Abstract
This study aimed to investigate the expression of secreted phosphoprotein 1 (SPP1) on lung cancer cells and explore its underlying mechanism on autophagy and apoptosis which effect the development of lung cancer cells. GSE19804 related to lung cancer cells was screened from Gene Expression Omnibus (GEO) database, and we screened the 47 pairs of differential expressed mRNAs in lung cancer cells and adjacent tissues using microarray analysis. The expression of the core gene SPP1 was detected by qRT-PCR and western-blot. The transfection efficiency of lung cancer cells was detected by qRT-PCR and the expression of transfected group was tested by western-blot. Cell proliferation after transfection was tested by MTT assay and plate cloning experiment. The apoptosis rate of each transfection group was detected by flow cytometry. We use western-blot to test protein expression of autophagy-related proteins Beclin-1, LC3-I, LC3-II and p62 of each transfected group. Through analysis of GSE19804,the heat map showed SPP1 was the highest expressed in tumor tissues. qRT-PCR and western-blot detected SPP1 expression in lung cancer tissues was higher than that in normal adjacent tissues and was significantly increased in lung cancer cell lines. After transfection with pcDNA3.1-SPP1 (p-SPP1 group), siRNA1-SPP1 (siRNA1 group) and siRNA2-SPP1 (siRNA2 group), showed different expression of SPP1. Up-regulation of SPP1 enhanced cell viability and promoted tumor cell proliferation, while knockdown of SPP1 inhibited tumor cell proliferation. From the results of apoptosis rate, SPP1 inhibited the tumor cell apoptosis. However, in normal lung cell, SPP1 had no effect on cell proliferation and apoptosis. And to test autophagy-related proteins, we found that overexpression of SPP1 inhibited autophagy. High expression of SPP1 inhibited autophagy and apoptosis to promote the development of small cell lung cancer cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Among all the cancers, lung cancer remains the highest death rate, accounting for about one-quarter (27%) of all cancer deaths [1]. To date, lung cancer still represents the leading cause of cancer-related deaths worldwide and non-small cell lung cancer (NSCLC) comprises about 85% of all types of lung cancer [2], with only a small number of patients achieving long-term survival. The specific mechanisms underlying lung cancer development and progression are still unclear. Although cigarette smoking and alcohol consumption are known to directly induce lung cancer, various other environmental risks, such as exposure to second-hand smoke, air pollution, and genetic susceptibility, are also involved in the development of lung cancer [3]. Indeed, multiple genetic and epigenetic changes have been shown to be associated with lung cancer development [4].
Autophagy has been known as a cellular catabolic pathway responsible for the modulation of numerous pathophysiological processes. Autophagy induction in response to stressful stimuli is involved in adaptation of the cells to various stressful conditions, such as shortage of nutrients and oxygen [5], or it can also cause dysregulated cell death [6]. Cancer cells in the tumor tissue are typical example exposed to the stressful conditions and growing evidences have suggested the critical role of autophagy in the regulation and/or progression of cancer. The role of autophagy in cancer progression is controversial. For example, cancer cells activate autophagy in response to stressful insults, allowing cells to mitigate damage and promote senescence that limit tumorigenesis, indicating a regulatory role of autophagy in cancer development [7]. However, many other recent studies have indicated autophagy as a cytoprotective adaptive response. Activation of oncogenes and inactivation of tumor suppressor genes in cancer cells enhance autophagy, thereby promoting survival of tumor cells. Furthermore, inhibition of autophagy by pharmacological (e.g. chloroquine or 3-methyladenine) or genetic interference (e.g., knocking down of essential proteins) accelerates cell death in conditions exposed to the cellular stress [8]. In addition, autophagy and apoptosis mutually negatively regulate each other. Autophagic process therefore contributes to the development of cancer [6, 9]. All these reports indicate that autophagy modulates cancer progression in a context-dependent manner.
In this study, the core gene SPP1 was screened for verifying its effect on the mechanism of lung cancer cells development. It was speculated that SPP1 might be used as a target of lung cancer cells after verifying the role of SPP1 in the occurrence and development of lung cancer cells for non-smoking female patients with early lung cancer cells and observing the correlation between SPP1 and autophagy and apoptosis. The proposition of this speculation based on small cell lung cancer cells was of great significance for patients.
