Abstract
MicroRNA-21 (miR-21) is overexpressed in a wide variety of cancers and has been related to cellular proliferation, apoptosis, and invasion; however, the function of miR-21 is unknown in oral tongue squamous cell carcinoma (OTSCC). The purpose of this study was to examine miR-21 expression in OTSCC, correlate it with clinicopathological factors, and investigate its contribution to OTSCC cell invasion. MiR-21 expression in 79 primary OTSCCs was evaluated using locked nucleic acid in situ hybridization, and correlation was examined with the clinicopathological factors. To determine the miR-21 target, we searched for molecular genes involved in tumor invasion using the commonly cited prediction program miRanda. In an OTSCC cell line, SCC25 cells, we further evaluated whether miR-21 contributes to cell invasiveness by blocking its expression with a specific knockdown LNA probe and confirmed the direct target by Matrigel invasion assay and Western blotting. MiR-21 overexpression was detected in 60 of 79 cases (75.9 %) and correlated with the pattern of invasion (P = 0.016). We selected DKK2 as a Wnt/antagonist involved in tumor invasion. MiR-21 overexpression was significantly correlated with the DKK2-/β-catenin- immunohistochemical phenotype. Knockdown of miR-21 significantly decreased the invasion potential of SCC25 cells with up-regulated DKK2. It was found that miR-21 is overexpressed and associated with tumor invasion in OTSCC, and that miR-21 promotes OTSCC cell invasion via the Wnt/β-catenin pathway by targeting DKK2 in vitro. These results suggest that miR-21 may be a potential therapeutic target for OTSCC treatment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Oral squamous cell carcinoma represents about 1–3 % of all human cancers, and is the 6th most frequent cancer in the world [1, 2]. Oral squamous cell carcinoma continues to show a poor prognosis and remains a lethal disease for more than 50 % of cases diagnosed annually [3]. In particular, its invasion potential is strongly related to its poor prognosis [4, 5]. Some investigators have demonstrated that a high malignancy grade of the deep invasive front had predictive value for neck regional lymph node metastasis [6] and prognosis [7, 8] in oral squamous cell carcinoma; therefore, elucidation of the mechanisms underlying invasion and a novel therapeutic strategy should be developed to improve the outcome of patients with oral squamous cell carcinoma.
MicroRNAs (miRNAs) are small non-coding RNAs of 18–25 nucleotides in length that have a crucial role in post-transcriptional regulation of gene expression [9]. By base-pairing to the 3′-untranslated region (3′-UTR) of target mRNAs, the mature miRNA is incorporated into the RNA-induced silencing complex (RISC) where it mediates gene expression by binding to target mRNA [9, 10]. Dysregulation of miRNAs in cancer has been shown to associate with various tumor characteristics and prognosis in a variety of cancers, including the oral cavity [11–13]. Several miRNAs have been functionally classified as proto-oncogenes or tumor suppressors, and contribute to tumor initiation and progression by promoting uncontrolled proliferation, favoring survival and/or promoting invasive behavior [11, 12].
In particular, microRNA-21 (miR-21) is the oncogenic miRNA upregulated in most types of cancer, including breast, colon, lung, pancreas, prostate, stomach, hepatocellular, ovarian, cervical, head and neck, and leukemia [14–16]. Recently, some authors have reported that miR-21 expression would be a useful biomarker associated with a worse outcome [17, 18]. Previous studies have shown that miR-21 may have a role in invasion and metastasis by several target molecules [19–22]; however, little is known as to how miR-21 affects the mechanism of OSCC invasion. Thus, in this study we focused on the correlation of miR-21 expression with tumor invasion, and the role of miR-21 in the biological behavior in OSCC. Finally, we showed that miR-21 promoted oral cancer invasion by down-regulating Wnt antagonist gene Dickkopf 2 (DKK2).
