Abstract
A total of 152 lactic acid bacteria (LAB) were isolated from pickles produced in the Ankara-Çubuk region. These isolates were clustered into eight groups on the basis of their phenotypic characteristics including cell morphology, CO2 production from glucose, growth at 10 and 45 °C, growth in 6.5 % NaCl, and growth at pH 9.6. API 50 CH carbohydrate fermentation test, 16S ribosomal RNA (rRNA) sequence analysis, and sodium dodecyl sulfate-acrylamide gel electrophoresis (SDS-PAGE) whole-cell protein profile analysis were also performed for precise identification of the isolates at the species level. Molecular identification revealed that the most prevalent LAB species involved in pickle fermentation were Pediococcus ethanolidurans (46 isolates, 30.3 %), Lactobacillus brevis (37 isolates, 24.3 %), Lactobacillus plantarum (37 isolates, 24.3 %), and Lactobacillus buchneri (15 isolates, 9.9 %). Other LAB were found in minor frequencies such as Pediococcus parvulus (8 isolates, 5.3 %), Lactobacillus namurensis (6 isolates, 3.9 %), Lactobacillus diolivorans (1 isolate, 0.7 %), Lactobacillus parabrevis (1 isolate, 0.7 %), and Enterococcus casseliflavus (1 isolate, 0.7 %). When results of phenotypic and genotypic identification methods were compared, differences in the species distribution of LAB associated with pickles were defined between the API and the 16S rRNA sequencing. The API 50 CHL test coincided with the 16S rRNA results in 71 out of the 152 tested isolates, indicating that API gave unreliable identification results. A clear correlation could not be found between the results of whole-cell SDS profiles and 16S rRNA sequencing. Therefore, molecular characterization by 16S rRNA sequencing was considered to be the most reliable method for identifying isolates. The results presented in this work provide insight in to the LAB population associated with traditional Çubuk pickles and constitute a LAB strain resource for further studies involving the development of starter cultures.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pickle, a traditional fermented vegetable product, is very popular in the Çubuk region of Ankara, Turkey. Pickling has an important role in Turkish lifestyle. The Municipality of Çubuk has obtained a patent for “Çubuk Pickles.” Pickle production relies typically on spontaneous fermentation by indigenous lactic acid bacteria (LAB) flora associated with vegetables, such as cucumber, cabbage, and green pepper. However, the sensory quality of the pickle products varies by the raw material and inherent microbial flora. Therefore, developing starter cultures for pickle production is crucial to standardize the fermentation and the quality of the final product. Although numerous lactic cultures are available for the dairy and meat industries, there are no commercial starters designed for pickle production. To our knowledge, no previous studies have been conducted to develop LAB starter cultures indigenous to Ankara-Çubuk pickles.
Phenotypic methods including morphological and physiological analysis, biochemical characterization by API 50 CHL system and protein profiling by SDS-PAGE, have been widely used for the identification of LAB. However, the main drawbacks of phenotypic methods are poor reproducibility and low discriminatory power that do not allow precise identification at species level (Temmerman et al. 2004; Markiewicz et al. 2010). Although protein profiling is considered to be more reliable for LAB identification, it gives poor discrimination of Lactobacillus groups (Temmerman et al. 2004). Since Lactobacillus species are known as the predominant in pickle fermentation, a genome-based method is needed to be performed in this study. Moreover, a combined phenotypic and genotypic approach could be preferred for reliable characterization.
It is known that technological and probiotic properties are strain dependent, and so, reliable identification of LAB is of great importance before selecting starter cultures appropriate for industrial use (Sánchez et al. 2004). DNA sequencing of the 16S ribosomal RNA (rRNA) genes has been largely used for the identification of LAB (Balcázar et al. 2007; Pogačić et al. 2010; Moraes et al. 2013).
There are undoubtedly numerous published reports dealing with the identification of LAB isolated from traditional fermented vegetables in different countries/regions, such as China, Japan, and India (Tamang et al. 2005; Endo et al. 2008; Kingston et al. 2010; Yu et al. 2012). However, it is known that the distribution of microbial community in fermented vegetable products varies by region. In spite of being very popular in Turkey, very limited information is available on LAB associated with Çubuk pickles. Therefore, the objective of this study was to characterize and identify the predominant lactic acid bacteria present in pickles from the Ankara-Çubuk region. 16S rRNA sequencing coupled with SDS-PAGE of whole-cell proteins and biochemical characterization by API 50 CHL test could provide the precise identification of LAB involved in pickle fermentation. This study constitutes the first step in selecting LAB starter cultures indigenous to the Ankara-Çubuk region in order to preserve autochthonous strains and characteristic sensorial properties of traditional pickles.
Materials and methods
Isolation of LAB
Fifty three of pickle samples (prepared by using cucumber, cabbage, pepper, tomato, carrot, bean, and eggplant separately or in mixtures) were randomly collected from 18 individual household, local markets, and small factories in the Ankara-Çubuk region. Samples were collected in sterile bottles and kept at 4 °C after transported to the laboratory. The pH of brine samples was measured by a calibrated pH meter (Mettler Toledo, SevenEasy pH, China). Titratable acidity, expressed as g lactic acid per 100 mL, was calculated by titrating the brine with 0.01 mol/L NaOH with 0.1 % (w/v) phenolphthalein as the indicator. The salt content of the brine, expressed as g NaCl per 100 mL, was determined by titration with 0.1 N AgNO3 using 5 % (w/v) potassium dichromate (K2CrO4) as the indicator.
