Abstract
Involvement of micro RNAs (miRNA) is currently the focus for cancer studies as they effect the post transcriptional expression of different genes. Let-7 family is among the firstly discovered miRNAs that play important role in cell proliferation and dysregulation leading to cell based diseases including cancer. Another family, miRNA-200 prevents transformation of cell to malignant form and tumor formation by interacting with epidermal mesenchymal transition (EMT). Similarly miRNA-125 controls apoptosis and proliferation by affecting multiple genes involved in transcription, immunological defense, resistance against viral and bacterial infections that ultimately leads to cell proliferation, metastasis and finally cancer. All of these micro RNAs are known to be either upregulated or downregulated in various cancers. Current review is focused to elaborate the role of these three families of micro RNAs on different genes that ultimately cause cancer. In conclusion we can say that the miRNAs discussed here are mostly downregulated in various cancers with some exceptions when upregulation of miRNA-125 may be attributed to cancer formation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
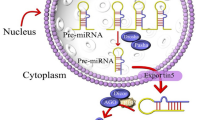
MicroRNAs (miRNA) are small non-coding, single stranded RNAs of about 20–25 nucleotides. They are generated from the primary microRNA transcript (pri-microRNA) which has many miRNAs. These primary miRNA transcripts are produced by RNA polymerase II, as separate transcriptional units or embedded within the introns of protein coding genes. In nucleus primary transcript is trimmed by the microprocessor complex containing the RNase III enzyme Drosha which is than exported to the cytoplasm by exportin 5. In the cytoplasm other protein complexes including DICER and TRBP further trim the pre-miRNA to produce the single-stranded mature miRNA, which subsequently incorporates into the RNA induced silencing complex (RISC). RISC bounded miRNA attaches with complementary RNA resulting in translational repression or mRNA degradation. These non coding RNAs were first studied in C. elegans in 1993 [1] and function in the regulation of several cellular processes, including cell differentiation, proliferation, apoptosis and metabolic homeostasis [2]. Deregulation of these small miRNAs lead to cancer and these may act as oncogene or tumor suppressor genes. Three of the most deregulated miRNAs belong to family of miRNA let-7, miRNA-200 and miRNA-125, which are frequently mutated in different cancers.
miRNA Let-7 Family
Let-7 miRNAs family was first discovered in nematodes and have 13 members located on nine different chromosomes. They are believed to be involved in controlling the timing of stem cell division, differentiation and proliferation. Therefore they play an important role in cell dependent diseases like cancer. It has been found in different researches that they are frequently downregulated in bladder, breast, colorectal, cervical, endometrium, head & neck, lung, ovary, prostate and renal cancers. These micro RNAs may act as therapeutic option for cancer treatment. They act as a regulator of normal cell differentiation and proliferation and inhibit the growth of tumor cells. The levels of let-7 are crucial for development of cell and they act directly on Ras genes via LIN28 [3]. Let-7 and LIN28 both have an inhibitory effect on each other but in embryonic cells let-7 also inhibits IL-6 that results in high levels of NFKB. NFKB along with c-Myc stimulates LIN28 creating higher levels of LIN28 in the cells. This increase in LIN28 leads to a marked decrease in let-7. At the same time Myc gene also inhibits let-7. Reduced levels of let-7 results in increased levels of RAS leading to cell proliferation (Fig. 1). On the other hand, in differentiating mature cells miRNA-125 and miRNA-200 inhibit LIN28 and this decrease in LIN28 leads to increased levels of let-7 causing further inhibition of LIN28. Highest levels of let-7 lead to decreased production of RAS gene and cause cell cycle, angiogenesis and cell adhesion to increase. Therefore under normal conditions miRNA let-7 act as tumor supressor gene and inhibit the activation of oncogenes that may lead to cancer cell formation. Let-7 also repress HMGA2 gene that is only present in embryonic tissue but undetectable in mature tissue [4]. Lower levels of let-7 leads to higher levels of HMGA2, more proliferation of cells which results in cancer cell formation.
Let-7 has been found to be downregulated in lung cancer patients as shown in many studies. It has also been found that post-operative survival time in patients is directly correlated with levels of let-7. Lung cancer patients with lower levels of let-7 survived for less time compared with high levels of let-7 patients. It is known that RAS genes control many signaling pathways and is upregulated in lung cancer. Previous work shows that let-7 directly control cell proliferation by negatively regulating RAS genes [5]. It was shown that down regulation of let 7 family members like let-7a, let-7c and let-7 g in lung cancer play an important role in the suppression of RAS protein [6]. Let-7 represses the expression by targetting the 3’UTR of RAS genes. Few members of the let-7 family are also found to be upregulated in lung cancer. For example, Let-7a-3 is methylated in normal lung tissue but its epigenetic activation leads to lung cancer initiation, ultimately let-7a-3 is upregulated in lung cancer tissues [7].
