Abstract
Huntington’s disease (HD) is an autosomal dominantly-inherited neurodegenerative disease, which is caused by CAG trinucleotide expansion in exon 1 of the Huntingtin (HTT) gene. Although HD is a rare disease, its monogenic nature makes it an ideal model in which to understand pathogenic mechanisms and to develop therapeutic strategies for neurodegenerative diseases. Clustered regularly-interspaced short palindromic repeats (CRISPR) is the latest technology for genome editing. Being simple to use and highly efficient, CRISPR-based genome-editing tools are rapidly gaining popularity in biomedical research and opening up new avenues for disease treatment. Here, we review the development of CRISPR-based genome-editing tools and their applications in HD research to offer a translational perspective on advancing the genome-editing technology to HD treatment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Huntington’s disease (HD) is an autosomal dominant neurodegenerative disease caused by a mutation in the Huntingtin (HTT) gene, which is localized on the short arm of chromosome 4 (4p16.3). The mutation is caused by the expansion of CAG trinucleotide repeats in exon 1 of HTT. In unaffected individuals, the number of CAG trinucleotide repeats in HTT varies from 6 to 35, whereas HD patients typically carry 40 or more CAG repeats [1,2,3,4]. The expanded CAG trinucleotide repeats are translated into a polyglutamine (polyQ) tract near the N-terminal region of the HTT protein, which renders the protein prone to misfold and aggregate. The mutant HTT protein confers gains of function that are neurotoxic, and the medium spiny neurons in the striatum are particularly vulnerable to such insults [5]. Therefore, the striatum is the most affected brain region in HD, but other regions, such as the cerebral cortex, globus pallidus, thalamus, subthalamic nucleus, substantia nigra, hypothalamus, and cerebellum can also be affected as the disease progresses. HD is a devastating disease, as most patients display characteristic symptoms such as chorea, motor dysfunction, psychiatric disturbance, and cognitive decline during middle age, which eventually leads to death in 15 to 20 years after the onset of symptoms [6]. Currently there is no effective treatment to halt or reverse the course of HD.
Since the HTT gene was identified as the causative gene for HD in 1993 [7], extensive efforts have been devoted to understanding HD pathogenesis, with the hope to eventually develop effective treatment options. Despite being a rare neurodegenerative disease, the monogenic background of HD makes it an ideal fit for such a challenging task: it is relatively easy to manipulate a single mutant gene to establish cellular and animal models; it is also quite straightforward that lowering HTT products should be able to alleviate neurotoxicity. Therefore, genome-editing technologies that can effectively manipulate individual genes, such as zinc finger nuclease (ZFN) and transcription activator-like effector nuclease (TALEN) [8,9,10], greatly facilitate HD-related research. This effect is much greater with the emergence of the latest technology, clustered regularly-interspaced short palindromic repeats (CRISPR) [11]. Here, we briefly review the development of CRISPR-based genome-editing technology. We also summarize the use of CRISPR in HD research from three aspects: generating animal models, studying disease mechanisms, and developing HD treatment strategies. Finally, we list the major innovations in CRISPR technology, and preview how to take advantage of such innovations in future HD research and treatment.
The Development of CRISPR Technology
The nucleotide sequence of the CRISPR system was first discovered by Ishino et al. in Escherichia coli in 1987 [12]. Since then, many similar sequences have been identified and given different names [13]. To avoid confusion related to the different nomenclatures, the acronym CRISPR was proposed in 2002 [14]. Although abundant CRISPR sequences have been identified, their biological function remained elusive. Subsequently, many studies have explored the potential function of CRISPR in cells. Most of the work has centered around adaptive immunity, as multiple research groups have independently reported that the spacers in CRISPR elements have high sequence homology with extrachromosomal elements [15,16,17,18]. These results triggered speculation that CRISPR may function as a protective mechanism against foreign genetic substances, such as phages and plasmids [19]. Indeed, most bacteria and archaea can incorporate the short invading DNA sequences into their own genome as CRISPRs [20,21,22], which helps them to acquire adaptive immunity to defend against the invading viruses [20]. In 2010, Garneau et al. revealed that the CRISPR-associated (Cas) protein, guided by short CRISPR RNAs, can specifically cleave bacteriophage and plasmid DNAs in a sequence-specific manner in vivo [23], thereby showing the great potential of the CRISPR/Cas system in genome editing. Two years later, the milestone study by Jinek et al. showed for the first time that the CRISPR/Cas9 system can be programmed to cut specific DNA sequences to generate double-stranded DNA breaks [24]. Since then, CRISPR-based genome-editing technology has begun to take off.
To date, more than 30 Cas proteins have been identified from different bacterial strains, and the number is rapidly expanding [25, 26], while CRISPR/Cas9 remains the major workhorse for genome editing in today’s research community. CRISPR/Cas9 consists of two parts, the Cas9 endonuclease and a synthetic single guide RNA (sgRNA). The sgRNA contains an ~20-nucleotide protospacer that pairs with the target DNA sequence, and a 3 to 6-nucleotide protospacer adjacent motif (PAM), which is indispensable for sgRNA-DNA hybridization [11]. In theory, CRISPR/Cas9 can edit any target DNA sequence by designing a proper protospacer sequence. Once the sgRNA recognizes the complementary sequences of the target gene, Cas9 binds to the sgRNA-DNA complex and cuts the DNA 3–4 nucleotides upstream of the correct PAM sequence. In eukaryotic cells, the cutting leads to a double-strand DNA break (DSB), which can be repaired by either homology-directed repair (HDR) or non-homologous end-joining (NHEJ). In most cases, the DSB is repaired by NHEJ, as it is a highly efficient but error-prone process. The random insertions or deletions caused by NHEJ can lead to frame-shift mutations or premature stop codons, so that a specific gene can be knocked out via this approach. Alternatively, HDR can precisely repair the DSB with a homologous DNA template. As a result, specific mutations can be introduced by providing a designed template to achieve gene knock-in (Fig. 1). Nonetheless, HDR efficiency is very low, especially in postmitotic cells, such as neurons [27, 28].