For colony counting and isolation of LAB, aliquots of brine samples were serially diluted with saline buffer containing 0.85 % (w/v) NaCl and plated on De Man-Rogosa-Sharpe (MRS) Agar (Merck, Germany) and M17 Agar (Merck, Germany). After incubation at 30 °C for 48–72 h, ten colonies showing different colony morphologies were picked from each plate of the highest dilution. Selected colonies were purified by using streak plate method and tested for cell morphology, Gram reaction, catalase, and oxidase activity. Gram-positive and catalase- and oxidase-negative cultures were considered to be presumptive LAB. The isolates were stored at −65 °C in 30 % (v/v) glycerol solution until identification analyses. Except where otherwise stated, the isolates were cultured at 30 °C for 48 h in MRS broth and/or MRS agar as basal media.
Phenotypic characterization of isolates
Phenotypic tests including cell morphology, CO2 production from glucose, growth at different temperatures (10, 15, and 45 °C), growth in 6.5 % NaCl, growth at pH 9.6, and ammonia production from arginine were conducted followed by further identification using the commercial API 50 CH carbohydrate fermentation test. Cell morphology of the isolates was determined using a phase contrast microscope. CO2 production from glucose was tested in MRS and M17 medium containing inverted Durham tubes. On the basis of the production of gas, cultures were considered as homofermentative/heterofermentative (Hitchener et al. 1982; Randazzo et al. 2004). Growth at different temperatures (10, 15, 45 °C) was examined after 72 h of incubation. Growth in 6.5 % NaCl was observed after 7 days of incubation at 30 °C in MRS broth containing 6.5 % NaCl as described by Chao et al. (2009). Growth at pH 9.6 was observed after 2 days of incubation at 30 °C in MRS broth which pH was adjusted to 9.6. Production of NH3 from arginine was determined according to Şimşek et al. (2006). Carbohydrate fermentation patterns were determined using API 50 CHL test strips according to manufacturer’s instructions (bioMérieux, France).
Genotypic characterization of isolates
SDS-PAGE whole-cell protein profile analysis
Whole-cell proteins were extracted according to De Angelis et al. 2001, Kizerwetter-Swida and Binek 2005, and Kim and Adachi 2007, and subsequently, sodium dodecyl sulfate (SDS)-acrylamide gel electrophoresis (PAGE) of the proteins was performed as described by Laemmli (1970). For the extraction of whole-cell proteins, cultures were incubated in MRS broth at 30 °C for 24–48 h. Cells were harvested and washed with 10 mL of distilled water by centrifugation at 4100g for 10 min. Washed cell pellets were resuspended in 100 μL lysozyme solution (BioChemica AppliChem GmbH, Germany) and incubated at 37 °C for 1 h vortexing every 10 min. Lysozyme-treated cells were then frozen at −20 °C and sonicated (VC 130, Sonics Vibra-Cell, USA) at 1000 amplitude for 1–2 min. After sonication, samples were mixed with 500 μL SDS-PAGE sample buffer (3.8 mL of distilled water, 1 mL of 0.5 mol/L Tris-HCl at pH 6.8, 0.8 mL of glycerol, 1.6 mL of 10 % (w/v) SDS, 0.8 mL of β-mercaptoethanol, 0.4 mL of 0.05 % (w/v) bromophenol blue), incubated at 95 °C for 5 min, and centrifuged at 14000g for 10 min. The supernatants were analyzed by SDS-PAGE with a vertical Mini-PROTEAN System (Bio-Rad Laboratories, USA). Briefly, 20 μL of supernatant samples were loaded onto 4–12 % (respective concentrations of stacking and separation gel) acrylamide gradient gels. Electrophoresis was performed using a power supply (EV231, Consort, Belgium) operated at 70 V through the stacking gel and 100 V through the separation gel for 2–3 h. The proteins were visualized by staining with Coomassie Brilliant blue G 250 dye solution (0.1 % (w/v) Coomassie Brilliant blue G 250 (Merck, Germany) 10 % (v/v) acetic acid, 50 % distilled water, 40 % ethanol) and were destained with the same solution without the addition of Coomassie Brilliant blue G 250. A standard protein solution (SM0661, Fermentas, EU), containing 14 proteins ranging in size from 10 kDa to 200 kDa, was used as a molar mass marker. Densitometry of SDS-PAGE gels was performed using Gel Logic 200 Imaging System (Kodak, USA). The similarity between protein band patterns of individual strains was estimated using the Dice coefficient, and dendrograms were derived from the similarity matrix by Unweighted Pair Group Method Using Average Linkage (UPGMA) cluster analysis that were performed using the statistiXL Version 1.8 software package.
16S rRNA gene sequences analysis
DNA extraction
Genomic DNA was isolated using a QIAGEN DNeasy Blood & Tissue Kit (Germany) in accordance with manufacturer’s instructions.