Let-7 has active role in breast cancer. Let-7a, let-7c, let-7d, let-7e, let-7f, let-7 g, let-7i has been found to be upregulated and let-7b is down regulated in case of breast cancer. Less differentiated cells lack let-7 and were reported to have more advanced stage of cancer [8]. Vice versa highly differentiated cells have higher levels of miRNA let-7 and less advance stage of cancer. It has been shown in a study that administration of let-7 is effective against breast cancer. Let −7 target many genes like ANG, CCND1–2, CDC25A, CDK4, CDK6, CYP19A1, DNA Polymerases, E2F5–6, ESR1, ESR2, FGF11, FGFR, GRB2, HMG2, IGF1, IGFR1, IL6, ITGB3, MAPK4–6, MMP2, MMP8, MYC, RAS, RB1, SKP2, TGFB, TGFBR, TP53, HMGA2 and H-RAS [9]. Breast tumor initiating cells (BTIC) show reduced expression of let-7 and its expressional levels increased with cellular differentiation [10]. In vivo administration of let-7 in BTIC resulted in less cell proliferation and metastasis, whereas silencing of let-7 in non-tumor initiating cells leads to increased ability to self-renewal. This study also proved that let-7 could be novel treatment option for breast cancer due to its effect on self-renewal ability of cancer cells. RAF inhibitory kinase protien (RIKP) is a tumor suppressor gene, that control breast cancer by targeting G protein coupled receptor kinase-2, NF-KB and MAPK signaling cascade [11]. RIKP is repressed by let-7 signaling in metastasis of breast cancer [12]. Reduced expressional levels of MAPK leads to low levels of LIN28 transcription and finally to decreased expression of let-7 targets. Therefore from these researches it became evident that lower levels of let-7 results in tumor suppressing effect of RKIP [13] whereas its upregulation causes reduced expression of HMGA2 which is associated with metastasis of breast cancer cells. Let-7 also exerts its effect through another pathway involving estrogen receptor alpha (ER alpha). Decreased levels of let-7 family members lead to increased ER alpha expression and consequently increased cell proliferation and decreased apoptosis [14]. Further studies on the molecular mechanisms behind let-7 activity in cancer would improve treatment plans for therapeutic applications of breast cancer.
miRNA-200
The human miR-200 family entails two paralogous clusters: miR-200a, miR-200b, and miR-429 located on chromosome 1p36 and miR-200c and miR-141 located on chromosome 12p13. Further division of these clusters is centered on the nucleotides 2–7 seed region, with miR-200b, 200c, and 429 grouped into one and miR-200a and 141 into other cluster. The variation between both clusters is only due to the nucleotide (U to C) at fourth position in the seed region [15]. miR-200bc/429 cluster modulates multidrug resistance of human cancer cell lines [16]. miR-200 family members also form a double negative feedback loop with transcription factors of the ZEB protein family, which controls the epithelial mesencymal transition (EMT) in tumorigenesis.
Urinary miR-200 family levels are repressed in patients with bladder cancer [17]. miR-200c exhibits weak expression in invasive tumors owing to promoter hypermethylation in comparison to non-invasive ones [18]. miR-200c/141 cluster is upregulated in bladder cancer. Consistently, microarray based comparison of miRNA expression pattern by Wiklund et al. (2011) in both invasive and superficial bladder tumors with normal urothelia also showed that miR-200c was upregulated in the bladder tumors as compared to normal urothelium [18]. In a microarray based miRNA study of tumors from ten bladder cancers and corresponding lymph node metastases, miR-200c along with other miR-200 family members were found to be downregulated [19]. Study and comparison of several bladder cancer cell lines displayed interdependence amongst epithelial phenotypes and high expression of miR-200b and miR-200c. miR-200c is majorly involved in epithelial-to-mesenchymal transition (EMT) in bladder cancer cells while loss of its expression has been linked to disease progression and poor outcome in patients carrying T1 stage bladder tumor [18]. Examination of human urothelial cells has also confirmed that methylation of miR-200b genes could serve as bladder cancer biomarker [20].
Decreased expression of miRNAs of the miR-200 family has been implicated in the growth and metastasis of breast cancer cells [21]. miR-200 regulates multifunctional phosphoglucose isomerase, which in turn mediates EMT, while silencing of miR-200 by ZEB1 via KISS1 stalling leads to augmented invasiveness of cancer cells [22]. miR-200c also regulates numerous EMT-related processes, curbs movement and breast cancer cell invasion by targeting actin-regulatory proteins PPM1F and FHOD1. miR-200 has been demonstrated to limit breast cancer metastasis in syngeneic mouse tumors via a moesin-dependent pathway [23]. miR-141 correlates positively with estrogen receptor, while miR-200c correlates negatively with HER2. This implies the dissimilar role of these miR-200 family members in different breast cancer sub types. miR-200c along with some other miRNAs is a potential biomarker for circulating tumor cell status analysis in metastatic breast cancer patients [24]. miR-200 levels in low amount have also been related to decreased survival while high expression levels proved beneficial in radiotherapy. Chen et al. (2013) reported downregulation of miR-200c in patients that do not respond to neoadjuvant chemotherapy as compared to the responsive ones [25].
