Abstract
The apoptosis-associated speck-like protein (ASC protein) plays a central role in eukaryotic innate immune response. Upon infection, multiple ASC molecules assemble into long filaments, which are fundamental for triggering the cellular defense mechanism by starting an inflammatory cascade with the activation of caspase-1. ASC is composed of two domains, the C-terminal caspase-recruitment domain, which is involved in the recruitment of the caspase, and the N-terminal PYRIN domain (PYD), which is responsible for the formation of the filament. Here we present the 13C and 15N chemical shift assignment for filaments formed by the PYD of mouse ASC, a 91-residue protein. The backbone between residues 4 and 84 is assigned without interruption. Also, 86 % of the sidechain resonances for this stretch are assigned. Residues 1–3 and 85–91 show unfavorable dynamics and are not observed. Secondary chemical-shift analysis shows the presence of six α-helices.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Biological context
Inflammasomes are multiprotein oligomers that are of fundamental importance for the innate immune system. The formation of these oligomers is triggered by the recognition of pathogen- and danger-associated molecular patterns (PAMPs and DAMPs) by specific, germ-line encoded pattern-recognition receptors (PRRs), and eventually leads to an inflammatory cascade. This cascade starts with the activation of caspase-1 and the maturation of the inflammatory cytokines Interleukin 1β (IL-1β) and Interleukin 18 (IL-18), leading eventually to cell death. It has been shown that the structures of inflammasome complexes are crucial for the efficiency of the immune response (Schroder and Tschopp 2010); moreover, malfunctions of such oligomers are related to human disease, such as cancer and autoimmune syndromes (Schroder and Tschopp 2010; Hu et al. 2013).
The signaling pathway from the PRRs to the activation of caspase-1 is mediated in many cases by the bipartite adaptor apoptosis-associated speck-like protein containing a CARD (ASC), which consists of an N-terminal PYD and a C-terminal CARD. In ASC-dependent inflammasomes, the PYD oligomerizes upon interaction with the receptor to form the so-called ASC speck, and the C-terminal CARD subsequently recruits the inflammatory caspase. The large ASC filament thus forms the structural core of the ASC-inflammasome (Masumoto et al. 1999).
The monomer structures of the human ASC-PYD (de Alba 2009), and full-length ASC (Liepinsh et al. 2003) have previously been obtained by solution-state NMR spectroscopy, and the interaction mode of PYDs in the inflammasome assembly was addressed (Vajjhala et al. 2012; Srimathi et al. 2008; Vajjhala et al. 2014; Moriya et al. 2005). Recently, the structure of the filaments of the human ASC-PYD has been solved at ∼3.8 Å resolution using cryo-electron microscopy (Lu et al. 2014). In this work we present the chemical-shift assignment for the filaments of the mouse PYD of ASC, as the basis for the structural study of the filaments. We show that the presence or the absence of the CARD does not influence the structure of the filaments formed by the PYD.
Methods and experiments
Expression, purification and filament formation
The full length mouse ASC protein (residues 1–193) and the ASC-PYD domain only (residues 1–91) was produced using a PET28a vector under the control of a T7 promoter. A GSGSLE linker and a His6-tag were added to the C-terminus of both protein constructs. Proteins were expressed in BL21(DE3) E. coli strains by growing the cultures at 37 °C to an OD600 of 0.8, inducing with 1 mM IPTG and harvesting after 4 h. [U-15N,13C]-labeling was obtained using 13C-glucose and 15NH4Cl. The cells were harvested by centrifugation and the pellet was resuspended in 50 mM phosphate buffer pH = 7.5, 300 mM NaCl, 0.1 mM Complete protease inhibitor (Roche). The obtained cells were incubated for 1 h at room temperature with DNase I and then sonicated on ice and centrifuged at 13,000 rpm at 4 °C for 30 min. The pellet was resuspended in 50 mM phosphate buffer pH = 7.5, 300 mM NaCl, 6 M guanidinium hydrochloride and centrifuged at 13,000 rpm at 4 °C for 30 min. The supernatant was incubated for 2 h at room temperature with pre-equilibrated Ni-NTA affinity resin (Thermo Scientific) and than passed through a plastic body column for gravity flow purification. The column was washed with 20 column volumes of resuspension buffer containing 20 mM imidazole and eluted with 3 column volumes of the same buffer with 500 mM imidazole. All these purification steps were carried out at 4 °C. Filaments were prepared by slow dilution into aqueous solution. The elution fraction from the NiNTA column was dialyzed overnight against 25 mM phosphate buffer pH = 7.5, 100 mM NaCl. The solution was then centrifuged at 13,000 rpm at 4 °C for 30 min yielding a gel-like pellet of ASC filaments that was resuspended in water for a wash step and stored at 4 °C. This same protocol also yielded samples for EM studies. Details thereof are discussed in a forthcoming publication.
