Abstract
Parkinson’s disease is amongst the most frequent and most devastating neurodegenerative diseases. It is tightly associated with the assembly of proteins into high-molecular weight protein species, which propagate between neurons in the central nervous system. The principal protein involved in this process is α-synuclein which is a structural component of the Lewy bodies observed in diseased brain. We here present the solid-state NMR sequential assignments of a new fibrillar form of this protein, the first one with a well-ordered and rigid N-terminal part.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Biological context
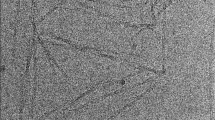
α-Synuclein is one of the major proteins involved in Parkinson’s disease. Aggregates of this normally soluble protein are found in Lewy Bodies which are associated to this disability (Brundin et al. 2010; Eller and Williams 2011). The superstructures of these aggregates, as seen by electron microcopy, are protein fibrils; the aggregation mechanism of α-synuclein, as well as the structure of the resulting fibrils, remain however unknown. Structural studies of α-synuclein fibrillar assemblies are challenging today as neither of the classical methods which allow to study protein structures at a molecular level, as X-ray crystallography or NMR in solution, are adequate tools for this task. Solid-state NMR has recently developed to become a powerful tool to reveal conformational details and atomic-resolution structures of insoluble and non-crystalline molecular assemblies (Van Melckebeke et al. 2010; Böckmann and Meier 2010; Renault et al. 2010; Debelouchina et al. 2010; Jehle et al. 2010; Wasmer et al. 2008). Initial structural studies by solid-state NMR of α-synuclein fibrils, or fragments thereof, have been performed by several groups (Heise et al. 2005, 2008; Kloepper et al. 2006, 2007; Vilar et al. 2008; Loquet et al. 2010) and 48 (form A) and 23 (form B) residues were sequentially assigned in ref (Heise et al. 2005) and 26 in ref (Kloepper et al. 2007). A comparison of the published spectra reveals the appearance of different polymorphs, as well as significant heterogeneity, in some individual samples indicative of differences in the conformation and packing of α-synuclein molecules within or between the different fibrils studied. Sample preparation is a crucial step in fibril structural studies, as structural differences or polymorphs can arise from different production, purification and assembly conditions. Extreme care has to be exercised when comparing results from samples of different sources or even different batches. We here present the sequential resonance assignments of an α-synuclein polymorph for which for the first time the N-terminal residues, postulated not to be part of the core in other studies (Vilar et al. 2008; Miake et al. 2002), are assigned and shown to be mainly in β-sheet conformation. This polymorph might show similarities to the B-form described by Heise et al. (2005) since for 8 residues the chemical shift assignments coincide with the here presented ones. The spectra also overlay well with those published for a yet unassigned polymorph (Loquet et al. 2010).
Methods and experiments
Protein expression and purification
Full-length a-synuclein was expressed in E. coli BL21 DE3 codon + cells (Stratagene). Cells were grown in LB medium to an Abs600nm of 0.8 and sedimented at 3,000g for 10 min in 1 l tubes. The pelleted bacteria were washed with 200 ml of M9 salt and spun at 3,000g for 10 min. The bacterial pellets were resuspended in half the volume where they originally grew of M9 media containing 1.75 g of 15NH4Cl, 2.5 g of U-13C glucose, 2 mM MgSO4, 0.1 mM CaCl2, 10 μg thiamine per litre of culture. Cells were grown for 30 min at 37°C and α-synuclein expression was induced by 0.5 mM IPTG for 3 h. The cells were then harvested by centrifugation (4,000g, 10 min), resupended into lysis buffer (10 mM Tris pH 7.5, 1 mM EDTA, 1 mM PMSF) and lysed by sonication. Cell extracts were clarified by centrifugation at 14,000g, 30 min. Purification was performed as previously described (Hansen et al. 2011).
Sample preparation
α-Synuclein was dialyzed 16 h against 5 mM Tris–HCl pH 7.5 in double-distilled H2O at 8°C. Fibrillation was achieved at 300 μM by shaking 1 ml solution aliquots at 37°C in an Eppendorf Thermomixer set at 600 rpm with a 3 mm orbital for 7 days. Fibrils were spun for 20 min at 20° C, 50,000 rpm on a TL100 ultracentrifuge (Beckman) using the TLA 100.4 rotor. The pelleted material was used to fill a ZrO2 3.2 mm rotor (Bruker) using a home made filling device (Böckmann et al. 2009) spun at 40,000 rpm for 16 h at 18°C in an SW60 TI rotor and an optima L90-K ultracentrifuge (Beckman).
NMR spectroscopy
We used a suite of 2D and 3D experiments, namely 2D DARR and DREAM, and 3D NCACB, CAN(CO)CA, NCOCA, CANCO, NCACO and CCC experiments to perform the assignment as described in detail in references (Habenstein et al., accepted, Schuetz et al. 2010) for sequential assignments. The sequential walk was achieved by connecting resonances from NCACO/NCACB, CANCO/CAN(CO)CA and NCOCA spectra. Sequential connections were also present in CCC spectra. Side-chain assignments were done using NCACB and CCC spectra. Full experimental details are given in Table S1. All assignment spectra were recorded on the same rotor. However, 2D DARR spectra were taken of four different preparations to check reproducibility of the sample preparation. All preparations yielded identical spectra.
