Abstract
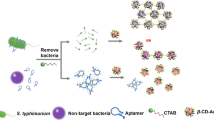
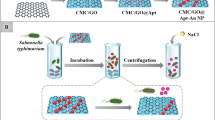
Salmonella typhimurium is one of the most common causes of food-associated disease. A colorimetric nanosensor was developed to detect S. typhimurium which was based on the gold nanoparticles’ (GNPs) color change effect. Fe3O4 magnetic nanoparticles (MNPs) and GNPs were synthesized separately. Two complementary sequences of the S. typhimurium target DNA were combined to Fe3O4 MNPs and GNPs to fabricate capture probes and signal probes. Fe3O4 MNPs could achieve the rapid separation and enrichment of target DNA. With the addition of S. typhimurium target DNA sequences, the sandwich-like structures were formed via the DNA hybridization recognition effect. The original good dispersion of GNPs was broken. GNPs showed different degrees of aggregation with different amount of S. typhimurium target DNA. The color changed from red, purple to blue which could be characterized by UV-Vis spectrophotometer. The absorbance spectra of GNPs red shifted constantly with the intensity ratio of A700/A521 changed regularly. There was a linear correlation between the ratio of A700/A521 and the amount of S. typhimurium target DNA. Thus, this was calculated for the basis of quantitative detection of S. typhimurium. This method is simple and rapid with high sensitivity and specificity which could be used to detect actual samples.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Salmonella typhimurium is a common anaerobic gram-negative bacterium which is widely distributed in the environment. Disease caused by S. typhimurium is one of the most common foodborne bacterial diseases around the world. It is also the most widespread zoonotic disease which is of important significance in public health (Arnold et al. 2011; Cheung et al. 2007; Magliulo et al. 2007). The traditional culture method for S. typhimurium detection is tedious and time-consuming which includes the sequential steps of pre-enrichment, selective enrichment, and selective differential plating (Patel et al. 2006). It is difficult to meet the needs of the current detection of foodborne bacteria. Therefore, researchers all over the world do a lot of researches for the rapid detection of S. typhimurium. With the gradually improving detection technology, a variety of detection methods have been reported such as immunofluorescence detection, immunodiffusion method, enzyme-linked immunoassay (ELISA), and latex agglutination test (Falkenhorst et al. 2013; Jain et al. 2012; Ma et al. 2014; Cucchi et al. 2013; Imre et al. 2013; Luo et al. 2014; Altinok et al. 2008). However, each method has its own drawbacks which limit the application.
Nanoparticles have attracted a wide application in biomedical fields, chemical fields, life sciences, and new materials owing to their specific characteristics such as surface effect, small size effect, and quantum size effect. Among which, gold nanoparticles (GNPs) have been extensively concerned for their simple preparation and many excellent optical properties (Currivan et al. 2013). Chloroauric acid is usually used as the raw material with the addition of a reducing agent to prepare GNPs. Various reducing agents, capping agents, and experimental conditions will result in the formation of GNPs with different sizes, morphologies, and aggregation degrees (Dai et al. 2002; Ojea-Jime et al. 2010). When the distance of GNPs gets closer to each other, the absorbance spectra red shifted and the color of the solution will be changed (Storhoff et al. 2004). Besides, GNPs also exhibit good biological compatibility with the interaction of biomacromolecules (Afonso et al. 2013; Feng et al. 2013; Wang and Zhang 2013). In addition, GNPs and sulfhydryl group have a strong bond and force. Therefore, biomolecules with thiol labeling could be combined to the surface of GNPs through Au–S bond for the preparation of nanoprobes and application in the detection of biological systems (Liu et al. 2013; Preechakasedkit et al. 2012).
With the increasing development of nanomaterials, magnetic nanomaterials (MNPs) have also attracted extensive attention due to their unique and super paramagnetic properties (Mascaraque et al. 2013; Tan et al. 2014; Wang et al. 2013a; Shokrollahi et al. 2014; Yan et al. 2014). MNPs obtain magnetism and are vulnerable to external magnetic field control. And the small size obtains with larger specific surface area. MNPs have been widely used in biomedical and life science applications with surface modification of various bioactive molecules (such as protein, antigen, and DNA sequence) through different functional groups (such as amino, carboxyl, hydroxyl, and mercapto groups) (Samoilova et al. 2014; Liu et al. 2014; Ma et al. 2009). MNPs could be easily separated from the other substances under the action of magnetic field. After the removal of the external magnetic field, MNPs can be dispersed in solution again. The unique properties of MNPs have important applications in the separation and rapid detection of biological molecules. Combined with the advantages of GNPs and MNPs, there are several related methods for the detection of proteins, bacteria, and so on (Yin et al. 2014; Wang et al. 2013b; Weidemaier et al. 2015).