Amplification and sequencing of the 16S rRNA
Amplification of bacterial 16S rRNA regions was carried out by using F27 (AGAGTTTGATCCTGGCTCAG) and R1492 (TACGGYTACCTTGTTACGACTT) primers. The reaction mixture consisted of 100 ng template DNA, 1× buffer, 25 mM MgCl, 0.2 mM dNTP, 0.4 pmol of each primer, and 1.25 U of Taq polymerase in a final volume of 50 μL (Lane 1991). PCR amplifying protocol was as follows: a primary denaturation cycle at 94 °C for 5 min, then 30 cycles of denaturation at 94 °C for 1 min, annealing at 55 °C for 1 min and elongation at 72 °C for 1 min.
Sequencing of the PCR fragments was conducted by a biotechnology company (Refgen Biotechnology, METU Technocity, Ankara) with the BigDye® Terminator v3.1 Cycle Sequencing Kit, using the ABI 3130xL Genetic Analyzer (Applied Biosystems). Databases were screened for similarities by using BLAST.
Nucleotide sequence accession numbers
The nucleotide sequences of the 16S rRNA genes from isolates MF556, MF553, MF548, MF535, MF531, MF495, MF494, MF493, MF489, MF460, MF458, MF404, MF400, MF398, MF380, MF378, MF377, MF376, MF357, MF356, MF354, MF352, MF343, MF322, MF314, MF310, MF308, MF305, MF303, MF293, MF291, MF286, MF282, MF279, MF278, MF276, MF275, MF272, MF271, MF270, MF269, MF268, MF266, MF265, MF253, MF252, MF251, MF250, MF249, MF245, MF244, MF243, MF242, MF241, MF240, MF239, MF237, MF236, MF235, MF233, MF232, MF231, MF230, MF229, MF228, MF226, MF225, MF224, MF223, MF221, MF219, MF215, MF213, MF212, MF210, MF205, MF202, MF201, MF198, MF196, MF195, MF194, MF192, MF187, MF185, MF183, MF180, MF179, MF178, MF177, MF176, MF175, MF172, MF171, MF169, MF167, MF163, MF161, MF159, MF158, MF156, MF152, MF150, MF148, MF147, MF146, MF143, MF141, MF138, MF136, MF128, MF126, MF124, MF118, MF117, MF115, MF114, MF112, MF111, MF107, MF105, MF104, MF103, MF102, MF99, MF92, MF90, MF87, MF86, MF83, MF82, MF78, MF68, MF67, MF63, MF60, MF57, MF56, MF55, MF50, MF48, MF34, MF33, MF31, MF28, MF12, MF11, MF4, MF1, MF14, MF69, and MF214 have been submitted to GenBank and given consecutive accession numbers KJ994358 to KJ994509.
Results and discussion
Chemical and microbiological characteristics of pickle samples
The pH of pickle brine samples ranged from 3.08 to 4.74 with an average of 3.43, and the titratable acidity of pickle brine samples ranged from 0.04 to 1.31 g/100 mL with an average of 0.48 g/100 mL (data not shown). No significant correlation was found between pH and titratable acidity (r = −0.5463). This may be due to the different buffering capacities of the brine samples from different vegetable origins. The salt content of the brine samples varied from 1.34 to 6.94 g/100 mL with an average of 3.86 g/100 mL. The wide variations in the chemical characteristics of the pickle samples may be attributed to different raw materials used in the preparation of collected pickle samples and sampling at different stages of the fermentation process. Another possible explanation for these results may be the different storage conditions in different sampling sites that might affect the natural microflora. These variations also indicate the unstandardized conditions of the industrial production. Accordingly, variable results for titratable acidity, pH, salt, and microbiological quality in pickle samples was also observed by Çon and Karasu (2009). The LAB counts on MRS Agar and M17 Agar plates ranged from <1.00 to 7.75 and from 2.52 to 7.94 log CFU/mL, respectively (data not shown). A total of 557 isolates growing on MRS and M17 Agar were obtained from pickle samples. One hundred and fifty two Gram-positive and catalase- and oxidase-negative isolates were regarded as presumptive LAB and were further characterized by phenotypic and genotypic methods.
Comparison of the classical biochemical methods and the API system
One hundred and fifty two presumptive LAB isolates that were randomly isolated from the fifty three pickle samples were classified into eight preliminary groups (groups A–H) on the basis of key phenotypic characteristics including cell morphology, CO2 production from glucose, growth at 10 and 45 °C, growth in 6.5 % NaCl, and growth at pH 9.6 (Table 1). This grouping was performed by using a classification scheme derived from taxonomic data on Axelsson 2004. The arginine hydrolysis test was also used as an identification tool (data not shown).
