Abstract
Rigidity sensing plays a fundamental role in multiple cell functions ranging from migration, to proliferation and differentiation (Engler et al., Cell 126:677–689, 2006; Lo et al., Biophys. J. 79:144–152, 2000; Wells, Hepatology 47:1394–1400, 2008; Zoldan et al., Biomaterials 32:9612–9621, 2011). During migration, single cells have been reported to preferentially move toward more rigid regions of a substrate in a process termed durotaxis. Durotaxis could contribute to cell migration in wound healing and gastrulation, where local gradients in tissue rigidity have been described. Despite the potential importance of this phenomenon to physiology and disease, it remains unclear how rigidity guides these behaviors and the underlying cellular and molecular mechanisms. To investigate the functional role of subcellular distribution and dynamics of cellular traction forces during durotaxis, we developed a unique microfabrication strategy to generate elastomeric micropost arrays patterned with regions exhibiting two different rigidities juxtaposed next to each other. After initial cell attachment on the rigidity boundary of the micropost array, NIH 3T3 fibroblasts were observed to preferentially migrate toward the rigid region of the micropost array, indicative of durotaxis. Additionally, cells bridging two rigidities across the rigidity boundary on the micropost array developed stronger traction forces on the more rigid side of the substrate indistinguishable from forces generated by cells exclusively seeded on rigid regions of the micropost array. Together, our results highlighted the utility of step-rigidity micropost arrays to investigate the functional role of traction forces in rigidity sensing and durotaxis, suggesting that cells could sense substrate rigidity locally to induce an asymmetrical intracellular traction force distribution to contribute to durotaxis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rigidity sensing plays a fundamental role in guiding the outcome of multiple dynamic cell behaviors including migration, proliferation, and differentiation.4,8,13,31,37 Single migrating cells have been reported to sense rigidity by preferentially migrating towards the more rigid region of the substrate in a process termed durotaxis.14,19 Durotaxis is potentially a prominent guidance cue for cell migration as many physiological situations where cell migration is important, such as wound healing and gastrulation, are characterized by local changes in environmental rigidity.2,36 In addition, pathologic conditions such as cancer and fibrosis are characterized by local increases in tissue rigidity, which may contribute to increased trafficking of fibroblasts and immune cells into the diseased foci.2,15 Despite its importance, detailed molecular and cellular understanding of durotaxis remains largely elusive.
Studying how matrix mechanics regulates durotaxis requires engineered substrates that present a reproducible and quantitatively well-defined substrate rigidity gradient. Prior studies documenting durotaxis have been performed by generating a rigidity gradient within synthetic hydrogels through varying the ratio of hydrogel monomer to cross-linker across a hydrogel substrate,13 using gradients of light to mediate photo-initiated PA crosslinking,28 using gradients of PA pre-polymer generated using microfluidic approaches,10,28 or simply applying a tangential strain in the direction away from a cell with a microneedle to locally pull PA gels.18,29 These studies have revealed the existence of durotaxis in different types of mechanosensitive adherent cells and shown that durotaxis is functionally correlated with substrate rigidity gradient magnitude and is mediated by actomyosin-mediated cellular traction forces and cell attachments to extracellular matrix proteins via focal adhesions (FAs).10,18,28,29 A recent study using composite materials containing local rigid adhesive islands grafted onto the surface of a non-adhesive polyacrylamide hydrogel has suggested that rigidity sensing may be dictated by material compliance across the cell length.9 Interestingly, a more recent study using high-resolution time-lapse traction force microscopy (TFM)18 has implicated that traction force fluctuation and distribution within single mature FAs might be important to regulate durotaxis.18 Together, these studies have established the importance of a substrate rigidity gradient in mediating directional cell migration and suggested that sensing substrate rigidity can be a local mechanotransductive molecular event occurring within individual FAs or a cellular response integrating rigidity signals across the whole cell body. However, whether or not cells sense gradients of rigidity within individual FAs or if rigidity is sensed globally between FAs during durotaxis has not been directly investigated.
