Abstract
An in planta transformation protocol for sorghum (Sorghum bicolor (L.) Moench) using shoot apical meristem of germinating seedlings is reported in this study. Agrobacterium tumefaciens strain, LBA4404 with pCAMBIA1303 vector and construct pCAMBIA1303TPS1 were individually used for transformation. Since, the transgene is integrated into the cells of already differentiated tissues, the T0 plants were chimeric and stable integration was observed in T1 generation. ß-Glucuronidase (GUS) expression in the seedlings and spikelets of emerging cob was the first indication of transformability in T0 generation which was further confirmed by PCR analysis using hpt and TPS1 gene-specific primers. Screening on 25 mg/L hygromycin combined with PCR analysis was used for selection of transformants in the T1 generation. Transformation efficiencies ranged between 34–38% and 26–34% using pCAMBIA1303 vector and construct pCAMBIA1303TPS1, respectively. Molecular characterization of the T2 transgenics using PCR, RT-PCR and Southern blot analyses further revealed the integration, expression and inheritance of the transgene. These results indicate the feasibility of the method to generate transgenics with pCAMBIA1303 vector and construct pCAMBIA1303TPS1. The abiotic stress tolerance of TPS1 transgenics developed in the present study was evident by the ability of the transformants to tolerate 200 mM NaCl as well as higher root growth and biomass.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sorghum (Sorghum bicolor (L.) Moench) is the fifth most important cereal crop in the world. It is cultivated mostly in semi-arid tropics which are characterized by unpredictable weather, limited and erratic rainfall, high temperatures and marginal lands with problems like salinity and nutrient deficiency (Sharma and Ortiz 2000; Maqbool et al. 2001). Traditional breeding has been the main avenue for crop improvement for many years in sorghum, but the success achieved so far is marginal (Girijashankar and Swathisree 2009). Molecular approaches can thus be used to complement genetic engineering as they offer direct access to a vast pool of useful genes (Tari et al. 2013).
Sorghum has been considered as one of the plant species which is difficult to manipulate through tissue culture and transformation (Grootboom et al. 2010). Both, particle bombardment (Casas et al. 1997; Tadesse et al. 2003; Maheswari et al. 2010) and Agrobacterium-mediated methods for genetic transformation of sorghum (Zhao et al. 2000; Howe et al. 2006; Liu and Godwin 2012) have been reported but with low transformation efficiencies ranging from 0.18–20.7%. These methods suffer from inherent disadvantages, i.e. require sterile conditions, long period of culture, somaclonal variations etc. which are due to the in vitro culture of plant cells (Supartana et al. 2005). Therefore, in planta transformation methods, which require no in vitro culture of plant cells, have become important in recalcitrant crops like sorghum.
In planta transformation is a tissue culture independent approach to obtain fertile transformed plants. In planta transformation reduces time, labour, cost and most importantly, it avoids somaclonal variations (Jaganath et al. 2014). Several economically and agriculturally important crops have been transformed by in planta transformation, such as buckwheat (Kojima et al. 2000), mulberry (Ping et al. 2003) kenaf (Kojima et al. 2004), rice (Supartana et al. 2005) and wheat (Supartana et al. 2006) using binary vector. In all these crops, Agrobacterium is directed towards either the apical meristem or the meristems of axillary buds, allowing them to grow into seedlings ex vitro. It has been reported that the transformation efficiency obtained through in planta transformation is much higher than the conventional tissue culture based transformation (Supartana et al. 2006).
Synthesis of osmoprotectants, is one of the mechanisms that plants have evolved for adaptation to water deficit. Osmoprotectants include amino acids, quaternary ammonium and tertiary sulphonium compounds, polyols and sugars (Rontein et al. 2002). Metabolic engineering of osmoprotectants is a well-known strategy for conferring abiotic stress tolerance in crop plants. Trehalose, an osmoprotectant, a nonreducing disaccharide of glucose, plays an important physiological role as an abiotic stress protectant including bacteria, yeast, invertebrates and resurrection plants (Delorge et al. 2014). Trehalose, which is formed by the engineering of TPS1 gene has been shown to stabilize dehydrated enzymes, proteins and lipid membranes efficiently as well as to protect biological structures from damage during desiccation. In S. cerevisiae trehalose-6-phosphate synthase (TPS) encoded by gene TPS1 catalyzes the transfer of glucose from UDP-glucose to glucose-6-phosphate to produce trehalose-6- phosphate plus UDP. Subsequently, trehalose-6-phosphate phosphatase (TPP) encoded by the TPS2 gene converts trehalose-6-phosphate to free trehalose (Cabib and Leloir 1958).