In this study, a new technique for the colorimetric detection of S. typhimurium is discussed. The specific oligonucleotide DNA sequence of S. typhimurium was used as the target. The two complementary sequences of target DNA were combined to Fe3O4 MNPs and GNPs, respectively, to fabricate capture probes and signal probes. Based on the DNA hybridization recognition technology, sandwich-like structures could be formed by target DNA, capture probes, and signal probes. Based on the optical distance-dependent properties of GNPs, the increased concentration of the target DNA could result in the changes of polymerization degree for GNPs. The absorbance spectra red shifted constantly. The color changed from red, purple to blue which could be observed by naked eyes. By means of the characterization of UV-Vis spectrophotometer, a rapid and sensitive method was developed for the detection of gram-negative bacterium S. typhimurium specific DNA sequences. It is expected to be applied in other biofunctionalized DNA sequence detection.
Materials and Methods
Materials
Sodium acetate trihydrate (CH3COONa), ferric chloride (FeCl3), chloroauric acid tetrahydrate (HAuCl4·4H2O), trisodium citrate (Na3C6H5O7), sodium chloride (NaCl), potassium chloride (KCl), disodium hydrogen phosphate (Na2HPO4·12H2O), potassium biphosphate (KH2PO4), potassium carbonate (K2CO3), ethylene glycol (C2H6O2), 1,6-hexanediamine (C6H16N2), ethanol anhydrous (CH3CH2OH), glutaraldehyde (C5H8O2), and sodium dodecyl sulfate (SDS, C12H25NaO4S) were purchased from the Sinopharm Chemical Reagent Co., Ltd. (Shanghai, China). Streptavidin was purchased from Sigma-Aldrich. S. typhimurium target DNA sequence and the complementary strand were synthesized by Shanghai Sangon Biological Science and Technology Company (Shanghai, China). The capture DNA sequence 1 was 5′-GAG CGT GCC TTA CCG ACG ATA-biotin-3′; the signal DNA sequence 2 was 5′-SH-ATA TCC ACG CAG GAA ATA ACA GGA CTT-3′; the target DNA sequence 3 was 5′-TAT CGT CGG TAA GGC ACG CTC AAT TGT CGT TAA AGT CCT GTT ATT TCC TGC GTG GAT AT-3′. The ultrapure water used in the experiments was prepared using a Millipore Direct-Q® 3 system (Merck Millipore, MA, USA) and had a resistivity of 18.2 MΩ cm.
Preparation of Streptavidin-Functionalized Fe3O4 MNPs
The amino-functionalized Fe3O4 MNPs were prepared according to the literature (Chen and Gao 2007) with some modifications. A solution of 6.5 g 1,6-hexanediamine, 2.0 g anhydrous sodium acetate, 1.0 g FeCl3, and 30 mL glycol was stirred vigorously at 50 °C until the formation of a transparent solution. The mixed solution was transferred to teflon-lined autoclave and reacted at 195 °C for 6 h. The lower black liquid was rinsed with water and ethanol using magnetic force for separation and dried at 50 °C to get the amino-functionalized Fe3O4 MNPs and stored at 4 °C for further use.
Two milligram amino-functionalized Fe3O4 MNP powder was added to 2 mL glutaraldehyde (5%) and reacted at 37 °C for 2 h. The uncombined glutaraldehyde was removed by rinsing with PBS for at least five times using magnetic force. Then, the Fe3O4 MNPs was added with 2 mL PBS and 250 μL streptavidin (1 mg/mL) and reacted at 4 °C for 12 h. Similarly, the uncombined streptavidin was removed by rinsing with PBS using magnetic force. The precipitation (streptavidin-functionalized Fe3O4 MNPs) was collected and used for further characterization.