In addition to PA gels, elastomeric micropost arrays made in polydimethylsiloxane (PDMS) have proven as a versatile tool to control substrate mechanics and report traction forces with a sub-nanonewton (nN) resolution for single adherent cells.5,20,23,33 Recent studies have further shown the possibility to generate PDMS micropost arrays with controlled rigidity profiles while monitoring live-cell traction forces during cell migration. The first PDMS micropost array with rigidity gradients was generated with discrete rigidity boundaries by changing post heights while keeping the post top surface coplanar.6 However, even through durotaxis was reported on the step-rigidity PDMS micropost array, this study did not report cellular traction forces during durotaxis. Further, the method employed to generate the step-rigidity PDMS micropost array required a complicated microfabrication process involving coating Si microscale hole arrays with a thick layer of SiO2 using plasma enhanced chemical vapor deposition (PECVD) before the surface of the wafer was planarized using chemical-mechanical polishing (CMP). Step changes in substrate rigidity have also been generated in PDMS micropost arrays by keeping the post height constant while spatially varying the post diameter.26 More recently, rigidity gradients have been generated within PDMS micropost arrays by gradually increasing the post diameter along one direction of the array.21 However, changing the post diameter in the PDMS micropost array affects cellular adhesive environment (e.g., the post diameter and density changes can directly affect the maximum size of individual FAs and FA organization, respectively) that can introduce additional microenvironmental signals to make interpretation of experimental findings difficult.
To assess whether cells sense substrate rigidity during durotaxis on a local or cellular length scale, herein we reported a novel microfabrication strategy that could generate PDMS micropost arrays with discrete step changes in rigidity without using complicated PECVD or CMP process (see “Methods” section). In order to keep the tips of the PDMS microposts coplanar, we altered micropost heights by changing the height of underlying base of the substrate. Cells seeded across post rigidity boundaries responded by migrating towards more rigid posts, indicating the presence of durotaxis on our step-rigidity PDMS micropost array. We then measured traction forces of cells bridging step-rigidity boundaries and observed that cells could generate asymmetrical strains and traction forces within each rigidity region, such that subcellular regions could behave analogously to cells wholly seeded on uniform substrates with that rigidity. In particular, traction forces of cells bridging step-rigidity boundaries were greater for subcellular regions on more rigid substrates. These results suggested that cells could sense substrate rigidity locally, and this caused an asymmetrical intracellular traction force distribution resulting in durotaxis.
Results
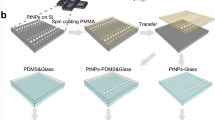
Negative masters for the step-rigidity PDMS micropost array were fabricated in silicon (Si) wafers using standard photolithography and a two-stage deep reactive-ion etching (DRIE) protocol (Fig. 1a). The first DRIE step was used to generate negative Si masters with a regular array of microscale cylindrical holes of a uniform depth across the Si substrate (Supporting Fig. 1a). The depth of microscale holes, which determined the post height in soft regions of the step-rigidity PDMS micropost array, was precisely controlled by monitoring DRIE etching time. Before the second DRIE, a thick photoresist layer was spin coated onto the negative Si master to fill completely the hole array before the second photolithography step to expose selected areas on the top surface of the Si master. Special care was taken to ensure photoresist filling the cylindrical holes remained after photolithography to protect the bottom surface of the holes during the second DRIE step (Supporting Fig. 1a). Owing to protection of photoresist for the bottom surface of the holes, the Si wafer was etched only from its top surface during the second DRIE step, with its etching depth controlled by monitoring DRIE etching time. The final step-rigidity PDMS micropost array was generated via replica molding using the Si negative master with silicone elastomer PDMS. In the final PDMS device, the selected surface regions of the Si substrate that were reduced in height during the second DRIE step became steps on which shorter PDMS posts were sitting, creating juxtaposed regions consisting of shorter (rigid) and taller (soft) PDMS microposts with their tops co-planar (Figs. 1b, 1c).