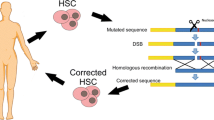
Schematic showing the genome-editing mechanism of CRISPR/Cas9. The designed single guide RNA (sgRNA) binds to its complementary sequence in the genome and recruits the Cas9 protein to that region. Cas9 utilizes its two distinct active motifs, RuvC and HNH, to generate site-specific nicks (3–4 nucleotides upstream of the PAM sequence) on the opposite DNA strands, causing a double-strand break (DSB). This DSB is repaired by two cellular mechanisms. The first is non-homologous end joining (NHEJ), which is efficient but error-prone. During the repair process, random nucleotide insertions or deletions are introduced near the DSB site, causing frame-shift mutations and gene knock-out. The other mechanism is homology directed repair (HDR), in which a donor DNA is used as a template to precisely repair the DSB. The donor DNA is specifically designed to achieve gene knock-in.
Using CRISPR Technology in HD Research
In the most common neurodegenerative diseases, such as Alzheimer’s disease (AD) and Parkinson’s disease (PD), ~90% of the cases are sporadic [29, 30], making it difficult to use genetic tools to model them. In contrast, HD is purely genetic, and is caused by a monogenic mutation. Early generations of genome-editing tools, such as ZFN and TALEN, have been successfully used in HD research [31,32,33]. Nonetheless, designing such tools requires special techniques, and is time-consuming and expensive. After the seminal work showing that CRISPR works in mammalian cells [34], CRISPR rapidly replaced ZFN and TALEN and became the favorite genome-editing tool in HD research. The major applications of CRISPR technology in HD research can be categorized into three aspects (Fig. 2).
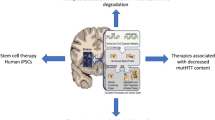
Summary of the major applications of CRISPR-based genome editing in Huntington’s disease (HD) research. CRISPR-base genome-editing technology is currently used in three areas of HD research: (1) establishing HD models, including isogenic cell lines, knock-in mouse models, and large animal models; (2) studying disease mechanisms, such as large-scale genetic screening and gene knock-out; (3) developing mutant HTT-lowering strategies.
Establishing Cellular and Animal Models
Disease models are essential for studying HD. Since HTT is a large gene spanning 180 kilobases and consisting of 67 exons, the conventional transgenic approach can only express DNA fragments that correspond to the neurotoxic N-terminal HTT proteins, which does not faithfully recapitulate HD conditions. In addition, overexpression of mutant proteins associated with the transgenic approach can lead to artificial or exaggerated pathological effects. Therefore, it is preferable to use a gene knock-in approach, so that mutant HTT toxicity can be studied under physiologically relevant conditions. CRISPR/Cas9, as a highly efficient genome-editing tool, greatly facilitates this process.
In 2014, CRISPR/Cas9 was first used to generate isogenic HD cellular models. By using CRISPR/Cas9 to induce DNA cutting in the region of exon 1 HTT and supplying a donor construct containing 97 CAG repeats, the researchers established human induced pluripotent stem cells (iPSCs) harboring 21, 72, or 97 CAG repeats in HTT [35]. Since then, several groups have used similar approaches to generate isogenic HD human iPSCs, and reported phenotypic abnormalities in such cells, including impaired neuronal differentiation, increased sensitivity to growth factor withdrawal, mitochondrial defects, and gene expression changes [36, 37]. These results suggest that isogenic HD cellular models are promising resources for mechanistic studies and drug screening.
Mouse models remain the primary platforms for HD-related research. Multiple lines of HD knock-in mice have already been established by different research groups [38,39,40,41]. Nonetheless, CRISPR/Cas9 offers more versatility in creating novel knock-in mouse models to test hypotheses related to HD pathogenesis. For example, a prevailing theory in HD research is the toxic fragment hypothesis, which means that full-length mutant HTT protein is proteolytically cleaved to generate N-terminal HTT fragments of different lengths that are neurotoxic [42, 43]. However, which N-terminal fragment(s) confers the most toxicity remains unknown. By using CRISPR/Cas9 to edit the HTT gene in the embryos of HD140Q knock-in mice, two knock-in mouse lines expressing different N-terminal HTT fragments (the first 96 or 571 amino-acids) have been established. Compared with full-length HD140Q mice, these two lines show similar neuropathology and disease progression, and they all contain a stable N-terminal mutant HTT fragment equivalent to exon 1 HTT, suggesting that exon 1 HTT is the key pathological form [44]. Another provocative hypothesis is that CAG repeat expansion in the HTT gene leads to repeat-associated non-AUG (RAN) translation to produce toxic peptides [45]. Via CRISPR/Cas9-mediated genome editing, a knock-in mouse model that expresses mutant HTT mRNA but not mutant HTT protein was established. RAN-translated products were not detected in this mouse model, nor were HD-related pathological changes, indicating that RAN translation does not play a major role in HD pathogenesis [46].
Knock-in mouse models are valuable resources for HD research, but they lack the overt neurodegeneration in the striatum, which is a typical hallmark in HD patients [47, 48]. In 2018, an HD knock-in pig model was established. By using CRISPR/Cas9 and a donor vector carrying human exon1 HTT with 150 CAG repeats, the researchers successfully knocked in mutant HTT in fetal pig fibroblasts. Somatic cell nuclear transfer was later applied to generate newborn HD knock-in pigs. This pig model displayed striking HD-like phenotypes and selective neurodegeneration in the striatum [49], supporting the rationale of using large animals to investigate HD pathogenesis and to explore potential therapeutics. Compared with the conventional method of homologous recombination, genome editing using CRISPR/Cas9 makes generating knock-in or knock-out animals much easier and faster, thereby turning the above research ideas into actual work.
Studying Disease Mechanisms
The plethora of disease models enabled extensive research on the pathogenic mechanisms of HD. A great number of biological pathways and individual genes have been linked to HD pathogenesis. In addition, as a large protein consisting of more than 3000 amino-acids, HTT is believed to interact with numerous proteins. Studies using different methods have identified hundreds of potential HTT-interacting proteins [50,51,52]. However, there are two major challenges to clarify HD pathogenic mechanisms: first, how to efficiently and reliably perform screening and second, how to find the key targets that are most relevant to HD pathogenesis. Because CRISPR/Cas9 can easily and efficiently knock out genes of interest, it has been rapidly adopted to address these issues.
