Abstract
The development of new lignocellulosic enzyme cocktails to digest raw material for the successful production of biofuels still represents great economic and scientific challenges. The aim of this study was the characterization of an aerobic consortium derived from the gut of the cockroach Nauphoeta cinerea (NaLC) under morphological, chemical, biochemical, and genetic views. Initially, the assembly of the consortium started by dissection and incubation of hindgut of N cinerea with medium containing sugarcane bagasse (BED) as carbon source. After 1 week of cultivation, the NaLC degraded about 55% of lignocellulosic material (mainly cellulose) and also promoted a significant decreased in the BED length. The biochemical analysis of consortium supernatants showed activities against Avicel, β-glucan, xylan, and CMC. The metagenomics analysis of 16S rDNA from NaLC showed 22 bacterial genomes with a predominance of Bacteroidetes (50.6%) and Proteobacteria (47.7%) with Flavobacterium spp. and Sphingomonas spp., respectively. The shotgun metagenomic analysis revealed the presence of several CAZymes, such as cellulases, hemicellulases, carbohydrate esterases, and enzymes with auxiliary activities. During the shotgun metagenomic analysis, various CAZymes organized as polysaccharide utilization loci (PUL) from Niabela sp. bacteria in NaLC were identified. It was also observed a new locus named here as LUL (lignin utilization locus) containing many genes as oxidases, dehydrogenases, peroxidases, and others that could be involved in lignin metabolization. So, this study described under different approaches an unprecedented aerobic consortium could be a useful resource of novel genes and enzymes involved in degradation of sugarcane bagasse.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The global demand for renewable energy sources such as wind, solar, and lignocellulosic biofuels to replace fossil fuels is very crescent [1]. Among the greatest challenges for the utilization of lignocellulosic biomass as a biofuel resource is the production of efficient and inexpensive enzymes that act on cellulose (around 35–50%), hemicellulose (around 20–35%), and lignin polymers (around 10–20%) of plant cell wall [2, 3]. For this reason, microorganisms from animal (including insect) guts that feed on lignocellulose could represent an interesting source of new enzymes specialized in lignocellulose breakdown. The gut of termites is considered the most efficient mini bioreactor in nature that promote the lignocellulose degradation by enzymes produced by termites and their intestinal microorganisms (more than 90% cellulose degradation). Considering the great phylogenetical similarities between termites and cockroaches [4, 5], these insects could be an interesting source of new enzymes involved in the digestion of lignocellulosic compounds. Cockroaches, like termites, digest lignocellulosic compounds through the cooperation of enzymes derived both from the cockroach itself, and its gut microbiota [3, 4]. Wharton & Wharton [5] and Elpidina et al. [6] reported enzymatic activities against proteins, starch, cellulose, and cellobiose in the midgut of the omnivorous cockroach Nauphoeta cinerea. The degradation of the lignocellulolytic material inside the gut the cockroaches is mainly made by microbial consortia [2].
Microbial consortia (natural or artificial) show several potential advantages for valuable product synthesis, lignocellulose utilization, bioremediation, and other biotechnological products in comparison to pure cultures. The synergies of different metabolic pathways of microorganisms from these consortia can result in more efficient degradation of substrates with a major spectrum of useful products for different sectors of biotechnology [7, 8]. Several research groups have assembled microbial consortia—from different sources, such as soil, intestine, rumen, and feces [9, 10] that are able to degrade lignocellulosic biomass. Thus, the development of microbial consortia in specific biomass represents a rich arsenal of genes/enzymes to supplement existing enzyme cocktails, as well as the transformation of fungus with an enzyme of metagenomic origin [9,10,11].
Metagenomic approaches allow to identify several new carbohydrate-active enzymes (CAZymes) originating from these different consortia. The organization of several CAZymes in a specific arrangement named PUL (polysaccharide utilization loci) has also been described, mainly in the Bacteroidetes genome [12]. The Bacteroidetes isolates from P. americana digestive tracts show the presence of several PUL and non-PUL-associated CAZyme-coding genes [12]. These genes could target starch, pectin, and/or hemicellulose. However, there has been no specific locus involved in lignin processing described so far.
In this study, an aerobic, lignocellulolytic microbial consortium derived from the intestinal microbiota of the cockroach Nauphoeta cinerea (NaLC) was developed, with the aim of identifying potential lignocellulolytic microorganisms and novel CAZymes. Using pretreated sugarcane bagasse as the exclusive carbon source, it was observed changes in the morphological and compositional aspects of the sugarcane bagasse after cultivation with the microbial consortium. The microbial community structure was investigated by high throughput sequencing of the 16S rRNA gene for prokaryotic diversity analysis and the Internal Transcribed Spacer 2 (ITS2) for fungal analysis. Furthermore, shotgun metagenomic and binning analyses were performed, revealing several CAZymes and microorganisms probably involved in the deconstruction of the pretreated sugarcane bagasse.
Materials and Methods
Enrichment of Nauphoeta Lignocellulolytic Consortium
The N. cinerea cockroach colony was maintained in glass aquariums exposed to natural temperature and photo regime conditions. The insects were fed exclusively with sugarcane bagasse pretreated by steam explosion for at least 14 days.
The sugarcane bagasse used in this study was pretreated by steam explosion followed by an alkaline delignification reaction [11]. The bagasse was ground in a Willey type mill (Pulverisette® 19—Fritsch) to a particle size of 2 mm, named here as BED. Bushnell Haas Broth commercial medium (BHB) [9] was supplemented with 0.5% (w/v) sugarcane bagasse as the only carbon source, kindly provided by the Brazilian Biorenewables National Laboratory (LNBR).
The initial inoculum to establish the microbial consortium consisted of a mixture of three intestines of the cockroach Nauphoeta cinerea in 100 mL of BHB supplemented with BED 0.5% (w/v). The culture was grown in Erlenmeyer flasks (250 mL), stirred at 120 RPM at 30 °C. The NaLC was maintained by continuous subculture inoculating 1 mL of supernatant transferred every 7 days into fresh BHB enrichment medium for at least 6 months, after that period the experiments were performed. The control group was generated in culture media incubated in the absence of NaLC.
Determination of BED Consumption by NaLC
After the 1st, 3rd, 5th, and 7th days of incubation, the culture medium was filtered through filter paper previously weighed. This material was placed in an oven at 50 °C for 48 h to eliminate water from the filter. Subsequently, the material was weighed, and the dry weight of the remaining sugarcane bagasse was compared to the control group. The control group consisted of the BED incubated in the absence of NaLC.