NMR sample preparation
The filaments so obtained were packed into 3.2 mm ZrO2 rotors (Bruker Biospin) by ultracentrifugation using a homemade filling device (Böckmann et al. 2009) in a SW41-T1 swing-out rotor in an optima L90-K ultracentrifuge (Beckmann). The sample was spun for 12 h, setting the spinning to 25,000 rpm at 4 °C. The drive tip of the rotor was finally sealed with epoxy glue (Araldit® blue) in order to prevent the dehydration of the samples.
Solid-state NMR spectroscopy
3D experiments were performed for the assignment, namely NCACB, NCACX, NCOCX and CANCO, as well as the relayed N(CO)CACB and CAN(CO)CA experiments for the assignment of the backbone and Cβ atoms of the protein (Schuetz et al. 2010) 2D NCA and DARR were used to check assignment consistency. Additional 3D CCC, N(CA)CBCX and 2D DARR were performed to continue the assignment of the side chains. All 3D spectra were recorded for 2–6 days each (Table 1). 20 ms DARR spectra on ASC-PYD and ASC full-length filaments were recorded for comparison.
All spectra were measured on a Bruker Avance II+ 850 MHz operating at a static magnetic field of 20 T. Experiments were performed with magic angle spinning (MAS) at 17 kHz with a Bruker 3.2 mm probe. The sample temperature was 12 °C, as determined from the water resonance (Gottlieb et al. 1997). Processing was done using TopSpin 3.1 (Bruker Biospin) by zero filling to no less than double the number of acquired points. The time domain was apodized using a squared cosine function (SSB 2.2–2.6). The spectra were analyzed and annotated with ccpNMR analysis 2.3 software (Fogh et al. 2002; Vranken et al. 2005; Stevens et al. 2011).
Secondary-structure analysis
Secondary chemical shifts were obtained by subtracting the random-coil shifts, as tabulated in (Wang and Jardetzky 2002). The analysis was done using the program TALOS+ (Shen et al. 2009).