Large-scale genetic screening is a powerful way to identify essential genes for HD toxicity. In 2020, an unbiased genome-wide genetic screening was performed in the mouse central nervous system, using CRISPR and shRNA lentivirus [53]. In wild-type mice, the researchers found that neurons are vulnerable to perturbations of synaptic function, autophagy, proteostasis, mRNA processing, and mitochondrial function. The same approach was later used to identify genes that are essential for neuronal viability in the presence of mutant HTT. Screening results using one HD transgenic mouse model and one HD knock-in mouse model revealed that genes involved in methylation-dependent chromatin silencing, dopamine signaling, and members of the Nme gene family are genetic modifiers of mutant HTT toxicity, suggesting these genes could be new targets for therapeutic interventions.
Manipulating the expression level of an individual gene and examining its influence on the neuropathology in HD models remain the gold standard to verify the large screening results. For example, several genes, including FAN1, RRM2B, and MLH1, have been identified as genetic modifiers through genome-wide association studies [54, 55]. CRISPR/Cas9 was used to generate knock-out mice for each gene. These knock-out mice were then crossed with HD knock-in mice so that the genetic modifying effect of an individual gene could be tested in vivo [56]. HAP1 was the first discovered protein that interacts with HTT [57]. Deletion of HAP1 via CRISRP/Cas9 in adult HD knock-in mice leads to selective neuronal loss in the striatum, and the neurotoxicity is mediated by Rhes, another HTT binding protein [58]. HspBP1 is an inhibitor of the carboxyl terminus of Hsp70-interacting protein, and functions as a regulator of protein quality control. Adeno-associated virus (AAV) carrying CRISPR/Cas9 was delivered to the striatum of HD knock-in mice to silence the expression of the HspBP1 gene, which reduced mutant HTT aggregates and attenuated neuropathology [59]. These are just a few examples of how CRISPR/Cas9 is enabling in-depth investigations of HD pathogenic mechanisms.
Developing Mutant HTT-lowering Strategies
Through more than 20 years of research, it is apparent that HD pathogenesis is quite complex. Mutant HTT causes abnormalities in a myriad of cellular functions [2], including dysregulation of transcription, impairment of protein homeostasis, mitochondrial dysfunction, defective vesicle transport, disrupted Ca2+ signaling, epigenetic-chromatin deregulation, and excitotoxicity (Fig. 3). It would be very challenging to rely on traditional chemical approaches to rectify most of these abnormalities. However, as a monogenic disease, eliminating mutant HTT products serves as the most straightforward therapeutic option. Substantial efforts have been devoted to developing mutant HTT-lowering therapies. Methods targeting mutant HTT mRNA, such as antisense oligonucleotide (ASO) and RNA interference (RNAi), are being actively pursued in both preclinical and clinical studies [60, 61]. Another promising approach is to use small-molecule compounds that specifically tether mutant HTT protein to the autophagosome for clearance [62]. By targeting mRNA or protein, these methods have successfully reduced mutant HTT production in an allele-specific or non-allele-specific manner, and have achieved therapeutic benefits in animal models of HD. Compared with these methods, CRISPR/Cas9 has one unique advantage: it targets the genome, so that one treatment should permanently inactivate mutant HTT expression.
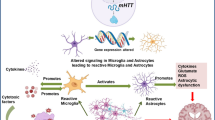
Summary of the major pathogenic mechanisms of Huntington’s disease (HD) currently identified. Multiple cellular functions are believed to be affected by the presence of mutant Huntingtin (HTT): (1) the N-terminal fragments of mutant HTT enter the nucleus, interact with selected transcription factors, such as specificity protein 1 (Sp1) and tumor protein P53, and disrupt their transcriptional activity; (2) mutant HTT causes epigenetic abnormalities and chromatin structural changes, possibly by directly binding to methyl-CpG binding protein 2 (MeCP2); (3) mutant HTT impairs the function of the proteasome ubiquitin system (UPS), which is the major mechanism for degrading misfolded proteins; (4) mutant HTT disrupts Ca2+ homeostasis and causes cytosolic Ca2+ overload; (5) mutant huntingtin triggers mitochondrial fragmentation and alters the mitochondrial proteome; (6) wild-type HTT is essential for axonal transport, whereas mutant HTT causes axonal transport defects; (7) mutant HTT inhibits glutamate uptake in glia, which leads to excitotoxicity; (8) mutant HTT has abnormal protein-protein interactions and affects the functions of individual proteins.
Multiple groups have already tested CRISPR/Cas9 genome editing in HD models (Table 1). Mutant HTT allele-specific deletion has been achieved by designing gRNAs targeting single nucleotide polymorphisms (SNPs) that only exist in the promoter region of the mutant allele [63, 64]. Because HD patients carry different SNPs in their genome, an array of sgRNAs needs to be designed, but they still cannot cover 100% of HD cases. Alternatively, sgRNAs spanning the CAG repeat were designed to remove the HTT protein in a non-allele-specific manner (Fig. 4). This strategy is backed by the findings that permanent removal of wild-type Htt in the mouse brain and temporary reduction of wild-type HTT in the non-human primate brain are well tolerated [65,66,67]. Indeed, non-allele-specific reduction of HTT by CRISPR/Cas9 AAV injection ameliorates neurotoxicity and behavioral deficits in an HD knock-in mouse model [68]. These studies provide proof-of-concept that CRISPR/Cas9 could be a potential therapy for HD. In addition, several groups have further optimized this process. For example, a smaller version of Cas9 (SaCas9) has been used in the R6/2 mouse model of HD and prolonged its lifespan [69]. Virus-mediated expression of the bacterial Cas9 protein alters the transcription of genes involved in neuronal functions. Tagging Cas9 with a fragment of the cell-cycle protein Geminin reduces Cas9 protein stability in neurons, and significantly alleviates the neurotoxicity of Cas9 [70]. A self-destructive version of Cas9 (KamiCas9) has also been designed to limit the duration of expression of Cas9 protein, which reduces off-target effects [71]. CRISPR/Cas9 has also been used to reduce mutant HTT expression in HD patient iPSCs and differentiated neural stem cells, and this approach ameliorated mitochondrial and redox modifications [72].