The introduction of exotic TPS encoding gene can bring about synthesis of trehalose-6-phosphate. The dephosphorylation of trehalose-6-phosphate can be catalyzed by nonspecific phosphoesterase which is familiar in plant cells. Engineering trehalose accumulation into agronomically important plants is a potential approach to increase drought and salinity tolerance (Penna 2003). Transgenic plants over expressing genes coding for enzymes of the trehalose biosynthetic pathway with improved drought tolerance have been reported both in model plants like tobacco (Holmstrom et al. 1996; Romero et al. 1997; Varalaxmi et al. 2012) and crop plants such as potato (Yeo et al. 2000), rice (Garg et al. 2002) and tomato (Cortina and Culianez-Macia 2005).
The present study aims to develop an in planta transformation protocol for sorghum which is highly recalcitrant in nature. Earlier attempts to develop in planta transformation protocol in crop plants (Supartana et al. 2005, 2006) have focussed on optimization using binary vectors. In the present work, efforts have also been made to develop transgenics with TPS1 gene for enhancing tolerance to abiotic stresses.
Materials and methods
Bacterial strain and plasmid constructs
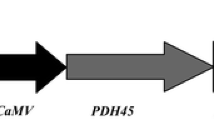
Agrobacterium tumefaciens strain LBA4404 harbouring pCAMBIA1303 vector and pCAMBIA1303TPS1 were used individually. pCAMBIA1303 is a binary vector having hygromycin resistance gene (hpt) as plant selection marker and ß-glucuronidase (gusA) and green fluorescent protein (mgfp5) as reporter genes. The TPS1 gene amplified from the strain MTCC no.174 was cloned into pTZ57R/T vector. The vector was digested with KpnI, BamHI and the resulting fragment was then sub-cloned into pRT100 for mobilization of promoter and terminator. The gene cassette was cloned into plant expression vector pCAMBIA1303 (Varalaxmi et al. 2012) and used for transformation experiments.
Transformation and generation of transgenics
One hundred µL culture of A. tumefaciens LBA4404 harbouring the vector pCAMBIA1303 and construct pCAMBIA1303TPS1 individually (grown for 14 h at 28°C) was inoculated into 10 mL YEM broth containing kanamycin (50 µg/mL) and grown at 28°C until an OD 600 of 0.8 was achieved. The cells were then centrifuged (5000 rpm/RT/10 min) and resuspended in 500 µL of fresh YEM medium and supplemented with acetosyringone (100 µM).
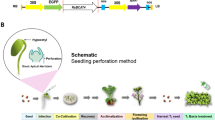
Sorghum seeds (S. bicolor (L.) Moench) cv. SPV462 were washed with Tween-20, rinsed 2–3 times with distilled water and then surface sterilized with 0.1% HgCl 2 for 10 min. Subsequently, seeds were washed 5–6 times with sterile distilled water and were aseptically germinated on wet cotton in culture bottles (50 seeds per bottle) in dark at 27°C for two days. At this stage, the meristem explants were around 2–3 mm and the root and shoot had emerged. Inoculation of the meristematic region with the resuspended Agrobacterium culture was carried out by using 1 mL sterile hypodermic syringes (1 mL B.D. disposable syringes, with 0.9 mm diameter needle). The inoculated seedlings were transferred to culture bottles layered with cotton at 15 seedlings per bottle and maintained at 27°C in light (85 µmol/m 2/s), under a 12 h photoperiod and were allowed to cocultivate for five days. Subsequently, the seedlings were immersed in sterile water containing 250 mg/L cefotaxime for 30 min with continuous shaking to wash off Agrobacterium cells, transferred to 16 cm diameter pots containing cocopit (10–15 seedlings per pot) under ambient conditions and allowed to grow for 15 days. The seedlings were then transferred into 30 × 30 cm plastic pots (three plants per pot) filled with 16 kg of sandy loamy soil (red soil : sand : farm yard manure in a ratio of 3 : 1 : 1) after acclimatization as mentioned above. The plants were selfed and grown to maturity (T0). Appropriate untransformed controls inoculated with water were maintained following all other procedures similar to transformants. Transformation experiments were carried out with pCAMBIA1303 vector and pCAMBIA1303TPS1, with 150 and 300 explants, respectively, and a few of the explants were subjected to GUS histochemical assay.