Group A was composed of rod-shaped and heterofermentative LAB. This group comprised obligately heterofermentative lactobacilli (group III) including Lactobacillus brevis, Lactobacillus buchneri, Lactobacillus fermentum, and Lactobacillus reuteri. Forty eight isolates belonged to group A. Group B contained 48 isolates that were rod shaped and homofermentative. This group comprised obligately homofermentative (group I) including Lactobacillus acidophilus, Lactobacillus delbrueckii, Lactobacillus helveticus, and Lactobacillus salivarius and facultatively heterofermentative (group II) including Lactobacillus casei, Lactobacillus curvatus, Lactobacillus plantarum, and Lactobacillus sakei. Group C was composed of heterofermentative cocci that were able to grow at 10 °C, but not at 45 °C. This group showed variable growth characteristic in 6.5 % NaCl and at pH 9.6. Possible members of this group were the genera Leuconostoc, Oenococcus, and Weissella. None of the tested isolates had exactly the same phenotypic characteristics with group C. Only three isolates were heterofermentative cocci, but had growth characteristics different from those of group C. Group D represented the Enterococcus genus. It was composed of homofermentative cocci that were able to grow at both 10 and 45 °C, at pH 9.6, and in 6.5 % NaCl. This group did not include any isolate. Group E comprised 53 isolates that were cocci shaped and homofermentative. Except for one isolate, none of the isolates in this group were able to grow at pH 9.6. Other characteristics in terms of growth at 10 and 45 °C and in 6.5 % NaCl were variable within Pediococcus group. Group F, containing no isolates, was ascribed to the genera Aerococcus and Tetragenococcus. This group was defined as being homofermentative, coccoid LAB growing at 10 °C, at pH 9.6, and in 6.5 % NaCl, but not at 45 °C. Groups G and H did not include any isolate. They shared the common characteristics of being homofermentative cocci and non-growing at pH 9.6, and in 6.5 % NaCl. Group G (Lactococcus, Vagococcus) could be distinguished from group H (Streptococcus, Pediococcus) by the ability to grow at 10 °C.
One hundred and fifty two isolates yielded four preliminary groups within the eight defined groups. The preliminary identification results obtained by classical biochemical methods were then compared with those of the API 50 CHL test strips. For the interpretation of API results, a percentage of similarity greater than 90 % was considered as acceptable for identification at species level. Otherwise, the results were taken as doubtful. Thirty two out of the 152 tested isolates yielded an uninterpretable or doubtful API profile.
In accordance with the results of classical phenotypic tests, the isolates belonging to group A were identified as L. brevis (n = 34) and L. buchneri (n = 8) through the API system. Additionally, two isolates were identified as Lactobacillus pentosus and one as L. salivarius although they produced gas from glucose. These two species are known as homofermentative lactobacilli (Axelsson 2004). Three isolates in this group yielded doubtful API profile.
As shown in Table 1, group B was a heterogeneous group of LAB. There was a consistency between the results of API and classical tests for the identification of 27 isolates in this group including L. plantarum (n = 25) and L. pentosus (n = 2). However, the API identification results of 16 isolates did not match those of classical biochemical tests. Thirteen isolates were identified as L. brevis by API although they did not produce gas from glucose. Three isolates that were rod shaped based on microscopic examination were identified as Pediococcus damnosus (isolate nos. MF215 and MF115) and Weissella confusa (isolate no. MF104) by API. However, some of the physiological characteristics including being obligate homofermentative non-growing at pH 9.6 and inability to hydrolyze arginine of isolate nos. MF215 and MF115 matched those of P. damnosus species (data not shown) (Axelsson 2004; Hutkins 2006). It was reported that Weissella strains could also be rod shaped. Moreover, isolate no. MF104 possessed a few phenotypic traits of the genus Weissella in terms of non-growing at 45 °C and pH 9.6, but it did not produce gas from glucose and did not grow at 10 °C (data not shown), in contrast to Weissella species (Axelsson 2004). An additional isolate was found to belong to Carnobacterium maltaromaticum. This isolate possessed some of the key characteristics of the genus Carnobacterium (data not shown) (Schillinger and Holzapfel 1995; Axelsson 2004; Hutkins 2006). Four isolates in this group yielded doubtful API profile.
Three isolates that tentatively belonged to group C were identified as L. brevis, L. plantarum, and L. buchneri by the use of API 50 CH. The traditional biochemical analyses did not give reliable identification results for these strains.
The isolates in group E that were presumptively assigned to Pediococcus were identified as L. acidophilus (n = 11), P. damnosus (n = 9), L. plantarum (n = 6), L. brevis (n = 1) and Pediococcus spp. (n = 1) due to their carbohydrate fermentation patterns by API 50 CH. Twenty-five isolates in this group gave doubtful API profile with a percentage of similarity below 90 %. Eighteen isolates belonging to group E were identified as L. acidophilus (n = 11), L. plantarum (n = 6), and L. brevis (n = 1) by API 50 CH although they appeared coccoid. This could be explained by morphological variations that could occur within some Lactobacillus species (Hammes and Hertel 2006).
Genotypic characterization
In order to corroborate the phenotypic characterization, the 16S rRNA sequencing of 152 isolates was also performed. The two selected primers, namely F27 and R1492, were used to amplify the 16S rRNA fragment of LAB isolates resulting in an amplicon of approximately 1600 bp in size (data not shown). Differences in the species distribution of LAB associated with pickles were defined between the results of API and 16S rRNA sequencing. Following API, the most abundant species was found to be L. brevis whereas P. ethanolidurans based on 16S rRNA sequencing. Table 2 summarizes the results of the 16S rRNA identification and compares them with the API identification results. The API 50 CHL test coincided with the 16S rRNA results in 71 out of the 152 tested isolates, indicating that API gave unreliable identification results. Consistency between these two methods was observed for L. brevis (n = 32), L. plantarum (n = 30), L. buchneri (n = 8), and Pediococcus spp. (n = 1). In general, the API test failed to identify Pediococcus species. Divergent identification results were obtained for the remaining 81 isolates.