Materials and Methods
Patients
Paraffin-embedded sections were obtained from biopsy specimens of 79 patients with squamous cell carcinoma of the tongue who underwent radical surgery in our department between April 1996 and March 2005. The tumor stage was classified according to the TNM classification of the International Union Against Cancer. Tumor histologic differentiation was defined according to the WHO classification. The pattern of invasion was classified according to Bryne’s classification [7].
Cell Line
The human tongue cancer cell line SCC25, obtained from the American Type Culture Collection (Manassas, VA), was cultured in a 1:1 mixture of Ham’s F-12/DMEM supplemented with 10 % fetal bovine serum at 37 °C in the presence of 5 % CO2.
In Situ MiR-21 Hybridization
Serial sections 4 μm thick were taken from the tissue blocks. Deparaffinized sections in xylene were rinsed in sterile H2O, pretreated with 15 μg/ml proteinase K for 8 min at 37 °C, washed three times with sterile H2O, submerged in 95 % ethanol for 1 min, and air-dried completely. Sections were then hybridized in humidified chambers overnight at 37 °C using a 20 nM locked nucleic acid (LNA) probe diluted with hybridization buffer (BioGenex, San Ramon, CA). Digoxigenin (DIG)-labeled miRCULY LNA miRNA detection probes (Exiqon, Vedbaek, Denmark) were used in this study. The sequences of the miR-21 probe and the scramble control probe as the negative control were 5′-TCAACATCAGTCTGATAAGCTA-3′ and 5′-CATTATGTCGGACAACTCAAT-3′, respectively. After incubation, sections were placed in room temperature 5x SSC. Stringent washing was performed at 37 °C: one in 5× SSC, twice in 1x SSC and twice in 0.2× SSC. Sections were blocked against unspecific binding of the detecting antibody using DIG wash and blocking reagent (DIG Wash and Block Buffer Set; Roche, Mannheim, Germany). Alkaline phosphatase (AP)-conjugated anti-DIG (Roche) was diluted 1:800 in blocking solution and incubated for 2 hs at 30 °C. The sections were washed with PBS, stained with 4-nitro-blue tetrazolium (NBT)/5-brom-4-chloro-3′-indolylphosphate (BCIP) substrate ready-to-use tablet (Roche), and counterstained with nuclear fast red (Vector Laboratories, Burlingame, CA). Finally, the sections were dehydrated through an increasing gradient of ethanol solutions and mounted with Eukitt mounting medium (VWR, Herlev, Denmark).
Each slide was scored by 2 blinded, independent pathologists. To detect miR-21, each TMA spot was scanned and scored based on hybridization intensity (0: negative, 1: weak, 2: strong) and the percent of positive epithelial cells detected (0: <1 %, 1: focal, 1 % to 50 %, 2: diffuse, greater than 50 %). An ISH score for each lesion was calculated by multiplying the intensity by the area [23]. This was classified as 0: negative vs. ≥1: positive, representing low vs. high miR-21 expression. This relatively simple, reproducible scoring method provided highly concordant results between independent evaluators.
MiR-21 Target Prediction
To determine the miR-21 target, we searched for molecular genes involved in tumor invasion using a commonly cited prediction program miRanda (http://www.microrna.org; August 2010 release), which is a comprehensive resource of microRNA target and expression profiles. Prediction scores were computed using the miRanda database of highly conserved target sites with good mirSVR scores.
Immunohistochemical Staining
Serial sections 4 μm thick were taken from the tissue blocks. Deparaffinized sections in xylene were soaked in 10 mM citrate buffer (pH 6) and placed in an autoclave at 121 °C for 5 min for antigen retrieval. Endogenous peroxidase was blocked using 0.3 % H2O2 in methanol for 30 min. Immunohistochemistry was performed by the EnVision method (EnVision+, DAKO, Glostrup, Denmark). The primary antibodies used were directed against E-cadherin (#4065; Cell Signaling Technology, Danvers, MA; 1:50 dilution), β-catenin (#9562; Cell Signaling Technology; 1:50 dilution), and DKK2 (ab38594; Abcam, Cambridge, UK; 1:50 dilution). The sections were incubated with the antibodies overnight at 4 °C. Reaction products were visualized by immersing the sections in diaminobenzidine (DAB) solution, and the samples were counterstained with Myer’s hematoxylin and mounted. The expression was evaluated by calculating the total immunostaining score as the product of the proportion score and the intensity score. The proportion score described the estimated fraction of positive-stained tumor cells (0: none, 1: <10 %, 2: 10–50 %, 3: 50–80 %, 4: >80 %). The intensity score represented the estimated staining intensity (0: no staining, 1: weak, 2: moderate, 3: strong). The total score ranged from 0 to 12. As described previously [24], positive expression was defined as a total score >4.