Characterization of T 0 transgenics
Transient assay for GUS gene expression:
Histochemical assay for transient GUS expression was performed according to Jefferson et al. (1987). Transient GUS expression was carried out in seedlings just before transferring it to small pots. The explants were incubated overnight at 37°C in a solution containing 50 mM sodium phosphate buffer pH 7.0, 0.1% Triton X-100, 10 mM EDTA and 1 mM 5-bromo-4- chloro-3-indolyl ß-D-glucuronide. Stable GUS expression was monitored in the spikelets of emerging cobs.
PCR analysis:
Leaf sample collected at the reproductive stage was used for molecular analysis. Genomic DNA was isolated following CTAB method (Doyle and Doyle 1990). PCR analysis was carried out on putative transgenics (T0) with pCAMBIA1303 vector using hpt gene-specific primers and pCAMBIA1303TPS1 using hpt and TPS1 gene-specific primers. PCR amplification was carried out using Gene AmpR PCR system 9700 (Applied Biosystems, Foster City, USA). The thermocycling conditions were initial denaturation at 94°C for 5 min followed by 30 cycles of 94°C for 30 s, 60°C for 30 s and 72°C for 45 s for hpt and 94°C for 30 s, 55°C for 30 s and 72°C for 45 s for TPS1 gene-specific primers and final extension at 72°C for 5 min. Genomic DNA from the untransformed control plant and plasmids pCAMBIA1303 and pCAMBIA1303TPS1 were used as negative and positive controls, respectively.
Characterization of T 1 transgenics
Screening on hygromycin and PCR analysis:
50 and 35 seeds from each of the selected three independent T0 transformants of pCAMBIA1303 vector (V4, V6 and V8) and pCAMBIA1303TPS1 (L1, L2 and L3) respectively, along with untransformed control were used. The seeds were surface sterilized and germinated on half-strength MS basal medium supplemented with 25 mg/L hygromycin, a concentration which is sublethal for control sorghum seeds (Maheswari et al. 2010). The plants which survived hygromycin selection pressure up to nine days were transferred to pots. DNA isolated from these plants was used for PCR analysis with hpt and TPS1 gene-specific primers, respectively.
Characterization of T 2 transgenics
PCR analysis:
The inheritance of the transgene in the progeny of selected three T 1 transformants of pCAMBIA1303 vector and pCAMBIA1303TPS1 was carried out using hpt and TPS1 gene-specific primers, respectively.
RT-PCR analysis
Two lines from each of the three independent transgenics of pCAMBIA1303 vector (V4-3-1, V4- 4-4, V6-5-3, V6-7-7, V8-5-2, V8-5-10) and pCAMBIA1303 TPS1 (L1-1-6, L1-2-9, L2-1-3, L2-2-4, L3-1-5, L3-2-7) were selected and total RNA was isolated using Trizol method (Chomczynski 1993). First strand cDNA synthesis was carried out using Qiagen First strand cDNA synthesis kit according to the manufacturer’s instructions. About 5 µg of total RNA was used for first-strand synthesis by incubating the reaction mixture at 37°C for 1 h. Second-strand synthesis was carried out using hpt and TPS1 gene-specific primers.