Materials and Methods
Gene Expression Omnibus (GEO) Microarray Data Processing
With “lung cancer cell” as keywords, 17,482 gene expression data were obtained from GEO database (http://www.nebi.nlm.nih.gov/geo), and “expression profiling by array” was selected. “GSE19804” microarray data set was screened, and the platform was GPL570: [HG-U133_Plus_2] Affymetrix Human Genome U133 Plus 2.0 Array. The included samples were 47 cases of stage I to II cancer tissues and paired adjacent tissues respectively. Differentially expressed miRNAs between normal and tumor were screened by SAMR (significance analysis of microarrays) package in R software, under the condition of log2FC (fold change) was over 2 and FDR (false discovery rate) less than 0.05.
Clinical Sample Collection
107 paired human lung cancer tissues and adjacent tissues were all obtained from patients with informed consent during Sep. 2016 to Dec. 2017. The protocols used in the study had been approved by the Ethic Committee of Qilu Hospital of Shandong University.
Cell Culture and Treatment
Lung cancer cell lines A549, H1975, NCI-H446, SK-MES-1 and human bronchial epithelial cells (BEAS-2B cells) were purchased from the American Type Culture Collection (ATCC, Philadelphia, PA, USA). Lung cancer cells were grown in RPMI 1640 (Invitrogen, Carlsbad, CA, USA) supplemented with 10% (v/v) fetal bovine serum (FBS; Invitrogen, Carlsbad, CA, USA) and 1% penicillin-streptomycin (Invitrogen, Carlsbad, CA, USA). BEAS-2B cells were grown in DMEM (Invitrogen, Carlsbad, CA, USA) supplemented with 10% FBS, 100 U/mL penicillin and 100 U/mL streptomycin (Invitrogen, Carlsbad, CA, USA). All cells were cultured in a humidified CO2 incubator at 37 °C.
Cell Transfection
pcDNA3.1 plasmid vectors, pcDNA3.1-SPP1, siRNA negative control, siRNA1-SPP1 and siRNA2-SPP1 were bought from GenePharma (Shanghai, China). The siRNA sequence of SPP1 listed in Table 1. NCI-H446 cells were seeded at 105 cells each well into 24-well plates in logarithmic growth phase and grown to 80% confluence. Cells were transfected with corresponding plasmids using the Lipofectamine 2000 (Life Technologies, Gaithersburg, MD, USA) and Opti-MEM serum-free medium (Invitrogen, Carlsbad, CA, USA) on the basis of the manufacturer’s recommendations. qRT-PCR was used for determining the transfection efficiency 48 h post transfection.
qRT-PCR
Initially, RNA was completely extracted using Trizol reagent (Beyotime, Shanghai, China) on the basis of the manufacturer’s guidelines. PrimeScript™ RT reagent kit (Takara, Tokyo, Japan) was used to oppositely transfer RNA into cDNA. qPCR reaction was accomplished using SYBRH Premix Ex Taq™ GC (Takara) and an Applied Biosystems 7500 real-time PCR system (Applied Biosystems, Foster City, CA, USA. β-actin mRNA was used for SPP1’s internal control and the process was done thrice. Samples were standardized to internal controls. And fold changes were obtained through the 2-ΔΔCT method.
Western Blot
Briefly, phosphate-buffered saline (PBS) were taken to wash cells and then cell were fostered with lysis buffer (Sangon Biotech, Co., Ltd., Shanghai, China). Protein was parted by SDS-PAGE and shifted to a PVDF membrane. Non-specific binding sites were blocked with TBST which contains 5% (w/v) non-fat dried milk. Next, the membrane was fostered with rabbit anti-SPP1 (Abcam, 1:200), rabbit anti-Beclin-1 (Abcam, 1:100), rabbit anti-LC3-I (Abcam, 1:200), rabbit anti-LC3-II (Abcam, 1:200), rabbit anti-p62 (Abcam, 1:200), rabbit anti-β-actin (Abcam, 1:200) at 4 °C all night long. Next, membranes were incubated overnight with a HRP-conjugated goat anti-rabbit IgG (1:10000, Abcam) for 1 h at 37 °C. Signals were observed with ECL chemiluminescence detection kit and semi-quantitated using ImageJ software. Equal protein loading was analyzed by β-actin.