PCR-based miRNA assay revealed tumor suppressor miR-200a to be considerably linked with overall patient survival [26] and expression profile of HeLa cells transfected with miR-200a proved that it could govern cancer phenotype by amending metastasis routes in cervical carcinoma [27].
miR-200 family is up-regulated in endometrial endometroid carcinomas [5]. Repression of miR-200 family results in EMT induction and high levels of miR-200c are related to poor survival. miR-200c hence, serves as a prognostic marker of overall survival in endometrial cancer [28].
miR-200c and other miR-200 family members modulate EMT and metastasis in the colon [19] with epigenetically regulated expression. High levels of miR-200a and miR-200c are associated with increased survival in colorectal cancer patients. miR-429 levels are directly allied to increased and disease-free survival [29]. Plasma and serum miR-200c levels canalso function as potential biomarkers for colorectal cancer prognosis/screening and predicting metastasis [30].
Expression of miR-200c in the metastatic lymph node area of head and neck squamous cell carcinoma tissues decreases. It also plays an important role in the regulation of stemness and EMT in head and neck tumor cells [31], but promoter hypermethylation of miR-200c targets (Zeb1/Zeb2) mask regulation of EMT and cell migration (Fig. 2) [32].
Tumor suppressive signatures of miR-200. a MiR-200 inhibits EMT by interacting with ZEB1/2 and the Notch pathway. b MiR-200 represses self-renewal and differentiation in CSCs. c MiR-200 is involved in the regulation of cell division and apoptosis [62]
Tellez and colleagues (2011) reported that decreased miR-200 is primary indication of lung cancer as expression of miR-200 modifies the tumor microenvironment to impede the processes of EMT and metastasis in reaction to extracellular signals [33]. miR-200 family is responsible for metastasis prevention and invasion in lung cancers [34]. High expression of tumor miR-200c has also been associated with poor prognosis and disrupted miR-200b expression with reversed chemoresistance in lung cancer patients [35].
miR-200 family members exhibit enhanced expression in ovarian cancer but are negligibly expressed at normal ovarian surface cells with expression critical to transition of benign to endometriosis-associated ovarian cancer [36]. Sequence variation among seed and non-seed regions of miR-200 family members is correlated with variation in the ability to induce mesenchymal-epithelial transition (MET) and drug sensitivity in ovarian cancer cells [37]. miR-200a, miR-200b and miR-200c overexpression can stimulate tumor development and may serve as markers to prophesize the patient survival and as therapeutic targets.
miR-200 controls EMT to MET and vice versa in prostate cancer metastasis is accompanied by elevated expression levels of the miR-200 family in case of MET. Cell invasion and G-protein Subunit-13 expression is regulated by miR-200a in prostate cancer cells. Plasma levels of miR-200c have been recognized as prospective biomarkers of contained prostate cancer distinction from metastatic castration resistant prostate cancer [38]. miR-200-family expression is low in renal cell carcinoma compared to non-cancerous tissues. This family is responsible for tumor suppression in renal cell carcinoma and when matched to its complementary proximal tubular epithelial kidney cells, appear dysregulated in renal cell cancer. miR-141 functions as a crucial manager of renal cell carcinoma metastasis and propagation by expression regulation of EphA2 [39].
miRNA-125 Family
MicroRNA 125 family consist of miR-125a, miR-125b-1 and miR-125b-2, which generate same products of discrete genes. MiRNA 125 family have been known to associate with different carcinomas and other diseases as either repressor or promoter. MiR-125a is located at 19q13, while miR-125b is confirmed to be transcribed from two loci located on chromosomes 11q23 (hsa-miR-125b-1) and 21q21 (hsa-miR-125b-2) [40].
Relationship between microRNA and malignant tumors had been explored widely and associated with diverse properties of different cancers types. MicroRNA-125, more importantly miRNA-125b in several cancer types have oncogenic potential caught up in down regulation of anti-apoptotic proteins. In this review role of miRNA-125 in different cancers was evaluated.
Study by Jiang and co-workers revealed that type 2 endometrial carcinoma were found to have higher tumorogenesis level when transfected with miRNA-125b in mice as compared to negative controls which indicated that upregulation of miRNA-125b promotes proliferation of disease [41]. Another group reported that miRNA-125a was downregulated in endometrioid endometrial carcinoma [42]. Translation of ERBB2 (proto oncogene) was inhibited by overexpression of miRNA-125 in endometeriod endometrial cancer. Over-expression of miR-125b in HEC1B cells inhibited in endometeriod endometrial cancer invasion and this inhibitory effect on HEC1B cells could be restored by miR-125b knock down [43].
In case of colorectal cancer high miR-125b expression group showed a greater incidence of advanced tumor size and tumor invasion compared to the low miR-125b expression group. MicroRNA-125b directly targets 3′ UTRs of Human p53 mRNAs along with its downregulating molecule p21 resulting in the reduced apoptosis and allowing tumor progression by inhibiting the activity of p53 [44]. In several other cancers over expression of miRNA-125 have tumor aggravating potential like in pancreatic cancer, prostate cancer, etc. According to Chen et al. (2013), miRNA-125 family acts as tumor suppressor in colorectal cancer but its target are not yet discovered [25]. Altered expression of miRNA-125b is directly related to colorectal cancer with brain metastasis. It was also confirmed from other studies that microRNAs were involved in tumor invasion and metastasis [45].