Southern blot analysis:
Southern blot analysis was carried out according to Sambrook and Russel (2001). The PCR amplified products of selected individual lines of three independent transgenics (T 2) of pCAMBIA1303 vector (V4-3-1, V6-5-3 and V8-5-2) and pCAMBIA1303TPS1 transgenics (L1-1-6, L2-1-3 and L3-1-5) using hpt and TPS1 gene-specific primers respectively were separated on a 0.8% agarose gel and were transferred onto the nylon membrane by capillary method. Hybridzation was performed at 55°C, overnight in a hybridization oven HB1000 hybridizer (UVP, Upland, USA). The 800 bp and 1431 bp hpt and TPS1 gene products were used as probes. Labelling and detection was carried out using Alkaphos Direct Labelling and Detection Kit (GE healthcare cat. no. RPN3680) according to the manufacturer’s instructions. GUS expression analysis in spikelets of the emerging cob of sorghum TPS1 transgenics (T 2) was also carried out.
Seed germination on 200 mM NaCl
Tolerance of sorghum TPS1 transgenics (T 1) to NaCl (200 mM) was tested. The seeds of TPS1 transgenics and untransformed control were surface-sterilized and inoculated onto MS basal medium supplemented with 200 mM NaCl. Visual observations on germination as well as shoot and root growth were recorded 15 d after inoculation. The root growth of the transgenics and untransformed control were recorded at harvest.
Results
Development of transgenics
Results of the experiments conducted on in planta transformation, generation of transgenics and molecular confirmations for stable integration and expression are presented below. Out of the total 150 seedlings that were subjected to Agrobacterium inoculation using pCAMBIA1303 vector, 128 plants survived the inoculation process and were transferred to transgenic glass house. Among them, 80 plants survived, attained maturity and set seed. Out of 300 seedlings which were subjected to Agrobacterium inoculation using the TPS1 gene construct, 204 plants survived the inoculation process and were transferred to the green house. Among them, 105 plants survived, attained maturity and set seed normally.
The feasibility of gene integration by in planta transformation protocol was ascertained based on GUS histochemical assay and PCR analysis. The blue colouration in the inoculated seedlings (figure 1, a–c) and spikelets of emerging cob (figure 1, d–f) of the pCAMBIA1303 vector and pCAMBIA1303TPS1 transgenics upon histochemical assay suggested the introduction and expression of the GUS gene. No such colouration was observed in untransformed control. The lines showing stable GUS expression were checked by PCR analysis. PCR amplification of leaf genomic DNA isolated from sorghum T0 transformants with pCAMBIA1303 vector and pCAMBIA1303TPS1 using hpt gene-specific primers had amplified 800 bp band corresponding to hygromycin resistance gene (figure 2, a–b). The 1431 bp band amplified using TPS1 gene-specific primers in pCAMBIA1303TPS1 transgenics also confirmed the presence of TPS1 gene (figure 2c). Only those lines which were positive for above analysis were chosen for further study in the subsequent T 1 generation.
PCR analysis of sorghum transgenics (T0). (a) pCAMBIA1303 vector using hpt gene-specific primers. (b&c) With construct pCAMBIA1303TPS1 using hpt and TPS1 gene-specific primers. Lane 1: a&c, 1 kb ladder; b, 1 kb plus ladder. Lane 2, positive control plasmid; lane 3, untransformed control; lanes 4–13, putative T 0 transgenics.
Integration and expression of the transgene in T 1 plants
Of the 50 seeds from each of the three independent primary transformants of pCAMBIA1303 vector screened on 25 mg/L hygromycin resulted in plants germinating from 22 seeds of line V4, 24 seeds of line V6 and 20 seeds of line V8. PCR analysis of the surviving plants using hpt gene-specific primers revealed that 18, 17 and 19 plants from each line respectively were found to be positive (figure 3a). Of the 35 seeds from each of the three independent TPS1 transgenic plants (T0) tested, the number of surviving seedlings on hygromycin were L1, 15; L2, 14; L3, 16. The number of lines positive to PCR analysis using TPS1 gene- specific primers were as follows: L1, 12; L2, 9; L3, 10 (figure 3b). Only these plants were grown to maturity and seed was collected. The transformation efficiencies were calculated and are provided in table 1.