As shown in Table 2, 14 isolates identified as L. brevis in group A were assigned to Lactobacillus. namurensis (6), L. buchneri (3), P. ethanolidurans (2), Lactobacillus diolivorans (1), L. plantarum (1), Lactobacillus parabrevis (1) by sequencing of the 16S rRNA gene. This result clearly indicates that the discrimination of these species by phenotypic methods is very difficult. In group A, isolates classified as L. pentosus and L. salivarius based on API were reidentified as L. brevis and L. buchneri by the 16S rRNA sequencing which was consistent with preliminary phenotypic identification. With regard to group B results, it seems that the API test failed to distinguish closely related species including L. brevis and L. buchneri as well as L. plantarum and L. pentosus. It has been reported that L. brevis and L. buchneri are phenotypically very similar, except that L. buchneri ferments melezitose (Pot et al. 1994). The results of both API and 16S rRNA sequencing coincided exactly in group C isolates. The group E isolates were mostly found to be Pediococcus genus (except for six isolates) which was supported by classical biochemical identification. In group E, the API identification of four isolates which were identified as L. plantarum was in agreement with 16S rRNA sequencing. It is worthy of note that Pediococcus species were incorrectly identified as L. acidophilus by the API 50 CHL test. In accordance with this result, Boyd et al. (2005) reported that some species of lactobacilli were erroneously identified as L. acidophilus.
In general, the discrepancy between the results of phenotypic and genotypic identification was in evidence. Even in microscopic examination, misleading morphological results were obtained. This could be explained by the extensive morphological heterogeneity within LAB. Remarkably, Lactobacillus is the most physiologically, biochemically diverse group among LAB (Hutkins 2006). Some heterofermentative lactobacilli may appear as cocci, which make the discrimination difficult (Pot et al. 1994).
It was revealed that a variety of LAB species were found in pickle samples. On the basis of 16S rRNA sequencing, the species distribution of LAB associated with pickle samples were as follows: P. ethanolidurans (46 isolates, 30.3 %), L. brevis (37 isolates, 24.3 %), L. plantarum (37 isolates, 24.3 %), L. buchneri (15 isolates, 9.9 %), Pediococcus parvulus (8 isolates, 5.3 %), L. namurensis (6 isolates, 3.9 %), L. diolivorans (1 isolate, 0.7 %), L. parabrevis (1 isolate, 0.7 %), and Enterococcus casseliflavus (1 isolate, 0.7 %). Unexpectedly, P. ethanolidurans was found to be the most predominant LAB species in pickles. Yu et al. (2012) reported that they isolated a small amount of P. ethanolidurans strains from naturally fermented pickles. Furthermore, salt-tolerant amylolytic LAB strain P. ethanolidurans was isolated from Japanese pickles (nuka-zuke) by Iuchi et al. (2012). In recent publications, P. ethanolidurans and L. buchneri were reported to be involved in the onset of secondary fermentations that lead to fermented cucumber spoilage (Franco et al. 2012; Franco and Pérez-Díaz 2012; Johanningsmeier and Mcfeeters 2013). Since the samples in this study were collected at the various stages of fermentation, it could be likely that some of the samples proceeded to secondary fermentation. Moreover, Pediococcus spp. along with L. plantarum, which are more acid-tolerant species, has generally been known to predominate at the end of the primary fermentation (Singh and Ramesh 2008; Franco and Pérez-Díaz 2012) and survive during storage (Franco and Pérez-Díaz 2012). Also, P. ethanolidurans isolated by Liu et al. (2006) was reported to be resistance to harsh conditions including high ethanol and low sugar concentrations. Although Leuconostoc mesenteroides is also considered to be the one of main LAB species present in vegetable fermentations, it could not be detected in any of the pickle samples either by the API test or the 16S rRNA sequence analysis. A possible explanation for this might be that Leuconostocs thrive during the initial stages of fermentation (Singh and Ramesh 2008; Kabak and Dobson 2011). As we collected the samples randomly without following a time-dependent basis to reflect the dynamic changes of LAB microflora during fermentation, it could be possible to miss some of the species involved in the early stage of fermentation. Following P. ethanolidurans, L. brevis and L. plantarum constituted the majority of the LAB species in our study. These two species have been reported to be the main LAB involved in vegetable fermentations including pickles, sauerkraut, olives, and kimchi (Hutkins 2006). Tamang et al. (2005) reported that the major strains of LAB