Knockdown of MiR-21 with Anti-Sense LNA Oligomers
miRCURY LNA knockdown probes for miR-21 (miRCURY LNA microRNA Power Inhibitor, has-miR-21, product sequence is CAACATCAGTCTGATAAGCT), and for control miRNA (miRCURY LNA microRNA Power Inhibitor control, Negative Control A, product sequence is GTGTAACACGTCTATACGCCCA) LNA probes as a negative control were purchased from Exiqon, Inc. Cells were transfected with LNA probes using Oligofectamine reagent (Invitrogen) according to the manufacturer’s protocol. The SCC25 tongue cancer cell line was used for this experiment. Briefly, 2.5 × 104 SCC25 cells were plated in each well of six-well plates and allowed to grow for 24 h (until they were approximately 50 % confluent). LNA probe was then transfected into cells using Oligofectamine reagent and serum-free medium. After 4-h incubation, serum-rich medium was added.
Real Time RT-PCR
Cells were harvested and total RNA was extracted 72 h after transfection as previously described [24]. Total RNA was isolated from transfected cells using TRIzol reagent (Invitrogen) according to the manufacturer’s protocol. Then, cDNA synthesis was performed using a Universal cDNA synthesis kit (Exiqon). cDNA served as a template for microRNA quantitative real-time PCR using the miRCURY LNA Universal RT microRNA PCR kit (Exiqon). Primers included Exiqon-validated miR-21 specific primer sets (hsa-miR-21 primer, target sequence is CAACACCAGUCGAUGGGCUGU) and GAPDH control primers (Life Technologies, Carlsbad, CA). qPCR assays were performed using the Mx3000P QPCR System (Agilent Technologies, Santa Clara, CA).
Western Blotting
Cells were harvested and protein was extracted 72 h after transfection as previously described [24]. Total cell lysates were purified using a Mammalian cell extraction kit (BioVision, Mountain View, CA). All subsequent manipulations were performed on ice. The protein concentration of each sample was measured with micro-BCA protein assay reagent (Pierce Chemical Co., Rockford, IL). The samples were denatured in SDS sample buffer and loaded onto12.5 % polyacrylamide gels. After electrophoresis, the proteins were transferred onto a polyvinylidine difluoride membrane and immunoblotted with DKK2 (ab38594; Abcam). The signal was detected using the horseradish peroxidase-conjugated secondary antibody (Amersham Biosciences, Piscataway, NJ), and then visualized using an ECL Kit (Amersham Pharmacia Biotech, Buckinghamshire, UK).
Invasion Assay
The BD BioCoat Matrigel Invasion Chamber (Becton Dickinson, Bedford, MA) contains an 8 μm pore size PET membrane with a Matrigel Basement Membrane Matrix. Twelve cell culture inserts and a 24-well multiwell companion plate were used for this experiment. Cells were knockdown miR-21, knockdown miR-21 negative control, and transfection reagent only (control), followed by incubation for 24 h. The cells were collected by tripsinization, followed by seeding in the internal chamber at 2 × 105 cells in medium containing 5 % FBS. The lower chamber was filled with medium containing 10 % FBS as a chemoattractant. Cells were incubated for 48 h at 37 °C in 5 % CO2 atmosphere. After incubation, the non-invading cells were removed from the upper surface of the membrane with a cotton-tipped swab. The cells on the lower surface of the membrane were stained with Diff-Quick stain, and then inserts were desiccated to air dry. The cells were counted under a microscope at 100× magnification. For the control cell count, cells that passed through a control chamber without Matrigel were counted. All experiments were completed in triplicate, and at least 4 fields/well were counted. The percentage of the cell count that passed through the Matrigel chamber to the control count that passed through a control chamber without Matrigel was calculated as the invasion index.