PCR analysis of the progeny of three independent sorghum transgenics (T 0). (a) pCAMBIA1303 vector transgenics (V4, V6 and V8) using hpt gene-specific primers. Lane 1, 1 kb ladder; lane 2, positive control plasmid; lane 3, untransformed control; lanes 4–16, T 1 transgenic lines. (b) pCAMBIA1303TPS1 transgenics (L1, L2 and L3) using TPS1 gene-specific primers. Lane 1, 1 kb ladder; lane 2, positive control plasmid; lane 3, untransformed control; lanes 4–10, T 1 transgenic lines.
Stability of transgene in T 2 plants
PCR analysis of pCAMBIA1303 vector and pCAMBIA1303- TPS1 transgenics (T 2) using hpt and TPS1 gene-specific primers amplified 800 and 1431 bp bands in all the lines tested confirming the stable integration of the hpt (figure 4a) and TPS1 genes (figure 4b). RT-PCR analysis further confirmed the integration and expression of hpt and TPS1 transgenes in all the selected lines (figure 5, a&b). No such amplification was found in untransformed control. Signal generated at the expected 800 and 1431 bp positions by Southern hybridization further confirmed the transgenic nature of pCAMBIA 1303 vector and pCAMBIA1303TPS1 transgenics, respectively (figure 5, c&d). A vector diagram showing various steps involved in in planta transformation protocol has been shown in figure 6.
PCR analysis of the sorghum transgenics (T2). (a) pCAMBIA1303 vector transgenics (V4, V6 and V8) using hpt gene-specific primers. Lane 1, 1 kb ladder; lane 2, positive control plasmid; lane 3, untransformed control; lanes 4–15, T 2 transgenic lines. (b) pCAMBIA1303TPS1 transgenics (L1, L2 and L3) using TPS1 gene-specific primers. Lane 1, 1 kb ladder; lane 2, positive control plasmid; lane 3, untransformed control; lanes 4–9, T 2 transgenic lines of L1; lanes 4–10, T 2 transgenic lines of L2; lanes 4–11, T 2 transgenic lines of L3.
(a,b) RT-PCR analysis of sorghum transgenics (T2). (a) pCAMBIA1303 vector transgenics using hpt gene-specific primers. Lane 1, 1 kb ladder; lane 2, positive control plasmid; lane 3, untransformed control; lane 4, V4-3-1; lane 5, V4-4-4; lane 6, V6-5-3; lane 7, V6-7-7; lane 8, V8-5-2; lane 9, V8-5-10. (b) pCAMBIA1303TPS1 transgenics using TPS1 gene-specific primers. Lane 1, 1 kb ladder; lane 2, positive control plasmid; lane 3, untransformed control; lane 4, L1-1-6; lane 5, L1-2-9; lane 6, L2-1-3; lane 7, L2-2-4; lane 8, L3-1-5; lane 9, L3-2-7. (c,d) Southern analysis of sorghum transgenics (T2). (c) pCAMBIA1303 vector transgenics using hpt gene-specific primers. Lane 1, positive control plasmid; lane 2, V4-3-1; lane 3, V6-5-3; lane 4, V8-5-2. (d) pCAMBIA1303TPS1 transgenics using TPS1 gene-specific primers. Lane 1, positive control plasmid; lane 2, L1-1-6; lane 3, L2-1-3; lane 4, L3-1-5.
GUS gene expression in the spikelets of the emerging cob of TPS1 transgenics further supported the integration and expression of the transgene (figure 7). When subjected to 200 mM NaCl treatment, seed germination as well as root and shoot growth of seedlings was higher in TPS1 transgenics when compared to untransformed control (figure 8). At maturity, higher root growth and biomass were observed in TPS1 transgenics when compared with untransformed control (figure 9).