Characterization of BED Degradation by NaLC
The chemical composition of the BED was performed by collecting 7th day growth samples. The control consisted of an incubation of non-inoculated medium under the same conditions as experimental groups. All experiments were performed in triplicate. Before analysis, the biomass was washed with sterile water to remove excess microbial biomass. The cellulose, hemicellulose, and lignin contents of the control and consortium samples were determined by an acid hydrolysis method and reported as a solid dry mass fraction [10]. The hydrolysate was characterized by high-performance anion exchange with pulsed amperometric detection (HPAEC-PAD) with a Dionex ICS-3000 system, using CarboPac PA1 column (4 × 250 mm) and CarboPac PA1 guard column (4 × 50 mm) to determine the concentration of cellulose and hemicellulose. Soluble lignin was determined from ultraviolet absorption (215 and 280 nm) of the hydrolysate. The insoluble lignin was determined as the solid faction from hydrolysis and subtracted from its ash content.
Enzyme Activity Analysis from NaLC Enrichment
For analysis of enzymatic activity, 1 mL of the culture supernatant was collected on the 3rd and 7th days. The final reaction volume was 100 μL, where 50 μL of the supernatant was mixed with 50 μL of pre-prepared 1% (w/v) solutions of different substrates as carboxymethylcellulose (CMC, Sigma), 1% (w/v) beta-glucan (Megazyme), 1% (w/v) beechwood xylan (Sigma), 1% (w/v) microcrystalline cellulose (Avicel®) (sigma), and 1% (w/v) xyloglucan from tamarind (Megazyme), among others diluted in 0.1 M phosphate buffer (pH 7.0). The reactions containing the supernatant from the enzymatic consortium and the substrates were incubated at 30 °C for 1 h. The amount of reducing sugars was determined using the 3,5-dinitrosalicylic acid (DNS) method [13]. As negative controls, the same procedure was performed, but with supernatant BED culture incubated in the absence of NaLC, and the residual sugar values subtracted from the results found for each substrate. Blank reactions were also made by incubating substrates in the absence of supernatants (both experimental and control groups), and the residual sugar values subtracted from the results found for each substrate.
Morphological Analyses by Electronic Microscopy
The analyses of dried BED by scanning electron microscopy (FEI Quanta 250) were performed by sampling on the 7th day of growth. The control consisted of a BED on the 7th day non-inoculated medium. ImageJ 1.33f, which is available online, was used to measure fiber areas of different lengths of sugarcane bagasse [14]. Environmental scanning electron microscopy micrographs were obtained with a non-commercial FEI Quanta 250 equipped with a Bruker QUANTAX EDS XFlash 6 detector. The samples were observed with an acceleration voltage of 15 kV, the current of the probe 80 pA, and the environmental distance between the sample surface and the second pressure limiting opening was 10 mm.
DNA Extraction and 16S rDNA and ITS2 Sequencing
Samples from the microbial consortium collected on the 7th day of cultivation were filtered using a nylon membrane (12 μm). The retained material was washed by centrifugation in 10 mL PBS, 14,000 g at 4 °C for 10 min. DNA extraction was performed using FastDNA® SPIN Kit for soil (MP Biomedicals), according to the manufacturer’s instructions.
The diversity and composition of bacterial communities were determined using the protocols of Caporaso et al. [15] and White et al. [16]. The pairs of oligonucleotide primers used in the amplification of the V4 region of the 16S rDNA gene and ITS2 were modified by the insertion of the illumina adapter sequence at the 5′ of each oligonucleotide, resulting in the following sequences, respectively: 515F_V4 5′- tcgtcggcagcgtcagatgtgtataagagacaggtgccagcmgccgcggtaa-3′, 806R_V4 5′-gtctcgtgggctcggagatgtgtataagagacagggactachvgggtwtctaat-3′ [15] and ITS3 (5′-tcgtcggcagcgtcagatgtgtataagagacagggcatcgatgaagaac-3′), ITS4 (5′- gtctcgtgggctcggagatgtgtataagagacagtcctccgcttattgatatg-3′) [16]. PCR was carried out with high fidelity Phusion polymerase (Thermo Scientific) using the following cycle: 2 min for 98 ºC (initialization and denaturing), followed by 98 °C for 30 s, 57 ºC for 40 s (annealing), 72 ºC for 2 min (extension) for 25 cycles, with a final elongation step at 72 °C for 10 min. The amplicon libraries were then prepared using Nextera DNA Sample Preparation Kit, following the manufacturer’s recommended protocol (Illumina). All amplicon libraries were performed in duplicate and finally pooled in equal molar quantities. The sequencing run was performed with a Miseq Reagent kit v2 (300 cycles) mixed with 15% PhiX control V3 on a Miseq instrument available at NGS sequencing facility at LNBR/CNPEM.
Diversity Analysis
Raw 16S rRNA reads from Illumina sequencing were quality checked using FASTQC (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/); the USEARCH merge pairs function was used to assemble pair-end reads and to remove low-quality sequences. Merged reads were further analyzed using the UPARSE pipeline [17]. Sequences were clustered into operational taxonomic units (OTU) based on a 97% identity using the USEARCH method. Chimeric sequences were identified and removed using the UCHIME method implemented with the cluster OTU function. The representative sequence from each OTU was assigned to taxonomic ranks using Ribosomal Database Project RDP Classifier [18] and UNITE Fungal ITS training sets [19]. The Chao1, ACE, Shannon, and Simpson indices, rarefaction curves, and sample coverage (Good’s coverage) were calculated by employing the Phyloseq and Vegan packages using R statistical software version 3.2. Nucleotide sequences were deposited in NCBI under the accession numbers: SRR12005466.
Metagenomic Analysis
An Illumina Nextera DNA library was prepared using total DNA from the 7th day of enrichment following the manufacturer’s specifications. Sequencing was performed in paired-end mode (2 × 100-bp reads) on an Illumina Hiseq 2500 instrument (Illumina, San Diego, CA, USA) available at NGS sequencing facility at LNBR/CNPEM. The de novo metagenome assembly was performed using the IDBA-UD software [20]. Gene prediction and automatic annotation were performed using Prokka [21]. Contigs shorter than 1000 nucleotides were excluded from downstream analysis. Nucleotide sequences were deposited in NCBI under the accession numbers: SRR11869874.
The metagenome de novo assembly was binned using CONCOCT [22] with default parameters to recover metagenome-assembled genomes (MAGs). The completeness and contamination ratios of the recovered genomes were assessed using CheckM [23] and the most likely taxonomic ranks of the recovered genomes were predicted by GTDB-Tk [24].
Functional annotation MAGs were done using Prokka. Further analysis for carbohydrate-active enzyme annotation was performed using the dbCAN2 HMM profiles [25], the carbohydrate-active enzymes database [26], and PFAM database [27]. Clusters of CAZyme genes were analyzed using CGC clusters from the dbCAN-seq database [28].
