Abstract
Multiple gene knockouts play an important role in metabolic engineering. The flanked homology length, homologous to the region adjacent to the target gene, of the knockout fragments has a great effect on the efficiency of multiple gene knockouts, whereas the existing gene knockout methods can only supply a very short homology. This article presents a strategy of easily extending homologous sequence based on the available strain library through one-step PCR amplification (the one-step PCR method). In this approach, the library of single gene mutants was used as the templates for PCR to amplify knockout fragments. Thus, the flanked homology can be extended as long as possible by designing primers upstream and downstream far from the target gene. Based on the one-step PCR method, we studied the effect of the homology length and the number of mutations on the efficiency of multiple gene knockouts. Our results indicated that the one-step PCR method permitted rapid and efficient construction of multiple mutants continuously or simultaneously, and a length of 200–300 bp homologous sequence was equal for multiple gene knockouts.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Serving as a fundamental molecular biology technique, gene knockout is widely used in many fields, such as functional genomics, reverse genetics, and metabolic engineering. Many strategies of gene knockout have been developed in Saccharomyces cerevisiae, Candida albicans, Bacillus subtilis, etc. (Baudin et al. 1993; Vagner et al. 1998; Wilson et al. 1999). For Escherichia coli, the λ Red-mediated homologous recombination system (Datsenko and Wanner 2000; Yu et al. 2000) is a general method for single gene disruption in the wild-type cells. Baba et al. (2006) even constructed a single gene mutation library of E. coli K-12 BW25113 of all the deletable genes employing the efficient single gene knockout approach developed by Datsenko and Wanner (2000). According to this method, gene knockout can be realized easily by introducing the DNA fragments that carry an Flp recombinase target (FRT)-flanked resistance gene and a very short DNA sequence of only 36–50 bp homologous to the regions adjacent to the target gene into the cells. Researches on metabolic engineering and synthetic biology demand global and systematic regulation of the cell physiology and metabolism. Therefore, it is usually necessary to generate multiple mutations in the same strain (Alper et al. 2005; Burgard and Maranas 2001).
The length of flanked DNA sequence, homologous to the region adjacent to the target gene, of the knockout fragments has a great effect on the recombination efficiency (Murphy 1998; Yu et al. 2000). Thereby, extending the length of homology is beneficial to gene deletion, especially to multiple gene deletions. Based on this, Derbise et al. (2003) reported a rapid and cloning-free method (the three-step PCR method) modified from the description of Horton et al. (1989) and Wach (1996) to extend the homologous sequence. But this method needs three PCR steps, which are too niggling, time consuming, and laborious, to construct the knockout fragments. The P1 phage-mediated transduction method described by Miller (1992) is now a general choice to inactivate multiple genes successively in one strain (Moon et al. 2011; Typas et al. 2008). However, P1 virus could bring uncertainty of the strains and cause cross-contamination of the experiments.
Here, we described a strategy for obtaining long enough homology to inactivate gene efficiently by one-step PCR amplification (the one-step PCR method). In this strategy, the single gene mutants were selected as the templates for PCR to amplify the knockout fragments which consisted of an antibiotic resistance gene and the flanking homology. The homology can be extended long enough by designing primers upstream and downstream far from the desired gene on one condition that the single gene mutant derived from the same species with the target strain. Based on the one-step PCR method, we studied the effect of the homology length and the number of FRT sites on the efficiency of multiple gene knockouts using the λ Red-mediated recombination method developed by Datsenko and Wanner (2000).
Materials and methods
Strains, plasmids, and primers
E. coli MG1655 (F − λ − ilvG− rfb-50 rph-1) was used as the host for gene deletion. The single gene mutants derived from MG1655 stocked in our laboratory (Table 1) were selected as the templates for PCR to amplify the fragments for multiple gene knockouts. Plasmids pKD46 (Datsenko and Wanner 2000) and pCP20 (Cherepanov and Wackernagel 1995) used to execute gene disruptions and eliminate antibiotic resistance were also stored in our laboratory. All primers used in this study were summarized in Table 2.
Construction of the knockout fragments
DNA fragments for gene knockout were constructed using the one-step PCR method as described below. In this strategy, the single gene mutants containing the antibiotic markers from the Keio collection or other resources were selected as the PCR templates rather than pKD3 or pKD4, which were the generally used template plasmids for the construction of the knockout fragments (Datsenko and Wanner 2000). A pair of 20–30 bp primers, 200–300 bp far from upstream and downstream of the target gene, respectively, was designed for the one-step PCR reaction (Fig. 1). The PCR reaction was performed under the condition of 1 cycle of pre-denaturation at 94 °C for 5 min, 30 cycles of denaturation at 94 °C for 45 s, annealing at 56 °C for 45 s, elongation at 72 °C for 2.5 min, and 1 cycle of post-elongation at 72 °C for 10 min. The obtained PCR product was an antibiotic resistance gene flanked by about 200–300 bp homology extensions at the both ends.