isolated from traditionally fermented vegetable products gundruk, sinki, khalpi, and inziangsang collected from the Northeast India were identified as L. brevis, L. plantarum, Pediococcus pentosaceus, Pediococcus acidilactici, and Leuconostoc fallax. L. plantarum has frequently been reported to be the predominant species in natural vegetable fermentations (Sánchez et al. 2000; Çetin 2011; Yu et al. 2012; Xiong et al. 2012) and could be isolated not only in the process of fermentation but also in the storage period (Tamminen et al. 2004). The predominance of L. plantarum in vegetable fermentations has generally been attributed to its distinct features including high resistance to acid and high production of lactic acid (Sánchez et al. 2000; Çon and Karasu 2009; Kabak and Dobson 2011; Xiong et al. 2012). In some previous studies, the presence of E. casseliflavus has been reported in raw cucumbers (Chen et al. 2012) as well as fermented cucumbers (Reina et al. 2005). P. parvulus has been associated with sauerkraut, fermented vegetables, fermented beans, beer, cider, and wine (Simpson and Taguchi 1995). Six isolates belonging to P. parvulus were isolated from the pickles of the Çubuk region in our study. As they are known to tolerate ethanol, it might also be possible to use these isolates for developing malolactic cultures in future investigations. L. namurensis was originally isolated from a traditional Belgian sourdough and belongs to the L. buchneri group, with Lactobacillus zymae, Lactobacillus acidifarinae, and Lactobacillus spicheri being the closest relatives (Scheirlinck et al. 2007). This species was also isolated from nukadoko, which is a fermented rice bran bed traditionally used in Japan for pickling vegetables (Sakamoto et al. 2011; Kato et al. 2014). L. namurensis was found to produce significant amounts of lactic acid, suggesting that it may contribute to the preservation of fermented products (Kato et al. 2014). L. diolivorans isolated from maize silage was proposed by Krooneman et al. (2002). L. diolivorans which belongs to L. buchneri group has obligately heterofermentative phenotype (Krooneman et al. 2002). L. buchneri has been known to convert lactic acid into 1,2-propanediol (Breidt et al. 2013), and subsequently, 2-propanediol is converted into propionate by L. diolivorans (Krooneman et al. 2002). Therefore, Zhang et al. (2010) suggested that propionate production by co-cultures of L. buchneri and L. diolivorans could be applied for bread preservation due to the antifungal capacity of the propionate. From this point of view, the L. buchneri and L. diolivorans isolates in our study could be used for similar purposes. Vancanneyt et al. (2006) reclassified two L. brevis strains which were isolated from cheese and wheat as L. parabrevis. Some of the described phenotypic characteristics of L. parabrevis, including being heterofermentative, growing at 15 °C and in 6.5 % NaCl, but not at 45 °C, were consistent with those of our isolate (data not shown).
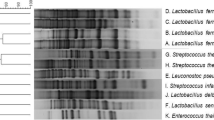
SDS-PAGE profiles
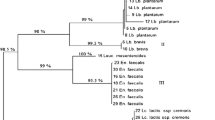
The comparison of whole-cell protein patterns obtained by highly standardized SDS-PAGE has proven to be a reliable method for the discrimination of closely related species (Pot et al. 1994; Vandamme et al. 1996; Hammes and Hertel 2006). Because of the lack of reference LAB strains in our laboratory, SDS-Page profiles of the isolates were used as an adjunct method rather than an identification tool. With this scope, cluster analysis was performed within the tested isolates. The UPGMA dendrogram obtained by whole-cell protein patterns is shown in Fig. 1. All isolates were clustered together at ca. 40 % similarity. When a similarity level of ca. 70 % was used, the isolates (with few exceptions, MF556, MF286, MF152, MF124) classified into six clusters. The isolates produced 1–21 protein bands of molar mass ranging from 7 to 250 kDa. Clusters I, II, and III totally contained the isolates of L. plantarum as identified by the 16S rRNA sequencing. In general, L. plantarum strains in our study give fewer blurry bands that make the discrimination ambiguous. Thus, L. plantarum species fell into all clusters. P. ethanolidurans and P. parvulus species mainly grouped into cluster IV. Cluster V comprised isolates belonging to L. brevis, L. buchneri, L. namurensis, L. diolivorans, L. parabrevis, P. ethanolidurans, P. parvulus, and E. casseliflavus. The high similarity level obtained between these species indicated that they cannot be distinguished by SDS-PAGE of whole-cell proteins. In summary, it is hard to find a correlation between the results of whole-cell SDS profiles and 16S rRNA sequencing without the use of reference LAB strains.