Transcription factor like TP53 engages with a variety of cancers including head and neck carcinomas, when activated leads to cell cycle arrest or apoptosis. Mitra et al. (2014) reported that miRNA-125b upregulation in head and neck cancer resulted in repression of TP53 leading to antiapoptosis [46]. Unusual activity of miRNA-125b was studied by Nakanishi et al. (2014) that expression of miRNA-125b-1 was suppressed in head and neck cancers, hyper methylation of its promoter was observed along with the finding that it regulates the extracellular signal–regulated kinase (ERK)–MAPK (mitogen-activated protein kinase) signalling pathway [47]. Decreased expression cause overexpression of gene TACSTD2 (36kD glycoprotein). This protein is overexpressed at late stages of epithelial carcinomas with low-to-no expression in normal tissues [47].
In prostrate cancer miRNA-125b behaves like onco-miRNA as it increases the cell proliferation and halts apoptosis. Relationship between androgen and miRNA was found in prostrate cancer that is an androgen-responsive element regulates miRNA-125b present within the promoter of the miR-125b-2 gene [48]. Upregulation of miRNA-125b resulted in androgen independent growth in prostrate cancer cells and apoptosis was inhibited by targeting apoptotic pathways like BAK1, BBC3, and p53. Amir et al. (2013) deliberated miRNA-125b target P53 dependent and independent apoptosis in prostate cancer [49]. In dependent pathway miRNA-125b interfere with P53 and its target genes P21 and Puma while in P53 independent process apoptosis was induced via the protein product of the ink4a/ARF locus, p14 (ARF).
Unique expression of miRNA-125 had been discovered in lung carcinomas. In some instances it acts as tumor suppressor and in other as oncogene by targeting different genes and pathways. Changed expression of miRNA-125b effect MTA1 which functions in regulating the persistent phenotype of lung cancer. Jiang, 2010 reported that hsa-miR-125a-3p and hsa-miR-125a-5p were downregulated in non-small cell lung cancer (NSCLC) but they have inverse relationship in migration and invasion of lung cancer cells [50]. According to the study miRNA-125a-5p act as tumor suppressor in lung cancer and was activated by the EGFR (growth factor) activation. Moreover negative correlation was found between miRNA-125a-5p and lung cancer metastasis which had inhibitory effect on cell proliferation, angiogenesis, cell motility, and invasion. According to another study miRNA-125a-5p was downregulated in lung cancer by activating p53 pathway that induces apoptosis [51].
MicroRNA-125 also has tumor suppressive activities in different cancers including breast, ovarian and bladder cancer [52]. One of the studies showed that in breast cancers, expression of miRNA-125a was inversely related with the HuR (a stress induced RNA binding protein). In normal cases expression of HuR is high in different cancers. Overexpression of miRNA-125a translationally downregulate HuR resulted in inhibition of cell growth and rates of apoptosis were increased [53]. miRNA both 125a and 125b were downregulated in breast cancer by mediating ERBB2 and ERBB3 pathway or by targeting ETS1 gene, hence showed tumor suppressive function [54].
Bladder cancer was suppressed by miRNA-125b by targeting E2F3 and it initiates apoptosis and inhibit cell proliferation [55]. Ichimi et al. (2009) reported the downregulation of miRNA-125b, which causes the suppression of bladder cancer. Genes downregulated in miRNA-125b were also identified as KRT7, S100A14, WWOX and LEPREI [56]. miRNA-125b was downregulated in bladder urothelial carcinomas so can be used as potential biomarker for treatment of cancer [57].
miRNA-125b is overexpressed in cervical cancer and target phosphoinositide 3-kinase catalytic subunit delta (PI3K/Akt/mTOR) signaling pathway that lead to decreased cell proliferation, apoptosis and suppression of tumor growth. A study was conducted with small cell carcinoma of the cervix (SCCC) which revealed that patients having advanced SCCC had miRNA-125b in suppressed state. Suppression of miRNA-125b had the propensity for poor prognosis as confirmed after Kaplan-Meier survival curves [55].
miRNA-125 exhibited dual role in renal cell cancer (oncogene as well as tumor suppressor). miRNA-125b can be used as biomarker in renal cell carcinomas as it was suppressed by downregulation of miRNA-125b [58]. According to one of the study overexpression of miRNA-125 led to the progression of renal clear cell carcinoma [59]. Another study reported that 4EBP1 pathway was targeted by miRNA-125b and miRNA-125a-5p, which resulted in progressive renal cell carcinoma.
In ovarian cancer miRNA-125b-1 also act as a diagnostic tool and biomarker for cure of ovarian cancer [47]. Downregulation of miRNA-125-b was supposed to be due to either somatic mutations or due to presence of mutations in the enzymes Dicer or Drosha. In 39% of ovarian cancer tissues analysed, decrease of 60 and 51% in the levels of Dicer and Drosha mRNAs respectively were recorded [60]. In human ovarian cancer cells EGFR pathway leading to transcriptional repression of a miRNA through an ETS factor was found to be conserved. miRNA regulation by growth factor signalling represents an additional way of regulating tumor progression. ARIDB3B a novel pathway of regulation of mesenchymal-to-epithelial transition by miR-125a in ovarian cancer was identified in mouse [61]. The repression of miR-125a releases ARID3B promoting a mesenchymal phenotype and contributing to disease progression.