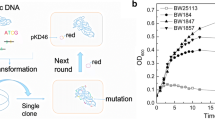
The strategy of easily extending homology flanked the target gene for gene replacement. The geneA mutant was used as the template for PCR to amplify knockout fragment and primers (P1 and P2) were designed upstream and downstream far from the target gene. Step 1 PCR amplification for obtaining the geneA knockout fragment with the flanking long homology. Step 2 Homologous recombination between the geneA knockout fragment and the target strain. Step 3 Obtaining of the correct recombinants with geneA mutation
Gene disruption
Gene disruption was executed following the λ Red recombination method developed by Datsenko and Wanner (2000). The correct recombinants were verified by colony PCR, and the recombination efficiency was defined as the ratio of the correct recombinants to all transformants picked out randomly (Baba et al. 2006).
Results and discussion
According to our experience, the homologous sequence length of 36–50 bp is relatively short for multiple gene deletions as it caused the lower recombination frequency and the high probability of false recombinants; while complete synthesis of primers containing long homology is expensive and fallible. Thus, we provided a strategy for extending long enough homology by one-step PCR method. With the presented Keio collection of 3,985 single gene mutants (Baba et al. 2006) or other sources as the templates, the knockout fragments with long homology can be amplified easily using the one-step PCR method. By employing this method, we studied the effect of the homology length and the number of FRT sites on the recombination efficiency. The plasmid pKD46 was used to supply Red recombinase for gene knockout and the plasmid pCP20 for eliminating antibiotic resistance.
Effect of the length of homologous sequence on recombination
By applying the one-step PCR method based on the available single gene mutant library, a large number of genes were deleted in the engineered E. coli which already had multiple FRT sites (Table 1). The statistical results indicated that the recombination efficiency was greatly improved with the extension of homologous sequence (Table 3). When 39-bp homologous sequence was used, the recombination efficiency was only 28.9 % and not stable (depends on the genes to be knocked out). This may be explained by that the short extensions increased probability of nonspecial recombination. The shorter the extensions are, the lower the specificity homologous to the sequence of the E. coli genome is. In addition, there are two FRT sites flanking the antibiotic resistance gene in the knockout fragment, and they can also recombine with the FRT sites which were left from previous gene replacement and eliminate antibiotic resistance in the target strain (Shen and Huang 1986), resulting in false recombination. The unstable recombination efficiency may due to the varied effect of the different gene to the cell metabolism (Baba et al. 2006). The mean efficiency increased to 70.8 % if we maintained the homologous sequence at 100 bp. With the extension of the homology length to 200–300 bp, the efficiency could reach 85.7 %. In the meantime, the difference of mutational frequency for different genes was also diminished apparently. Therefore, we concluded that the length of homology extensions had a great effect on recombination; the recombination efficiency can be easily improved through the extension of homologous sequence.
Effect of FRT sites of E. coli on recombination
According to the Datsenko method, the number of the FRT sites remained in the genome DNA reflected the number of mutations. Our previous study showed that FRT sites left in E. coli strain may lower the gene knockout efficiency, especially when short homology extensions were used. The more FRT sites are left in the genome of E. coli, the lower knockout efficiency was. Therefore, by using the PCR products with 200–300 bp homology extensions, which were generated from the single gene knockout strains, we compared the gene knockout efficiency with the engineered E. coli which had different numbers of FRT sites and got high efficiencies of gene disruption (Table 4). When wild-type E. coli was used as the host strain, the knockout efficiency reached to 98.8 %. Despite the recombinant efficiency decrease in the engineered E. coli which had one to three FRT sites, the knockout efficiency could nearly reach to 86.9 %. Even in E. coli which had five to seven FRT sites, the mean efficiency was still over 85.0 %. This also indicated that homology extensions of 200–300 bp are enough for the gene knockout in the multiple gene mutants.
Simultaneous gene knockout using long homology extensions
The one-step PCR method is not only applicable to knockout many genes continuously in one strain, but also able to knockout two or more genes simultaneously in the same strain if different selectable markers are used. We tried to simultaneously knockout genes iclR and pta in E. coli MG1655 following the one-step PCR method. Although only five recombinants were grown out on the double resistant antibiotic-containing plates, all the grown recombinants performed correct recombination of the two genes, resulting in a high frequency of the double deletion. We also tried genes ldhA and ptsG in E. coli MG1655 and all the three recombinants obtained were correct constructs.
Multiple gene knockouts play an important role in metabolic engineering and synthetic biology, whereas the Red homologous recombination method developed by Datsenko and Wanner (2000) is suitable for single gene knockout in the wild-type strains. For multiple gene deletions, our results indicated that the homologous sequence used in this method was short, and the recombination efficiency was lower and unstable when short homologous extension was used. Fortunately, our strategy can solve this problem easily by the one-step PCR method of extending homologous sequence based on the single gene mutants.