Patterned PDMS micropost arrays to spatially regulate substrate rigidity. (a) Schematic of the microfabrication process to generate spatially patterned PDMS micropost arrays. Drawings were not to scale. The fabrication process involved standard photolithography, deep reactive-ion etching (DRIE), and replica molding of PDMS. (b) Cross-sectional scanning electron microscopy (SEM) images showing microfabricated silicon hole array masters with spatially patterned hole depths L as indicated. The silicon holes had a diameter d of 2.4 μm, with a center-to-center spacing of 4 μm. (c) SEM images of PDMS micropost arrays with spatially patterned post heights and rigidity
To determine if cells would undergo durotaxis when encountering a discrete step change in substrate rigidity, migratory directions of single NIH 3T3 fibroblasts initially seeded at the rigidity boundary on the step-rigidity PDMS micropost array were analyzed. To ensure the cells to make a binary decision between two different rigidities, the step-rigidity PDMS micropost array was microcontact printed with 10 μm wide stripes of fibronectin (FN) perpendicular to the rigidity boundary, such that NIH 3T3 fibroblasts could only migrate away from the rigidity boundary. Patterning of FN stripe on the step-rigidity PDMS micropost array precluded gradual turning-based cellular migratory behaviors, but permitted reversal of migration direction (180°) and binary statistical analysis. The migration direction of NIH 3T3 fibroblasts attaching at the rigidity boundary was then recorded and analyzeds (Figs. 2a, 2b). The initial migration direction of NIH 3T3 fibroblasts attaching at the rigidity boundary on the step-rigidity PDMS micropost array was biased toward the more rigid posts when the difference between post rigidities was sufficiently large (Fig. 2c). NIH 3T3 fibroblasts landing on the 15/39 nN/μm rigidity boundary migrated towards the stiffer posts 55% of the time (n = 79), while cells landing on the 7.5/40 nN/μm boundary migrated towards the stiffer posts 69% of the time (n = 71). By representing the cell decision as a Bernoulli trial we determined that migration direction of NIH 3T3 fibroblasts was statistically biased towards more rigid posts on the 7.5/40 nN/μm step-rigidity PDMS micropost array (student’s t test, p = 0.000984) but was indistinguishable from random cell migration on the 15/39 nN/μm PDMS micropost array (student’s t test, p = 0.3143). For NIH 3T3 fibroblasts that initially seeded on the soft or only on the rigid PDMS posts (not near the rigidity boundary) and then migrated toward the rigidity boundary, we observed that cells did not change migration direction or speed as a function of crossing rigidity boundaries. That is, for cells already polarized in a particular direction and migrating, durotaxis was not observed across any of the rigidity boundaries studied in this work. This suggests that durotaxis in this setting proceeds when cells polarize during the initial stage of migration, while cells already migrating in one direction on the stripes, perhaps because they are polarized to a high degree, have difficulty repolarizing in response to the durotactic cue. Many components of the cell migration machinery polarize in migrating cells, including myosin27, FAs and associated traction forces. However, on substrates with step-rigidity whether traction forces polarized within cells remained unclear.
Substrate rigidity guided initial cell polarization and migratory direction. (a) Percentage of NIH/3T3 fibroblasts migrating towards more rigid posts on step-rigidity PDMS micropost arrays with different step changes of post rigidity (for 40/7.5 nN/μm: n = 71, for 39/15 nN/μm: n = 79). Single NIH/3T3 fibroblasts were initially seeded on the rigidity boundary on the step-rigidity PDMS micropost arrays. Asterisk denotes statistically significantly difference from cell random migration (dashed line) according to a Bernoulli trial (p = 0.000984). (b) Representative field of view of initial attachment of NIH/3T3 fibroblasts to step-rigidity PDMS micropost arrays. Dashed box indicates a region used to generate the kymograph in c. (c) Kymograph depicting a single NIH/3T3 fibroblast as it attached to the step-rigidity PDMS micropost array at the rigidity boundary, spread, polarized and migrated towards more rigid posts. Dashed line indicates the leading edge of the cell
To test whether traction forces polarized on step-rigidity substrates, we functionalized the step-rigidity PDMS micropost array with circular FN islands using microcontact printing (Figs. 3a, 4a). Single NIH 3T3 fibroblasts or human mesenchymal stem cells (hMSCs) were then plated and allowed to spread (Figs. 3b, 3c). Symmetric, circular cell spreading on these islands precluded asymmetric cell shape from guiding cell polarity.11,25 As reported previously for adherent cells seeded on PDMS micropost arrays of uniform rigidities, post deflection was greatest on soft posts and decreased as post rigidity increased (Figs. 3d, 4c).5 This post deflection corresponded to an average force per post that increased from softer posts to more rigid ones (Figs. 3d, 4d). Interestingly, for cells at the rigidity boundary (in contact with both rigid and soft posts) on the step-rigidity PDMS micropost array, the average post deflection and average force per post were as expected based on the local rigidity of the post the cell was in contact with (Figs. 3d, 4b–4d). For instance, cells deflected rigid posts to the same extent whether the entire cell was contact with only rigid posts or with both rigid and soft posts (Fig. 4c). Thus, cells appeared to be able to locally respond to the rigidity of each post, and such response seemed to be independent of other regions of the cell. Interestingly, this led to a polarization in traction force magnitudes across the cell, directly indicating that traction forces might be part of the polarization machinery in cells on step-rigidity substrates, and suggesting that traction force polarization might underlie durotaxis.