Preparation of GNPs
GNPs were prepared with some modifications as described by Grabar et al. (1996). First, 4.2 mL HAuCl4·4H2O (1%, w/w) and 95.8 mL ultrapure water were added to a flask with three necks. The mixture was heated to boil until 10 min under uniform magnetic stirring with oil bath. Then, 10 mL sodium citrate (1%, w/w) was rapidly injected and reacted for another 15 min. The obtained wine red solution was GNPs. The resulting GNPs were purified by three times of centrifugation (10,000 rpm, 25 min) and were redispersed in 40 mL of ultrapure water. The GNPs were stored at 4 °C for further use and characterization.
Preparation of Signal Probes and Capture Probes
Twenty microliter thiolated signal DNA 2 with gradient dilutions was added to 0.5 mL GNPs and reacted at 37 °C for 24 h. Five milligram SDS was added and shaken for a while. Then, a certain amount of NaCl was added and aged for another 24 h at 37 °C. The mixture was purified by three times of centrifugation (10,000 rpm, 10 min) to get thiolated signal DNA 2-modified GNPs (signal probes).
Twenty microliter biotin-labeled capture DNA 1 with gradient dilutions was dissolved in 2 mL PBS. Then, 1 mg of streptavidin-functionalized Fe3O4 MNPs was added and reacted at 37 °C for 12 h. The resulting solution was purified using magnetic force, and the precipitation was collected as biotin-labeled capture DNA 1-modified MNPs (capture probes).
Colorimetric Detection Procedure
Twenty microliter target DNA sequence 3 with gradient dilutions was dissolved in 2 mL PBS, and 1 mg of capture probes was added and reacted at 37 °C for 12 h. The mixture was purified using magnetic force, and the precipitation was collected. Then, 2 mL of signal probes was added and reacted at 37 °C for another 12 h. The resulting solution was characterized by UV-Vis spectrophotometer.
Activation, Cultivation, and Plate Counting of S. typhimurium
The S. typhimurium were inoculated into the LB liquid medium and cultivated under shaking at 37 °C for 12-h enrichment. The enriched bacterial was centrifuged at 5800 r/min for 10 min (25 °C), and the supernatant was discarded. The precipitate was washed with PBS (0.1 M, pH 7.4) three times and resuspended in PBS. The absorbance was measured at 600 nm. The collection was centrifuged and diluted repeatedly to optical density (OD) value 0.12. This preparation was used as the original S. typhimurium sample. Then, the original bacterium was diluted to eight concentrations along a gradient from 10−1 to 10−8 using physiological saline. One hundred microliter of 10−5, 10−6, and 10−7 S. typhimurium was coated on the solid LB agar plates. Each plate was coated with three parallel boards. After cultivation at 37 °C for 12 h, the colonies were counted for the calculation of the S. typhimurium sample (cfu/mL).
The Extraction of S. typhimurium Target DNA and Detection
The DNA extraction kit was used for the target DNA extraction of different dilutions of S. typhimurium. Then, after a water bath at 95 °C for 5 min and 0 °C for 5 min, the single-stranded DNA as the target DNA was produced. This was used to do the colorimetric detection, and the results were compared to the plate counting method.
Recovery Experiments for Milk Sample
In this experiment, the commercial milk was used as realistic samples for recovery experiments in the detection of S. typhimurium. Gradient dilutions of S. typhimurium were added to the milk sample. Then, the colorimetric detection method was conducted to calculate the detectable amount of S. typhimurium. The results were compared with the traditional plate counting method, and the recovery rate was calculated.
Results and Discussion
Principle for Colorimetric Detection
The colorimetric detection process is shown in Fig. 1. Biotinylated capture DNA 1 was modified at the surface of streptavidin-functionalized Fe3O4 MNPs. Thiolated signal DNA 2 was decorated to GNPs using covalent attraction between thiol group and GNPs. When the target DNA sequences were added, the unique sandwich structure was formed on account of the base complementation pairing rule. The color of GNPs has a characteristic of light-distance dependence. When the distances between GNPs get closer, the absorbance spectra red shift and the color of GNPs is changed. With the addition of gradient dilutions of S. typhimurium target DNA sequences, GNPs show different aggregation states which reflect in color changes. This could be characterized by UV-Vis spectrophotometer.