Statistical Analysis
Statistical analyses were performed using StatMateIV (AtmsCo., Tokyo, Japan). The categorical data were assessed by Fisher’s exact test. Continuous data are given as the mean ± standard deviation. Data sets were examined by one-way analysis of variance (ANOVA) followed by Scheffe’s post-hoc test. The disease-specific survival rate was calculated using the Kaplan-Meier method. Significance was evaluated using the log-rank test. P values less than 0.05 were considered significant.
Results
Correlation of MiR-21 Overexpression and Clinicopathological Features
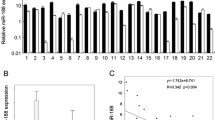
MiR-21 were expressed in the cytoplasm as violet by ISH, and were expressed in almost all cancer tissues rather than normal tissues (Fig. 1). MiR-21 tended to be especially highly expressed in the invasive front of cancer tissues. MiR-21 overexpression was detected in 60 of 79 cases (75.9 %) and correlated with the pattern of invasion (P = 0.016), but not age, gender, T stage, N stage, or pathological differentiation (Table 1).
Although patients without miR-21 overexpression showed a weak trend toward better survival, there was no significant relationship between miR-21 overexpression and survival (Fig. 2).
Determination of MiR-21 Target Gene
We used miRanda as a source in this study. For homosapiens, we found many target mRNAs from miR-21. The target mRNAs detected from the site contained many incorrect targets. We selected DKK2 as a Wnt/antagonist involved in tumor invasion. Moreover, DKK2 showed the highest mirSVR score (−0.72) among molecules involved in the Wnt pathway.
Relationship Between MiR-21 Overexpression and E-Cadherin, β-Catenin, or DKK2
Membranous immunostaining with E-cadherin and β-catenin was stronger in normal epithelium adjacent to the cancer tissue compared with the corresponding cancer areas (Fig. 3a). β-catenin and DKK2 immunostaining in cancer tissues were reversely correlated with miR-21 overexpression (Table 2; P < 0.001, respectively).
a Representative E-cadherin and β-catenin immunostaining (100×). Membranous immunostaining with E-cadherin and β-catenin was stronger in normal epithelium adjacent to cancer tissue compared with the corresponding cancer areas. b Representative miR-21 ISH, DKK2 and β-catenin immunostaining (100×). DKK2 and β-catenin immunostaining was reduced in areas with miR-21 overexpression
DKK2 expression was recognized in the cytoplasm of normal tissues, and loss of DKK2 expression was observed in cancer cells. In particular, DKK2 immunostaining was reduced in the area with miR-21 overexpression. In addition, strong staining of miR-21 was correlated with the reduced staining of DKK2 and β-catenin (Fig. 3b).
It was found that the miR-21 status was significantly correlated with the DKK2/β-catenin immunohistochemical phenotype (Table 3, P < 0.001). Almost all combined DKK2-/β-catenin- cases showed miR-21 overexpression.
Knockdown of MiR-21 Significantly Decreased the Invasion Potential of SCC25 Cells
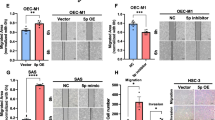
To determine the effect of miR-21 knockdown on the invasion potential, we transfected SCC25 cells with miR-21 knockdown probe, mock, and scrambled control probe. Mock and scrambled control probe transfection had no effect on miR-21 expression. A clear reduction in miR-21 was observed with the miR-21 knockdown probe (Fig. 4a), and was analyzed using the Matrigel invasion assay. Transfection with the miR-21 knockdown probe significantly decreased the invasive cells in SCC25 (P <0.01, Fig. 4b). The mean invasion index of cells transfected with the miR-21 knockdown probe was 12.5 %, whereas the means of the invasion index of cells treated with mock and transfected with the scrambled control probe were 58.2 % and 53.3 %, respectively.