Subcellular quantifications of micropost deflections and traction forces for single human mesenchymal stem cells (hMSCs) seeded on step-rigidity PDMS micropost arrays patterned with circular FN islands of 100 μm diameter. (a) A low-magnification transmitted light image showing the step-rigidity PDMS micropost array. (b) Transmitted light image showing single hMSCs spreading on patterned circular adhesive islands of FN on the step-rigidity PDMS micropost array. Individual hMSCs were entirely on rigid and soft posts, or spanning across post rigidity boundaries. (c) Fluorescent image showing Δ9-DiI stained PDMS microposts beneath a single hMSC spanning the post rigidity boundary (Soft: 15 nN/μm, Medium: 21.5 nN/μm). (d) Colorimetric and vector maps showing subcellular distributions of micropost displacement (left column), corresponding traction forces (middle column) and corresponding traction force vectors (right column) of three representative single hMSCs that were entirely on soft (top; 15 nN/μm) or rigid (middle; 21.5 nN/μm) posts, or spanning the rigidity boundary (bottom; 21.5/15 nN/μm)
Cells that span the rigidity boundary on the step-rigidity PDMS micropost array developed different levels of local traction forces based on the local rigidity of the microposts. (a) Schematic representation of single-rigidity and step-rigidity micropost arrays patterned with circular FN islands. Soft: 15 nN/μm, Medium: 21.5 nN/μm, Rigid: 39 nN/μm. (b) Histogram of force per post for single hMSCs seeded on single-rigidity (left column) and step-rigidity micropost arrays (right column). (c&d) Average post deflection and force per post for single hMSCs seeded on single-rigidity (under the wholecell bar) and step-rigidity (under the subcellular bar) micropost arrays. In c & d, xx/yy + zz means that the portion of a cell on the xx side of a yy + zz rigidity boundary was analyzed. Error bars means ± standard deviations. P-values calculated using two-side unpaired t test. *P < 0.05
Methods
Fabrication of Micromolded PDMS Micropost Arrays
Silicon masters of the step-rigidity PDMS micropost arrays were defined and etched into a Si wafer using photolithography and DRIE in a cleanroom environment. A 5× reduction step-and-repeat projection stepper (Nikon NSR2005i9, Nikon Precision Inc., CA) was used for patterning the first layer of thin photoresist, Shipley SPR700-1.0 (Shipley Company, L.L.C., Marlborough, MA). Then DRIE was performed using C4F8 and SF6 etching chemistry with an inductively-coupled plasma (ICP) deep reactive-ion etcher (passivation: C4F8 (35 sccm), platen power (60 W); etch: SF6 (70 sccm), platen power (80 W); ICP Deep Trench Etching Systems, Surface Technology Systems, Newport, UK) to etch cylindrical hole arrays into the Si wafer. Depth of the cylindrical holes was controlled by monitoring DRIE etching time. After stripping photoresist with a Piranha solution (4:1 H2SO4/H2O2, 120 °C, 10 min), the Si wafer was spin coated with thick photoresist (AZ 4620, ~10 μm; Hoechst Celanese Corp., Somerville, NJ). Special care was taken to ensure complete filling of the thick photoresist in the cylindrical holes. The photoresist-coated wafer was then exposed using a mask contact aligner (Electronic Visions EV620, EV Group Inc., AZ) for patterning rigidity stripes on the cylindrical hole array. The following photoresist development step was monitored closely under an inspection microscope to ensure that exposed photoresist on the wafer was completely dissolved in the photoresist developer, and more importantly, photoresist filling the cylindrical holes were remained in the holes to protect the bottom of the holes during the following DRIE step. A second DRIE was performed to define the rigidity stripes on the cylindrical hole array (Supporting Fig. 1b). Because of protection of photoresist in the cylindrical holes, the Si wafer was etched away only from its top surface, and the bottoms of the holes were protected from etching. Finally, the thick photoresist was stripped with a Piranha solution, and the silicon master was silanized with (tridecafluoro-1,1,2,2,-tetrahydrooctyl)-1-trichlorosilane (United Chemical Technologies, Bristol, PA) for 4 h under vacuum to aid subsequent release of the negative PDMS (Sylgard 184, Dow-Corning, Midland, MI) template (see below) from the silicon master.