Assignment and data deposition
The α-synuclein sample studied here reveals a well-resolved 2D 13C–13C spectrum with narrow lines, shown in Fig. 1 (extract of the aliphatic and carbonyl regions). Assignments are given for crosspeaks corresponding to directly bonded carbons (in the aliphatic region only below the diagonal). A representative plane of the 3D NCACB, NCACO, CANCO and NCOCA experiments is shown in Fig. 2. The observed line width in the spectra is mostly comprised between 0.5 and 1 ppm. The lines are not quite as narrow as in HET-s(218–289) (Siemer et al. 2005), but allow for an assignment using the afore mentioned 3D methods. Sequential assignments could be achieved for 85% of residues 1–97 and for 74% of all carbon atoms and 82% backbone amide nitrogen atoms. The chemical shifts have been deposited in the BMRB under the accession number 17498. The C-terminal portion including residues 98–140 seems to be flexible, as no signals could be attributed to this region. However, all amino-acid types present in the C-terminal region could be observed in the scalar-coupling based 1H–13C INEPT spectrum (see Supplementary Figure S1). For the stretch ranging from residue 44–57, unambiguous sequential assignments were difficult due to lower signal/noise or spectral overlap. Figure S2 compares the intensities of resonances of some assigned residues with “weak” peaks tentatively assigned to residues in this stretch. Tentative assignments are not considered in the following and were not submitted to the BMRB.
2D 13C–13C DARR Spectrum of α-synuclein fibrils recorded with a mixing time of 20 ms at a magnetic field of 14.1 Tesla. The data was zero-filled and apodized in both dimensions using a 0.3 shifted squared sine bell function. The spectrum was processed using NMRPipe (Delaglio et al. 1995). All one-bond correlations are labeled (in the aliphatic region only below the diagonal)
Planes of 3D NCACO, CANCO, NCOCA and NCACB spectra. For experimental details see Table S1. The data were analyzed using the CCPN software (Fogh et al. 2002). Assignments of the peaks are given. Grey labels mean that the maximum of the corresponding peak is at a different N-shift, therefore leading to weaker signals in the plane presented
Assigned atoms are shown in the assignment graph in Figure S3. Most resolved and strong cross peaks in a 20 ms 2D DARR spectrum can be explained based on this assignment. Most weaker peaks in this spectrum are due to sequential contacts and can also be explained by the assignments.
The secondary chemical shifts for the assigned residues are given in Fig. 3. One can see that the protein mostly displays secondary chemical shifts typical of β-sheet conformation. The assigned regions do not correspond to the regions previously assigned by (Heise et al. 2005, 2008) for both A- and B-forms, who found the first 37 N-terminal residues statically disordered. Here, we observe most of the N-terminal residues of the protein up to residue 97, with an exception being residues 44–57, which are potentially less structurally homogeneous or more mobile. The structured regions observed in our sample correspond however to the regions identified in the presence of detergent to be α-helical (Bussell and Eliezer 2003). In the moment we prefer not to add another structural model to the flurry of already existing ones, but note that the sequential assignment of α-synuclein presents a first step towards a high-resolution structure of this protein, which seems perfectly feasible.
Secondary chemical shifts (Wishard and Sykes 1994) of the sequentially assigned residues of α-synuclein. The grey bars for residues 32–34 stand for the secondary chemical shifts of the doubled stretch. Three negative shifts in a row are indicative for β-sheet secondary structure
References
Böckmann A, Meier BH (2010) Prions: en route from structural models to structures. Prion 4:72–79
Böckmann A, Gardiennet C, Verel R, Hunkeler A, Loquet A, Pintacuda G, Emsley L, Meier BH, Lesage A (2009) Characterization of different water pools in solid-state NMR protein samples. J Biomol NMR 45:319–327
Brundin P, Melki R, Kopito R (2010) Prion-like transmission of protein aggregates in neurodegenerative diseases. Nat Rev Mol Cell Biol 11:301–307
Bussell R Jr, Eliezer D (2003) A structural and functional role for 11-mer repeats in a-synuclein and other exchangeable lipid binding proteins. J Mol Biol 329:763–778
Debelouchina GT, Platt GW, Bayro MJ, Radford SE, Griffin RG (2010) Intermolecular alignment in beta(2)-microglobulin amyloid fibrils. J Am Chem Soc 132:17077–17079