Statistical Analysis
The statistical set of the GraphPad Prism 6 program (GraphPad Software, San Diego, CA) was used for data and image analysis. The results were expressed as an average of 3 independent repetitions. The experimental data microscopic images and enzyme activity data were submitted to ANOVA and the means compared with the Tukey test using a 95% confidence interval. For metagenomics data, the statistical analysis was made using specific bioinformatic programs.
Result and Discussion
The Mobilization of Sugarcane Bagasse by NaLC
The pretreatment of lignocellulosic biomass is considered an essential step in increasing the accessibility of lignocellulose to enzymatic hydrolysis [1]. The initial chemical composition of raw sugarcane bagasse was 41.3% cellulose, 29.3% hemicellulose, and 23.9% lignin. The steam explosion pretreatment process promoted partial solubilization and consequent detachment of hemicellulose, reducing the initial content from 29% to just 7.9% (in remaining solids), and a relative increase of cellulose content (by percentage) in the treated sugarcane bagasse (Table 1). The chemical composition of sugarcane bagasse (BED) incubated with no bacteria consortia (control groups) showed little modifications when compared to non-incubated BED. These differences may be explained by the release of microspheres in the culture medium that were not collected for chemical analysis.
Due to the delignification step, the lignin content of BED also decreased to 18.2%. The pretreatment of sugarcane (with steam explosion followed by delignification) results in partial hydrolysis of hemicelluloses and concomitant lignin oxidation. This process also generates defibrillation of cellulose, allowing access to hydrolytic enzymes for subsequent fermentation [1].
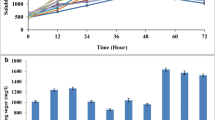
The analysis of BED degradation by the NaLC was performed over 7 days, showing a high consumption rate from the 1st to the 3rd day of cultivation, during which around 35% of the biomass was consumed compared to the control (Fig. 1A). However, from the 3rd to the 7th day, the rate of degradation decelerated. In general, 55% degradation of the lignocellulosic content by NaLC occurred (Fig. 1A), where it was mainly the cellulosic fraction that decreased from 75.7 to 47.7% after 1 week of incubation (Table 1).
Lignocellulose degradation profile of NaLC. A Determination of the dry mass loss of the sugarcane bagasse. B The extracellular enzymatic activities of the supernatant lignocellulolytic microbial consortia (NaLC), gray bars activity after BED incubation for 3 days and black bars for 7 days. Values are the mean ± SD (standard deviation) from three independent experiments; NS indicates a non-significant difference at p = 0.05. Xyg, xyloglucanase; AVCase, avicelase; BG, B-glucanase; Xn, xylanase
Wongwilaiwalin et al. [2] showed efficient degradation of filter paper (54.7%), sugarcane bagasse (59.4%), and rice straw (75.3%) through a thermophilic microbial consortium bred from sugarcane bagasse compost. Although it is very difficult to compare the results of different studies, several factors such as the microorganisms’ source, substrate composition, and the pretreatment and culture conditions which influenced the rates of fiber degradation by different consortia.
In order to correlate biomass degradation with the enzymatic activities, cellulases and hemicellulases of NaLC were evaluated. Enzymatic activities were detected in the supernatant of 3rd day cultivation samples for β-glucanase and xylanase (Fig. 1B). The NaLC supernatant showed low activity against β-glucan, xylan, avicel, and CMC, and showed no activity against pectin, xyloglucan or arabinoxylan.
The breakdown of insoluble and complex polysaccharides requires a close interaction between the enzyme and the substrate as well as the cooperation of multiple enzymes to improve hydrolysis [3]. Anaerobic microorganisms (such as those of intestinal microbiota) degrade the lignocellulolytic material through a multienzymatic complex, called a cellulosome. This structure has great spatial proximity to and high affinity with the lignocellulolytic surface, facilitating the synergistic action of hydrolytic enzymes upon substrates [29]. In contrast, the aerobic microorganisms degrade lignocellulose by secreting several soluble cellulases and hemicellulases [29].
It was also demonstrated some morphological changes in BED after the 7th day of cultivation with NaLC. The structure of BED showed a significant decrease in the thickness of the fibers observed by scanning electron microscopy (Fig. 2A–C). In addition, there was also observed the presence of microspheres with varied sizes in both control and experimental groups, but these structures were generally 3 × larger in BED incubated with NaLC, although less common (Fig. 2D, E). The distribution, size, and abundance of these microspheres vary, depending on the type of pretreatment [30]. The enlargement of microspheres formed by incubation with NaLC may be caused by the fusion of smaller spheres.
Morphological analysis sugarcane bagasse treated with NaLC. General view of the sample showing fibers and pseudo-lignin after incubation in control (no bacteria-CTL) (A) and NaLC (B) at 7 days. The insets show the amplification of pseudo-lignin in the both groups. Measures from fiber areas control (black bars) and bacteria consortia (NaLC) (grey bars) by 7 days (C). Measures of diameter and frequency of fibrous microspheres by scanning electronic microscope of sugarcane bagasse incubated in control (black bars) and NaLC (gray bars) at 7 days (D and E, respectively). Values are the mean ± SD (standard deviation) from three independent experiments. Asterisk (*) indicates a significant difference where *p < 0.05 and ****p < 0.0001
According to Shinde et al. [30], these microspheres show similar properties with lignin and are named pseudo-lignin. Several studies have reported the formation of pseudo-lignin due to pretreatment with low pH and/or high temperatures, such as pretreatment with steam explosion [30, 31]. This is the first time that formation of these structures has been observed due to microbial consortia. The composition and quantity of pseudo-lignin vary significantly depending on the type of biomass and pretreatment conditions used. One mechanism involved in the formation of pseudo-lignin is the condensation between lignin and furfural/hydroxymethyl furfural (HMF) derived from the dehydration of pentose and hexose. HMF and furfural can also produce other aromatic compounds which may give rise to pseudo-lignin via polymerization/polycondensation reactions [30, 31]. Another factor that probably contributes to the formation of pseudo-lignin is the cultivation condition of NaLC under high aeration conditions. Hu et al. [31] studied the role of oxygen in the formation of pseudo-lignin and showed that an atmosphere of enriched oxygen significantly increases the formation of these microspheres. An important aspect is that pseudo-lignin could bind lignocellulolytic enzymes/microbes in vitro, affecting negatively the enzymatic hydrolysis of polysaccharides [30, 31], which might explain the low enzymatic activity of NaLC showed in Fig. 1B.