Assignment and data deposition
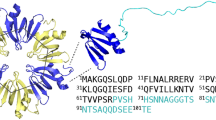
The [13C–13C] and [15N–13C] correlation NMR spectra of the mouse ASC PYD filaments feature narrow lines (line width of cross peaks on the order of 0.5 ppm), which is a clear indication of a homogeneous sample. The filaments showed no polymorphism, as shown in Fig. 1: all the peaks present in the 20 ms DARR (Fig. 1a) and in the NCA (Fig. 1b) were assigned to a single nucleus, with the exception of two additional weak threonine signals present in the DARR. Most likely, these two signals arise from Thr 87 and Thr 88, but no nitrogen resonance and no connectivity information could be obtained from the different spectra. Figure 2 compares the 20 ms DARR (Fig. 2a) and the NCA (Fig. 2b) measured on the filaments of the isolated PYD (blue) with the full length ASC (red). The very good coincidence of the resonances in these spectra indicates that the rigid part of the filaments is composed only by the PYD, while the CARD shows dynamic behavior. Moreover, the presence of the CARD does not influence the structure of the PYD in the filaments. All measurements for sequential assignments were thus performed on samples of filaments of the isolated ASC PYD. Using 3D methods (Schuetz et al. 2010; Habenstein et al. 2011), all backbone atoms from residues 3–85 could be assigned. The relayed 3D experiments where no evolution of the C′ was recorded [e.g. N(CO)CACB] were fundamental to reduce the ambiguities in some stretches of the protein because of the larger shift dispersion of the Cβ compared to C′. For the CO-CA transfer, whose efficiency is important for these experiments, we have used the efficient band selective homonuclear CO-CA cross polarization (Chevelkov et al. 2013). In Fig. 3, a representative strip plot of the sequential assignment is shown, using a combination of the relayed 3D experiments with an NCACO, in which equally side-chain resonances can be observed. A total of about 80 % of all side-chain carbon nuclei were assigned with additional 2D and 3D spectra, including CCC (Fig. 4). The non-assigned nuclei were either not visible in the spectra due to local dynamics (as for the side chains of 5 Arg, 25 Met Cε, 70 Met Cε) or not assignable due to spectral overlap (as for 6, 51, 54 Asp Cε). The chemical shifts have been deposited in the BMRB database under the accession number 26550. The first three and the last six residues were only visible in some of the dipolar-transfer-based NMR spectra and could not be sequentially assigned with certainty. We performed an INEPT-type [15N–1H] spectrum of the ASC PYD filaments to investigate the flexibility of these stretches, but it was devoid of peaks (data not shown). We concluded that these residues experience dynamics on the intermediate time scale, and are thus line-broadened below the detection limit using either solution- or solid-state NMR methods. In Fig. 5, an overview of the assignment is shown. The secondary chemical shifts derived from the chemical shifts reported here using a TALOS+ analysis is shown in Fig. 6, and points to the presence of six α-helices, at positions 3–14, 17–29, 41–46, 49–59, 62–76, and 80–84. These locations are similar to those observed in the monomeric human ASC PYD using solution-state NMR, where helices were defined including residues 3–14, 17–26, 41–46, 49–59, 62–76, 80–89 (Liepinsh et al. 2003).
Solid-state NMR spectra of uniformly 13C, 15N labeled mouse ASC-PYD filaments acquired at a magnetic field of 20 T and MAS at a frequency of 17 kHz. a 2D 13C–13C DARR spectrum with a mixing time of 20 ms. b NCA spectrum. The sequence-specific assignments indicated were derived from the sequential backbone walk in 3D spectra as shown in Fig. 3
Strips extracted from the 3D CCC spectrum of ASC-PYD filaments, used for the assignment of the sidechains. In a CCC spectrum, polarization transfers are selected to mainly display Cα resonances in the first dimension, Cβ in the second and the remaining side chains in the third dimension. In a aromatic sidechains for Tyr 59 are shown, in b sidechains for different Ile and in c for Leu 32
Secondary chemical shifts analysis of labeled filaments of ASC-PYD. For glycine ∆δCα is plotted and the corresponding bars are plotted in gray. Four positive secondary shifts in a row are indicative of α-helix structure. Blue squares are drawn for TALOS+ predicted α-helix regions. Only residues without warning and with small dispersion of predictions were included
The secondary-structure elements determined by the secondary chemical shifts will be used in a forthcoming publication, in combination with high-resolution cryo electronmicroscopy data, to determine a de-novo structure of the ASC PYD filament.
