BH, Lindborg SR, Schacht AL (2010) How to improve R&D productivity: the pharmaceutical industry’s grand challenge. Nat Rev Drug Discov 9:203–214
Paulsen Bda S, De Moraes Maciel R, Galina A, Souza Da Silveira M, Dos Santos Souza C, Drummond H, Nascimento Pozzatto E, Silva H Jr, Chicaybam L, Massuda R, Setti-Perdigao P, Bonamino M, Belmonte-De-Abreu PS, Castro NG, Brentani H, Rehen SK (2012) Altered oxygen metabolism associated to neurogenesis of induced pluripotent stem cells derived from a schizophrenic patient. Cell Transpl 21:1547–1559
Paulsen Bda S, Cardoso SC, Stelling MP, Cadilhe DV, Rehen SK (2014) Valproate reverts zinc and potassium imbalance in schizophrenia-derived reprogrammed cells. Schizophr Res 154:30–35
Quinn JF, Raman R, Thomas RG, Yurko-Mauro K, Nelson EB, Van Dyck C, Galvin JE, Emond J, Jack CR Jr, Weiner M, Shinto L, Aisen PS (2010) Docosahexaenoic acid supplementation and cognitive decline in Alzheimer disease: a randomized trial. JAMA 304:1903–1911
Qureshi ZP, Seoane-Vazquez E, Rodriguez-Monguio R, Stevenson KB, Szeinbach SL (2011) Market withdrawal of new molecular entities approved in the United States from 1980 to 2009. Pharmacoepidemiol Drug Saf 20:772–777
Ren Y, Jiang H, Hu Z, Fan K, Wang J, Janoschka S, Wang X, Ge S, Feng J (2015) Parkin mutations reduce the complexity of neuronal processes in iPSC-derived human neurons. Stem Cells 33:68–78
Robinton DA, Daley GQ (2012) The promise of induced pluripotent stem cells in research and therapy. Nature 481:295–305
Ryan SD, Dolatabadi N, Chan SF, Zhang X, Akhtar MW, Parker J, Soldner F, Sunico CR, Nagar S, Talantova M, Lee B, Lopez K, Nutter A, Shan B, Molokanova E, Zhang Y, Han X, Nakamura T, Masliah E, Yates JR 3rd, Nakanishi N, Andreyev AY, Okamoto S, Jaenisch R, Ambasudhan R, Lipton SA (2013) Isogenic human iPSC Parkinson’s model shows nitrosative stress-induced dysfunction in MEF2-PGC1alpha transcription. Cell 155:1351–1364
Sala L, Yu Z, Ward-Van Oostwaard D, Van Veldhoven JP, Moretti A, Laugwitz KL, Mummery CL, Ap IJ, Bellin M (2016) A new hERG allosteric modulator rescues genetic and drug-induced long-QT syndrome phenotypes in cardiomyocytes from isogenic pairs of patient induced pluripotent stem cells. EMBO Mol Med 8:1065–1081
Sanchez-Danes A, Richaud-Patin Y, Carballo-Carbajal I, Jimenez-Delgado S, Caig C, Mora S, Di Guglielmo C, Ezquerra M, Patel B, Giralt A, Canals JM, Memo M, Alberch J, Lopez-Barneo J, Vila M, Cuervo AM, Tolosa E, Consiglio A, Raya A (2012) Disease-specific phenotypes in dopamine neurons from human iPS-based models of genetic and sporadic Parkinson’s disease. EMBO Mol Med 4:380–395
Sareen D, Ebert AD, Heins BM, Mcgivern JV, Ornelas L, Svendsen CN (2012) Inhibition of apoptosis blocks human motor neuron cell death in a stem cell model of spinal muscular atrophy. PLoS ONE 7:e39113
Sharma A, Marceau C, Hamaguchi R, Burridge PW, Rajarajan K, Churko JM, Wu H, Sallam KI, Matsa E, Sturzu AC, Che Y, Ebert A, Diecke S, Liang P, Red-Horse K, Carette JE, Wu SM, Wu JC (2014) Human induced pluripotent stem cell-derived cardiomyocytes as an in vitro model for coxsackievirus B3-induced myocarditis and antiviral drug screening platform. Circ Res 115:556–566
Soares FA, Sheldon M, Rao M, Mummery C, Vallier L (2014) International coordination of large-scale human induced pluripotent stem cell initiatives: wellcome trust and ISSCR workshops white paper. Stem Cell Rep 3:931–939
Soga M, Ishitsuka Y, Hamasaki M, Yoneda K, Furuya H, Matsuo M, Ihn H, Fusaki N, Nakamura K, Nakagata N, Endo F, Irie T, Era T (2015) HPGCD outperforms HPBCD as a potential treatment for Niemann-Pick disease type C during disease modeling with iPS cells. Stem Cells 33:1075–1088
Stadtfeld M, Nagaya M, Utikal J, Weir G, Hochedlinger K (2008) Induced pluripotent stem cells generated without viral integration. Science 322:945–949
Takahashi K, Yamanaka S (2006) Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126:663–676
Takahashi K, Tanabe K, Ohnuki M, Narita M, Ichisaka T, Tomoda K, Yamanaka S (2007) Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131:861–872
Takayama K, Morisaki Y, Kuno S, Nagamoto Y, Harada K, Furukawa N, Ohtaka M, Nishimura K, Imagawa K, Sakurai F, Tachibana M, Sumazaki R, Noguchi E, Nakanishi M, Hirata K, Kawabata K, Mizuguchi H (2014) Prediction of interindividual differences in hepatic functions and drug sensitivity by using human iPS-derived hepatocytes. Proc Natl Acad Sci USA 111:16772–16777