Taxonomic Analysis of NaLC
Large-scale sequencing is considered a powerful tool for the study of complex microbial consortia, allowing for the identification of genes, pathways, and genomes involved in plant biomass conversion [3]. In this study, it was evaluated the microbial diversity and composition of the NaLC using 16S rDNA and ITS2. The analysis of the eukaryotic profile based on the ITS2 did not reveal sequences related to fungal species. In addition, the visualization of bagasse by optical and electronic microscopy after NaLC incubation did not reveal the presence of yeasts and/or hyphae (data not shown). The low nutritional quality of the medium with sugarcane bagasse as the sole carbon source probably explains the absence of fungi.
Regarding 16S rDNA sequence analysis, 62.04 quality sequences were obtained, and these were used for diversity analysis (see Supplementary Figure 1), which resulted in the identification of 22 OTUs. Based on the rarefaction curve, which formed a plateau, there was found that depth of sequencing was enough to estimate diversity and richness of this bacterial community (Figure 3A). The NaLC community was dominated by bacteria only and the most abundant phyla in the community were Bacteroidetes (50.6%) and Proteobacteria (47.7%), less representative phyla were Firmicutes (1.5%) and Acidobacteria (<1%) (Figure 3B). The microbial composition in NaLC resulted in the predominance of two phyla, Bacteroidetes and Proteobacteria. These results were similar to other data from aerobic bacterial communities from environments that degrade lignocellulosic compounds from different biomasses, including sugarcane bagasse [32]. However, under anaerobic conditions, several studies on lignocellulolytic microbial enrichment have shown that Firmicutes is the most abundant phylum [32].
The Bacteroidetes phylum consists of gram-negative bacteria, and their members colonize different habitats, including the gastrointestinal tract of animals [14, 32]. This phylum produces a great variety of hydrolytic enzymes, which are also involved in the processing of complex polysaccharides [32], through the genomic organization of CAZymes in polysaccharide utilization loci (PUL) [12]. Among the Bacteroidetes of NaLC, the representatives of the genus Flavobacterium sp. had an impressive representation (41.8%) (Fig. 3B), which comprises several aerobic cellulolytic bacteria [33].
The phylum of Proteobacteria had a representation of 47.7% in the NaLC and is considered the most extensive and varied among the phyla of bacteria. This phylum has great diversity in morphological and metabolic terms, including autotrophic and heterotrophic, aerobic, and anaerobic [34]. This group is considered to be efficient in the degradation of several aromatic compounds, aliphatic components, and others [34]. In insects, several metagenomic studies have showed that Bacteroidetes, Proteobacteria, Firmicutes, and Actinobacteria are the predominant phyla in the gut of diverse insect orders [35, 36].
Among the Proteobacteria, Sphingomonas sp. and Sphingopyxis sp. had a relative abundance of 18.2% and 4.5% respectively (Fig. 3C). Sphingomonas and Sphingopyxis are widespread in nature and can be obtained from various sources such as water, soil, living organisms, and even human skin. According to Mnich et al. [37] and Shin et al. [38], many Sphingomonas strains produce lignin-degrading enzymes and some species of Sphingopyxis are capable of expressing lignocellulolytic hydrolases.
The phylum Firmicutes is found in several environments, such as the gastrointestinal tract of animals, especially under anaerobic and/or micro aerobic conditions [35]. They are able to degrade recalcitrant components [32], but in data of this study, Firmicutes represented only a small percentage of the analyzed sequences (Fig. 3B). This fact may be related to the aerobic condition of consortia cultivation.
It was also found that the Acidophilic phylum that has already been observed in several environments, such as soils, hot springs, active sludge, and lake sediments [39]. These microorganisms are considered important lignocellulose degrading agents. The genomic analysis of this group showed the presence of several genes involved in the degradation of complex plant polysaccharides. These microorganisms can be cultivated in nitrogen-poor substrates and withstand considerable levels of anti-nutritional compounds such as those found in sugarcane [39].
The Profile of CAZyme Genes in NaLC
To evaluate the composition of genes encoding CAZymes in the NaLC, the metagenomic DNA on the 7th day of cultivation with pretreated sugarcane bagasse was sequenced. The de novo metagenome assembly was performed using the software IDBA_UD and 148.50 open reading frames (ORFs) were identified for most of these genes (about 87%) predicted to encode proteins with sequences of more than 100 amino acids [see Supplementary Fig. 2]. In order to identify putative genes encoding enzymes involved in breaking and modifying carbohydrates, the predicted ORFs were annotated using a dbCAN HMM-based search.
Among the identified CAZyme classes, genes encoding glycoside hydrolases (GHs) represented the most abundant class, followed by glycosyl transferases (GTs), carbohydrates esterases (CEs), while less prevalent classes where those of carbohydrate binding modules (CBMs), auxiliary enzymes (AAs), and polysaccharides lyases (PLs) (Fig. 4A).
Functional classification of identified CAZymes. A The open reading frames (ORFs) of genes encoding proteins classified according to their functions using the dbCAN HMM profiles for automated carbohydrate-active enzyme annotation. B The taxonomic annotation assigned by Ghost Koala annotation software from identified CAZymes (18,954 proteins). C (GH) glycoside hydrolases represented by 20 families with the highest number annotated. D (CE) Carbohydrate esterase. E (CBM) Carbohydrate binding modules are (CBM) represented by 20 families with the highest number annotated. F (AA) Auxiliary activities
The taxonomic profile of CAZymes revealed the predominance of sequences belonging to the Bacteroidetes and Proteobacteria phylum, indicating that these taxa are probably primary agents in the deconstruction of sugarcane bagasse by NaLC (Fig. 4B), which appears to correlate with the results from 16S rDNA sequence analysis. The specific participation of these NaLC genes, during sugarcane bagasse degradation, is under investigation using the metatranscriptome approach (data not shown).
The polysaccharide lyases (EC 4.2.2.-) are a group of enzymes that cleave uronic acid-containing polysaccharides through an elimination mechanism. The amount of PLs is generally low and always less than that of GHs, the most likely explanation for this observation is that the substrates of PLs represent only a small proportion of all carbohydrate polymers [40]. Therefore, the present study will focus only on the most abundant CAZyme in the breakdown and modification of carbohydrates, which are GHs, CEs, CBMs, and AAs.
Glycoside hydrolases (EC 3.2.1.-) are a group of enzymes which hydrolyze the glycosidic bond between two or more carbohydrates or between a carbohydrate and a non-carbohydrate moiety [41]. Several GH families comprised of classical cellulolytic enzymes such as endoglucanases (GH5 and GH9) and β-glucosidases (GH1 and GH3) were identified in NaLC and are among the most abundant (Fig. 4C). However, genes encoding for cellobiohydrolase family GH7 have not been identified, which was predicted as this family has not yet been found in bacteria or archaea, and only one gene was found for GH6. The family GH23 was highly represented among the total GHs found in NaLC, followed by GH13 and GH43 (Fig. 4C). The glycosyl hydrolases of the GH23 family are known as lysozyme type G, peptidoglycan lyases and chitinases, which represent a group of enzymes that hydrolyze polysaccharides [42]. The enzymes of GH23 family have been classified as debranching enzymes that could act in polysaccharide mobilization [43]. Bras et al. [42] identified in Clostridium thermocellum as endoglucanase whose active site exhibited a conserved structure with enzymes of the GH23 family. The GH13 family, known as the α-amylase family, contains enzymes with different activities and substrate specificities. According to Li et al. [44], these proteins were identified in several metagenomes from different environments.