Controversial functions of miR-125 had been studied in different cancers (Fig. 3). In some cases it act as tumor suppressor, reducing tumor proliferation and metastasis by targeting/ downregulating oncogenes and in other scenario it has tumor enhancing property by altering the apoptotic pathways. Same is the case with the widely studied member of family miR-125, miR-125b which is also a double edge miRNA. With the extensive study of miR-125 target genes of this family has been identified, which give insight into the exact role of this microRNA in cellular processes that are related with the cancer diagnosis and treatment. Aberrant expression of miRNA-125 in different cancers allows utilizing these microRNAs as biomarker for early diagnosis of cancers and other diseases.
MiRNA-125b associated with tumor suppressive signalling [57]
In conclusion this review showed that let-7 family of micro RNAs are downregulated and are primarily involving LIN28 and starting a cascade of reactions leading to cancer. Down regulation of miRNA-200 affects the genes that control transition of epithelial to mesenchymal cells, whereas, abnormal expression of miRNA-125 leads to cancer by exerting its negative role on many genes leading to loss of apoptosis and cell proliferation.
Change history
29 April 2017
An erratum to this article has been published.
References
Lee RC, Feinbaum RL, Ambros V (1993) The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75(5):843–854
Calin GA, Croce CM (2006) MicroRNA signatures in human cancers. Nat Rev Cancer 6(11):857–866. doi:10.1038/nrc1997
Iliopoulos D, Hirsch HA, Struhl K (2009) An epigenetic switch involving NF-kappaB, Lin28, let-7 MicroRNA, and IL6 links inflammation to cell transformation. Cell 139(4):693–706. doi:10.1016/j.cell.2009.10.014
Lee YS, Dutta A (2007) The tumor suppressor microRNA let-7 represses the HMGA2 oncogene. Genes Dev 21(9):1025–1030. doi:10.1101/gad.1540407
Lee H, Han S, Kwon CS, Lee D (2016) Biogenesis and regulation of the let-7 miRNAs and their functional implications. Protein & cell 7(2):100–113. doi:10.1007/s13238-015-0212-y
Kumar MS, Erkeland SJ, Pester RE, Chen CY, Ebert MS, Sharp PA, Jacks T (2008) Suppression of non-small cell lung tumor development by the let-7 microRNA family. Proc Natl Acad Sci U S A 105(10):3903–3908. doi:10.1073/pnas.0712321105
Brueckner B, Stresemann C, Kuner R, Mund C, Musch T, Meister M, Sultmann H, Lyko F (2007) The human let-7a-3 locus contains an epigenetically regulated microRNA gene with oncogenic function. Cancer Res 67(4):1419–1423. doi:10.1158/0008-5472.CAN-06-4074
Shell S, Park SM, Radjabi AR, Schickel R, Kistner EO, Jewell DA, Feig C, Lengyel E, Peter ME (2007) Let-7 expression defines two differentiation stages of cancer. Proc Natl Acad Sci U S A 104(27):11400–11405. doi:10.1073/pnas.0704372104
Barh D, Malhotra R, Ravi B, Sindhurani P (2010) MicroRNA let-7: an emerging next-generation cancer therapeutic. Curr Oncol 17(1):70–80
Yu F, Yao H, Zhu P, Zhang X, Pan Q, Gong C, Huang Y, Hu X, Su F, Lieberman J, Song E (2007) Let-7 regulates self renewal and tumorigenicity of breast cancer cells. Cell 131(6):1109–1123. doi:10.1016/j.cell.2007.10.054
Ariazi EA, Brailoiu E, Yerrum S, Shupp HA, Slifker MJ, Cunliffe HE, Black MA, Donato AL, Arterburn JB, Oprea TI, Prossnitz ER, Dun NJ, Jordan VC (2010) The G protein-coupled receptor GPR30 inhibits proliferation of estrogen receptor-positive breast cancer cells. Cancer Res 70(3):1184–1194. doi:10.1158/0008-5472.CAN-09-3068
Piskounova E, Polytarchou C, Thornton JE, LaPierre RJ, Pothoulakis C, Hagan JP, Iliopoulos D, Gregory RI (2011) Lin28A and Lin28B inhibit let-7 microRNA biogenesis by distinct mechanisms. Cell 147(5):1066–1079. doi:10.1016/j.cell.2011.10.039
Yun J, Frankenberger CA, Kuo WL, Boelens MC, Eves EM, Cheng N, Liang H, Li WH, Ishwaran H, Minn AJ, Rosner MR (2011) Signalling pathway for RKIP and let-7 regulates and predicts metastatic breast cancer. EMBO J 30(21):4500–4514. doi:10.1038/emboj.2011.312
Dangi-Garimella S, Yun J, Eves EM, Newman M, Erkeland SJ, Hammond SM, Minn AJ, Rosner MR (2009) Raf kinase inhibitory protein suppresses a metastasis signalling cascade involving LIN28 and let-7. EMBO J 28(4):347–358. doi:10.1038/emboj.2008.294