MTT Assay
NCI-H446 cells were plated in 96-well plates. After transfection for 48 h, the culture medium was discarded and 20 μl (5 mg/ml) 3-(4, 5-dimethylthiazol-2-yl)-2, 5-diphenyltetrazolium bromide (MTT) solution was added to each well. After culturing for 4 h at 37 °C, 100 μl dimethylsulfoxide (DMSO) was added to terminate the reaction. The absorbance at 490 nm was measured using a microplate reader.
Annexin V-FITC / PI Double Staining Flow Cytometry
The experiment was carried out according to the Apoptosis Detection Kit (Invitrogen, # BMS500FI-100). 48 h after transfection, cells were digested with trypsin without EDTA, and the cells were centrifuged at 1000 rpm and centrifuged for 5 min. Cells were washed with cold PBS twice, then centrifuged them at 1000 rpm for 5 min. 5 μl of Annexin V-FITC dye was added into 200 μl of Binding Buffer. Then they were mixed well, and reacted in room temperature and dark for 15 min. 5 μl of PI dye was added for reaction. Flow cytometry was performed within 1 h and Cell Quest software was used for analysis.
Statistical Analysis
Data were represented with the mean ± SD and analyzed by GraphPad Prism 6.0 software (GraphPad Software, La Jolla, CA, USA). Two groups were compared by Student’s t test. Multiple groups were compared by one-way ANOVA analysis. All experiments were independently performed at least three times. P value less than 0.05 was considered statistically significant.
Results
Analysis of GEO19804
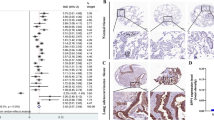
We selected 47 cases of stage I to II cancer tissues and paired adjacent tissues as objects of the study from 120 samples obtained from GSE19804 of NCBI. Differentially expressed mRNAs were screened by SAMR (significance analysis of microarrays) package in R software, under the condition of log2FC (fold change) was over 2 and FDR (false discovery rate) less than 0.05. In the above conditions, the top 10 mRNA levels were selected and obviously distributed in heat map (Fig. 1). And we found that SPP1 was the highest expressed mRNA in top 10 mRNA. Based on previous studies, SPP1 had showed something to do with tumor growth. And the results we got in heat map hinted that SPP1 were in close contact with tumor growth. So, we chose SPP1 as our research gene in the following experiments.
Analysis of GSE19804. There were a total of 120 samples in the GSE19804, the 47 cases of stage I to II cancer tissues and paired adjacent tissues were selected as the research object. Differentially expressed mRNA was screened with the Fold change value greater than 2 and P < 0.05 as screening conditions. The cluster heat map showed the top 10 mRNA levels and found that SPP1 was the highest expressed in cancer tissues
SPP1 Was Observably High Expressed in Lung Cancer Cell Lines and Tissues
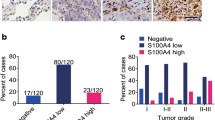
Total RNA was extracted from lung cancer and adjacent tissues of 107 patients, of whom the clinical characteristics were listed in Table 2. Expression of SPP1 was evidently increased in 107 lung cancer tissues, compared with adjacent tissues detected by qRT-PCR (P < 0.01, Fig. 2a). Besides, protein expression of SPP1 was remarkably high in 107 lung cancer tissues compared with adjacent tissues detected by qRT-PCR (P < 0.01, Fig. 2b) And taking many factors into consideration, we randomly chose 10 samples to do western-blot (Fig. 2c), using β-actin as internal control (Fig. 2c). The correlation between SPP1 expression and clinic-pathological features in lung cancer was listed in Table 3. There was no significant association between SPP1 level and age, gender or tumor size, while patients with high expression of SPP1 are more potential to get an advanced clinical stage (P < 0.05). After transfection with SPP1, we could see different degree expression of SPP1 in different cell lines. The expression levels of SPP1 was remarkably high in lung carcinoma cell lines (A549, H1975, NCI-H446 and SK-MES-1 cells) compared with human normal lung epithelial cells (BEAS-2B cell) (Fig. 2d) and found that SPP1 expression was the highest in NCI-H446 cells in four cancer cells. So, we chose NCI-H446 cells as our experimental subjects in the following experiments to explore the way SPP1 affect the oncogenesis.