Schematic showing sgRNAs targeting the HTT gene used in previous studies. Different sgRNA designs have been adopted in previous studies. To allele-specifically silence mutant HTT, sgRNAs covering HD patient-specific SNPs are located either in the 5’UTR [64] or in intron 3 [63] (red). In the non-allele-specific approach, dual sgRNAs are designed to target sequences spanning the CAG repeats in exon 1 HTT [68, 69, 72] (green), or a single sgRNA upstream of exon 1 HTT is used [71] (purple).
Nonetheless, major concerns need to be addressed before CRISPR can be used as a therapeutic approach for HD. It remains technically challenging to effectively deliver CRISPR to the HD patient’s brain, so that better delivery vehicles that can cross the blood-brain barrier and diffuse to a broad brain region are highly desirable. The off-target effects of CRISPR, which means that CRISPR induces DNA-cutting in unwanted chromosomal locations, need to be carefully examined using more stringent sequencing methods. Moreover, the long-term cellular reactions to HTT removal and the abnormal responses caused by the exogenous Cas9 protein, such as immune activation [73, 74], need to be tested, preferably in non-human primates. It is noteworthy that recently two highly-anticipated HTT-lowering candidates based on ASO failed in late-stage clinical trials. What is more confounding is that one of the candidates even worsened patient outcome measures compared with the placebo [61]. Such failure highlights the importance of correct dosage and timing for using ASOs to treat HD. Considering the effect of CRISPR is permanent, there would be less concern about timing, as CRISPR could be delivered to pre-symptomatic patients. Nonetheless, dosage remains a major issue. Given that HTT-lowering caused by CRISPR is irreversible, extra precautions need to be exercised in advancing CRISPR-based therapeutic methods for HD.
Innovations of CRISPR Technology
In recent years, CRISPR technology has evolved so fast that novel innovations are published almost daily. Many of these innovations have yet been tested in HD research but hold great promise (Table 2). For instance, Cpf1 or Cas12a was identified from Francisella novicida U112, and belongs to the Class 2, type V CRISPR system [75]. Compared with the commonly-used Cas9 (SpCas9), Cpf1 possesses some unique features. First, it is smaller, making it easier to package into viral vectors. Second, Cpf1 recognizes the TTN or TTTN PAM sequence, which is different from the NGG PAM sequence of Cas9, thereby offering more flexibility in targeting different genetic loci. Third, Cpf1 cleaves DNA and introduces a staggered DSB with 4- to 5-nucleotide overhangs, allowing for the incorporation of designed sequences, whereas Cas9 generates a blunt-ended DSB. Last, Cpf1 allows for multiplexed genome editing, as a single crispr RNA (crRNA) array can target multiple loci in the genome [76]. These features make Cpf1 an ideal alternative to Cas9 to satisfy specific research needs.
SpCas9 and SaCas9 are large proteins consisting of 1368 and 1053 amino-acids, respectively. A constant endeavor is to find smaller Cas9 orthologues that offer more flexibility in viral packaging. For example, CjCas9, derived from Campylobacter jejuni, is composed of 984 amino-acid residues [77]; Cas12f (also known as Cas14), identified from uncultivated archaea, is a family of nucleases that are composed of 400–700 amino-acids [78,79,80]. These variants have been tested in human cells or mouse models, with editing efficiency and specificity similar to SpCas9 [81,82,83,84], thereby paving the way to test their uses in the central nervous system.
CRISPR/Cas9 does not have to cleave DNA. Instead, nuclease-null Cas9 (dCas9) has been engineered to fuse with various elements to further expand its applications. In 2016, RNA-targeting Cas9 (RCas9) was created by fusing dCas9 with GFP, together with a modified sgRNA scaffold that preferentially targets RNA but not the encoding DNA. RCas9 does not affect mRNA abundance or protein translation, but enables the tracking of endogenous mRNAs of interest in living cells [85]. In 2017, the same group further modified RCas9 by fusing dCas9 with the PIN RNA endonuclease domain from SMG6 (PIN-dCas9). PIN-dCas9 was directed to target mRNAs containing repeat expansions, including CUG, CCUG, CAG, and GGGGCC; it successfully cleaved and eliminated all the repeat expansion mRNAs [86]. Because CUG, CCUG, CAG, and GGGGCC repeat expansions are responsible for myotonic dystrophy type 1, myotonic dystrophy type 2, polyglutamine diseases, and frontotemporal dementia/amyotrophic lateral sclerosis (c9FTD/ALS) respectively, PIN-dCas9 has the potential to treat many genetic diseases caused by microsatellite expansions in different genomic loci. Nonetheless, it remains to be tested whether PIN-dCas9 can effectively work in vivo, and how long the effects last, considering PIN-dCas9 targets RNA, not DNA.
Gene expression can also be controlled by modulating transcriptional activity. To this end, a myriad of transcription regulators have been fused with dCas9 to achieve transcriptional activation or repression [87]. For example, the VP64 transcriptional activation domain has been used to activate the transcription of endogenous human genes, whereas fusion of the KRAB transcriptional repression domain from human KOX1 to dCas9 has been shown to silence transcription [88, 89]. Epigenetic editors such as DNA methyltransferase 3 alpha has been fused to dCas9 to achieve site-specific DNA methylation and transcriptional repression at several human promoters [90]. Likewise, fusion of ten-eleven translocation methylcytosine dioxygenase 1 leads to DNA demethylation and targeted upregulation of gene transcription [91]. In addition, many histone modifiers, including histone acetyltransferase p300 and histone deacetylase HDAC3, have been used to change the histone landscape at selected genomic loci, and resulted in gene activation or repression, respectively [92, 93]. These efforts created a full CRISPR arsenal to alter gene expression, which can be readily tested in HD to lower mutant HTT transcription. Indeed, a zinc finger-KRAB fusion protein (ZFP-TF) has already been created to specifically repress mutant HTT transcription. Treatment with ZFP-TF successfully ameliorates the neuropathology and behavioral deficits in multiple lines of HD mice [94]. It is possible that a similar approach using CRISPR technology could achieve comparable or even better outcomes.