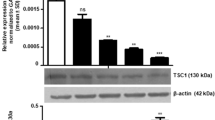
a Effect of miR-21 knockdown probe on the expression of miR-21 was evaluated by RT-PCR of SCC25 cells. Cells were transfected for 72 h with Mock (transfection reagent alone), control probe, or miR-21 knockdown probe. b Knockdown of miR-21 significantly decreased the invasion index in SCC25 cells. Data are presented as the means of three separate experiments, each performed in triplicate. Bars, SD. *P < 0.01, compared with Mock and control probe, respectively. c Effect of miR-21 knockdown probe on the expression of DKK2. Western blot shows DKK2 expression
Knockdown of MiR-21 Significantly Decreased the Level of Total Cellular DKK2 in SCC25 Cells
We assessed the effect of miR-21 knockdown on the total cellular content of DKK2. Transfection with the miR-21 knockdown probe significantly decreased the level of total cellular DKK2 in SCC25 cells (Fig. 4c).
Discussion
MiR-21 is a key oncomir that is up-regulated in several cancers, including breast, pancreas, lung, gastric, prostate, colon, and head and neck cancers [14–22]. Some researchers have demonstrated that miR-21 overexpression was involved in many characteristics of cancer, including proliferation, invasiveness, metastasis, and chemo-sensitivity [17–21]. MiR-21 negatively regulates several targets and impacts carcinogenesis. In our study using in situ hybridization, we showed that the level of miR-21 expression was significantly higher in OTSCC tissues than adjacent normal oral tissues. Furthermore, miR-21 overexpression was significantly associated with the invasion pattern of OTSCC; however, miR-21 overexpression demonstrated no relationship with age, gender, TN stage, or differentiation. These results implied that miR-21 overexpression might play critical roles in OTSCC tumorigenesis and tumor invasiveness. A systemic review and meta-analysis showed that miR-21 overexpression did indeed result in the poor survival of patients with a variety of carcinomas [16]. Li et al. [25] reported that miR-21 would be a clinically relevant independent prognostic factor in OTSCC. In our ISH analyses, patients without miR-21 overexpression showed a weak trend toward better survival, although the statistical difference was not significant.
MiR-21 overexpression has been linked to increased tumor invasion [19–22, 26–30]. Recent studies indicated that several molecules, including phosphatase and tensin homolog deleted on chromosome ten (PTEN) [21, 29, 30], programmed cell death 4 (PDCD4) [19, 20], and matrix metalloproteinases inhibitors RECK [22], were targets of miR-21, suggesting that miR-21 is an important oncogenic miRNA closely related to tumor invasion. Han et al. [30] demonstrated that miR-21 can regulate the cell invasiveness of breast cancer via AKT and ERK1/2 pathways by targeting PTEN. Moreover, Asangani et al. [19] reported that tumor suppressor PDCD4 could be negatively regulated by miR-21 at the post-transcriptional level via a specific target site within the 3′-UTR in colorectal cancer; however, the effect of miR-21 on tumor invasiveness in OTSCC remains to be elucidated.
In this study, we showed that miR-21 overexpression is correlated with the DKK2-/β-catenin- immunohistochemical phenotype in OTSCC; therefore, we suggested that DKK2 expression would be down-regulated by miR-21 in OTSCC. The DKK family is composed of four members, DKK1-DKK4, which act as inhibitors of Wnt/β-catenin signaling [31–33]. Among the DKK family, DKK2 is a putative Wnt/β-catenin signaling inhibitor that is generally down-regulated in human cancers, including ovarian cancer and renal cancer [33, 34]. DKK2 can function as either a Wnt agonist or antagonist, depending on the cellular context and the expressed amount of its binding partner low-density lipoprotein receptor-related protein 6 (LRP6) and its cofactor Kremen 2 [32]. Wnt/β-catenin signaling, transduced by LRP6 and Frizzled receptor complexes, leads to nuclear translocation of β-catenin and its interaction with TCF/LEF factors to regulate transcription [35]. Recently, some researchers reported that DKK2 inhibits tumor progression or invasion through the Wnt/β-catenin signaling pathway [33, 34]. Our results in this study suggested that miR-21inhibits tumor suppressor DKK2, activates the Wnt/β-catenin pathway and promotes OTSCC invasion.