Uhlmann S, Zhang JD, Schwager A, Mannsperger H, Riazalhosseini Y, Burmester S, Ward A, Korf U, Wiemann S, Sahin O (2010) miR-200bc/429 cluster targets PLCgamma1 and differentially regulates proliferation and EGF-driven invasion than miR-200a/141 in breast cancer. Oncogene 29(30):4297–4306. doi:10.1038/onc.2010.201
Zhu W, Xu H, Zhu D, Zhi H, Wang T, Wang J, Jiang B, Shu Y, Liu P (2012) miR-200bc/429 cluster modulates multidrug resistance of human cancer cell lines by targeting BCL2 and XIAP. Cancer Chemother Pharmacol 69(3):723–731. doi:10.1007/s00280-011-1752-3
Wang G, Chan ES, Kwan BC, Li PK, Yip SK, Szeto CC, Ng CF (2012) Expression of microRNAs in the urine of patients with bladder cancer. Clinical genitourinary cancer 10(2):106–113. doi:10.1016/j.clgc.2012.01.001
Wiklund ED, Bramsen JB, Hulf T, Dyrskjot L, Ramanathan R, Hansen TB, Villadsen SB, Gao S, Ostenfeld MS, Borre M, Peter ME, Orntoft TF, Kjems J, Clark SJ (2011) Coordinated epigenetic repression of the miR-200 family and miR-205 in invasive bladder cancer. Int J Cancer 128(6):1327–1334. doi:10.1002/ijc.25461
Baffa R, Fassan M, Volinia S, O'Hara B, Liu CG, Palazzo JP, Gardiman M, Rugge M, Gomella LG, Croce CM, Rosenberg A (2009) MicroRNA expression profiling of human metastatic cancers identifies cancer gene targets. J Pathol 219(2):214–221. doi:10.1002/path.2586
Kohler CU, Bryk O, Meier S, Lang K, Rozynek P, Bruning T, Kafferlein HU (2013) Analyses in human urothelial cells identify methylation of miR-152, miR-200b and miR-10a genes as candidate bladder cancer biomarkers. Biochem Biophys Res Commun 438(1):48–53. doi:10.1016/j.bbrc.2013.07.021
Radisky DC (2011) miR-200c at the nexus of epithelial-mesenchymal transition, resistance to apoptosis, and the breast cancer stem cell phenotype. Breast cancer research : BCR 13(3):110. doi:10.1186/bcr2885
Teng Y, Mei Y, Hawthorn L, Cowell JK (2014) WASF3 regulates miR-200 inactivation by ZEB1 through suppression of KISS1 leading to increased invasiveness in breast cancer cells. Oncogene 33(2):203–211. doi:10.1038/onc.2012.565
Li X, Roslan S, Johnstone CN, Wright JA, Bracken CP, Anderson M, Bert AG, Selth LA, Anderson RL, Goodall GJ, Gregory PA, Khew-Goodall Y (2014) MiR-200 can repress breast cancer metastasis through ZEB1-independent but moesin-dependent pathways. Oncogene 33(31):4077–4088. doi:10.1038/onc.2013.370
Madhavan D, Zucknick M, Wallwiener M, Cuk K, Modugno C, Scharpff M, Schott S, Heil J, Turchinovich A, Yang R, Benner A, Riethdorf S, Trumpp A, Sohn C, Pantel K, Schneeweiss A, Burwinkel B (2012) Circulating miRNAs as surrogate markers for circulating tumor cells and prognostic markers in metastatic breast cancer. Clinical cancer research : an official journal of the American Association for Cancer Research 18(21):5972–5982. doi:10.1158/1078-0432.CCR-12-1407
Chen Y, Sun Y, Chen L, Xu X, Zhang X, Wang B, Min L, Liu W (2013) miRNA-200c increases the sensitivity of breast cancer cells to doxorubicin through the suppression of E-cadherin-mediated PTEN/Akt signaling. Mol Med Rep 7(5):1579–1584. doi:10.3892/mmr.2013.1403
Hu X, Schwarz JK, Lewis JS Jr, Huettner PC, Rader JS, Deasy JO, Grigsby PW, Wang X (2010) A microRNA expression signature for cervical cancer prognosis. Cancer Res 70(4):1441–1448. doi:10.1158/0008-5472.CAN-09-3289
Pedroza-Torres A, Lopez-Urrutia E, Garcia-Castillo V, Jacobo-Herrera N, Herrera LA, Peralta-Zaragoza O, Lopez-Camarillo C, De Leon DC, Fernandez-Retana J, Cerna-Cortes JF, Perez-Plasencia C (2014) MicroRNAs in cervical cancer: evidences for a miRNA profile deregulated by HPV and its impact on radio-resistance. Molecules 19(5):6263–6281. doi:10.3390/molecules19056263
Torres A, Torres K, Pesci A, Ceccaroni M, Paszkowski T, Cassandrini P, Zamboni G, Maciejewski R (2013) Diagnostic and prognostic significance of miRNA signatures in tissues and plasma of endometrioid endometrial carcinoma patients. Int J Cancer 132(7):1633–1645. doi:10.1002/ijc.27840
Diaz-Martin J, Diaz-Lopez A, Moreno-Bueno G, Castilla MA, Rosa-Rosa JM, Cano A, Palacios J (2014) A core microRNA signature associated with inducers of the epithelial-to-mesenchymal transition. J Pathol 232(3):319–329. doi:10.1002/path.4289