References
Axelsson L (2004) Lactic acid bacteria: classification and physiology. In: Salminen S, von Wright A, Ouwehand A (eds) Lactic acid bacteria: microbiological and functional aspects, 3rd edn. Marcel Dekker, New York
Balcázar JL, de Blas I, Ruiz-Zarzuela I, Vendrell D, Gironés O, Muzquiz JL (2007) Sequencing of variable regions of the 16S rRNA gene for identification of lactic acid bacteria isolated from the intestinal microbiota of healthy salmonids. Comp Immunol Microbiol Infect Dis 30:111–118
Boyd MA, Antonio MAD, Hillier SL (2005) Comparison of API 50 CH strips to whole-chromosomal DNA probes for identification of Lactobacillus species. J Clin Microbiol 43(10):5309–5311
Breidt F, Medina E, Wafa D, Pérez-Díaz I, Franco W, Huang H-Y, Johanningsmeier SD, Kim JH (2013) Characterization of cucumber fermentation spoilage bacteria by enrichment culture and 16S rDNA cloning. J Food Sci 78(3):M470–M476
Çetin B (2011) Production of probiotic mixed pickles (Turşu) and microbiological properties. Afr J Biotechnol 10(66):14926–14931
Chao S-H, Wu R-J, Watanabe K, Tsai Y-C (2009) Diversity of lactic acid bacteria in suan-tsai and fu-tsai, traditional fermented mustard products of Taiwan. Int J Food Microbiol 135:203–210
Chen Y, Wu H, Lo H, Lin W, Hsu W, Lin C, Lina P, Yanagida F (2012) Isolation and characterisation of lactic acid bacteria from jiang-gua (fermented cucumbers), a traditional fermented food in Taiwan. J Sci Food Agric 92:2069–2075
Çon AH, Karasu N (2009) Determination of antagonistic starter cultures for pickle and olive fermentation processes. Czech J Food Sci 27(3):185–193
De Angelis M, Corsetti A, Tosti N, Rossi J, Corbo MR, Gobbetti M (2001) Characterization of Non-starter lactic acid bacteria from Italian ewe cheeses based on phenotypic, genotypic, and cell wall protein analyses. Appl Environ Microbiol 67(5):2011–2020
Endo A, Mizuno H, Okada S (2008) Monitoring the bacterial community during fermentation of sunki, an unsalted, fermented vegetable traditional to the Kiso area of Japan. Lett Appl Microbiol 47:221–226
Franco W, Pérez-Díaz IM (2012) Role of selected oxidative yeasts and bacteria in cucumber secondary fermentation associated with spoilage of the fermented fruit. Food Microbiol 32:338–344
Franco W, Pérez-Díaz IM, Johanningsmeier SD, McFeeters RF (2012) Characteristics of spoilage-associated secondary cucumber fermentation. Appl Environ Microbiol 78(4):1273–1284
Hammes WP, Hertel C (2006) The genera Lactobacillus and Carnobacterium. In: Dworkin M, Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E (eds) The Prokaryotes A Handbook on the Biology of Bacteria, 3rd edn. Springer Science + Business Media, New York, pp 320–403
Hitchener BJ, Egan AF, Rogers PJ (1982) Characteristics of lactic acid bacteria isolated from vacuum-packaged beef. J Appl Bacteriol 52(1):31–37
Hutkins RW (2006) Microbiology and technology of fermented foods. Blackwell Publishing, USA
Iuchi A, Haruguchi S, Mongkolthanaruk W, Arima J, Nagase M, Khanh HQ, Ichiyanagi T, Yamaguchi T, Shimomura N, Aimi T (2012) Characterization of novel amylase from amylolytic lactic acid bacteria Pediococcus ethanolidurans isolated from Japanese pickles (nuka-zuke). Food Sci Technol Res 18(6):861–867
Johanningsmeier SD, McFeeters RF (2013) Metabolism of lactic acid in fermented cucumbers by Lactobacillus buchneri and related species, potential spoilage organisms in reduced salt fermentations. Food Microbiol 35:129–135
Kabak B, Dobson ADW (2011) An introduction to the traditional fermented foods and beverages of Turkey. Crit Rev Food Sci Nutr 51:248–260
Kato K, Toh H, Sakamoto N, Mori K, Tashiro K, Hibi N, Sonomoto K, Nakayamaa J (2014) Draft genome sequence of Lactobacillus namurensis Chizuka 01, isolated from nukadoko, a pickling bed of fermented rice bran. Genome Announc 2(1):e01263–13. doi:10.1128/genomeA. 01263-13
Kim S-Y, Adachi Y (2007) Biological and genetic classification of canine intestinal lactic acid bacteria and bifidobacteria. Microbiol Immunol 51(10):919–928
Kingston JJ, Radhika M, Roshini PT, Raksha MA, Murali HS, Batra HV (2010) Molecular characterization of lactic acid bacteria recovered from natural fermentation of beet root and carrot Kanji. Indian J Microbiol 50:292–298
Kizerwetter-Swida M, Binek M (2005) Selection of potentially probiotic Lactobacillus strains towards their inhibitory activity against poultry enteropathogenic bacteria. Pol J Microbiol 54(4):287–294
Krooneman J, Faber F, Alderkamp AC, Elferink SJHWO, Driehuis F, Cleenwerck I, Swings J, Gottschal JC, Vancanneyt M (2002) Lactobacillus diolivorans sp. nov., a 1,2-propanediol-degrading bacterium isolated from aerobically stable maize silage. Int J Syst Evol Microbiol 52:639–646
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. John Wiley and Sons, Chichester, pp 115–147
Liu L, Zhang B, Tong H, Dong X (2006) Pediococcus ethanolidurans sp. nov., isolated from the walls of a distilled-spirit-fermenting cellar. Int J Syst Evol Microbiol 56:2405–2408