Because our data using clinical samples suggested that miR-21 overexpression is related to the tumor invasion of OTSCC by targeting DKK2, we further evaluated whether miR-21 contributes to cell invasiveness by blocking its expression with a specific knockdown LNA probe. We observed that miR-21 knockdown decreased the invasion potential with up-regulated DKK2 in OTSCC cells. These results supported that miR-21 promotes OTSCC cell invasion via the Wnt/β-catenin pathway by targeting DKK2 (Fig. 5). In our study, we identified DKK2 as a marked miR-21 target because DKK2 showed the highest mirSVR score among molecules involved in the Wnt pathway. The MirSVR score is a new machine-learning method for ranking microRNA target sites by their down-regulation score, is calibrated to correlate with down-regulation, and can be interpreted as the empirical probability of target inhibition, leading to an intuitive choice of score threshold [36]. On the other hand, previous studies reported that miR-21 promotes tumor invasion through the β-catenin/STAT3 pathway by targeting RECK [22, 37]; therefore, not only the miR-21/DKK2/Wnt/β-catenin pathway has been implicated in the promotion of tumor invasion. Therefore, further studies are necessary to elucidate the molecular mechanism by which miR-21 modulates the Wnt/β-catenin pathway to promote OTSCC invasion.
In summary, we found that miR-21 is overexpressed and associated with tumor invasion in OTSCC, and that miR-21 promotes OTSCC cell invasion via the Wnt/β-catenin pathway by targeting DKK2 in vitro. These results suggest that miR-21 may be a potential therapeutic target for OTSCC treatment.
Abbreviations
- miR-21:
-
MicroRNA-21
- OTSCC:
-
Oral tongue squamous cell carcinoma
- RISC:
-
RNA-induced silencing complex
- DKK2:
-
Dickkopf 2
- LNA:
-
Locked nucleic acid
- DIG:
-
Digoxigenin
- AP:
-
Alkaline phosphatase
- NBT:
-
4-nitro-blue tetrazolium
- BCIP:
-
5-brom-4-chloro-3′-indolylphosphate
- ANOVA:
-
Analysis of variance
- PTEN:
-
Phosphatase and tensin homolog deleted on chromosome ten
- PDCD4:
-
Programmed cell death 4
References
Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D (2011) Global cancer statistics 2011. CA Cancer J Clin 61:69–90
Warnakulasuriya S (2009) Global epidemiology of oral and oropharyngeal cancer. Oral Oncol 45:309–316
Rogers NS, Brown SJ, Woolgar AJ, Lowe D, Magennis P, Shaw JR, Sutton D, Errington D, Vaughan D (2009) Survival following primary surgery for oral cancer. Oral Oncol 45:201–211
Yanamoto S, Kawasaki G, Yoshitomi I, Mizuno A (2002) P53, mdm2, and p21 expression in oral squamous cell carcinomas: relationship with clinicopathologic factors. Oral Surg Oral Med Oral Pathol Oral Radiol Endod 94:593–600
Sessions GD, Lenox J, Spector JG, Chao C, Chaudry AO (2003) Analysis of treatment results for base of tongue cancer. Laryngoscope 113:1252–1261
Kurokawa H, Zhang M, Matsumoto S, Yamashita Y, Tomoyose T, Tanaka T, Fukuyama H, Takahashi T (2005) The high prognostic value of the histologic grade at the deep invasive front of tongue squamous cell carcinoma. J Oral Pathol Med 34:329–333