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Eller M, Williams DR (2011) Review: alpha-synuclein in Parkinson disease and other neurodegenerative disorders. Clin Chem Lab Med 49:403–408
Fogh R, Ionides J, Ulrich E, Boucher W, Vranken W, Linge JP, Habeck M, Rieping W, Bhat TN, Westbrook J, Henrick K, Gilliland G, Berman H, Thornton J, Nilges M, Markley J, Laue E (2002) The CCPN project: an interim report on a data model for the NMR community. Nat Struct Biol 9:416–418
Habenstein B, Wasmer C, Bousset L, Sourigues Y, Schütz A, Loquet A, Meier BH, Melki R, Böckmann A (accepted) De novo solid-state NMR sequential resonance assignments of the 33 kDa C-terminal domain of the Ure2 prion. J Biomol NMR
Hansen C, Angot E, Bergstrom AL, Steiner JA, Pieri L, Paul G, Outeiro TF, Melki R, Kallunki P, Fog K, Li JY, Brundin P (2011) alpha-Synuclein propagates from mouse brain to grafted dopaminergic neurons and seeds aggregation in cultured human cells. J Clin Invest 121:715–725
Heise H, Hoyer W, Becker S, Andronesi OC, Riedel D, Baldus M (2005) Molecular-level secondary structure, polymorphism, and dynamics of full-length alpha-synuclein fibrils studied by solid-state NMR. Proc Natl Acad Sci USA 102:15871–15876
Heise H, Celej MS, Becker S, Riedel D, Pelah A, Kumar A, Jovin TM, Baldus M (2008) Solid-state NMR reveals structural differences between fibrils of wild-type and disease-related A53T mutant alpha-synuclein. J Mol Biol 380:444–450
Jehle S, Rajagopal P, Bardiaux B, Markovic S, Kuhne R, Stout JR, Higman VA, Klevit RE, van Rossum BJ, Oschkinat H (2010) Solid-state NMR and SAXS studies provide a structural basis for the activation of alphaB-crystallin oligomers. Nat Struct Mol Biol 17:1037–1042
Kloepper KD, Woods WS, Winter KA, George JM, Rienstra CM (2006) Preparation of alpha-synuclein fibrils for solid-state NMR: expression, purification, and incubation of wild-type and mutant forms. Protein Expr Purif 48:112–117
Kloepper KD, Hartman KL, Ladror DT, Rienstra CM (2007) Solid-state NMR spectroscopy reveals that water is nonessential to the core structure of alpha-synuclein fibrils. J Phys Chem B 111:13353–13356
Loquet A, Giller K, Becker S, Lange A (2010) Supramolecular interactions probed by 13C–13C solid-state NMR spectroscopy. J Am Chem Soc 132:15164–15166
Miake H, Mizusawa H, Iwatsubo T, Hasegawa M (2002) Biochemical characterization of the core structure of alpha-synuclein filaments. J Biol Chem 277:19213–19219
Renault M, Cukkemane A, Baldus M (2010) Solid-state NMR spectroscopy on complex biomolecules. Angew Chem Int Ed Engl 49:8346–8357
Schuetz A, Wasmer C, Habenstein B, Verel R, Greenwald J, Riek R, Böckmann A, Meier BH (2010) Protocols for the sequential solid-state NMR spectroscopic assignment of a uniformly labeled 25 kDa protein: HET-s(1–227). Chembiochem 11:1543–1551
Siemer AB, Ritter C, Ernst M, Riek R, Meier BH (2005) High-resolution solid-state NMR spectroscopy of the prion protein HET-s in its amyloid conformation. Angew Chem Int Ed Engl 44:2441–2444
Van Melckebeke H, Wasmer C, Lange A, Ab E, Loquet A, Böckmann A, Meier BH (2010) Atomic-resolution three-dimensional structure of HET-s(218–289) amyloid fibrils by solid-state NMR spectroscopy. J Am Chem Soc 132:13765–13775
Vilar M, Chou HT, Luhrs T, Maji SK, Riek-Loher D, Verel R, Manning G, Stahlberg H, Riek R (2008) The fold of alpha-synuclein fibrils. Proc Natl Acad Sci USA 105:8637–8642
Wasmer C, Lange A, Van Melckebeke H, Siemer AB, Riek R, Meier BH (2008) Amyloid fibrils of the HET-s(218–289) prion form a beta solenoid with a triangular hydrophobic core. Science 319:1523–1526
Wishard DS, Sykes BD (1994) The 13C chemical-shift index: a simple method for the identification of protein secondary structure using 13 chemical-shift data. J Biomol NMR 4:171–180
Acknowledgments
This work was supported by the Agence Nationale de la Recherche (ANR-07-PCVI-0013-03, ANR-PCV08 321323 and ANR08-PCVI-0022-02), the ETH Zurich, the Swiss National Science Foundation (Grant 200020_124611), the Era-Net Neuron (project MIPROTRAN, ANR-08-NEUR-001-01) and the Centre National de la Recherche Scientifique. We also acknowledge support from the European Commission under the Seventh Framework Programme (FP7), contract Bio-NMR 261863.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Julia Gath, Birgit Habenstein and Luc Bousset have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gath, J., Habenstein, B., Bousset, L. et al. Solid-state NMR sequential assignments of α-synuclein. Biomol NMR Assign 6, 51–55 (2012). https://doi.org/10.1007/s12104-011-9324-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-011-9324-3