Discussion
Application of transformation technology to sorghum has been impeded by its recalcitrant nature and lower transformation efficiencies (Grootboom et al. 2010). In the present study, a tissue culture independent in planta transformation protocol was used to develop transformants. The most important requirement of the gene transfer approaches is high transformation frequency and efficient plant recovery which is low in case of conventional methods (Birch 1997). In the optimization of the in planta transformation protocol, the results indicated that the survival and recovery were higher in three-day-old explants compared to two-day-old explants. Further, a cocultivation period of five days allowed the optimal inoculation process and also helped in the establishment of the explants. No incubation of the explants in the Agrobacterium culture was carried out, as it is well known that with the increase in the infection time the explants turn brown and necrotic (Keshamma et al. 2008). Most of the explants survived the process of inoculation and a survival frequency of 85% was noted. About 80 primary transformants survived from a total of 150 seedlings. A success rate of 53% was achieved (figure 6). The developmental morphology of these plants was similar to that of seed-derived plants and they flowered normally and produced mature viable seeds.
In planta transformation targets the Agrobacterium to the wounded apical meristem of the differentiated seed embryo. It transfers the gene into the genome of diverse cells which are already destined to develop into specific organs and also to the meristematic cells still to be differentiated, hence, the T0 plants will be chimeric. The transgene integrated into the genome of the already differentiated cells could determine only the phenotype of the lower part of mature T0 plants and not that of the upper part of the mature plants. In contrast, transgene integrated into the genome of yet undifferentiated cell in the meristem, which was destined to become an apical meristem of mature plants could contribute to the production of a shoot apex and hence to the phenotype of mature T0 plants (Supartana et al. 2005). Thus, the transformants of the T1 and subsequent generation would be true transformants.
The analysis of T0 plants was carried out to assess the transformants. The blue colouration in the infected seedlings showed GUS expression indicating the transgene integration. GUS histochemical analysis of the reproductive structures can provide leads about the possibility of generating stable T1 transformants. GUS expression in the spikelets of the emerging cobs clearly indicated that some of the T1 seeds from these plants would produce stable transformants. PCR analysis of T0 plants using hpt and TPS1 gene-specific primers further supported the integration of the transgene.
As the number of primary transformants (T0) produced by this method is large, a highly efficient screening protocol is needed to select putative transformants. Screening of the putative transformants on selectable marker hygromycin (25 mg/L) revealed that 42–50% of the seedlings survived were transplanted to transgenic glass house. Further, these plants were analysed by PCR analysis using hpt and TPS1 gene-specific primers to eliminate the chances of escapes. About 34–38% and 26–34% of the seedlings confirmed gene integration in case of pCAMBIA1303 vector and pCAMBIA1303TPS1, respectively. Higher transformation efficiency in case of pCAMBIA1303 vector was observed when compared to pCAMBIA1303TPS1, indicating difficulty in generating the transgenics with gene construct. Similar in planta transformation efficiencies with binary vector based on PCR analysis has also been reported in other crops (Supartana et al. 2005, 2006).
In the present study, the transformation efficiencies obtained in sorghum (26–38%) are higher than that reported using conventional methods (up to 20.7%) such as particle bombardment (Casas et al. 1997; Tadesse et al. 2003; Maheswari et al. 2010) and Agrobacterium mediated approaches (Zhao et al. 2000; Howe et al. 2006; Liu and Godwin 2012). Also, most of the reports on genetic transformation in sorghum have used immature embryo explants which are season-dependent, difficult to handle and time-consuming (Gurel et al. 2009; Grootboom et al. 2010). During fertilization, in planta Agrobacterium-mediated transformation of flowering panicles has been reported by Elkonin et al. (2009) but only 1.4–2.2% survival on kanamycin has been reported.
The progeny of these plants in T2 generation showed the amplification of the transgene in all the selected plants showing transgene inheritance. RT-PCR and Southern analyses further confirmed the expression and integration of the transgene. GUS expression analysis of the spikelets of the emerging cobs of TPS1 transgenics further confirmed the inheritance of the transgene. The above analysis clearly confirmed in planta development of sorghum transgenics with pCAMBIA1303 vector and TPS1 gene.
In an effort to ascertain the role of TPS1 gene in conferring abiotic stress tolerance, seed germination assay of the sorghum TPS1 transgenics (T1) was carried out. Seed germination as well as root and shoot growth of seedlings were higher in transgenics of sorghum when challenged with 200 mM NaCl. At maturity, higher root growth and biomass of TPS1 transgenics were observed when compared to untransformed control. Higher root growth of TPS1 transgenics was earlier reported in tobacco (Romero et al. 1997; Varalaxmi et al. 2012), rice (Garg et al. 2002) and tomato (Cortina and Culianez-Macia 2005).