Preparation of Fe3O4 MNPs and Surface Modification
Figure 2 shows the TEM image of the amino-functionalized Fe3O4 MNPs and its infrared (IR) spectrum. The particle size was about 40 nm. The strong IR band at 3413.3 and 1627.6 cm−1 from the amino-functionalized nanoparticles matched well with those from free 1,6-hexadiamine, indicating the existence of the free –NH2 group on the amino-functionalized nanoparticles.
Streptavidin was used for the modification of Fe3O4 MNPs for further DNA sequence connection. Results are shown in Fig. 3. In order to get the optimal streptavidin concentration, different volumes (50, 100, 150, 200, 250, 300, 350, 400 μL) of streptavidin (1 mg/mL) were added to react with 2 mL Fe3O4 MNPs (1 mg/mL). By calculating the differences of absorbance intensity before and after modification, we can see that the differences almost achieve maximum when the volume was 250 μL. So, the optimal streptavidin (1 mg/mL) addition amount was 250 μL.
Preparation of GNPs
Figure 4 is the TEM image and UV-visible absorption spectrum of GNPs. It shows that the prepared GNPs have an absorption peak at approximately 521 nm and the size is ~15 nm.
Optimization of Experimental Conditions
In this study, thiol group-labeled signal DNA 2 was connected to the GNPs via the strong affinity of the thiol group and gold to construct signal probes. When the amount of DNA sequence increased, some basic group of oligonucleotide could adsorb to the surface of GNPs which might influence the formation of Au–S chemical bond. The addition of electrolyte NaCl solution could do the desorption treatment of basic group. It is reported by Demers et al. (2000) that saline ions could weaken the electrostatic interaction of DNA sequences adsorbed on the surface of GNPs. Then, the DNA sequence may stand up on the GNP surface which increases the connection density. Therefore, the fixing amount of signal DNA 2 increases. However, the excessive addition of saline ions may result in the aggregation of GNPs accompanied by the change of color. This will influence the colorimetric detection result. So, the optimal concentration of NaCl is a key factor for this experiment.
When the concentration of NaCl increased from 50 to 200 mM, the absorption intensity decreased slightly and the color of GNPs remained wine red. While the concentration further increased to 250 mM, the color of GNPs changed to purple which could be observed by naked eyes. And the absorption intensity decreased obviously along with the red shift of the absorption peak position which is depicted in Fig. 5a. This indicated that the GNPs were aggregated. Figure 5b shows the binding rate of DNA sequences to GNPs according to the different amount addition of NaCl. The binding rate was calculated by the absorption intensity of DNA at 260 nm measured before and after modification to GNPs. With the amount of NaCl increased from 50 to 200 mM, the binding rate also increased significantly. After that, the binding rate changed slowly. Combined with the results of Fig. 5a, b, the optimal concentration of NaCl was 200 mM.
a UV-visible absorption spectra of GNPs with the addition of different concentrations of NaCl solution (0.05, 0.10, 0.15, 0.20, 0.25 M (1–5)). The inset is the photograph of the corresponding GNP solution. b The binding rate of signal DNA linked to GNPs with the treatment of different concentrations of NaCl
Analytical Performance
A series of concentrations of target DNA sequence 3 were detected under the optimal experimental conditions. Results are shown in Fig. 6. With the increased amount of DNA 3, the absorbance spectra red shifted constantly. The absorbance intensity at 521 nm gradually decreased, while the absorbance intensity at 700 nm increased (Fig. 6a). The ratio of A700/A521 was calculated as the basis for quantitative analysis. As depicted in Fig. 6b, there was a good linear correlation between the ratio of A700/A521 and the amount of DNA 3 ranged from 1 to 1500 pM. The linear correlation equation obtained was y = 0.007x + 0.2535 (R 2 = 0.9983). And the statistical analysis revealed that the detection limit of target DNA 3 was 0.8 pM. The detection limit is based on the calculation formula D = 3N/S (N is the standard deviation of blank sample signal. S is the slope of standard curve).
Specificity
Some other DNA sequences were used to do the specificity detection. These included the single base mutation chain, five-base mutation chain, random control chain, Staphylococcus aureus target DNA sequences, Vibrio parahemolyticus target DNA sequences, Bacillus cereus target DNA sequences, and Shigella dysenteriae target DNA sequences. The concentration of the DNA sequences was maintained at 1 nM. Experimental results shown in Fig. 7 clearly showed that the ratio of A700/A521 for the other DNA sequences was much lower than that of the S. typhimurium target DNA sequence.