Genes from the GH43 family that encode hemicellulases have also been found in NaLC, including xylanases (GH43 and also families GH8 and GH5) that randomly cleave the main chain of xylan and β-xylosidase (GH43 and also families GH3 and GH1) that cleave non-reducing ends-releasing xylose monomers, as well as mannosidases (GH2) and α-L-arabinofuranosidases (GH43) (Fig. 4C). Genes from this family, as well as from GH5, are often identified in several microbial communities and are divided into 37 (GH43) and 51 (GH5) subfamilies, which differ in structure and function [45, 46].
The fourth most abundant family among the GHs in the NaLC consortium was the GH3 family (Fig. 4C); this family also appeared among the most abundant in several metagenomes analyzed, including the human gut, mouse, termite, and bovine rumen [44]. In the termite gut, 69 modules corresponding to the catalytic domain of GH3 were identified, suggesting its participation in the final metabolism of oligosaccharides into simple sugars [47].
The second most abundant class of CAZymes found in NaLC was the CEs, a group of enzymes that remove ester substituents from glycan chains, helping GHs in the conversion of biomass [48]. Among the most abundant families found in NaLC were CE1 and CE4, described as acetyl xylan esterases, directly involved in hemicellulose modification [29] (Fig. 4D). These data are similar to that observed by Liu et al. [49], where CE1 and CE4 were one of the most abundant families in the gut microbiota of the higher termite Globitermes brachycerastes. Another well-represented family was CE10 (carboxyl esterase), but these enzymes are no longer considered carbohydrate esterases and their information is no longer updated in the CAZy database.
In addition to the GHs and CEs, non-catalytic modules (CBMs) were also abundant in the NaLC community. CBMs recognize and bind to carbohydrates, improving the catalytic efficiency of GHs in the degradation of polysaccharides [29]. The most abundant family was CBM50, which binds to peptidoglycan and chitin (Fig. 4E) [48]. CBM50 can be found associated with 6 families of GHs, including GH23 (the most abundant among the GHs of NaLC), that could explain the predominance of these CBMs in NaLC.
The class of auxiliary activities (AAs) is involved in the degradation of lignocellulose through oxidative pathways (Fig. 4F) [50]. AA genes were also found among the NaLC, such as genes encoding enzymes from the AA2 (peroxidases), AA4 (vanillyl alcohol oxidase), and AA6 (1,4-benzoquinone reductase) families. Among the AAs involved in the degradation of glucose-based polymers, it was found genes encoding for AA3 (cellobiose dehydrogenases/glucose oxidase) and AA7 (glucooligosaccharide oxidase). Genes from families AA3 and AA7 appear to be associated with both lignin and cellulose degradation [50].
The enzymes from AA3, AA4, AA6, and AA7 families have been associated to eukaryotic cells, mainly fungal species [50]. However, lignin degradation by AA families produced by some bacteria has also been recently described [51]. But the role of these AA enzymes in bacteria is not well understood.
Several Cazyme Genes in NaLC are Organized as PUL
The metagenome assembly used a database for the recovery of microbial genomes throughout the contig binning strategy. This analysis resulted in the recovery of 22 MAGs, being mostly of high-quality, with more than 70% completeness and < 5% contamination, which is in agreement with the total number of OTUs found in 16S rDNA sequence analysis. Among the MAGs, 16 genomes were recovered with more than 90% completeness and 14 genomes were classified at the genus level [see Supplementary Fig. 3]. The MAGs were analyzed using dbCAN database for the identification of CAZymes and CAZyme gene clusters. It was assembled in 30 contigs with a genome size of 5.78 Mb and was classified as belonging to the gram-negative genus Niabella sp., family Chitinophagaceae from the Bacteroidetes phylum.
Bacteria from Bacteroidetes are found in many ecosystems and they usually have several CAZymes involved in the degradation of polysaccharides. They degrade complex glycans outside the cell and fully convert them to monosaccharides in the periplasmic space, avoiding competition for the substrate with other bacteria [52]. The Bacteroidetes microorganism organizes its genes involved in metabolism of specific polysaccharides in specific loci encoding for different proteins such as transporters, hydrolases, and regulators in the same portion of the genome [12]. In the gut of cockroach Periplaneta americana, some Bacteroidetes isolates showed (at their genomes) PUL with typical markers as SusD cell surface glycan binding proteins, SusC-like/TonB-dependent transporters, and transcriptional regulators that include extracytoplasmic (ECT) sigma/anti-sigma factors systems [12].
The specific arrangement of these genes is called a PUL and is widely disseminated in bacteria from bovine rumen, human distal intestine, and ocean environments [53]. Despite being identified in the intestinal microbiota of termites/cockroaches, the presence/participation of PUL in the deconstruction of lignocellulose remains poorly investigated. Therefore, a bioinformatics protocol was run for the identification of PUL in the genome of Niabella sp., using the presence of SusC and SusD gene pairs (transporter genes) as markers [54], and several PUL were identified in the genome of Niabella sp. from NaLC (see supplementary Fig. 4). Supplementary Fig. 5 shows the CAZyme prediction of the MAGs at the phylum level, revealing that the majority of CAZymes identified were GHs, for which 451 genes were found from Bacteroidetes MAGs, 210 from Proteobacteria, 121 from Verrucomicrobia, 79 from Acidobacteria, and 16 from Planctomycetes. Genes encoding CEs, PLs, and AAs were also found in the MAGs, with the majority of them belonging to the Bacteroidetes phylum. Among the recovered genomes, MAG 34 showed the highest number of annotated CAZymes, a total of 193 genes. It is a high-quality MAG with 99.5% genome completeness and 0.5% contamination [see Figure Supplementary 3].