The step-rigidity PDMS micropost array was generated by replica-molding as previously described.33,34 Briefly, PDMS prepolymer was poured over the Si master, degassed under vacuum, cured at 110 °C for 20 h, and peeled off the master. When peeling induced collapse of PDMS microposts, we regenerated the PDMS micropost array by sonication in 100% ethanol for 30 s followed by dry-release with liquid CO2 using a critical point dryer (Samdri®-PVT-3D, Tousimis, Rockville, MD).
Cell Culture and Reagents
Human mesenchymal stem cells (hMSCs, Lonza, Walkersville, MD) were maintained in a growth medium (GM) consisting of low-glucose Dulbecco’s modified Eagle’s medium (DMEM, Invitrogen Corporation, Carlsbad, CA) supplemented with 10% fetal bovine serum (Atlanta Biological, Atlanta, GA), 0.3 mg/mL l-glutamine, 100 units/mL penicillin, and 100 μg/ml streptomycin. NIH 3T3 fibroblasts were maintained in DMEM supplemented with 10% bovine serum and 1% l-glutamine. Fresh 0.005% (vol/vol) trypsin-EDTA in PBS was used to re-suspend hMSCs and NIH 3T3 fibroblasts before seeding them as single cells on the PDMS micropost array.
Culture of Cells on PDMS Microposts
The PDMS micropost array was prepared for cell attachment using microcontact printing as previously described.34 Briefly, patterned PDMS stamps were generated via replica-molding. Stamps were immersed in 50 μg/mL fibronectin (BD Biosciences, San Jose, CA) in distilled (DI) water for 1 h, washed three times in DI water, and blown dry under nitrogen. Fibronectin coated stamps were placed in conformal contact with UV ozone-treated, surface-oxidized PDMS microposts (ozone cleaner; Jelight, Irvine, CA), to facilitate fibronectin transfer from the stamp to the PDMS micropost array. PDMS microposts were labeled by incubating with 5 μg/mL of Δ9-DiI (Invitrogen, Carlsbad, CA) in DI water at room temperature for 1 h. Following microcontact printing, protein adsorption to all PDMS surfaces not coated with fibronectin was prevented by incubating in 0.1–1% Pluronics F127 NF (BASF, Ludwigshafen, Germany) in DI water for 30 min at room temperature. hMSCs were seeded as single cells onto the PDMS micropost array and then allowed to spread overnight before other assays. NIH 3T3 fibroblasts were seeded as single cells on the PDMS micropost array and allowed to attach and spread for 15 min at which point time-lapse images were taken every 5 min for 16 h.
Quantification of Traction Forces
Quantitative analysis of subcellular level traction forces was performed as previously described.34 Briefly, the micromolded PDMS micropost arrays underlying live hMSCs were imaged with a 40×, 1.3 NA, oil immersion objective (EC Plan NEOFLUAR) and a CCD camera (AxioCam HRM, Carl Zeiss MicroImaging) attached to an inverted microscope (Zeiss Axiovert 200 M, Carl Zeiss MicroImaging, Thornwood, NY). The microscope was enclosed in a live cell incubator (In Vivo Scientific, Saint Louis, MO) to maintain the experimental environment at 37 °C and 5% CO2. For each cell, fluorescent images of the Δ9-DiI-stained PDMS microposts were acquired at two different focal planes. The top image was acquired at the focal plane passing through the top surfaces of the microposts, while the bottom one approximately 1 μm above the base of the microposts. These two images were analyzed with a custom-developed MATLAB program (Mathworks, Natick, MA) to calculate the traction forces. The centroids of the cross sections of the microposts in both the top and bottom images were determined by the localized thresholding algorithm (LT) described previously. The non-zero total net (vector sum of) forces arise as an error in this method because we do not constrain net force to be zero, in contrast to other traction methods that need this constraint to calculate forces. A small error in the deflection of each post arises from the experimental determination of position of each post tip, on the order of tens of nanometers using a Gaussian fit method, and this error propagates with the sum of forces on the order of square root of the number of posts measured under each cell.