Compared with the three-step PCR method and the P1 virus-mediated transduction, our strategy has many advantages: simple, high efficiency, easy to manipulate, more direct and certain, and suitable for multiple gene deletions successively or simultaneously in the same strain. Both the one-step PCR method and the P1 transduction-mediated multiple gene knockouts method need the single gene knockout strain to be a template or a donor strain; fortunately, the Keio collection (Baba et al. 2006) makes it feasible. However, the P1 transduction-mediated procedure required that the donor strains and recipients strain should be the same species; otherwise, mistakes may be brought in. Moreover, the transduction frequencies of different genes are far different, related to the gene position (Cooper and Helmstetter 1968; Masters and Broda 1971). For our strategy, single gene mutants from different sources can be also used as the template on one condition that the DNA sequence which flanked the target gene is homologous. Hence, this strategy can be also introduced into other microbes as long as there are available mutants as the templates for PCR. The recombinant efficiency for our strategy largely lies on the length of the homology, but it can be extended easily as long as possible through the one-step PCR method.
References
Alper H, Jin YS, Moxley JF, Stephanopoulos G (2005) Identifying gene targets for the metabolic engineering of lycopene biosynthesis in Escherichia coli. Metab Eng 7:155–164
Baba T, Ara T, Hasegawa M, Takai Y, Okumura Y, Baba M, Datsenko KA, Tomita M, Wanner BL, Mori H (2006) Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol Syst Biol 2:2006.0008
Baudin A, Ozier-Kalogeropoulos O, Denouel A, Lacroute F, Cullin C (1993) A simple and efficient method for direct gene deletion in Saccharomyces cerevisiae. Nucleic Acids Res 21:3329–3330
Burgard AP, Maranas CD (2001) Probing the performance limits of the Escherichia coli metabolic network subject to gene additions or deletions. Biotechnol Bioeng 74:364–375
Cherepanov PP, Wackernagel W (1995) Gene disruption in Escherichia coli: TcR and KmR cassettes with the option of Flp-catalyzed excision of the antibiotic-resistance determinant. Gene 158:9–14
Cooper S, Helmstetter CE (1968) Chromosome replication and the division cycle of Escherichia coli B/r. J Mol Biol 31:519–540
Datsenko KA, Wanner BL (2000) One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci 97:6640–6645
Derbise A, Lesic B, Dacheux D, Ghigo JM, Carniel E (2003) A rapid and simple method for inactivating chromosomal genes in Yersinia. FEMS Immunol Med Microbiol 38:113–116
Horton RM, Hunt HD, Ho SN, Pullen JK, Pease LR (1989) Engineering hybrid genes without the use of restriction enzymes: gene splicing by overlap extension. Gene 77:61–68
Masters M, Broda P (1971) Evidence for the bidirectional replications of the Escherichia coli chromosome. Nat New Biol 232:137–140
Miller JH (1992) A short course in bacterial genetics: a laboratory manual and handbook for Escherichia coli and related bacteria. Cold Spring Harbor Laboratory Press, New York
Moon TS, Clarke EJ, Groban ES, Tamsir A, Clark RM, Eames M, Kortemme T, Voigt CA (2011) Construction of a genetic multiplexer to toggle between chemosensory pathways in Escherichia coli. J Mol Biol 406:215–227
Murphy KC (1998) Use of bacteriophage lambda recombination functions to promote gene replacement in Escherichia coli. J Bacteriol 180:2063–2071
Shen P, Huang HV (1986) Homologous recombination in Escherichia coli: dependence on substrate length and homology. Genetics 112:441–457
Typas A, Nichols RJ, Siegele DA, Shales M, Collins SR, Lim B, Braberg H, Yamamoto N, Takeuchi R, Wanner BL, Mori H, Weissman JS, Krogan NJ, Gross CA (2008) A tool-kit for high-throughput, quantitative analyses of genetic interactions in E. coli. Nat Methods 5:781–787
Vagner V, Dervyn E, Ehrlich SD (1998) A vector for systematic gene inactivation in Bacillus subtilis. Microbiology 144:3097–3104
Wach A (1996) PCR-synthesis of marker cassettes with long flanking homology regions for gene disruptions in S. cerevisiae. Yeast 12:259–265
Wilson RB, Davis D, Mitchell AP (1999) Rapid hypothesis testing with Candida albicans through gene disruption with short homology regions. J Bacteriol 181:1868–1874
Yu D, Ellis HM, Lee EC, Jenkins NA, Copeland NG, Court DL (2000) An efficient recombination system for chromosome engineering in Escherichia coli. Proc Natl Acad Sci 97:5978–5983
Acknowledgments
This research was financially supported by a grant from the National Natural Science Foundation of China (31070092) and a grant of the National Basic Research Program of China (2011CB707405).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, M., Gu, P., Kang, J. et al. Extending homologous sequence based on the single gene mutants by one-step PCR for efficient multiple gene knockouts. Folia Microbiol 57, 209–214 (2012). https://doi.org/10.1007/s12223-012-0111-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-012-0111-z