The CRISPR system has also been designed to precisely edit the genome through the invention of base editors, which are chimeric dCas9 proteins fused with DNA deaminase enzymes. To date, two types of base editor have been widely used: one is cytosine base editors (CBEs), which use the rat APOBEC1 cytidine deaminase domain to catalyze the conversion of C·G base-pairs to T·A base-pairs; the other one is adenine base editors, which use the TadA adenine deaminase to convert A·T base-pairs to G·C base-pairs [95, 96]. The base editors rely on base excision and DNA mismatch repair to achieve genome editing. More importantly, these repair mechanisms are active in post-mitotic cells such as neurons [97], thereby making these editors suitable for applications in the brain. By substituting single nucleotides, the base editors can create stop codons to terminate protein translation [98]. In 2020, this approach was tested in a mouse model of ALS. Intrathecal injection of AAVs encoding CBE significantly reduced the expression of mutant SOD1, a causative gene for ALS, leading to protection of motor neurons and reduction of muscle atrophy [99].
The above innovations could potentially push CRISPR-based therapies further into clinical applications. First, the large size of SpCas9 has always been a headache for viral packaging, and is known to elicit a myriad of cellular responses in mammalian cells. These more compact variants of nucleases could easily fit into the small AAV genome and potentially cause fewer host responses after delivery. Second, the permanent inactivation of gene expression by genome editing is a major safety concern. In contrast, transcriptional repressors or epigenetic modulators can silence gene expression without cutting chromosomes. Third, another drawback of CRISPR/Cas9 is the random mutations introduced by DNA DSB and NHEJ. Base editors that precisely change the genetic sequence enable genome editing in a controllable fashion.
Concluding Remarks
Technological advancement has always been essential to biological and biomedical research. Former technological breakthroughs, such as molecular biology techniques and large-scale sequencing, have profoundly changed the ways of conducting scientific research. CRISPR-based genome editing has the potential to play a similar role. The rapid development of CRISPR technology has particularly benefited HD research. Being a rare, monogenic neurodegenerative disease, HD can serve as a vanguard in testing these innovative technologies, and such experiences can bring valuable insights to understand or even treat other diseases as well. We are hopeful that CRISPR, together with other technological advances, will bring a cure for the devastating HD in the near future.
References
Lieberman AP, Shakkottai VG, Albin RL. Polyglutamine repeats in neurodegenerative diseases. Annu Rev Pathol 2019, 14: 1–27.
Zuccato C, Valenza M, Cattaneo E. Molecular mechanisms and potential therapeutical targets in Huntington’s disease. Physiol Rev 2010, 90: 905–981.
Lu SY, Lu BX. Degeneration versus development: Hunting-out the D-unit of Huntington’s disease. Neurosci Bull 2021, 37: 757–760.
Cheng HR, Li XY, Yu HL, Xu M, Zhang YB, Gan SR. Correlation between CCG polymorphisms and CAG repeats during germline transmission in Chinese patients with Huntington’s disease. Neurosci Bull 2020, 36: 811–814.
Orr HT, Zoghbi HY. Trinucleotide repeat disorders. Annu Rev Neurosci 2007, 30: 575–621.
Bates GP, Dorsey R, Gusella JF, Hayden MR, Kay C, Leavitt BR, et al. Huntington disease. Nat Rev Dis Primers 2015, 1: 15005.
A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington's disease chromosomes. The Huntington's Disease Collaborative Research Group. Cell 1993, 72: 971–983. doi: https://doi.org/10.1016/0092-8674(93)90585-e.
Carroll D. Genome engineering with targetable nucleases. Annu Rev Biochem 2014, 83: 409–439.
Hockemeyer D, Soldner F, Beard C, Gao Q, Mitalipova M, DeKelver RC, et al. Efficient targeting of expressed and silent genes in human ESCs and iPSCs using zinc-finger nucleases. Nat Biotechnol 2009, 27: 851–857.
Gaj T, Gersbach CA, Barbas CF III. ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol 2013, 31: 397–405.
Vachey G, Déglon N. CRISPR/Cas9-mediated genome editing for Huntington’s disease. Methods Mol Biol 2018, 1780: 463–481.
Ishino Y, Shinagawa H, Makino K, Amemura M, Nakata A. Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. J Bacteriol 1987, 169: 5429–5433.
Mojica FJ, Ferrer C, Juez G, Rodríguez-Valera F. Long stretches of short tandem repeats are present in the largest replicons of the Archaea Haloferax mediterranei and Haloferax volcanii and could be involved in replicon partitioning. Mol Microbiol 1995, 17: 85–93.
Jansen R, Embden JD, Gaastra W, Schouls LM. Identification of genes that are associated with DNA repeats in prokaryotes. Mol Microbiol 2002, 43: 1565–1575.
Horvath P, Barrangou R. CRISPR/Cas, the immune system of bacteria and Archaea. Science 2010, 327: 167–170.
Bolotin A, Quinquis B, Sorokin A, Ehrlich SD. Clustered regularly interspaced short palindrome repeats (CRISPRs) have spacers of extrachromosomal origin. Microbiology (Reading) 2005, 151: 2551–2561.
Pourcel C, Salvignol G, Vergnaud G. CRISPR elements in Yersinia pestis acquire new repeats by preferential uptake of bacteriophage DNA, and provide additional tools for evolutionary studies. Microbiology (Reading) 2005, 151: 653–663.
Mojica FJM, Díez-Villaseñor C, García-Martínez J, Soria E. Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J Mol Evol 2005, 60: 174–182.
Makarova KS, Grishin NV, Shabalina SA, Wolf YI, Koonin EV. A putative RNA-interference-based immune system in prokaryotes: Computational analysis of the predicted enzymatic machinery, functional analogies with eukaryotic RNAi, and hypothetical mechanisms of action. Biol Direct 2006, 1: 7.
Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 2007, 315: 1709–1712.