Discussion
Rigidity sensing and durotaxis are mediated by actomyosin-mediated cellular traction forces and integrin-based FAs connecting adherent cells to extracellular environment.2,22 Thus, being able to measure traction forces while cells are simultaneously interacting with substrates of different rigidities is a fundamental step to understanding the mechanism underlying durotaxis. However, current methods of generating rigidity gradients in synthetic hydrogels make it technically difficult to ascertain where the cell is in the rigidity gradient or the precise values of local mechanical properties of the hydrogel where the cell is sitting.10,13,28 While using the PDMS micropost array can overcome such problem, to date the only PDMS micropost arrays with patterned rigidity profiles or gradients either have involved a complex microfabrication protocol without reporting traction forces,7 or have varied post rigidity by changing the post diameter,21,26 making it difficult to decouple the effects of FA size and organization from effects imparted by substrate rigidity alone.32
Here, we present the use of a step-rigidity micropost array substrate for studying the asymmetrical traction force distribution across the whole cell area during durotaxis. Our fabrication strategy to generate the step-rigidity PDMS micropost array does not involve complicated PECVD or CMP process, making this tool more easily accessible for studies of mechanobiology and rigidity sensing. Cells that span the rigidity boundary on the step-rigidity PDMS micropost array developed different levels of traction forces based on the local rigidity of the microposts (Figs. 3, 4). In our studies, both traction forces and post deflections depended on the micropost rigidity, suggesting that rigidity sensing in adherent cells might be a combination of both stress- and strain-mediated intracellular molecular events (Fig. 4). Patterned adherent cells bridging the rigidity boundary on the step-rigidity PDMS micropost array showed stronger traction forces on the rigid posts and weaker tractions on the softer ones, supporting that rigidity sensing might be primarily a local cellular event that could be integrated across the whole cell to guide coordinated cell behaviors such as directional cell migration.5,18
In this work we directly demonstrated for the first time that cells spanning a substrate of different rigidities could exert greater traction forces on the more rigid side of the substrate. Rigid substrates are known to elicit a myriad of effects on cell mechanics. For example, rigid substrates can provide greater resistance of substrate deformation to cellular traction forces causing the cell to increase its cytoskeletal tension and become mechanically more rigid.24,35 While it has long been appreciated that cells can migrate up a rigidity gradient, the ability to analyze this behavior has been technically challenging.13 The subcellular traction force gradient observed in this work supports the previous hypothesis that durotaxis may be a result of a feedback mechanism whereby higher traction forces on the more rigid portion of the substrate bias migration by pulling the cell in that direction.
Although our studies were performed on cells in isolation, we posit that in the presence of neighboring cells, a cell integrates polarizing mechanotransductive cues from not only the substrate but also neighboring cells. Cell–cell contact triggers polarization of cells, and physical force can be transmitted through cell–cell contacts. Experimental force application through a cell–cell adhesion receptor can stimulate polarization and directed migration.1,3,12,16,30 Recently, a study on collective migration in epithelial sheets has suggested that substrate rigidity enhances transmission of mechanical force signals transmitted through cell–cell adhesions.17 We suggest that cells integrate mechanical cues from both the substrate (in the form of rigidity) and neighboring cells (in the form of physical forces exerted at cell–cell contacts) as needed in a given migration context.
References
Desai, R. A., L. Gao, S. Raghavan, W. F. Liu, and C. S. Chen. Cell polarity triggered by cell-cell adhesion via E-cadherin. J. Cell Sci. 122:905–911, 2009.
Discher, D. E., D. J. Mooney, and P. W. Zandstra. Growth factors, matrices, and forces combine and control stem cells. Science 324:1673–1677, 2009.
Dupin, I., E. Camand, and S. Etienne-Manneville. Classical cadherins control nucleus and centrosome position and cell polarity. J. Cell Biol. 185:779–786, 2009.
Engler, A. J., S. Sen, H. L. Sweeney, and D. E. Discher. Matrix elasticity directs stem cell lineage specification. Cell 126:677–689, 2006.