Specificity result for the detection of a S. typhimurium target DNA, b single base mutation chain, c five-base mutation chain, d random control chain, e S. aureus target DNA sequences, f V. parahemolyticus target DNA sequences, g B. cereus target DNA sequences, and h S. dysenteriae target DNA sequences
The Extraction and Detection of S. typhimurium Target DNA
In addition to the known sequence of S. typhimurium target DNA, the DNA extraction kit was used for the target DNA extraction of S. typhimurium. The obtained sequence was used as the target DNA to do the colorimetric detection. Results were compared with the plate counting method. As depicted in Fig. 8, the ratio of A700/A521 showed good linear correlation to the amount of S. typhimurium obtained by plate counting method ranging from 30 to 8600 cfu/mL. The linear equation is y = 0.0001x + 0.3083 (R 2 = 0.9962). And the statistical analysis revealed that the detection limit of S. typhimurium was 23 cfu/mL. The detection limit is based on the calculation formula D = 3N/S (N is the standard deviation of blank sample signal. S is the slope of standard curve).
Compared with the traditional culture method for S. typhimurium, this method is fast and free from pre-enrichment treatment. This method combined with the advantages of both MNPs and GNPs. The MNPs have the fast separation and enrichment properties. And the optical properties of signal GNPs will be gradually changed according to the aggregation extent. The color changes could be used as a qualitative and quantitative detection basis which is simple for the analysis of pathogenic bacterium.
Milk Sample Detection
The utility of the colorimetric detection for S. typhimurium was examined using milk sample obtained from a supermarket. The sample was tested using the new method and the classical plate counting methods. The analytical results are shown in Table 1. The results obtained using the colorimetric detection method were similar to those obtained using the plate counting method. The recoveries were between 90.7 and 102.2%, indicating good accuracy of the proposed test for S. typhimurium detection. There is no complicated step for the pretreatment of milk sample. This colorimetric method is expected for the field test of S. typhimurium.
Conclusion
The colorimetric detection of S. typhimurium using Fe3O4 MNPs and GNPs was investigated. Fe3O4 MNPs were immobilized with capture DNA to fabricate capture probe. GNPs were modified with signal DNA to fabricate signal probe. The capture probe and signal probe could be connected to S. typhimurium target DNA via the DNA base complementary matching principle. MNPs possess good magnetic characteristics, which make it easier to separate the product from the mixture during the experiment. The quantitative detection of S. typhimurium target DNA was achieved by the characterization of UV-Vis spectra of GNPs. In addition, this developed method was successfully used to analyze milk samples, and there was no significant difference between a classical plate counting method and the developed method. So, it has the potential for wide use in the detection of other foodborne pathogenic bacteria in food samples.
References
Afonso AS, Perez-Lopez B, Faria RC, Mattoso LHC, Hernández-Herrero M, Roig-Sagués AX, Maltez-da CM, Merkoci A (2013) Electrochemical detection of Salmonella using gold nanoparticles. Biosens Bioelectron 40(1):121–126
Altinok I, Capkin E, Kayis S (2008) Development of multiplex PCR assay for simultaneous detection of five bacterial fish pathogens. Vet Microbiol 131:332–338
Arnold ME, Carrique-Mas JJ, McLaren I, Davies RH (2011) A comparison of pooled and individual bird sampling for detection of Salmonella in commercial egg laying flocks. Prev Vet Med 99:176–184
Chen ZT, Gao L (2007) Synthesis and magnetic properties of CoFe2O4 nanoparticles by using PEG as surfactant additive. Mater Sci Eng B-Solid State Mater Adv Technol 141:82–86