SPP1was highly expressed in lung cancer tissues. aSPP1 was in a higher expression level in lung cancer tissues by qRT-PCR. **P < 0.01, compared with adjacent tissues. b、c Western blot results revealed that SPP1 was increased in lung cancer tissues. **P < 0.01, compared with adjacent tissues. d Expression levels of SPP1 in lung carcinoma cell lines (A549, H1975, NCI-H446 and SK-MES-1 cells) and human normal lung epithelial cells (BEAS-2B cell). *P < 0.05, **P < 0.01 compared with BEAS-2B group
SPP1 Promotes Small Cell Lung Cancer Cell Proliferation
NCI-H446 lung cancer cells were transfected with negative control (NC group), pcDNA3.1-SPP1 (p-SPP1 group), siRNA1-SPP1 (siRNA1 group) and siRNA2-SPP1 (siRNA2 group). Through establishing different grade of SPP1 expression group, we measured SPP1’s impact on lung cancer cell proliferation. The relative expression of SPP1 was measured by qRT-PCR as well as western blot. The expression of SPP1 mRNA and protein was obviously down-regulated by siRNA1-SPP1 and siRNA2-SPP1 in lung cancer cells, whereas the expression of SPP1 was significantly facilitated in pcDNA3.1-SPP1 group (P < 0.01, Fig. 3a, b). SPP1 in pcDNA3.1-SPP1 group was overexpressed, and was inhibited in siRNA1-SPP1 and siRNA2-SPP1 group. MTT assay was performed in order to investigate the effect of SPP1 on lung cancer cell growth. The result showed that the capacity of cell proliferation with SPP1 overexpression was obviously higher than the control group in lung cancer cell line (P < 0.01), which suggesting that SPP1 overexpression could promote cell growth to a certain extent (Fig. 3c). And the low expression of SPP1 in siRNA1-SPP1 and siRNA2-SPP1 group showed that the capacity of cell proliferation was inhibited, compared with NC group. Plate clone assay revealed that the number of cell proliferation was high in p-SPP1 group, while the number of cell proliferation was decreased in siRNA1 group and siRNA2 group (Fig. 3d, e). Both experiment consistently suggested that SPP1 overexpression would promoted the cell proliferation in lung cancer cells.
SPP1Promotes Lung Cancer Cell Proliferation. a Negative control (NC group), pcDNA3.1-SPP1 (p-SPP1 group), siRNA1-SPP1 (siRNA1 group) and siRNA2-SPP1 (siRNA2 group) were transfected into NCI-H446 lung cancer cells respectively. SPP1 was down-regulated in cell lines after transfected with siRNA1-SPP1 and siRNA2-SPP1 and was up-regulated in cell lines after transfected with pcDNA3.1-SPP1 by qRT-PCR. **P < 0.01, compared with NC group. b Western blot showed protein expression of SPP1 was facilitated in p-SPP1 group and was decreased in siRNA1 group and siRNA2 group.**P < 0.01, compared with NC group. c MTT assay detected the cell viability of p-SPP1 group was enhanced while that was inhibited in siRNA1group and siRNA2 group, *P < 0.05, compared with NC group. d、e Plate clone assay revealed that the number of cell proliferation was high in p-SPP1 group, while the number of cell proliferation was decreased in siRNA1 group and siRNA2 group, **P < 0.01 compared with NC group
SPP1 Had no Effect on Bronchial Epithelial Cell Proliferation
BEAS-2B cells were transfected with negative control (NC group), pcDNA3.1-SPP1 (p-SPP1 group), siRNA1-SPP1 (siRNA1 group) and siRNA2-SPP1 (siRNA2 group) to measure SPP1’s impact on normal cell proliferation. The relative expression of SPP1 was measured by qRT-PCR and western blot. The expression of SPP1 mRNA and protein was obviously down-regulated by siRNA1-SPP1 and siRNA2-SPP1 in human bronchial epithelial cells, whereas the expression of SPP1 was significantly facilitated in pcDNA3.1-SPP1 group (P < 0.01, Fig. 4a, b). SPP1 in pcDNA3.1-SPP1 group was overexpressed and was inhibited in siRNA1-SPP1 and siRNA2-SPP1 group. MTT assay was carried out to explore the effect of SPP1 on bronchial epithelial cell growth. The result showed that the proliferation capacity of cell with SPP1 overexpression had no statistic difference compared with the control group (P < 0.01, Fig. 4c). And the low expression of SPP1 in siRNA1-SPP1 and siRNA2-SPP1 group showed the similar result compared with NC group. Plate clone assay and flow cytometry revealed that the number of cell proliferation and apoptosis rate among p-SPP1 group, siRNA1 group and siRNA2 group had no statistic difference (Fig. 4d, e).