Fu, J., et al. Mechanical regulation of cell function with geometrically modulated elastomeric substrates. Nat. Methods 7:733–736, 2010.
Ghassemi, S., et al. Fabrication of elastomer pillar arrays with modulated stiffness for cellular force measurements. J. Vac. Sci. Technol. B 26:2549–2553, 2008.
Ghassemi, S., et al. Fabrication of elastomer pillar arrays with modulated stiffness for cellular force measurements. J. Vac. Sci. Technol. B 26:2549–2553, 2008.
Gilbert, P. M., et al. Substrate elasticity regulates skeletal muscle stem cell self-renewal in culture. Science 329:1078–1081, 2010.
Hoffecker, I. T., W-h Guo, and Y-l Wang. Assessing the spatial resolution of cellular rigidity sensing using a micropatterned hydrogel-photoresist composite. Lab. Chip 11:3538–3544, 2011. doi:10.1039/c1lc20504h.
Isenberg, B. C., P. A. DiMilla, M. Walker, S. Kim, and J. Y. Wong. Vascular smooth muscle cell durotaxis depends on substrate stiffness gradient strength. Biophys. J. 97:1313–1322, 2009.
Jiang, X., D. A. Bruzewicz, A. P. Wong, M. Piel, and G. M. Whitesides. Directing cell migration with asymmetric micropatterns. Proc. Natl Acad. Sci. U.S.A. 102:975–978, 2005. doi:10.1073/pnas.0408954102.
Liu, Z., et al. Mechanical tugging force regulates the size of cell-cell junctions. Proc. Natl Acad. Sci. U.S.A. 107:9944–9949, 2010.
Lo, C. M., H. B. Wang, M. Dembo, and Y. L. Wang. Cell movement is guided by the rigidity of the substrate. Biophys. J. 79:144–152, 2000.
Lo, C. M., et al. Nonmuscle myosin IIb is involved in the guidance of fibroblast migration. Mol. Biol. Cell 15:982–989, 2004.
Lopez, J. I., I. Kang, W. K. You, D. M. McDonald, and V. M. Weaver. In situ force mapping of mammary gland transformation. Integr. Biol. (Camb) 3:910–921, 2011.
Maruthamuthu, V., B. Sabass, U. S. Schwarz, and M. L. Gardel. Cell-ECM traction force modulates endogenous tension at cell-cell contacts. Proc. Natl Acad. Sci. U.S.A. 108:4708–4713, 2011.
Ng, M. R., A. Besser, G. Danuser, and J. S. Brugge. Substrate stiffness regulates cadherin-dependent collective migration through myosin-II contractility. J. Cell Biol. 199:545–563, 2012.
Plotnikov, S. V., A. M. Pasapera, B. Sabass, and C. M. waterman. force fluctuations within focal adhesions mediate ECM-rigidity sensing to guide directed cell migration. Cell 151:1513–1527 (2012).
Raab, M., et al. Crawling from soft to stiff matrix polarizes the cytoskeleton and phosphoregulates myosin-II heavy chain. J. Cell Biol. 199:669–683, 2012.
Saez, A., A. Buguin, P. Silberzan, and B. Ladoux. Is the mechanical activity of epithelial cells controlled by deformations or forces? Biophys. J. 89:L52–L54, 2005.
Sochol, R. D., A. T. Higa, R. R. R. Janairo, S. Li, and L. W. Lin. Unidirectional mechanical cellular stimuli via micropost array gradients. Soft Matter 7:4606–4609, 2011.
Sun, Y., C. S. Chen, and J. Fu. Forcing stem cells to behave: a biophysical perspective of the cellular microenvironment. Annu. Rev. Biophys. 41:519–542, 2012. doi:10.1146/annurev-biophys-042910-155306.
Sun, Y., L. T. Jiang, R. Okada, and J. Fu. UV-modulated substrate rigidity for multiscale study of mechanoresponsive cellular behaviors. Langmuir 28:10789–10796.
Tee, S. Y., J. Fu, C. S. Chen, and P. A. Janmey. Cell shape and substrate rigidity both regulate cell stiffness. Biophys. J. 100:L25–L27, 2011.