Cheung PY, Kwok KK, Kam KM (2007) Application of BAX system, tecra unique Salmonella test, and a conventional culture method for the detection of Salmonella in ready-to-eat and raw foods. J Appl Microbiol 103:219–227
Cucchi MEC, Marcoppido G, Trangoni MD, Cravero SL (2013) Detection of fiber-digesting bacteria in the forestomach contents of llamas (Lama glama) by PCR. Rev Argent Microbiol 45(3):147–149
Currivan S, Connolly D, Paull B (2013) Production of polymer monolithic capillary columns with integrated gold nano-particle modified segments for on-capillary extraction. Microchem J 111:32–39
Dai XH, Tan YW, Xu J (2002) Formation of gold nanoparticles in the presence of o-anisidine and the dependence of the structure of poly (o-anisidine) on synthetic conditions. Langmuir 18(23):9010–9016
Demers LM, Mirkin CA, Mucic RC, Reynolds RA, Letsinger RL, Elghanian R, Viswanadham G (2000) A fluorescence-based method for determing the surface coverage and hybridization efficiency of thiol-capped oligonucleotides bound to gold thin flims and nanoparticles. Anal Chem 72:5535–5541
Falkenhorst G, Ceper TH, Strid MA, Mølbak K, Krogfelt KA (2013) Serological follow-up after non-typhoid Salmonella infection in humans using a mixed lipopolysaccharide ELISA. Int J Med Microbiol 303(8):533–538
Feng R, Zhang Y, Li H, Wu D, Xin XD, Zhang S, Yu HQ, Wei Q, Du B (2013) Ultrasensitive electrochemical immunosensor for zeranol detection based on signal amplification strategy of nanoporous gold films and nano-montmorillonite as labels. Anal Chim Acta 758(3):72–79
Grabar KC, Smith PC, Musick MD, Davis JA, Walter DG, Jackson MA (1996) Natan, kinetic control of interparticle spacing in Au colloid-based surfaces: rational nanometer-scale architecture. J Am Chem Soc 118(5):1148–1153
Imre A, Bukovinszki A, Margaret A (2013) Gene expression analysis of Salmonella enterica SPI in macrophages indicates differences between serovars that induce systemic disease from those normally causing enteritis. Vet Microbiol 167(3–4):675–679
Jain S, Chattopadhyay S, Jackeray R, Abid CKVZ, Kohli GS, Sinqh H (2012) Highly sensitive detection of Salmonella typhi using surface aminated polycarbonate membrane enhanced-ELISA. Biosens Bioelectron 31(1):37–43
Liu CC, Yeung CY, Chen PH, Yeh MK, Hou SY (2013) Salmonella detection using 16S ribosomal DNA/RNA probe-gold nanoparticles and lateral flow immunoassay. Food Chem 141(3):2526–2532
Liu M, Wang Z, Zong S, Chen H, Zhu D, Wu L, Hu G, Cui Y (2014) SERS detection and removal of mercury(II)/silver(I) using oligonucleotide-functionalized core/shell magnetic silica sphere@Au nanoparticles. ACS App Mater Inter 6(10):7371–7379
Luo R, Li YH, Lin XJ (2014) A colorimetric assay method for invA gene of Salmonella using DNAzyme probe self-assembled gold nanoparticles as single tag. Sens Actuator B-Chem 198(31):87–93
Ma LL, Feldman MD, Larson TA, Tam JM, Paranjape AS, Cheruku KK, Tam JO, Ingram DR, Paramita V, Villard JW, Jenkins JT, Wang TY, Clarke GD, Asmis R, Sokolov K, Chandrasekar B, Milner TE, Johnston KP (2009) Small multifunctional nanoclusters (nanoroses) for targeted cellular imaging and therapy. ACS Nano 3(9):2686–2696
Ma K, Deng Y, Bai Y, Xu DX, Chen EN, Wu HJ, Li BM, Gao LJ (2014) Rapid and simultaneous detection of Salmonella, Shigella, and Staphylococcus aureus in fresh pork using a multiplex real-time PCR assay based on immunomagnetic separation. Food Control 42:87–93
Magliulo M, Simoni P, Guardigli M, Michelini E, Luciani M, Lelli R, Roda A (2007) A rapid multiplexed chemiluminescent immunoassay for the detection of Escherichia coli O157:H7, Yersinia enterocolitica, Salmonella typhimurium, and Listeria monocytogenes pathogen bacteria. J Agric Food Chem 55:4933–4939
Mascaraque A, Rodriguez de la Fuente O, Gonzalez-Barrio MA (2013) From surfaces to magnetic properties: special section dedicated to Juan Rojo. J Phys-Condens Matter 25(48):480301