Andersson AF, Banfield JF. Virus population dynamics and acquired virus resistance in natural microbial communities. Science 2008, 320: 1047–1050.
Brouns SJJ, Jore MM, Lundgren M, Westra ER, Slijkhuis RJH, Snijders APL, et al. Small CRISPR RNAs guide antiviral defense in prokaryotes. Science 2008, 321: 960–964.
Garneau JE, Dupuis MÈ, Villion M, Romero DA, Barrangou R, Boyaval P, et al. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 2010, 468: 67–71.
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 2012, 337: 816–821.
Makarova KS, Wolf YI, Koonin EV. Classification and nomenclature of CRISPR-cas systems: Where from here? CRISPR J 2018, 1: 325–336.
Adli M. The CRISPR tool kit for genome editing and beyond. Nat Commun 1911, 2018: 9.
Nishiyama J, Mikuni T, Yasuda R. Virus-mediated genome editing via homology-directed repair in mitotic and postmitotic cells in mammalian brain. Neuron 2017, 96: 755-768.e5.
Mao ZY, Bozzella M, Seluanov A, Gorbunova V. DNA repair by nonhomologous end joining and homologous recombination during cell cycle in human cells. Cell Cycle 2008, 7: 2902–2906.
Bekris LM, Yu CE, Bird TD, Tsuang DW. Genetics of Alzheimer disease. J Geriatr Psychiatry Neurol 2010, 23: 213–227.
Klein C, Westenberger A. Genetics of Parkinson’s disease. Cold Spring Harb Perspect Med 2012, 2: a008888.
Mittelman D, Moye C, Morton J, Sykoudis K, Lin YF, Carroll D, et al. Zinc-finger directed double-strand breaks within CAG repeat tracts promote repeat instability in human cells. Proc Natl Acad Sci U S A 2009, 106: 9607–9612.
Garriga-Canut M, Agustín-Pavón C, Herrmann F, Sánchez A, Dierssen M, Fillat C, et al. Synthetic zinc finger repressors reduce mutant huntingtin expression in the brain of R6/2 mice. Proc Natl Acad Sci U S A 2012, 109: E3136–E3145.
Fink KD, Deng P, Gutierrez J, Anderson JS, Torrest A, Komarla A, et al. Allele-specific reduction of the mutant huntingtin allele using transcription activator-like effectors in human Huntington’s disease fibroblasts. Cell Transplant 2016, 25: 677–686.
Cong L, Ran FA, Cox D, Lin SL, Barretto R, Habib N, et al. Multiplex genome engineering using CRISPR/Cas systems. Science 2013, 339: 819–823.
An MC, O'Brien RN, Zhang NZ, Patra BN, de la Cruz M, Ray A, et al. Polyglutamine disease modeling: Epitope based screen for homologous recombination using CRISPR/Cas9 system. PLoS Curr 2014, https://doi.org/10.1371/currents.hd.0242d2e7ad72225efa72f6964589369a
Malankhanova T, Suldina L, Grigor’eva E, Medvedev S, Minina J, Morozova K, et al. A human induced pluripotent stem cell-derived isogenic model of Huntington’s disease based on neuronal cells has several relevant phenotypic abnormalities. J Pers Med 2020, 10: 215.
Xu XH, Tay Y, Sim B, Yoon SI, Huang YH, Ooi J, et al. Reversal of phenotypic abnormalities by CRISPR/Cas9-mediated gene correction in Huntington disease patient-derived induced pluripotent stem cells. Stem Cell Reports 2017, 8: 619–633.
Menalled LB, Kudwa AE, Miller S, Fitzpatrick J, Watson-Johnson J, Keating N, et al. Comprehensive behavioral and molecular characterization of a new knock-in mouse model of Huntington’s disease: zQ175. PLoS One 2012, 7: e49838.
Menalled LB, Sison JD, Dragatsis I, Zeitlin S, Chesselet MF. Time course of early motor and neuropathological anomalies in a knock-in mouse model of Huntington’s disease with 140 CAG repeats. J Comp Neurol 2003, 465: 11–26.
Lin CH, Tallaksen-Greene S, Chien WM, Cearley JA, Jackson WS, Crouse AB, et al. Neurological abnormalities in a knock-in mouse model of Huntington’s disease. Hum Mol Genet 2001, 10: 137–144.
Heng MY, Duong DK, Albin RL, Tallaksen-Greene SJ, Hunter JM, Lesort MJ, et al. Early autophagic response in a novel knock-in model of Huntington disease. Hum Mol Genet 2010, 19: 3702–3720.
Lee CYD, Cantle JP, Yang XW. Genetic manipulations of mutant huntingtin in mice: New insights into Huntington’s disease pathogenesis. FEBS J 2013, 280: 4382–4394.
Ross CA, Tabrizi SJ. Huntington’s disease: From molecular pathogenesis to clinical treatment. Lancet Neurol 2011, 10: 83–98.
Yang HM, Yang S, Jing L, Huang LX, Chen LX, Zhao XX, et al. Truncation of mutant huntingtin in knock-in mice demonstrates exon1 huntingtin is a key pathogenic form. Nat Commun 2020, 11: 2582.
Bañez-Coronel M, Ayhan F, Tarabochia AD, Zu T, Perez BA, Tusi SK, et al. RAN translation in Huntington disease. Neuron 2015, 88: 667–677.
Yang S, Yang HM, Huang LX, Chen LX, Qin ZH, Li SH, et al. Lack of RAN-mediated toxicity in Huntington’s disease knock-in mice. Proc Natl Acad Sci U S A 2020, 117: 4411–4417.
Crook ZR, Housman D. Huntington’s disease: Can mice lead the way to treatment? Neuron 2011, 69: 423–435.
Levine MS, Cepeda C, Hickey MA, Fleming SM, Chesselet MF. Genetic mouse models of Huntington’s and Parkinson’s diseases: Illuminating but imperfect. Trends Neurosci 2004, 27: 691–697.
Yan S, Tu ZC, Liu ZM, Fan NN, Yang HM, Yang S, et al. A huntingtin knockin pig model recapitulates features of selective neurodegeneration in Huntington’s disease. Cell 2018, 173: 989-1002.e13.