Toiyama Y, Hur K, Tanaka K, Inoue Y, Kusunoki M, Boland CR, Goel A (2014) Serum miR-200c is a novel prognostic and metastasis-predictive biomarker in patients with colorectal cancer. Ann Surg 259(4):735–743. doi:10.1097/SLA.0b013e3182a6909d
Tu HF, Lin SC, Chang KW (2013) MicroRNA aberrances in head and neck cancer: pathogenetic and clinical significance. Current opinion in otolaryngology & head and neck surgery 21(2):104–111. doi:10.1097/MOO.0b013e32835e1d6e
Tamagawa S, Beder LB, Hotomi M, Gunduz M, Yata K, Grenman R, Yamanaka N (2014) Role of miR-200c/miR-141 in the regulation of epithelial-mesenchymal transition and migration in head and neck squamous cell carcinoma. Int J Mol Med 33(4):879–886. doi:10.3892/ijmm.2014.1625
Tellez CS, Juri DE, Do K, Bernauer AM, Thomas CL, Damiani LA, Tessema M, Leng S, Belinsky SA (2011) EMT and stem cell-like properties associated with miR-205 and miR-200 epigenetic silencing are early manifestations during carcinogen-induced transformation of human lung epithelial cells. Cancer Res 71(8):3087–3097. doi:10.1158/0008-5472.CAN-10-3035
Kundu ST, Byers LA, Peng DH, Roybal JD, Diao L, Wang J, Tong P, Creighton CJ, Gibbons DL (2016) The miR-200 family and the miR-183 ~ 96 ~ 182 cluster target Foxf2 to inhibit invasion and metastasis in lung cancers. Oncogene 35(2):173–186. doi:10.1038/onc.2015.71
Feng B, Wang R, Chen LB (2012) Review of miR-200b and cancer chemosensitivity. Biomedicine & pharmacotherapy = Biomedecine & pharmacotherapie 66(6):397–402. doi:10.1016/j.biopha.2012.06.002
Suryawanshi S, Vlad AM, Lin HM, Mantia-Smaldone G, Laskey R, Lee M, Lin Y, Donnellan N, Klein-Patel M, Lee T, Mansuria S, Elishaev E, Budiu R, Edwards RP, Huang X (2013) Plasma microRNAs as novel biomarkers for endometriosis and endometriosis-associated ovarian cancer. Clinical cancer research : an official journal of the American Association for Cancer Research 19(5):1213–1224. doi:10.1158/1078-0432.CCR-12-2726
Jabbari N, Reavis AN, McDonald JF (2014) Sequence variation among members of the miR-200 microRNA family is correlated with variation in the ability to induce hallmarks of mesenchymal-epithelial transition in ovarian cancer cells. Journal of ovarian research 7:12. doi:10.1186/1757-2215-7-12
Watahiki A, Wang Y, Morris J, Dennis K, O'Dwyer HM, Gleave M, Gout PW, Wang Y (2011) MicroRNAs associated with metastatic prostate cancer. PLoS One 6(9):e24950. doi:10.1371/journal.pone.0024950
Chen X, Wang X, Ruan A, Han W, Zhao Y, Lu X, Xiao P, Shi H, Wang R, Chen L, Chen S, Du Q, Yang H, Zhang X (2014) miR-141 is a key regulator of renal cell carcinoma proliferation and metastasis by controlling EphA2 expression. Clinical cancer research : an official journal of the American Association for Cancer Research 20(10):2617–2630. doi:10.1158/1078-0432.CCR-13-3224
Rodriguez A, Griffiths-Jones S, Ashurst JL, Bradley A (2004) Identification of mammalian microRNA host genes and transcription units. Genome Res 14(10A):1902–1910. doi:10.1101/gr.2722704
Jiang L, Huang Q, Chang J, Wang E, Qiu X (2011) MicroRNA HSA-miR-125a-5p induces apoptosis by activating p53 in lung cancer cells. Exp Lung Res 37(7):387–398. doi:10.3109/01902148.2010.492068
Yanokura M, Banno K, Iida M, Irie H, Umene K, Masuda K, Kobayashi Y, Tominaga E, Aoki D (2015) MicroRNAS in endometrial cancer: recent advances and potential clinical applications. EXCLI J 14:190–198. doi:10.17179/excli2014-590
Shang C, Lu YM, Meng LR (2012) MicroRNA-125b down-regulation mediates endometrial cancer invasion by targeting ERBB2. Medical science monitor : international medical journal of experimental and clinical research 18(4):BR149–BR155
Nishida N, Yokobori T, Mimori K, Sudo T, Tanaka F, Shibata K, Ishii H, Doki Y, Kuwano H, Mori M (2011) MicroRNA miR-125b is a prognostic marker in human colorectal cancer. Int J Oncol 38(5):1437–1443. doi:10.3892/ijo.2011.969
Li W, Duan R, Kooy F, Sherman SL, Zhou W, Jin P (2009) Germline mutation of microRNA-125a is associated with breast cancer. J Med Genet 46(5):358–360. doi:10.1136/jmg.2008.063123
Mitra S, Mukherjee N, Das S, Das P, Panda CK, Chakrabarti J (2014) Anomalous altered expressions of downstream gene-targets in TP53-miRNA pathways in head and neck cancer. Scientific reports 4:6280. doi:10.1038/srep06280