To the best of our knowledge, this is the first report of developing a tissue culture independent protocol for sorghum. In summary, here, we present the development of an efficient in planta transformation protocol for sorghum and also the successful generation of transgenics with TPS1 gene with enhanced abiotic stress tolerance.
References
Birch R. G. 1997 Plant transformation: problems and strategies for practical application. Ann. Rev. Plant Phys. Plant Mol. Biol. 48, 297–326.
Cabib E. and Leloir L. 1958 The biosynthesis of trehalose phosphate. J. Biochem. 231, 259–275.
Casas A. M., Kononowicz A. K., Haan T. G., Zhang L., Tomes D. T., Bressan R. A. and Hasegawa P. M. 1997 Transgenic sorghum plants obtained after microprojectile bombardment of immature inflorescences. In Vitro Cell Dev. Biol. Plant 33, 92–100.
Chomczynski P. 1993 A reagent for the single-step simultaneous isolation of RNA, DNA and proteins from cell and tissue samples. Biotechniques 15, 532–537.
Cortina C. and Culianez-Macia F. A. 2005 Tomato abiotic stress tolerance enhanced by trehalose biosynthesis. Plant Sci. 169, 75–82.
Delorge I., Janiak M., Carpentier S. and Dijck P. V. 2014 Fine tuning of trehalose biosynthesis and hydrolysis as novel tools for the generation of abiotic stress tolerant plants. Front. Plant Sci. 5, 1–9.
Doyle J. J. and Doyle J. L. 1990 Isolation of plant DNA from fresh tissue. Focus 12, 13–15.
Elkonin L. A., Ravin N. V., Leshko E. V., Volokhina I. V., Chumakov M. I. and Skryabin K. G. 2009 Inplanta agrobacterial transformation of sorghum plants. Biotekhnologiya 1, 23–30.
Garg A. K., Kim J. K., Ranwala A. P., Choi Y. D., Kochian L. V. and Wu R. J. 2002 Trehalose accumulation in rice plants confers high tolerance levels to different abiotic stresses. Proc. Natl. Acad. Sci. USA 99, 15898–15903.
Girijashankar V. and Swathisree V. 2009 Genetic transformation of Sorghum bicolor. Physiol. Mol. Biol. Plants 15, 1–15.
Grootboom A. W., Mkhonza N. L., O’Kennedy M. M., Chakauya E., Kunert K. and Chikwamba R. K. 2010 Biolistic mediated sorghum (Sorghum bicolor (L.) Moench) transformation via mannose and bialaphos based selection systems. Int. J. Bot. 6, 89–94.
Gurel S., Gurel E., Kaur R., Wong J., Meng L., Tan H. -Q. and Lemaux P. G. 2009 Efficient, reproducible Agrobacterium-mediated transformation of sorghum using heat treatment of immature embryos. Plant Cell Rep. 28, 429–444.
Holmstrom K. O., Maentyla E. E., Welin B., Mandal A., Palva E. T., Tunnela O. E. and Londesborough J. 1996 Drought tolerance in tobacco. Nature 379, 683–684.
Howe A., Sato S., Dweikat I., Fromm M. and Clemente T. 2006 Rapid and reproducible Agrobacterium-mediated transformation of sorghum. Plant Cell Rep. 25, 784–791.
Jaganath B., Subramanyam K., Subramanian M., Karthik S., Elayaraja D., Uday Kumar R. et al. 2014 An efficient inplanta transformation of Jatropha curcus (L.) and multiplication of transformed plants through in vivo grafting. Protoplasma 251, 591–601.
Jefferson R. A., Kavnagh T. A. and Bevan M. 1987 GUS fusions: ß-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J. 6, 3907–3910.
Keshamma E., Rohini S., Rao K. S., Madhusudan B. and Udaya kumar M. 2008 Tissue culture-independent inplanta transformation strategy: an Agrobacterium tumifaciens-mediated gene transfer method to overcome recalcitrance in cotton (Gossypium hirsutum L.) J. Cotton Sci. 12, 264–272.