However, unlike PUL for carbohydrates, gene clusters involved in lignin mobilization have not been previously described. Supplementary Fig. 6 shows a specific arrangement of some genes that suggest the presence of a putative lignin utilization locus – LUL. In this case, the SusC and SusD gene pairs were surrounded by genes encoding an AA2 peroxidase, NADH dehydrogenase, and a MnSOD (manganese superoxide dismutase) gene. The catalytic cycle of lignin-modifying peroxidases (AA2) is started with hydrogen peroxide (H2O2) provided by the AA3 family [55], the most abundant AA family in the NaLC, shown in Fig. 4 F. In addition to the peroxidase-based mechanism, some oxidases and reductases can also promote a redox cycle for lignin modification [56]. The detection of NADH dehydrogenase in the genome of Niabella sp. suggests participation of this nucleotide in redox balance during lignin mobilization. The occurrence of lignin degradation by the Fenton reaction, which consists of the reaction between Fe2+ with H2O2, generating hydroxyl radicals with high redox potential [57] and can attack lignin, cannot be ruled out. Moreover, located next to the AA2 gene, an acyl carrier gene was identified, suggesting involvement in the transport of hydrophobic intermediates during lignin oxidation. In addition, integrating this LUL, it was observed a MnSOD gene that was recently described as a ligninase in the genome of Sphingobacterium sp.—T2, also from Bacteroidetes phylum—which would be a new function since it was believed to act only against oxidative stress [57].
This putative new locus (LULs) in Bacteroidetes genomes opens several important questions about co-localization, co-regulation, genomic architecture, sensing/sequestration of monomers/products, enzymatic oxidation, and transport of oligo/monomers of lignin (or derivatives) during lignin mobilization. The organization of these genes in LULs could also give new insights into biomass mobilization by microorganisms.
Conclusion
This study showed that the lignocellulolytic microbial consortium derived from the gut microbiota of the cockroach Nauphoeta cinerea was able to consume 55% of the sugarcane bagasse, which was the only carbon source. The 16S and ITS2 analyses showed that the microbial community of NaLC was composed only of bacteria, mainly from phyla Bacteroidetes and Proteobacteria. The shotgun metagenomic analysis showed that this enrichment consortium had a diversified reservoir of genes encoding for CAZymes (mainly GHs and CEs). It was also observed the presence of a novel locus named here LULs that encodes several enzymes probably involved in lignin mobilization (mainly AA2 peroxidase, NADH dehydrogenase, and a MnSOD gene). Finally, the enzymatic repertoire detailed here showed the potential of microbial communities from the insect gut as a reservoir of novel genes and microorganisms that efficiently decompose pretreated lignocellulosic biomass.
Data Availability
All data generated or analyzed during this study are included in its Additional files.
Abbreviations
- AAs:
-
Auxiliary enzymes
- Avicel:
-
Microcrystalline cellulose
- Avicelase:
-
Enzyme that break avicel
- BED:
-
Pre-treated sugarcane bagasse
- BHB:
-
Bushnell Haas Broth commercial medium
- CAZymes:
-
Carbohydrate-active enzymes
- CBMs:
-
Carbohydrate binding modules
- CEs:
-
Carbohydrates esterases
- CMC:
-
Carboxymethylcellulose
- CMCase:
-
Enzyme that break carboxymethylcellulose
- CNPEM:
-
Centro Nacional de Pesquisa em Energia e Materiais
- DNS:
-
3,5-Dinitrosalicylic acid
- DUFs:
-
Domains of unknown function
- GHs:
-
Glycoside hydrolases
- GTs:
-
Glycosyl transferases
- HMF:
-
Furfural/hydroxymethyl furfural
- H2O2 :
-
Hydrogen peroxide
- HPAEC-PAD:
-
High-performance anion exchange with pulsed amperometric detection
- ITS:
-
Internal Transcribed Spacer 2
- LNBR:
-
Brazilian Biorenewables National Laboratory
- LULs:
-
Lignin utilization loci
- MAGs:
-
Metagenome-assembled genomes
- MnSOD:
-
Manganese superoxide dismutase
- NaLC:
-
Consortium derived from the gut of Nauphoeta cinerea
- ORFs:
-
Open reading frames
- OTU:
-
Operational taxonomic units
- PUL:
-
Polysaccharide utilization loci
References
SH Mood AH Golfeshan M Tabatabaei GS Jouzani GH Najafi M Gholami M Ardjmand 2013 Lignocellulosic biomass to bioethanol, a comprehensive review with a focus on pretreatment Renew Sustain Energy Rev 27 77 93 https://doi.org/10.1016/j.rser.2013.06.033
S Wongwilaiwalin U Rattanachomsri T Laothanachareon L Eurwilaichitr Y Igarashi V Champreda 2010 Analysis of a thermophilic lignocellulose degrading microbial consortium and multi-species lignocellulolytic enzyme system Enzyme Microb Technol 47 283 290 https://doi.org/10.1016/j.enzmictec.2010.07.013
Sun J, Zhou XJ (2011) Utilization of lignocellulose- feeding insects for viable biofuels: an emerging and promising area of entomological science. In: Liu T., Kang L. (eds) Recent Advances in Entomological Research. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-17815-3_25
C Schauer CL Thompson A Brune 2012 The bacterial community in the gut of the cockroach Shelfordella lateralis reflects the close evolutionary relatedness of cockroaches and termites Appl Environ Microbiol 78 2758 2767 https://doi.org/10.1128/AEM.07788-11
DRA Wharton ML Wharton 1965 The cellulase content of various species of cockroach J Insect Physiol 11 1401 1405 https://doi.org/10.1016/0022-1910(65)90177-0
EN Elpidina KS Vinokurov VA Gromenko YA Rudenskaya YE Dunaevsky DP Zhuzhikov 2001 Compartmentalization of proteinases and amylases in Nauphoeta cinerea midgut Arch Insect Biochem Physiol 48 206 216 https://doi.org/10.1002/arch.10000
LL Jiang JJ Zhou CS Quan ZL Xiu 2017 Advances in industrial microbiome based on microbial consortium for biorefinery Bioresour Bioprocess 4 1 11 https://doi.org/10.1186/s40643-017-0141-0
C Xu H Yu 2021 Insights into constructing a stable and efficient microbial consortium Chin J Chem Eng 30 112 120 https://doi.org/10.1016/j.cjche.2020.12.012
LD Bushnell HF Haas 1941 The utilization of certain hydrocarbons by microorganisms J Bacteriol 41 653 673
ER Gouveia RTD Nascimento AM Souto-Maior GJM Rocha De 2009 Validação de metodologia para a caracterização química de bagaço de cana-de-açúcar Quim Nova 32 1500 1503 https://doi.org/10.1590/S0100-40422009000600026
GJM Rocha AR Gonçalves BR Oliveira EG Olivares CEV Rossell 2012 Steam explosion pretreatment reproduction and alkaline delignification reactions performed on a pilot scale with sugarcane bagasse for bioethanol production Ind Crop Prod 35 274 279 https://doi.org/10.1016/j.indcrop.2011.07.010
A Vera-Ponce de León BC Jahnes J Duan LA Camuy-Vélez ZL Sabree 2020 Cultivable, host-specific Bacteroidetes symbionts exhibit diverse polysaccharolytic strategies Appl Environ Microbiol 86 8 1 24 https://doi.org/10.1128/AEM.00091-20
G Miller R Blum W Glennon A Burton 1960 Measurement of carboxymethylcellulase activity Anal Biochem 132 127 132 https://doi.org/10.1016/0003-2697(60)90004-X
Rasband WS (1997) Image J. Bethesda, MD National Institutes of Health. http://imagej.nih.gov/ij.