Bryne M, Koppang SH, Lilleng R, Kjaerheim A (1992) Malignacy grading of the deep invasive margins of oral squamous cell carcinoma has high prognostic value. J Pathol 166:375–381
Bryne M, Jenssen N, Boysen M (1995) Histologic grading in the deep invasive front of T1 and T2 glottic squamous cell carcinomas has high prognostic value. Virchows Arch 427:277–281
Bartel PD (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Hannon JG, Rossi JJ (2004) Unlocking the potential of the human genome with RNA interference. Nature 431:371–378
Esquela-Kerscher A, Slack JF (2006) Oncomirs-microRNAs with a role in cancer. Nat Rev Cancer 6:259–269
Calin AG, Croce MC (2006) MicroRNA signatures in human cancers. Nat Rev Cancer 6:857–866
Wu HB, Xiong PX, Jia J, Zhang FW (2011) MicroRNAs: new actors in the oral cancer scene. Oral Oncol 47:314–319
Volinia S, Calin AG, Liu GC, Ambs S, Cimmino A, Petrocca F, Visone R, Iorio M, Roldo C, Ferracin M, Prueitt LR, Yanaihara N, Lanza G, Scarpa A, Vecchione A, Negrini M, Harris CC, Croce MC (2006) A microRNA expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci USA 103:2257–2261
Krichevsky MA, Gabriely G (2009) MiR-21: a small multi-faceted RNA. J Cell Mol Med 13:39–53
Fu X, Han Y, Wu Y, Zhu X, Lu X, Mao F, Wang X, He X, Zhao Y, Zhao Y (2011) Prognostic role of microRNA-21 in various carcinomas: a systematic review and meta-analysis. Eur J Clin Invest 41:1245–1253
Yan XL, Huang FX, Shao Q, Huang YM, Deng L, Wu LQ, Zeng XY, Shao YJ (2008) MicroRNA miR-21 overexpression in human breast cancer is associated with advanced clinical stage, lymph node metastasis and patient poor prognosis. RNA 14:2348–2360
Hwang HJ, Voortman J, Giovannetti E, Steinberg MS, Leon GL, Kim YT, Funel N, Park KJ, Kim AM, Kang HG, Kim WS, Del Chiaro M, Peters JG, Giaccone G (2010) Identification of microRNA-21 as a biomarker for chemoresistance and clinical outcome following adjuvant therapy in resectable pancreatic cancer. PLoS ONE 5:e10630
Asangani AI, Rasheed AS, Nikolova AD, Leupold HJ, Colburn HN, Post S, Allgayer H (2008) MicroRNA-21 (miR-21) post-transcriptionally downregulates tumor suppressor Pdcd4 and stimulates invasion, intravasation and metastasis in colorectal cancer. Oncogene 27:2128–2136
Reis PP, Tomenson M, Cervigne KN, Machado J, Jurisica I, Pintilie M, Sukhai AM, Perez-Ordonez B, Grenman R, Gilbert WR, Gullane JP, Irish CJ, Kamel-Reid S (2010) Programmed cell death 4 loss increases tumor cell invasion and is regulated by miR-21 in oral squamous cell carcinoma. Mol Cancer 9:238
Zhang GB, Li FJ, Yu QB, Zhu GZ, Liu YB, Yan M (2012) MicroRNA-21 promotes tumor proliferation and invasion in gastric cancer by targeting PTEN. Oncol Rep 27:1019–1026
Han L, Yue X, Zhou X, Lan MF, You G, Zhang W, Zhang LK, Cheng QJ, Yu ZS, Pu YP, Jiang T, Kang SC (2012) MicroRNA-21 expression is regulated by β-catenin/STAT3 pathway and promotes glioma cell invasion by direct targeting RECK. CNS Neurosci Ther 18:573–583
Li T, Li SR, Li HY, Zhong S, Chen YY, Zhang MC, Hu MM, Shen JZ (2012) MiR-21 as an independent biochemical recurrence predictor and potential therapeutic target for prostate cancer. J Urol 187:1466–1472