Culver BP, Savas JN, Park SK, Choi JH, Zheng SQ, Zeitlin SO, et al. Proteomic analysis of wild-type and mutant huntingtin-associated proteins in mouse brains identifies unique interactions and involvement in protein synthesis. J Biol Chem 2012, 287: 21599–21614.
Goehler H, Lalowski M, Stelzl U, Waelter S, Stroedicke M, Worm U, et al. A protein interaction network links GIT1, an enhancer of huntingtin aggregation, to Huntington’s disease. Mol Cell 2004, 15: 853–865.
Shirasaki DI, Greiner ER, Al-Ramahi I, Gray M, Boontheung P, Geschwind DH, et al. Network organization of the huntingtin proteomic interactome in mammalian brain. Neuron 2012, 75: 41–57.
Wertz MH, Mitchem MR, Pineda SS, Hachigian LJ, Lee H, Lau V, et al. Genome-wide in vivo CNS screening identifies genes that modify CNS neuronal survival and mHTT toxicity. Neuron 2020, 106: 76-89.e8.
Genetic Modifiers of Huntington’s Disease (GeM-HD) Consortium. Identification of genetic factors that modify clinical onset of Huntington's disease. Cell 2015, 162: 516–526.
Lee JM, Chao MJ, Harold D, Abu Elneel K, Gillis T, Holmans P, et al. A modifier of Huntington’s disease onset at the MLH1 locus. Hum Mol Genet 2017, 26: 3859–3867.
Loupe JM, Pinto RM, Kim KH, Gillis T, Mysore JS, Andrew MA, et al. Promotion of somatic CAG repeat expansion by Fan1 knock-out in Huntington’s disease knock-in mice is blocked by Mlh1 knock-out. Hum Mol Genet 2020, 29: 3044–3053.
Li XJ, Li SH, Sharp AH, Nucifora FC, Schilling G, Lanahan A, et al. A huntingtin-associated protein enriched in brain with implications for pathology. Nature 1995, 378: 398–402.
Liu Q, Cheng SY, Yang HM, Zhu LY, Pan YC, Jing L, et al. Loss of Hap1 selectively promotes striatal degeneration in Huntington disease mice. Proc Natl Acad Sci U S A 2020, 117: 20265–20273.
Zhao T, Hong Y, Yin P, Li SH, Li XJ. Differential HspBP1 expression accounts for the greater vulnerability of neurons than astrocytes to misfolded proteins. Proc Natl Acad Sci U S A 2017, 114: E7803–E7811.
Drouet V, Ruiz M, Zala D, Feyeux M, Auregan G, Cambon K, et al. Allele-specific silencing of mutant huntingtin in rodent brain and human stem cells. PLoS One 2014, 9: e99341.
Kingwell K. Double setback for ASO trials in Huntington disease. Nat Rev Drug Discov 2021, 20: 412–413.
Li ZY, Wang C, Wang ZY, Zhu CG, Li J, Sha T, et al. Allele-selective lowering of mutant HTT protein by HTT-LC3 linker compounds. Nature 2019, 575: 203–209.
Shin JW, Kim KH, Chao MJ, Atwal RS, Gillis T, MacDonald ME, et al. Permanent inactivation of Huntington’s disease mutation by personalized allele-specific CRISPR/Cas9. Hum Mol Genet 2016, 25: 4566–4576.
Monteys AM, Ebanks SA, Keiser MS, Davidson BL. CRISPR/Cas9 editing of the mutant huntingtin allele in vitro and in vivo. Mol Ther 2017, 25: 12–23.
Grondin R, Kaytor MD, Ai Y, Nelson PT, Thakker DR, Heisel J, et al. Six-month partial suppression of Huntingtin is well tolerated in the adult rhesus striatum. Brain 2012, 135: 1197–1209.
McBride JL, Pitzer MR, Boudreau RL, Dufour B, Hobbs T, Ojeda SR, et al. Preclinical safety of RNAi-mediated HTT suppression in the rhesus macaque as a potential therapy for Huntington’s disease. Mol Ther 2011, 19: 2152–2162.
Wang GH, Liu XD, Gaertig MA, Li SH, Li XJ. Ablation of huntingtin in adult neurons is nondeleterious but its depletion in young mice causes acute pancreatitis. Proc Natl Acad Sci U S A 2016, 113: 3359–3364.
Yang S, Chang RB, Yang HM, Zhao T, Hong Y, Kong HE, et al. CRISPR/Cas9-mediated gene editing ameliorates neurotoxicity in mouse model of Huntington’s disease. J Clin Invest 2017, 127: 2719–2724.
Ekman FK, Ojala DS, Adil MM, Lopez PA, Schaffer DV, Gaj T. CRISPR-Cas9-mediated genome editing increases lifespan and improves motor deficits in a Huntington’s disease mouse model. Mol Ther Nucleic Acids 2019, 17: 829–839.
Yang S, Li SH, Li XJ. Shortening the half-life of Cas9 maintains its gene editing ability and reduces neuronal toxicity. Cell Rep 2018, 25: 2653-2659.e3.
Merienne N, Vachey G, de Longprez L, Meunier C, Zimmer V, Perriard G, et al. The self-inactivating KamiCas9 system for the editing of CNS disease genes. Cell Rep 2017, 20: 2980–2991.
Lopes C, Tang Y, Anjo SI, Manadas B, Onofre I, de Almeida LP, et al. Mitochondrial and redox modifications in Huntington disease induced pluripotent stem cells rescued by CRISPR/Cas9 CAGs targeting. Front Cell Dev Biol 2020, 8: 576592.
Chew WL, Tabebordbar M, Cheng JKW, Mali P, Wu EY, Ng AHM, et al. A multifunctional AAV-CRISPR-Cas9 and its host response. Nat Methods 2016, 13: 868–874.
Wang D, Mou HW, Li SY, Li YX, Hough S, Tran K, et al. Adenovirus-mediated somatic genome editing of pten by CRISPR/Cas9 in mouse liver in spite of Cas9-specific immune responses. Hum Gene Ther 2015, 26: 432–442.