SPP1had no effect on bronchial epithelial cell proliferation. a Negative control (NC group), pcDNA3.1-SPP1 (p-SPP1 group), siRNA1-SPP1 (siRNA1 group) and siRNA2-SPP1 (siRNA2 group) were transfected into BEAS-2B cell respectively. SPP1 was down-regulated in cell lines after transfected with siRNA1-SPP1 and siRNA2-SPP1 and was up-regulated in cells after transfected with pcDNA3.1-SPP1 by qRT-PCR. **P < 0.01, compared with NC group. b Western blot showed protein expression of SPP1 was facilitated in p-SPP1 group and was decreased in siRNA1 group and siRNA2 group.**P < 0.01, compared with NC group. c MTT assay detected the cell viability in p-SPP1 group, siRNA1group and siRNA2 group, compared with NC group. d Plate clone assay revealed the number of cell proliferation in p-SPP1 group, siRNA1 group and siRNA2 group. e Flow Cytometer revealed the apoptosis rate of cell in p-SPP1 group, siRNA1 group and siRNA2 group
SPP1 Promotes Apoptosis and Autophagy
Another way involve to the cancer cell growth was the death of cancer cells. As is known to us all, apoptosis and autophagy are the two maintain ways regulate cell death. The apoptosis rate of p-SPP1 group was down-regulated, while the apoptosis rate of siRNA1 group and siRNA2 group was up-regulated detected by flow cytometry (Fig. 5a). Down-regulate of apoptosis rate when SPP1 was overexpressed could prove that SPP1 inhibited cancer cell apoptosis, which means decreased the death of cancer cells. When it comes to another way, western blot was used to detect the expression of autophagy-related proteins Beclin-1, LC3-I, LC3-II and p62 in each transfection group to explain the autophagy level. Upregulation of SPP1 decreased the protein expression of Beclin-1 and LC3-II/LC3-I, while promoted p62 protein expression. Downregulation of SPP1 increased Beclin-1 and LC3-II/LC3-I protein expression, but reduced p62 protein expression (Fig. 5b).
SPP1promoted autophagy and apoptosis in lung cancer cells. a The apoptosis rate of p-SPP1 group was down-regulated, while the apoptosis rate of siRNA1 group and siRNA2 group was up-regulated detected by flow cytometry, **P < 0.01 compared with NC group. b Western blot was used to detect the expression of autophagy-related proteins Beclin-1, LC3-I, LC3-II and p62 in each transfection group. **P < 0.01 compared with NC group
Discussion
In the study, through analyzing the microarray GSE19804, we found the top ten mRNA levels in tumor tissue. SPP1 was the highest expressed gene of the mRNAs in tumor tissues. For better illustration, we used qRT-PCR to detect the expression levels of SPP1 and its proteins in 107 lung cancer tissues and adjacent tissues. And 10 cases were randomly selected for western-blot to detect protein expression levels. For a more detailed explanation, we chose four lung cancer cell lines and one normal lung cell line to examine the expression of SPP1 in lung cancer cell lines. SPP1 was highly expressed in lung cancer cells detected by qRT-PCR compared with normal lung cells. All of these experiments illustrate the high expression of SPP1 in lung cancer cells. Therefore, may be SPP1 overexpression could promote the development of lung cancer cells. The NCI-H446 cell line with the highest SPP1 expression in four lung cancer cell lines was selected and transfected with negative control (NC), pcDNA3.1-SPP1 (p-SPP1), siRNA1-SPP1 SPP1 (siRNA2 group). The expression of SPP1 was detected by qRT-PCR and western-blot. In the p-SPP1 group, SPP1 expression was promoted, and in the siRNA1 group and the siRNA2 group, SPP1 expression was suppressed. MTT assay found that SPP1 overexpression could enhance the cell viability of NCI-H446 cells. And the plate clone assay showed that SPP1 overexpression could increase the number of cell proliferation. The experiments verified that SPP1 overexpression could promote the development of lung cancer cells.
Lung cancer is one of the most common types of cancer worldwide and is associated with a poor 5-year relative survival rate [10]. The initial treatment of patients hinges on the detection of targetable molecular drivers, now challenges were posed on finding more effective diagnosis and prognosis markers [11]. Accordingly, the role of SPP1 as a tumor promoter was preferred in the present study. SPP1 was highly expressed in lung cancer tissues and cells and promoted cell growth, inhibited cell apoptosis and autophagy at the same time.
