Abstract
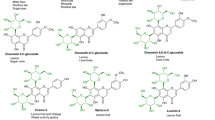
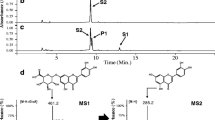
Flavonoids glycosylated with UDP-glucuronic acid and UDP-xylose are spatially distributed in nature. To produce these glycosides, E. coli was engineered to overexpress biosynthetic gene clusters of UDP-sugars (galU from E. coli K12, UDP-glucose dehydrogenase (calS8), and UDP-glucuronic acid decarboxylase (calS9) from Micromonospora echinospora spp. calichensis). Flavonoids were glycosylated by overexpression of the glycosyltransferase gene (atGt-5) from Arabidopsis thaliana. Finally, metabolically engineered host E. coli (US89Gt-5) was generated. Production of flavonoid glycosides was observed in a biotransformation system consisting of flavonoids (naringenin and quercetin) exogenously fed to host cells. The glycosylated derivatives 7-O-glucuronyl naringenin (m/z+ 449), 7-O-xylosyl naringenin (m/z+ 405), and 7-O-glucuronyl quercetin (m/z+ 479) were detected and confirmed by ESI-MS/MS, ESI-MS/MS and LC/MS-MS analysis, respectively.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Mao, Z., H. D. Shin, and R. R. Chen (2006) Engineering the E. coli UDP-glucose synthesis pathway for oligosaccharide synthesis. Biotechnol. Prog. 22: 369–374.
Liang, H. -R., P. Vuorela, and H. Vuorela (1997) Isolation and immunomodulatory effect of flavonol glycosides from Epimedium hunanense. Planta Med. 63: 316–319.
Thorson, J. S., T. J. Hosted Jr, J. Jiang, J. B. Biggins, and J. Ahlert (2001) Nature’s carbohydrate chemists: The enzymatic glycosylation of bioactive bacterial metabolities. Curr. Org. Chem. 5: 139–167.
Li, D., J. H. Park, J. -T. Park, C. S. Park, and K. H. Park (2004) Biotechnological production of highly soluble daidzein glycosides using Thermotoga maritima maltosyltransferase. J. Agric. Food Chem. 52: 2561–2567.
Ruffing, A. and R. R. Chen (2006) Metabolic engineering of microbes for oligosaccharide and polysaccharide synthesis. Microb. Cell Fact. 5: 25.
Stobiecki, M. (2000) Application of mass spectrometry for identification and structural studies of flavonoid glycosides. Photochem. 54: 237–256.
Mendez, C. and J. A. Salas (2001) Altering the glycosylation pattern of bioactive compounds. Trends Biotechnol. 19: 449–456.
Walsh, C. T. (2002) Combinatorial biosynthesis of antibiotics: Challenges and opportunities. ChemBioChem. 3: 124–134.
Winkel-Shirley, B. (2001) Flavonoid biosynthesis: A colorful model for genetics, biochemistry, cell biology and biotechnology. Plant Physiol. 126: 485–493.
Mackenzie, P. I., I. S. Owens, B. Burchell, K. W. Bock, A. Bairoch, A. Belanger, S. Fournel-Gigleux, M. Green, D. W. Hum, T. Iyanagi, D. Lancet, P. Louisot, J. Magdalou, J. R. Chowdhury, J. K. Ritter, H. Schachter, T. R. Tephly, K. F. Tipton, and D. W. Nebert (1997) The UDP glycosyltransferase gene superfamily: Recommended nomenclature update based on evolutionary divergence. Pharmacogenetics 7: 255–269.
Kapitonov, D. and R. K. Yu (1999) Conserved domains of glycosyltransferases. Glycobiol. 9: 961–978.
Paquette, S., B. L. Moller, and S. Bak (2003) On the origin of family 1 plant glycosyltransferase. Phytochem. 62: 399–413.
Simkhada, D., T. J. Oh, B. B. Pageni, H. C. Lee, K. Liou, and J. K. Sohng (2009) Characterization of CalS9 in the biosynthesis of UDP-xylose and production of xylosyl-attached hybrid compound. Appl. Microbiol. Biotechnol. 83: 885–895.
Kurumbang, N. P., K. Liou, and J. K. Sohng (2009) Biosynthesis of paromamine derivatives in engineered Escherichia coli by heterologous expression. J. Appl. Microbiol. 108: 1780–1788.
Cane, D. E., C. T. Walsh, and C. Khosla (1998) Harnessing the biosynthetic code: Combinations, permutations, and mutations. Science 282: 63–68.
White-Phillip, J., C. J. Thibodeaux, and H. W. Liu (2009) Enzymatic synthesis of TDP-deoxysugars. Methods Enzymol. 459: 521–544.
Lamb, S. and G. Wright (2005) Accessorizing natural products: Adding to nature’s toolbox. Proc. Natl. Acad. Sci. USA. 102: 519–520.
Thorson, J. S., W. A. Barton, D. Hoffmeister, C. Albermann, and D. B. Nikolov (2004) Strucutre-based enzyme engineering and its impact on in vitro glycorandomization. ChemBioChem. 5: 16–25.
Boue, M. S., H. C. Carter-Wientjes, Y. B. Shih, and E. T. Cleveland (2003) Identification of flavone aglycones and glycosides in soybean pods by liquid chromatography-tandem mass spectrometry. J. Chromatogr. A. 991: 61–68.
Chen, L. J., D. E. Games, J. Jones, and H. Kidwell (2003) Separation and identification of flavonoids in an extract from the seeds of Oroxylum indicum by CCC. J. Liq. Chromatogr. Relat. Technol. 26: 1623–1636.
Ma, Y. L., Q. M. Li, H. Van den Heuvel, and M. Claeys (1997) Characterization of flavone and flavonol aglycones by collisioninduced dissociation tandem mass spectrometry. Rapid Commun. Mass Spectrum. 11: 1357–1364.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Simkhada, D., Kurumbang, N.P., Lee, H.C. et al. Exploration of glycosylated flavonoids from metabolically engineered E. coli . Biotechnol Bioproc E 15, 754–760 (2010). https://doi.org/10.1007/s12257-010-0012-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12257-010-0012-4