References
Böckmann A, Gardiennet C, Verel R, Hunkeler A, Loquet A, Pintacuda G, Emsley L, Meier BH, Lesage A (2009) Characterization of different water pools in solid-state NMR protein samples. J Biomol NMR 45:319–327
Chevelkov V, Shi C, Fasshuber HK, Becker S, Lange A (2013) Efficient band-selective homonuclear CO-CA cross-polarization in protonated proteins. J Biomol NMR 56:303–311
de Alba E (2009) Structure and interdomain dynamics of apoptosis-associated speck-like protein containing a CARD (ASC). J Biol Chem 284:32932–32941
Fogh R, Ionides J, Ulrich E, Boucher W, Vranken W, Linge JP, Habec M, Rieping W, Bhat TN, Westbrook J, Henrick K, Gilliland G, Berman H, Thornton J, Nilges M, Markley J, Laue E (2002) The CCPN project: an interim report on a data model for the NMR community. Nat Struct Biol 9(6):416–418
Gottlieb H, Kotlyar V, Nudelman A (1997) NMR chemical shifts of common laboratory solvents as trace impurities. J Org Chem 62:7512
Habenstein B, Wasmer C, Bousset L, Sourigues Y, Schütz A, Loquet A, Meier BH, Melki R, Böckmann A (2011) Extensive de novo solid-state NMR assignment of the 33 kDa C-terminal domain of the Ure2 prion. J Biomol NMR 51:235–243
Hu Z, Yan C, Liu P, Huang Z et al (2013) Crystal structure of NLRC4 reveals its autoinhibition mechanism. Science 341:172–175
Liepinsh E, Barbals R, Dahl E, Sharipo A et al (2003) The death-domain fold of the ASC PYRIN domain, presenting a basis for PYRIN/PYRIN recognition. J Mol Biol 332:1155–1163
Lu A, Magupalli VG, Ruan J, Yin Q et al (2014) Unified polymerization mechanism for the assembly of ASC-dependent inflammasomes. Cell 156:1193–1206
Masumoto J, Taniguchi S, Ayukawa K, Sarvotham H et al (1999) ASC, a novel 22-kDa protein, aggregates during apoptosis of human promyelocytic leukemia HL-60 cells. J Biol Chem 274:33835–33838
Moriya M, Taniguchi S, Wu P, Liepinsh E et al (2005) Role of charged and hydrophobic residues in the oligomerization of the PYRIN domain of ASC. Biochemistry 44:575–583
Schroder K, Tschopp J (2010) The inflammasomes. Cell 140:821–832
Schuetz A, Wasmer C, Habenstein B, Verel R, Greenwald J, Riek R, Böckmann A, Meier BH (2010) Protocols for the sequential solid-state NMR assignment of a uniformly labelled 25 kDa protein: HET-s(1-227). Chembiochem 11:1543–1551
Shen Y, Delaglio F, Cornilescu G, Bax A (2009) TALOS+: a hybrid method for predicting protein backbone torsion angles from NMR chemical shifts. J Biomol NMR 44:213–223
Srimathi T, Robbins SL, Dubas RL, Chang H et al (2008) J Biol Chem 283:15390–15398
Stevens TJ, Fogh RH, Boucher W, Higman VA, Eisenmenger F, Bardiaux B, van Rossum B-J, Oschkinat H, Laue ED (2011) A software framework for analysing solid-state MAS NMR data. J Biomol NMR 51(4):437–447
Vajjhala PR, Mirams RE, Hill JM (2012) Multiple binding sites on the pyrin domain of ASC protein allow self-association and interaction with NLRP3 protein. J Biol Chem 287:41732–41743
Vajjhala PR, Kaiser S, Smith SJ, Ong QR et al (2014) Crystal structure of NALP3 protein pyrin domain (PYD) and its implications in inflammasome assembly. J Biol Chem 289:23504–23519
Vranken W, Boucher W, Stevens T, Fogh R, Pajon A, Llinas P, Ulrich E, Markley J, Ionides J, Laue E (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59(4):687–696
Wang Y, Jardetzky O (2002) Probability-based protein secondary structure identification using combined NMR chemical-shift data. Protein Sci 11(4):852–861
Acknowledgments
This work was supported SNF 200020_146757, 200020_159707 and ANR-12-BS08-0013-01, ANR-14-CE09-0024B.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Ravotti, F., Sborgi, L., Cadalbert, R. et al. Sequence-specific solid-state NMR assignments of the mouse ASC PYRIN domain in its filament form. Biomol NMR Assign 10, 107–115 (2016). https://doi.org/10.1007/s12104-015-9647-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-015-9647-6