Autophagy plays a major role in maintaining cell quality control and homeostasis but under extreme conditions autophagic signaling can be recruited to activate cell death pathways. Thus, autophagy can act either as a tumor suppressor or as a tumor maintenance factor [12, 13]. In the last years, autophagy has gained relevance as a mechanism that, acting in concert with the intrinsic pathway of apoptosis, controls tissue homeostasis and cell survival [14, 15]. Autophagy can be triggered by different causes of environmental stress such us nutritional deficiencies, hypoxia or exposure to high temperatures [16]. This process involves the degradation of cellular components through lysosomal machinery and is thought to be required for normal turnover of cellular components, especially in response to starvation [14, 15]. The interaction between apoptosis and autophagy is complicated and depends on the cellular context. Autophagy can act both as an alternative death pathway to apoptosis and as stress adaptation mechanism to avoid cell death [17]. In the study, western blot was used to detect the relationship of autophagy-related proteins Beclin-1, LC3-I, LC3-II and p62 and SPP1. The first identified mammalian autophagy-related gene, Beclin1, regulates the activity of autophagy particularly in the initiation phase by combining with different ligands [18]. LC3 has been identified to be the most widely used specific marker of autophagy initiation, which is first synthesized as its precursor, then cut up into its cytosolic form, LC3-I, which can be processed into LC3-II [19]. Autophagosome is essential for the elongation stage of the phagophore membrane and LC3-II specifically binds to it. To test the amount of autophagosome can be measured by the expression of LC3-II or LC3-II/LC3-I [20]. So we investigate expression of LC3-II/LC3-I. In addition, P62 is negatively correlated with autophagy activity, reflecting the degradative capability of autophagy and the intensity of autophagic flux [21]. Observable changes of autophagy-related proteins showed that upregulation of SPP1 decreased the protein expression of Beclin-1 and LC3-II/LC3-I, while promoted p62 protein expression. These markers of autophagy told us SPP1 inhibited the autophagy. The more complex transduction pathways associated with autophagy under the influence of SPP1 and whether autophagy-related proteins can regulate lung cancer cells or not need further study.
Nevertheless, limitations in this report are to be taken into consideration. For example, we identified SPP1 could promote the development of lung cancer cells by inhibiting autophagy and apoptosis, but upstream mechanism of SPP1 was not explored. In addition, the relationship between SPP1 and survival time of patient with lung cancer needs investigation by future experiments.
Conclusion
In conclusion, we detected SPP1 expression was highly expressed in lung cancer tissues as well as cell lines. High expression of SPP1 was capable to significantly promote the development of lung cancer cells via inhibiting the autophagy and apoptosis.
References
Nanavaty P, Alvarez MS, Alberts WM (2014) Lung cancer screening: advantages, controversies, and applications. Cancer Control 21:9–14. https://doi.org/10.1177/107327481402100102
Hasan N, Kumar R, Kavuru MS (2014) Lung cancer screening beyond low-dose computed tomography: the role of novel biomarkers. Lung 192:639–648. https://doi.org/10.1007/s00408-014-9636-z
Malhotra J, Malvezzi M, Negri E, La Vecchia C, Boffetta P (2016) Risk factors for lung cancer worldwide. Eur Respir J 48:889–902. https://doi.org/10.1183/13993003.00359-2016
Zhu J, Zeng Y, Xu C, Qin H, Lei Z, Shen D, Liu Z, Huang JA (2015) Expression profile analysis of microRNAs and downregulated miR-486-5p and miR-30a-5p in non-small cell lung cancer. Oncol Rep 34:1779–1786. https://doi.org/10.3892/or.2015.4141
He C, Klionsky DJ (2009) Regulation mechanisms and signaling pathways of autophagy. Annu Rev Genet 43:67–93. https://doi.org/10.1146/annurev-genet-102808-114910