Nakanishi H, Taccioli C, Palatini J, Fernandez-Cymering C, Cui R, Kim T, Volinia S, Croce CM (2014) Loss of miR-125b-1 contributes to head and neck cancer development by dysregulating TACSTD2 and MAPK pathway. Oncogene 33(6):702–712. doi:10.1038/onc.2013.13
Sun YM, Lin KY, Chen YQ (2013) Diverse functions of miR-125 family in different cell contexts. J Hematol Oncol 6:6. doi:10.1186/1756-8722-6-6
Amir S, Ma AH, Shi XB, Xue L, Kung HJ, Devere White RW (2013) Oncomir miR-125b suppresses p14(ARF) to modulate p53-dependent and p53-independent apoptosis in prostate cancer. PLoS One 8(4):e61064. doi:10.1371/journal.pone.0061064
Jiang L, Huang Q, Zhang S, Zhang Q, Chang J, Qiu X, Wang E (2010) Hsa-miR-125a-3p and hsa-miR-125a-5p are downregulated in non-small cell lung cancer and have inverse effects on invasion and migration of lung cancer cells. BMC Cancer 10:318. doi:10.1186/1471-2407-10-318
Hsieh TH, Hsu CY, Tsai CF, Long CY, Wu CH, Wu DC, Lee JN, Chang WC, Tsai EM (2015) HDAC inhibitors target HDAC5, upregulate microRNA-125a-5p, and induce apoptosis in breast cancer cells. Molecular therapy : the journal of the American Society of Gene Therapy 23(4):656–666. doi:10.1038/mt.2014.247
Gonzalez-Vallinas M, Breuhahn K (2016) MicroRNAs are key regulators of hepatocellular carcinoma (HCC) cell dissemination-what we learned from microRNA-494. Hepatobiliary surgery and nutrition 5(4):372–376. doi:10.21037/hbsn.2016.05.07
Guo X, Wu Y, Hartley RS (2009) MicroRNA-125a represses cell growth by targeting HuR in breast cancer. RNA Biol 6(5):575–583
Wang S, Huang J, Lyu H, Lee CK, Tan J, Wang J, Liu B (2013) Functional cooperation of miR-125a, miR-125b, and miR-205 in entinostat-induced downregulation of erbB2/erbB3 and apoptosis in breast cancer cells. Cell Death Dis 4:e556. doi:10.1038/cddis.2013.79
Huang L, Luo J, Cai Q, Pan Q, Zeng H, Guo Z, Dong W, Huang J, Lin T (2011) MicroRNA-125b suppresses the development of bladder cancer by targeting E2F3. Int J Cancer 128(8):1758–1769. doi:10.1002/ijc.25509
Ichimi T, Enokida H, Okuno Y, Kunimoto R, Chiyomaru T, Kawamoto K, Kawahara K, Toki K, Kawakami K, Nishiyama K, Tsujimoto G, Nakagawa M, Seki N (2009) Identification of novel microRNA targets based on microRNA signatures in bladder cancer. Int J Cancer 125(2):345–352. doi:10.1002/ijc.24390
Banzhaf-Strathmann J, Edbauer D (2014) Good guy or bad guy: the opposing roles of microRNA 125b in cancer. Cell communication and signaling : CCS 12:30. doi:10.1186/1478-811X-12-30
Wang H (2016) Predicting MicroRNA biomarkers for cancer using phylogenetic tree and microarray analysis. Int J Mol Sci 17(5). doi:10.3390/ijms17050773
Osanto S, Qin Y, Buermans HP, Berkers J, Lerut E, Goeman JJ, van Poppel H (2012) Genome-wide microRNA expression analysis of clear cell renal cell carcinoma by next generation deep sequencing. PLoS One 7(6):e38298. doi:10.1371/journal.pone.0038298
Merritt WM, Lin YG, Han LY, Kamat AA, Spannuth WA, Schmandt R, Urbauer D, Pennacchio LA, Cheng JF, Nick AM, Deavers MT, Mourad-Zeidan A, Wang H, Mueller P, Lenburg ME, Gray JW, Mok S, Birrer MJ, Lopez-Berestein G, Coleman RL, Bar-Eli M, Sood AK (2008) Dicer, Drosha, and outcomes in patients with ovarian cancer. N Engl J Med 359(25):2641–2650. doi:10.1056/NEJMoa0803785
Takebe T, Enomura M, Yoshizawa E, Kimura M, Koike H, Ueno Y, Matsuzaki T, Yamazaki T, Toyohara T, Osafune K, Nakauchi H, Yoshikawa HY, Taniguchi H (2015) Vascularized and complex organ buds from diverse tissues via mesenchymal cell-driven condensation. Cell Stem Cell 16(5):556–565. doi:10.1016/j.stem.2015.03.004
Feng X, Wang Z, Fillmore R, Xi Y (2014) MiR-200, a new star miRNA in human cancer. Cancer Lett 344(2):166–173. doi:10.1016/j.canlet.2013.11.004
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
None.
Additional information
An erratum to this article is available at https://doi.org/10.1007/s12253-017-0235-1.
Rights and permissions
About this article
Cite this article
Masood, N., Yasmin, A. Entangling Relation of Micro RNA-let7, miRNA-200 and miRNA-125 with Various Cancers. Pathol. Oncol. Res. 23, 707–715 (2017). https://doi.org/10.1007/s12253-016-0184-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12253-016-0184-0