Théry, M., et al. Anisotropy of cell adhesive microenvironment governs cell internal organization and orientation of polarity. Proc. Natl Acad. Sci. U.S.A. 103:19771–19776, 2006. doi:10.1073/pnas.0609267103.
Trichet, L., et al. Evidence of a large-scale mechanosensing mechanism for cellular adaptation to substrate stiffness. Proc. Natl Acad. Sci. U.S.A. 109:6933–6938, 2012.
Vicente-Manzanares, M., K. Newell-Litwa, A. I. Bachir, L. A. Whitmore, and A. R. Horwitz. Myosin IIA/IIB restrict adhesive and protrusive signaling to generate front–back polarity in migrating cells. J. Cell Biol. 193:381–396, 2011. doi:10.1083/jcb.201012159.
Vincent, L. G., Y. S. Choi, B. Alonso-Latorre, J. C. del Álamo, and A. J. Engler. Mesenchymal stem cell durotaxis depends on substrate stiffness gradient strength. Biotechnol. J. 8:472–484, 2013. doi:10.1002/biot.201200205.
Wang, H.-B., M. Dembo, S. K. Hanks, and Y-l Wang. Focal adhesion kinase is involved in mechanosensing during fibroblast migration. Proc. Natl Acad. Sci. U.S.A. 98:11295–11300, 2001. doi:10.1073/pnas.201201198.
Weber, G. F., M. A. Bjerke, and D. W. DeSimone. A mechanoresponsive cadherin-keratin complex directs polarized protrusive behavior and collective cell migration. Dev. Cell 22:104–115, 2012.
Wells, R. G. The role of matrix stiffness in regulating cell behavior. Hepatology 47:1394–1400, 2008.
Weng, S., and J. Fu. Synergistic regulation of cell function by matrix rigidity and adhesive pattern. Biomaterials 32:9584–9593, 2011.
Yang, M. T., J. Fu, Y.-K. Wang, R. A. Desai, and C. S. Chen. Assaying stem cell mechanobiology on microfabricated elastomeric substrates with geometrically modulated rigidity. Nat. Protoc. 6:187–213, 2011.
Yang, M. T., N. J. Sniadecki, and C. S. Chen. Geometric considerations of micro- to nanoscale elastomeric post arrays to study cellular traction forces. Adv. Mater. 19:3119–3123, 2007.
Yeung, T., et al. Effects of substrate stiffness on cell morphology, cytoskeletal structure, and adhesion. Cell Motil. Cytoskelet. 60:24–34, 2005.
Zhou, J., H. Y. Kim, and L. A. Davidson. Actomyosin stiffens the vertebrate embryo during crucial stages of elongation and neural tube closure. Development 136:677–688, 2009.
Zoldan, J., et al. The influence of scaffold elasticity on germ layer specification of human embryonic stem cells. Biomaterials 32:9612–9621, 2011.
Acknowledgments
We acknowledge financial support from the National Institutes of Health (EB00262, HL73305, and GM74048), the Penn Institute for Regenerative Medicine, the Nano/Bio Interface Center, and the Center for Musculoskeletal Disorders of the University of Pennsylvania. J. F. was partially funded by the American Heart Association Postdoctoral Fellowship, and R. D. was supported by a National Science Foundation Fellowship. We thank Pan Mao for assistance in scanning electron microscopy. The M.I.T. Microsystems Technology Laboratories is acknowledged for support in cleanroom fabrication.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Associate Editor Michael R. King oversaw the review of this article.
Mark T. Breckenridge and Ravi A. Desai contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supporting Figure 1. (a) Cross-sectional SEM images showing the negative Si master after the first DRIE step with a regular array of cylindrical, vertical holes of a uniform depth across the Si substrate (i). ii shows the negative Si master after the second photolithography where exposed photoresist on the Si wafer was completely dissolved but photoresist filling the cylindrical holes were remained in the holes. (b) Tilted top-view SEM images showing the negative step-rigidity Si master with different magnifications as indicated.
Rights and permissions
About this article
Cite this article
Breckenridge, M.T., Desai, R.A., Yang, M.T. et al. Substrates with Engineered Step Changes in Rigidity Induce Traction Force Polarity and Durotaxis. Cel. Mol. Bioeng. 7, 26–34 (2014). https://doi.org/10.1007/s12195-013-0307-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12195-013-0307-6