JG Caporaso CL Lauber WA Walters D Berg-Lyons J Huntley N Fierer SM Owens J Betley L Fraser M Bauer N Gormley JA Gilbert G Smith R Knight 2012 Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms ISME J 6 1621 1624 https://doi.org/10.1038/ismej.2012.8
White TJ, Bruns S, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protocols: a guide to methods and applications. In: Innis MA, Gelfand DH editors. London: Academic Press 315–322.
RC Edgar 2013 UPARSE: highly accurate OTU sequences from microbial amplicon reads Nat Methods 10 996 998 https://doi.org/10.1038/nmeth.2604
Q Wang GM Garrity JM Tiedje JR Cole 2007 Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy Appl Environ Microbiol 73 5261 5267 https://doi.org/10.1128/AEM.00062-07
RH Nilsson L Tedersoo M Ryberg E Kristiansson M Hartmann M Unterseher 2015 A comprehensive, automatically updated fungal ITS sequence dataset for reference-based chimera control in environmental sequencing efforts Microbes Environ 30 145 150 https://doi.org/10.1264/jsme2.ME14121
F Peng P Peng F Xu RC Sun 2012 Fractional purification and bioconversion of hemicelluloses Biotechnol Adv 30 879 903 https://doi.org/10.1016/j.biotechadv.2012.01.018
T Seemann 2014 Prokka: rapid prokaryotic genome annotation Bioinformatics 30 2068 2069 https://doi.org/10.1093/bioinformatics/btu153
J Alneberg BS Bjarnason I Bruijn M Schirmer J Quick UZ Ijaz L Lahti NJ Loman AF Anderson C Quince 2014 Binning metagenomic contigs by coverage and composition Nat Methods 11 1144 1146 https://doi.org/10.1038/nmeth.3103
DH Parks M Imelfort CT Skennerton P Hugenholtz GW Tyson 2015 CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes Genome Res 25 1043 1055 https://doi.org/10.1101/gr.186072.114
PA Chaumeil AJ Mussig P Hugenholtz DH Parks 2019 GTDB-Tk: a toolkit to classify genomes with the Genome Taxonomy Database Bioinformatics 36 1925 1927 https://doi.org/10.1093/bioinformatics/btz848
Y Yin X Mao J Yang X Chen F Mao Y Xu 2012 dbCAN: a web resource for automated carbohydrate-active enzyme annotation Nucleic Acids Res 40 445 451 https://doi.org/10.1093/nar/gks479
V Lombard GH Ramulo E Drula PM Coutinho B Henrissat 2014 The carbohydrate-active enzymes database (CAZy) in 2013 Nucleic Acids Res 42 490 495 https://doi.org/10.1093/nar/gkt1178
S El-Gebali J Mistry A Bateman SR Eddy A Luciani SC Potter M Qureshi LJ Richardson GA Salazar A Smart ELL Sonnhammer L Hirsh L Paladin D Piovesan SCE Tosatto RD Finn 2019 The Pfam protein families’ database in 2019 Nucleic Acids Res 8 427 432 https://doi.org/10.1093/nar/gky995
H Zhang T Yohe L Huang S Entwistle P Wu Z Yang PK Busk Y Xu Y Yin 2018 dbCAN2: a meta server for automated carbohydrate-active enzyme annotation Nucleic Acids Res 46 95 101 https://doi.org/10.1093/nar/gky418
ME Himmel Q Xu Y Luo SY Ding R Lamed EA Bayer 2010 Microbial enzyme systems for biomass conversion: emerging paradigms Biofuels 1 323 341 https://doi.org/10.4155/bfs.09.25
SD Shinde X Meng R Kumar AJ Ragauskas 2018 Recent advances in understanding the pseudolignin formation in a lignocellulosic biorefinery Green Chem 20 2192 2205 https://doi.org/10.1039/C8GC00353J
F Hu S Jung AJ Ragauskas 2012 Pseudo-lignin formation and its impact on enzymatic hydrolysis Bioresour Technol 117 7 12 https://doi.org/10.1016/j.biortech.2012.04.037
P Kanokratana W Mhuantong T Laothanachareon S Tangphatsornruang L Eurwilaichitr K Pootanakit V Champreda 2013 Phylogenetic analysis and metabolic potential of microbial communities in an industrial bagasse collection site Microb Ecol 66 322 334 https://doi.org/10.1007/s00248-013-0209-0
D Lednická J Mergaert MC Cnockaert J Swings 2011 Isolation and identification of cellulolytic bacteria involved in the degradation of natural cellulosic fibres Syst Appl Microbiol 23 292 299 https://doi.org/10.1016/S0723-2020(00)80017-X
K Kersters De vos P, Gillis M, Swings J, Vandamme P, Stackebrandt E, 2006 Introduction to the Proteobacteria Prokaryotes 5 3 37 https://doi.org/10.1007/0-387-30745-1_1
L Auer A Lazuka D Sillam-Dusses E Miambi M O’Donohue G Hernandez Raquet 2017 Uncovering the potential of termite gut microbiome for lignocellulose bioconversion in anaerobic batch bioreactors Front Microbiol 8 2623 https://doi.org/10.3389/fmicb.2017.02623
S Jang Y Kikuchi 2020 Impact of the insect gut microbiota on ecology, evolution, and industry Curr Opin Insect Sci 41 33 39 https://doi.org/10.1016/j.cois.2020.06.004
E Mnich R Vanholme P Oyarce S Liu F Lu G Goeminne B Jørgensen 2017 Degradation of lignin b-aryl ether units in Arabidopsis thaliana expressing LigD, LigF and LigG from Sphingomonas paucimobilis SYK-6 Plant Biotechnol J 15 581 593 https://doi.org/10.1111/pbi.12655
KC Shin DK Oh 2014 Characterization of a novel recombinant β-glucosidase from Sphingopyxis alaskensis that specifically hydrolyzes the outer glucose at the C-3 position in protopanaxadiol-type ginsenosides J Biotechnol 172 30 37
NL Ward JF Challacombe PH Janssen B Henrissat PMM Coutinho G Wu DH Xie M Haft J Sait RD Badger B Barabote TS Bradley LM Brettin D Brinkac T Bruce SC Creasy TM Daugherty 2009 Three genomes from the phylum Acidobacteria provide insight into the lifestyles of these microorganisms in soils Appl Environ Microbiol 75 2046 2056 https://doi.org/10.1128/AEM.02294-08
V Lombard T Bernard C Rancurel H Brumer PM Coutinho B Henrissat 2010 A hierarchical classification of polysaccharide lyases for glycogenomics Biochem J 432 3 437 444 https://doi.org/10.1042/BJ20101185
G Davies B Henrissat 1995 Structures and mechanisms of glycosyl hydrolases Structure 3 853 859 https://doi.org/10.1016/S0969-2126(01)00220-9