Markiewicz LH, Biedrzycka E, Wasilewska E, Bielecka M (2010) Rapid molecular identification and characteristics of Lactobacillus strains. Folia Microbiol 55(5):481–488
Moraes PM, Perin LM, Júnior AS, Nero LA (2013) Comparison of phenotypic and molecular tests to identify lactic acid bacteria. Braz J Microbiol 44(1):109–112
Pogačić T, Kelava N, Zamberlin Š, Dolenčić-Špehar I, Samaržija D (2010) Methods for culture-independent identification of lactic acid bacteria in dairy products. Food Technol Biotechnol 48(1):3–10
Pot B, Ludwig W, Kersters K, Schleifer K-H (1994) Taxonomy of lactic acid bacteria. In: De Vuyst L, Vandamme EJ (eds) Bacteriocins of lactic acid bacteria. Microbiology, genetics and applications. Springer Science + Business Media, New York, pp 13–90
Randazzo CL, Restuccia C, Romano AD, Caggia C (2004) Lactobacillus casei, dominant species in naturally fermented Sicilian green olives. Int J Food Microbiol 90:9–14
Reina LD, Breidt F Jr, Fleming HP, Kathariou S (2005) Isolation and selection of lactic acid bacteria as biocontrol agents for nonacidified, refrigerated pickles. J Food Sci 70(1):M7–M11
Sakamoto N, Tanaka S, Sonomoto K, Nakayama J (2011) 16S rRNA pyrosequencing-based investigation of the bacterial community in nukadoko, a pickling bed of fermented rice bran. Int J Food Microbiol 144:352–359
Sánchez I, Palop L, Ballesteros C (2000) Biochemical characterization of lactic acid bacteria isolated from spontaneous fermentation of ‘Almagro’ eggplants. Int J Food Microbiol 59:9–17
Sánchez I, Seseña S, Palop LL (2004) Polyphasic study of the genetic diversity of lactobacilli associated with ‘Almagro’ eggplants spontaneous fermentation, based on combined numerical analysis of randomly amplified polymorphic DNA and pulsed-field gel electrophoresis patterns. J Appl Microbiol 97:446–458
Scheirlinck I, Meulen RV, Schoor AV, Cleenwerck I, Huys G, Vandamme P, Vuyst LD, Vancanneyt M (2007) Lactobacillus namurensis sp. nov., isolated from a traditional Belgian sourdough. Int J Syst Evol Microbiol 57:223–227
Schillinger U, Holzapfel WH (1995) The genus Carnobacterium. In: Wood BJB, Holzapfel WH (eds) The genera of lactic acid bacteria, 1st edn. Chapman & Hall, UK, pp 307–324
Simpson WJ, Taguchi H (1995) The genus Pediococcus, with notes on the genera Tetratogenococcus and Aerococcus. In: Wood BJB, Holzapfel WH (eds) The genera of lactic acid bacteria, 1st edn. Chapman & Hall, UK, pp 125–172
Şimşek Ö, Çon AH, Tulumoğlu Ş (2006) Isolating lactic starter cultures with antimicrobial activity for sourdough processes. Food Control 17:263–270
Singh AK, Ramesh A (2008) Succession of dominant and antagonistic lactic acid bacteria in fermented cucumber: insights from a PCR-based approach. Food Microbiol 25:278–287
Tamang JP, Tamang B, Schillinger U, Franz CMAP, Gores M, Holzapfel WH (2005) Identification of predominant lactic acid bacteria isolated from traditionally fermented vegetable products of the Eastern Himalayas. Int J Food Microbiol 105:347–356
Tamminen M, Joutsjoki T, Sjöblom M, Joutsen M, Palva A, Ryhänen E-L, Joutsjoki V (2004) Screening of lactic acid bacteria from fermented vegetables by carbohydrate profiling and PCR–ELISA. Lett Appl Microbiol 39:439–444
Temmerman R, Huys G, Swings J (2004) Identification of lactic acid bacteria: culture-dependent and culture-independent methods. Trends Food Sci Technol 15:348–359
Vancanneyt M, Naser SM, Engelbeen K, Wachter MD, Meulen RV, Cleenwerck I, Hoste B, Vuyst LD, Swings J (2006) Reclassification of Lactobacillus brevis strains LMG 11494 and LMG 11984 as Lactobacillus parabrevis sp. nov. Int J Syst Evol Microbiol 56:1553–1557
Vandamme P, Pot B, Gillis M, De Vos P, Kersters K, Swings J (1996) Polyphasic taxonomy, a consensus approach to bacterial systematic. Microbiol Rev 60(2):407–438
Xiong T, Guan Q, Song S, Hao M, Xie M (2012) Dynamic changes of lactic acid bacteria flora during Chinese sauerkraut fermentation. Food Control 26:178–181
Yu J, Gao W, Qing M, Sun Z, Wang W, Liu W, Pan L, Sun T, Wang H, Bai N, Zhang H (2012) Identification and characterization of lactic acid bacteria isolated from traditional pickles in Sichuan, China. J Gen Appl Microbiol 58:163–172
Zhang C, Brandt MJ, Schwab C, Gänzle MG (2010) Propionic acid production by cofermentation of Lactobacillus buchneri and Lactobacillus diolivorans in sourdough. Food Microbiol 27:390–395
Acknowledgments
This research was supported by the Scientific and Technological Research Council of Turkey (Project No. 108O491).
Author information
Authors and Affiliations
Corresponding author
Additional information
Simel Bağder Elmacı and Mehmet Tokatlı contributed equally to this work.
Rights and permissions
About this article
Cite this article
Bağder Elmacı, S., Tokatlı, M., Dursun, D. et al. Phenotypic and genotypic identification of lactic acid bacteria isolated from traditional pickles of the Çubuk region in Turkey. Folia Microbiol 60, 241–251 (2015). https://doi.org/10.1007/s12223-014-0363-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-014-0363-x