Yanamoto S, Kawasaki G, Yoshitomi I, Iwamoto T, Hirata K, Mizuno A (2007) Clinicopathologic significance of EpCAM expression in squamous cell carcinoma of the tongue and its possibility as a potential target for tongue cancer gene therapy. Oral Oncol 43:869–877
Li J, Huang H, Sun L, Yang M, Pan C, Chen W, Wu D, Lin Z, Zeng C, Yao Y, Zhang P, Song E (2009) MiR-21 indicates poor prognosis in tongue squamous cell carcinomas as an apoptosis inhibitor. Clin Cancer Res 15:3998–4008
Moriyama T, Ohuchida K, Mizumoto K, Yu J, Sato N, Nabae T, Takahata S, Toma H, Nagai E, Tanaka M (2009) MicroRNA-21 modulates biological functions of pancreatic cancer cells including their proliferation, invasion, and chemoresistance. Mol Cancer Ther 8:1067–1074
Zheng J, Xue H, Wang T, Jiang Y, Liu B, Li J, Li Y, Wang W, Zhang B, Sun M (2011) MiR-21 downregulates the tumor suppressor p12CDK2AP1 and stimulates cell proliferation and invasion. J Cell Biochem 112:872–880
Ren J, Zhu D, Liu M, Sun Y, Tian L (2010) Downregulation of miR-21 modulates Ras expression to promote apoptosis and suppress invasion of laryngeal squamous cell carcinoma. Eur J Cancer 46:3409–3416
Liu LZ, Wang H, Liu J, Wang XZ (2013) MicroRNA-21 (miR-21) expression promotes growth, metastasis, and chemo- or radioresistance in non-small cell lung cancer cells by targeting PTEN. Mol Cell Biochem 374:35–45
Han M, Liu M, Wang Y, Chen X, Xu J, Sun Y, Zhao L, Qu H, Fan Y, Wu C (2012) Antagonism of miR-21 reverses epithelial-mesenchymal transition and cancer stem cell phenotype through AKT/ERK1/2 inactivation by targeting PTEN. PLoS ONE 7:e39520
Krupnik EV, Sharp DJ, Jiang C, Robison K, Chickering WT, Amaravadi L, Brown ED, Guyot D, Mays G, Leiby K, Chang B, Duong T, Goodeari DA, Gearing PD, Sokol YS, MaCarthy AS (1999) Functional and structural diversity of the human Dickkopf gene family. Gene 238:301–313
Mao B, Niehrs C (2003) Kremen2 modulates Dickkopf2 activity during Wnt/LRP6 signaling. Gene 302:179–183
Zhu J, Zhang S, Gu L, Di W (2012) Epigenetic silencing of DKK2 and Wnt signal pathway components in human ovarian carcinoma. Carcinogenesis 33:2334–2343
Hirata H, Hinoda Y, Nakajima K, Kawamoto K, Kikuno N, Kawakami K, Yamamura S, Ueno K, Majid S, Saini S, Ishii N, Dahiya R (2009) Wnt antagonist gene DKK2 is epigenetically silenced and inhibits renal cancer progression through apoptotic and cell cycle pathways. Clin Cancer Res 15:5678–5687
Clvers H, Nusse R (2012) Wnt/β-catenin signaling and disease. Cell 149:1192–1205
Betel D, Koppal A, Agius P, Sander C, Leslie C (2010) Comprehensive modeling of microRNA targets predicts functional non-conserved and non-canonical sites. Genome Biol 11:R90
Yang HC, Yue J, Fan M, Pfeffer ML (2010) IFN induces miR-21 through a signal transducer and activator of transcription 3-dependent pathway as a suppressive negative feedback on IFN-induced apoptosis. Cancer Res 70:8108–8116
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kawakita, A., Yanamoto, S., Yamada, Si. et al. MicroRNA-21 Promotes Oral Cancer Invasion via the Wnt/β-Catenin Pathway by Targeting DKK2. Pathol. Oncol. Res. 20, 253–261 (2014). https://doi.org/10.1007/s12253-013-9689-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12253-013-9689-y