Ojea-Jime I, Romero FM, Basts NG, Puntes V (2010) Small gold nanoparticles synthesized with sodium citrate and heavy water: insights into the reaction mechanism. J Phys Chem C 114:1800–1804
Patel JR, Bhagwat AA, Sanglay GC, Solomon MB (2006) Rapid detection of Salmonella from hydrodynamic pressure-treated poultry using molecular beacon real-time PCR. Food Microbiol 23:39–46
Preechakasedkit P, Pinwattana K, Dungchai W, Siangproh W, Chaicumpa W, Tongtawe P, Chailapakul O (2012) Development of a one-step immunochromatographic strip test using gold nanoparticles for the rapid detection of Salmonella typhi in human serum. Biosens Bioelectron 31(1):562–566
Samoilova N, Tikhonov V, Krayukhina M, Yamskov L (2014) Interpolyelectrolyte complexes of maleic acid copolymers and chitosan for stabilization and functionalization of magnetite nano- and microparticles. J Appl Polym Sci 131(1):39663
Shokrollahi H, Khorramdin A, Isapour G (2014) Magnetic resonance imaging by using nano-magnetic particles. J Magn Magn Mater 369:176–183
Storhoff JJ, Lucas A, Garimella V, Patno T, Muller UR (2004) Homogeneous detection of unamplified genomic DNA sequences based on colorimetric scatter of gold nanoparticle probes. Nat Biotechnol 22:883–887
Tan C, Gao N, Deng Y, Deng J, Zhou SQ, Lia J, Xin XY (2014) Radical induced degradation of acetaminophen with Fe3O4 magnetic nanoparticles as heterogeneous activator of peroxymonosulfate. J Hazard Mater 276:452–460
Wang XL, Zhang XL (2013) Electrochemical co-reduction synthesis of graphene/nano-gold composites and its application to electrochemical glucose biosensor. Electrochim Acta 112(1):774–782
Wang BX, Yin YC, Liu CJ, Yu SS, Chen KZ (2013a) Synthesis of flower-like BaTiO3/Fe3O4 hierarchically structured particles and their electrorheological and magnetic properties. Dalton Trans 42(27):10042–10055
Wang CH, Chang CJ, Wu JJ, Lee GB (2013b) An integrated microfluidic device utilizing vancomycin conjugated magnetic beads and nanogold-labeled specific nucleotide probes for rapid pathogen diagnosis. Nanomed-Nanotechnol Biol Med 10(4):809–818
Weidemaier K, Carruthers E, Curry A, Kuroda M, Fallows E, Thomas J, Sherman D, Muldoon M (2015) Real-time pathogen monitoring during enrichment: a novel nanotechnology-based approach to food safety testing. Int J Food Microbiol 198:19–27
Yan J, Liu Y, Wang Y, Xu XW, Lu Y, Pan YJ, Guo FF, Shi DL (2014) Effect of physiochemical property of Fe3O4 particle on magnetic lateral flow immunochromatographic assay. Sens Actuator B-Chem 197:129–136
Yin HQ, Jia MX, Shi LJ, Liu J, Wang R, Lv MM, Ma YY, Zhao X, Zhang JG (2014) Evaluation of a novel ultra-sensitive nanoparticle probe-based assay for ricin detection. J Immunotoxicol 11(3):291–295
Acknowledgments
This work was supported by the NSFC (31401665, 21375049), National S&T Support Program of China (2015BAD17B02), National Natural Science Foundation of Jiangsu Province (BK20161479), the Educational Commission of Jiangsu Province (16KJB150043), Qing Lan Project of Jiangsu Province, and the program of “Collaborative Innovation Center of Food Safety and Quality Control in Jiangsu Province.”
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Xiaoyuan Ma declares that she has no conflict of interest. Liangjing Song declares that she has no conflict of interest. Yu Xia declares that he has no conflict of interest. Caiyun Jiang declares that she has no conflict of interest. Zhouping Wang declares that he has no conflict of interest.
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed Consent
Not applicable.
Rights and permissions
About this article
Cite this article
Ma, X., Song, L., Xia, Y. et al. A Novel Colorimetric Detection of S. typhimurium Based on Fe3O4 Magnetic Nanoparticles and Gold Nanoparticles. Food Anal. Methods 10, 2735–2742 (2017). https://doi.org/10.1007/s12161-017-0819-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12161-017-0819-0