Telias M, Segal M, Ben-Yosef D (2013) Neural differentiation of Fragile X human embryonic stem cells reveals abnormal patterns of development despite successful neurogenesis. Dev Biol 374:32–45
Turetsky T, Aizenman E, Gil Y, Weinberg N, Shufaro Y, Revel A, Laufer N, Simon A, Abeliovich D, Reubinoff BE (2008) Laser-assisted derivation of human embryonic stem cell lines from IVF embryos after preimplantation genetic diagnosis. Hum Reprod 23:46–53
Urbach A, Bar-Nur O, Daley GQ, Benvenisty N (2010) Differential modeling of fragile X syndrome by human embryonic stem cells and induced pluripotent stem cells. Cell Stem Cell 6:407–411
Wainger BJ, Kiskinis E, Mellin C, Wiskow O, Han SS, Sandoe J, Perez NP, Williams LA, Lee S, Boulting G, Berry JD, Brown RH Jr, Cudkowicz ME, Bean BP, Eggan K, Woolf CJ (2014) Intrinsic membrane hyperexcitability of amyotrophic lateral sclerosis patient-derived motor neurons. Cell Rep 7:1–11
Wakayama T, Perry AC, Zuccotti M, Johnson KR, Yanagimachi R (1998) Full-term development of mice from enucleated oocytes injected with cumulus cell nuclei. Nature 394:369–374
Williams EC, Zhong X, Mohamed A, Li R, Liu Y, Dong Q, Ananiev GE, Mok JC, Lin BR, Lu J, Chiao C, Cherney R, Li H, Zhang SC, Chang Q (2014) Mutant astrocytes differentiated from Rett syndrome patients-specific iPSCs have adverse effects on wild-type neurons. Hum Mol Genet 23:2968–2980
Wilmut I, Schnieke AE, Mcwhir J, Kind AJ, Campbell KH (1997) Viable offspring derived from fetal and adult mammalian cells. Nature 385:810–813
Woltjen K, Michael IP, Mohseni P, Desai R, Mileikovsky M, Hamalainen R, Cowling R, Wang W, Liu P, Gertsenstein M, Kaji K, Sung HK, Nagy A (2009) PiggyBac transposition reprograms fibroblasts to induced pluripotent stem cells. Nature 458:766–770
Yaffe K (2010) Treatment of Alzheimer disease and prognosis of dementia: time to translate research to results. JAMA 304:1952–1953
Yagi T, Ito D, Okada Y, Akamatsu W, Nihei Y, Yoshizaki T, Yamanaka S, Okano H, Suzuki N (2011) Modeling familial Alzheimer’s disease with induced pluripotent stem cells. Hum Mol Genet 20:4530–4539
Yoshida M, Kitaoka S, Egawa N, Yamane M, Ikeda R, Tsukita K, Amano N, Watanabe A, Morimoto M, Takahashi J, Hosoi H, Nakahata T, Inoue H, Saito MK (2015) Modeling the early phenotype at the neuromuscular junction of spinal muscular atrophy using patient-derived iPSCs. Stem Cell Rep 4:561–568
Yu J, Vodyanik MA, Smuga-Otto K, Antosiewicz-Bourget J, Frane JL, Tian S, Nie J, Jonsdottir GA, Ruotti V, Stewart R, Slukvin II, Thomson JA (2007) Induced pluripotent stem cell lines derived from human somatic cells. Science 318:1917–1920
Yu J, Hu K, Smuga-Otto K, Tian S, Stewart R, Slukvin II, Thomson JA (2009) Human induced pluripotent stem cells free of vector and transgene sequences. Science 324:797–801
Yu D, Swaroop M, Wang M, Baxa U, Yang R, Yan Y, Coksaygan T, Detolla L, Marugan JJ, Austin CP, Mckew JC, Gong DW, Zheng W (2014) Niemann-pick disease type C: induced pluripotent stem cell-derived neuronal cells for modeling neural disease and evaluating drug efficacy. J Biomol Screen 19:1164–1173
Yusa K, Rad R, Takeda J, Bradley A (2009) Generation of transgene-free induced pluripotent mouse stem cells by the piggyBac transposon. Nat Method 6:363–369
Zhang S, Chen S, Li W, Guo X, Zhao P, Xu J, Chen Y, Pan Q, Liu X, Zychlinski D, Lu H, Tortorella MD, Schambach A, Wang Y, Pei D, Esteban MA (2011) Rescue of ATP7B function in hepatocyte-like cells from Wilson’s disease induced pluripotent stem cells using gene therapy or the chaperone drug curcumin. Hum Mol Genet 20:3176–3187
Acknowledgements
This research was supported by the Bio & Medical Technology Development Program of the National Research Foundation (NRF) funded by the Ministry of Science, ICT & Future Planning (2012M3A9C6050368).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author declares that there is no conflict of interest.
Rights and permissions
About this article
Cite this article
Suh, W. A new era of disease modeling and drug discovery using induced pluripotent stem cells. Arch. Pharm. Res. 40, 1–12 (2017). https://doi.org/10.1007/s12272-016-0871-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12272-016-0871-0