JLA Bras A Cartmell AL Carvalho G Vérze EA Bayer Y Vazana MA Correia JA Prates S Ratnaparkhe AB Boraston MJ Romão CM Fontes HJ Gilbert 2011 Structural insights into a unique cellulase fold and mechanism of cellulose hydrolysis PNAS 108 5237 5242 https://doi.org/10.1073/pnas.1015006108
L Wang G Zhang H Xu H Xin Y Zhang 2019 Metagenomic analyses of microbial and carbohydrate-active enzymes in the rumen of holstein cows fed different forage-to-concentrate ratios Front Microbiol 10 649 https://doi.org/10.3389/fmicb.2019.00649
LL Li SR McCorkle S Monchy S Taghavi D Lelie van der 2009 Bioprospecting metagenomes: glycosyl hydrolases for converting biomass Biotechnol Biofuels 2 10 https://doi.org/10.1186/1754-6834-2-10
H Aspeborg PM Coutinho Y Wang H Brumer B Henrissat 2012 Evolution, substrate specificity and subfamily classification of glycoside hydrolase family 5 (GH5) BMC Evol Biol 12 186 https://doi.org/10.1186/1471-2148-12-186
K Mewis N Lenfant V Lombard B Henrissat 2016 Dividing the large glycoside hydrolase family 43 into subfamilies: a motivation for detailed enzyme characterization Appl Environ Microbiol 82 1686 1692 https://doi.org/10.1128/AEM.03453-15
F Warnecke P Luginbühl N Ivanovna M Ghassemian TH Richardson JT Stege 2007 Metagenomic and functional analysis of hindgut microbiota of a wood-feeding higher termite Nature 450 560 565 https://doi.org/10.1038/nature06269
BL Cantarel PM Coutinho C Rancurel T Bernard V Lombard B Henrissay 2009 The carbohydrate-active enzymes database (CAZy): an expert resource for glycogenomics Nucleic Acids Res 37 233 238 https://doi.org/10.1093/nar/gkn663
N Liu H Li MG Chevrette L Zhang L Cao H Zhou X Zhou Z Zhou PB Pope CR Currie Y Huang Q Wang 2019 Functional metagenomics reveals abundant polysaccharide degrading gene clusters and cellobiose utilization pathways within gut microbiota of a wood-feeding higher termite ISME J 13 104 117 https://doi.org/10.1038/s41396-018-0255-1
A Levasseur E Drula V Lombard PM Coutinho B Henrissat 2013 Expansion of the enzymatic repertoire of the CAZy database to integrate auxiliary redox enzymes Biotechnol Biofuels 6 41 https://doi.org/10.1186/1754-6834-6-41
JP Silva ARP Ticona PRV Hamann BF Quirino EF Noronha 2021 Deconstruction of lignin: from enzymes to microorganisms Molecules 26 8 2299 https://doi.org/10.3390/molecules26082299
N Terrapon V Lombard HJ Gilbert B Henrissat 2015 Automatic prediction of polysaccharide utilization loci in Bacteroidetes species Bioinformatics 31 5 647 655 https://doi.org/10.1093/bioinformatics/btu716
JM Grondin K Tamura G Déjean DW Abbott H Brumer 2017 Polysaccharide utilization loci: fueling microbial communities J Bacteriol 199 e00860 e916 https://doi.org/10.1128/JB.00860-16
MH Foley DW Cockburn NM Koropatkin 2016 The Sus operon: a model system for starch uptake by the human gut Bacteroidetes Cell Mol Life Sci 73 2603 2617 https://doi.org/10.1007/s00018-016-2242-x
L Sützl CVFP Laurent AT Abrera G Schütz R Ludwig D Haltrich 2018 Multiplicity of enzymatic functions in the CAZy AA3 family Appl Microbiol Biotechnol 102 2477 2492 https://doi.org/10.1007/s00253-018-8784-0
L Lin Y Cheng Y Pu S Sun X Li M Jin EA Pierson DC Gross BE Dale SY Dai AJ Ragauskas JS Yuan 2016 Systems biology-guided biodesign of consolidated lignin conversion Green Chem 18 20 5536 5547 https://doi.org/10.1039/C6GC01131D
GMM Rashid CR Taylor Y Liu X Zhang D Rea V Fülöp TDH Bugg 2015 Identification of manganese superoxide dismutase from Sphingobacterium sp. T2 as a novel bacterial enzyme for lignin oxidation ACS Chem Biol 10 2286 2294 https://doi.org/10.1021/acschembio.5b00298
Acknowledgements
We would like to express our gratitude to David Forrest for editorial assistance.
Funding
This work was supported by grants from the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES), Fundação Carlos Chagas Filho de Amparo à Pesquisa do Estado do Rio de Janeiro (FAPERJ), Fundação de Amparo à Pesquisa do Estado de São Paulo (Fapesp). Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), and INCT-Entomologia Molecular.
Author information
Authors and Affiliations
Contributions
D.B.C. maintained bacteria consortia in laboratory conditions, performed experiments, analyzed data, edited figures, created the line drawing, and drafted the manuscript; D.A.P. maintained bacteria consortia in laboratory conditions, performed experiments, and analyzed data; G.F.P. performed the bioinformatics analysis of the article; J.C. performed and supervised the experiments of enzymatic activities; S.C.R. performed the chemical analysis of the article; A.G. performed the chemical and statistical analysis of the article; A.P.S. performed the chemical and statistical analysis of the article; M.B. performed the chemical and statistical analysis of the article; R.R. performed the sequencing experiments and its analysis; L.M.Z. performed the sequencing experiments and its analysis; F.M.S. designed, provided financial support, analyzed data; J.P.L.C. conceived, designed, and coordinated the study, analyzed data, and reviewed the manuscript; E.A.M conceived, designed, and coordinated the study, provided financial support, analyzed data, and reviewed the manuscript. All authors gave final approval for publication.
Corresponding author
Ethics declarations
Ethics Approval and Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Carvalho, D.B., Paixão, D.A., Persinoti, G.F. et al. Degradation of Sugarcane Bagasse by Cockroach Consortium Bacteria. Bioenerg. Res. 15, 1144–1156 (2022). https://doi.org/10.1007/s12155-021-10363-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12155-021-10363-4