Zetsche B, Gootenberg JS, Abudayyeh OO, Slaymaker IM, Makarova KS, Essletzbichler P, et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 2015, 163: 759–771.
Zetsche B, Heidenreich M, Mohanraju P, Fedorova I, Kneppers J, DeGennaro EM, et al. Multiplex gene editing by CRISPR-Cpf1 using a single crRNA array. Nat Biotechnol 2017, 35: 31–34.
Kim E, Koo T, Park SW, Kim D, Kim K, Cho HY, et al. In vivo genome editing with a small Cas9 orthologue derived from Campylobacter jejuni. Nat Commun 2017, 8: 14500.
Harrington LB, Burstein D, Chen JS, Paez-Espino D, Ma EB, Witte IP, et al. Programmed DNA destruction by miniature CRISPR-Cas14 enzymes. Science 2018, 362: 839–842.
Xiao RJ, Li Z, Wang SK, Han RJ, Chang LF. Structural basis for substrate recognition and cleavage by the dimerization-dependent CRISPR-Cas12f nuclease. Nucleic Acids Res 2021, 49: 4120–4128.
Takeda SN, Nakagawa R, Okazaki S, Hirano H, Kobayashi K, Kusakizako T, et al. Structure of the miniature type V-F CRISPR-Cas effector enzyme. Mol Cell 2021, 81: 558-570.e3.
Koo T, Lu-Nguyen NB, Malerba A, Kim E, Kim D, Cappellari O, et al. Functional rescue of dystrophin deficiency in mice caused by frameshift mutations using Campylobacter jejuni Cas9. Mol Ther 2018, 26: 1529–1538.
Lee JY, Jang YJ, Bae JH, Lee YH, Bae HS, Kim S, et al. Efficient and specific generation of knockout mice using Campylobacter jejuni CRISPR/Cas9 system. Biochem Biophys Rep 2020, 22: 100752.
Bigelyte G, Young JK, Karvelis T, Budre K, Zedaveinyte R, Djukanovic V, et al. Miniature type V-F CRISPR-Cas nucleases enable targeted DNA modification in cells. Nat Commun 2021, 12: 6191.
Kim DY, Lee JM, Moon SB, Chin HJ, Park S, Lim Y, et al. Efficient CRISPR editing with a hypercompact Cas12f1 and engineered guide RNAs delivered by adeno-associated virus. Nat Biotechnol 2022, 40: 94–102.
Nelles DA, Fang MY, O’Connell MR, Xu JL, Markmiller SJ, Doudna JA, et al. Programmable RNA tracking in live cells with CRISPR/Cas9. Cell 2016, 165: 488–496.
Batra R, Nelles DA, Pirie E, Blue SM, Marina RJ, Wang H, et al. Elimination of toxic microsatellite repeat expansion RNA by RNA-targeting Cas9. Cell 2017, 170: 899-912.e10.
Goell JH, Hilton IB. CRISPR/cas-based epigenome editing: Advances, applications, and clinical utility. Trends Biotechnol 2021, 39: 678–691.
Maeder ML, Linder SJ, Cascio VM, Fu YF, Ho QH, Joung JK. CRISPR RNA-guided activation of endogenous human genes. Nat Methods 2013, 10: 977–979.
Gilbert LA, Larson MH, Morsut L, Liu ZR, Brar GA, Torres SE, et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 2013, 154: 442–451.
Amabile A, Migliara A, Capasso P, Biffi M, Cittaro D, Naldini L, et al. Inheritable silencing of endogenous genes by hit-and-Run targeted epigenetic editing. Cell 2016, 167: 219-232.e14.
Liu XS, Wu H, Ji X, Stelzer Y, Wu XB, Czauderna S, et al. Editing DNA methylation in the mammalian genome. Cell 2016, 167: 233-247.e17.
Hilton IB, D’Ippolito AM, Vockley CM, Thakore PI, Crawford GE, Reddy TE, et al. Epigenome editing by a CRISPR-Cas9-based acetyltransferase activates genes from promoters and enhancers. Nat Biotechnol 2015, 33: 510–517.
Kwon DY, Zhao YT, Lamonica JM, Zhou ZL. Locus-specific histone deacetylation using a synthetic CRISPR-Cas9-based HDAC. Nat Commun 2017, 8: 15315.
Zeitler B, Froelich S, Marlen K, Shivak DA, Yu Q, Li D, et al. Allele-selective transcriptional repression of mutant HTT for the treatment of Huntington’s disease. Nat Med 2019, 25: 1131–1142.
Gaudelli NM, Komor AC, Rees HA, Packer MS, Badran AH, Bryson DI, et al. Programmable base editing of A·T to G·C in genomic DNA without DNA cleavage. Nature 2017, 551: 464–471.
Komor AC, Kim YB, Packer MS, Zuris JA, Liu DR. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 2016, 533: 420–424.
Iyama T, Wilson DM III. DNA repair mechanisms in dividing and non-dividing cells. DNA Repair 2013, 12: 620–636.
Kuscu C, Parlak M, Tufan TR, Yang JK, Szlachta K, Wei XL, et al. CRISPR-STOP: Gene silencing through base-editing-induced nonsense mutations. Nat Methods 2017, 14: 710–712.
Lim CKW, Gapinske M, Brooks AK, Woods WS, Powell JE, et al. Treatment of a mouse model of ALS by in vivo base editing. Mol Ther 2020, 28: 1177–1189.
Kolli N, Lu M, Maiti P, Rossignol J, Dunbar GL. CRISPR-Cas9 mediated gene-silencing of the mutant huntingtin gene in an in vitro model of Huntington’s disease. Int J Mol Sci 2017, 18: 754.
Funding
This review was supported by the National Key R&D Program of China (2021YFA0805200), the National Natural Science Foundation of China (31970954, 81901289 and 31872779) and the Guangdong Key Laboratory of Non-human Primate Research (2020B121201006).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Qin, Y., Li, S., Li, XJ. et al. CRISPR-Based Genome-Editing Tools for Huntington’s Disease Research and Therapy. Neurosci. Bull. 38, 1397–1408 (2022). https://doi.org/10.1007/s12264-022-00880-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12264-022-00880-3