Nikoletopoulou V, Markaki M, Palikaras K, Tavernarakis N (2013) Crosstalk between apoptosis, necrosis and autophagy. Biochim Biophys Acta 1833:3448–3459. https://doi.org/10.1016/j.bbamcr.2013.06.001
Yang ZJ, Chee CE, Huang S, Sinicrope F (2011) Autophagy modulation for cancer therapy. Cancer Biol Ther 11:169–176. https://doi.org/10.4161/cbt.11.2.14663
Mathew R, Karantza-Wadsworth V, White E (2007) Role of autophagy in cancer. Nat Rev Cancer 7:961–967. https://doi.org/10.1038/nrc2254
Gordy C, He YW (2012) The crosstalk between autophagy and apoptosis: where does this lead? Protein Cell 3:17–27. https://doi.org/10.1007/s13238-011-1127-x
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A (2015) Global cancer statistics, 2012. CA Cancer J Clin 65:87–108. https://doi.org/10.3322/caac.21262
Heymann JJ, Bulman WA, Swinarski D, Pagan CA, Crapanzano JP, Haghighi M, Fazlollahi L, Stoopler MB, Sonett JR, Sacher AG, Shu CA, Rizvi NA, Saqi A (2017) PD-L1 expression in non-small cell lung carcinoma: comparison among cytology, small biopsy, and surgical resection specimens. Cancer 125:896–907. https://doi.org/10.1002/cncy.21937
Gozuacik D, Kimchi A (2004) Autophagy as a cell death and tumor suppressor mechanism. Oncogene 23:2891–2906. https://doi.org/10.1038/sj.onc.1207521
Ogier-Denis E, Codogno P (2003) Autophagy: a barrier or an adaptive response to cancer. Biochim Biophys Acta 1603:113–128
Marino G, Niso-Santano M, Baehrecke EH, Kroemer G (2014) Self-consumption: the interplay of autophagy and apoptosis. Nat Rev Mol Cell Biol 15:81–94. https://doi.org/10.1038/nrm3735
Marino G, Lopez-Otin C (2004) Autophagy: molecular mechanisms, physiological functions and relevance in human pathology. Cell Mol Life Sci 61:1439–1454. https://doi.org/10.1007/s00018-004-4012-4
Zhang M, Jiang M, Bi Y, Zhu H, Zhou Z, Sha J (2012) Autophagy and apoptosis act as partners to induce germ cell death after heat stress in mice. PLoS One 7:e41412. https://doi.org/10.1371/journal.pone.0041412
Maiuri MC, Le Toumelin G, Criollo A, Rain JC, Gautier F, Juin P, Tasdemir E, Pierron G, Troulinaki K, Tavernarakis N, Hickman JA, Geneste O, Kroemer G (2007) Functional and physical interaction between Bcl-X(L) and a BH3-like domain in Beclin-1. EMBO J 26:2527–2539. https://doi.org/10.1038/sj.emboj.7601689
Wirawan E, Lippens S, Vanden Berghe T, Romagnoli A, Fimia GM, Piacentini M, Vandenabeele P (2012) Beclin1: a role in membrane dynamics and beyond. Autophagy 8:6–17. https://doi.org/10.4161/auto.8.1.16645
Tanida I, Ueno T, Kominami E (2004) LC3 conjugation system in mammalian autophagy. Int J Biochem Cell Biol 36:2503–2518. https://doi.org/10.1016/j.biocel.2004.05.009
Kabeya Y, Mizushima N, Ueno T, Yamamoto A, Kirisako T, Noda T, Kominami E, Ohsumi Y, Yoshimori T (2000) LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. EMBO J 19:5720–5728. https://doi.org/10.1093/emboj/19.21.5720
Ichimura Y, Komatsu M (2010) Selective degradation of p62 by autophagy. Semin Immunopathol 32:431–436. https://doi.org/10.1007/s00281-010-0220-1
Acknowledgments
This study was supported by Shandong Province Science and Technology Public Relations (No.2012GGE27122) and Science and Technology Program Project of Jinan (No. 201704093).
Funding
This study was funded by Shandong Province Science and Technology Public Relations (No.2012GGE27122) and Science and Technology Program Project of Jinan (No. 201704093).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Ethical Approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of Qilu Hospital of Shandong University.
Informed Consent
Informed consent was obtained from all individual participants included in the study.
Rights and permissions
About this article
Cite this article
Liu, H., Wei, S., Zhang, L. et al. Secreted Phosphoprotein 1 Promotes the Development of Small Cell Lung Cancer Cells by Inhibiting Autophagy and Apoptosis. Pathol. Oncol. Res. 25, 1487–1495 (2019). https://doi.org/10.1007/s12253-018-0504-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12253-018-0504-7