Kojima M., Arai Y., Iwase N., Shiratori K., Shioiri H. and Nozue M. 2000 Development of a simple and efficient method for transformation of buckwheat plants (Fugopyrum esculentum) using Agrobacterium tumefaciens. Biosci. Biotechnol. Biochem. 64, 845–847.
Kojima M., Shioiri H., Nogawa M., Nozue M., Matsumoto D., Wada A. et al. 2004 Inplanta transformation of kenaf plants (Hibiscus cannabinus var. aokawa no.3) by Agrobacterium tumefaciens. J. Biosci. Bioeng. 98, 136–139.
Liu G. and Godwin I. D. 2012 Highly efficient sorghum transformation. Plant Cell Rep. 31, 32–37.
Maheswari M., Varalaxmi Y., Vijayalakshmi A., Yadav S. K., Sharmila P., Venkateswarlu B. et al. 2010 Metabolic engineering using mtlD gene enhances tolerance to water deficit and salinity in sorghum. Biol. Plant 54, 647–652.
Maqbool S. B., Devi P. and Sticklen M. B. 2001 Biotechnology: Advances for the genetic improvement of sorghum (Sorghum bicolor (L.) Moench). In Vitro Cell Dev. Biol. Plant 37, 504– 515.
Penna S. 2003 Building stress tolerance through over-producing trehalose in transgenic plants. Trends Plant Sci. 8, 355– 357.
Ping L. X., Nogawa M., Nozue M., Makita M., Takeda M., Bao L. and Kojima M. 2003 Inplanta transformation of mulberry trees (Morus alba L.) by Agrobacterium tumefaciens. J. Insect Biotechnol. Sericol. 72, 177–184.
Romero C., Belles J. M., Vaya J. L., Serrano R. and Culianez-Macia F. A. 1997 Expression of the yeast trehalose-6-phosphate synthase gene in transgenic tobacco plants, pleiotropic phenotypes include drought tolerance. Planta 201, 293–297.
Rontein D., Basset G. and Hanson A. D. 2002 Metabolic engineering of osmoprotectant accumulation in plants. Metab. Eng. 4, 49–56.
Sambrook J. and Russel D. W. 2001 Molecular cloning: a laboratory manual, 3rd edition, vol. 2. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, USA.
Sharma K. K. and Ortiz R. 2000 Program for the application of genetic transformation for crop improvement in the semi-arid tropics. In Vitro Cell Dev. Biol. Plant 36, 83–92.
Supartana P., Shimizu T., Shioiri H., Nogawa M., Nozue M. and Kojima M. 2005 Development of simple and efficient inplanta transformation method for rice (Oryza sativa L.) using Agrobacterium tumefaciens. J. Biosci. Bioeng. 100, 391– 397.
Supartana P., Shimizu T., Nogawa M., Shioiri H., Nakijima T., Haramoto N. et al. 2006 Development of simple and efficient inplanta transformation method for wheat (Triticum aestivum L.) using Agrobacterium tumefaciens. J. Biosci. Bioeng. 100, 162–170.
Tadesse Y., Sagi L., Swennen R. and Jacobs M. 2003 Optimization of transformation conditions and production of transgenic sorghum (Sorghum bicolor) via microparticle bombardment. Plant Cell Tiss. Org. Cult. 75, 1–18.
Tari I., Laskay G., Takacs Z. and Poor P. 2013 Response of sorghum to abiotic stresses: a review. J. Agron. Crop Sci. 199, 264–274.
Varalaxmi Y., Reddy L. A., Yadav S. K. and Maheswari M. 2012 Molecular cloning, characterization and functional validation of TPS1 gene isolated from S. cerevisiae. Res. J. Biotechnol. 7, 175–179.
Yeo E. T., Kwon H. B., Han S. E., Lee J. T., Ryu J. C. and Byun M. O. 2000 Genetic engineering of drought resistant potato plants by introduction of the trehalose-6-phosphate synthase (TPS1) gene from Saccharomyces cerevisiae. Mol. Cells 10, 263–268.
Zhao Z.Y., Cai T., Tagliani L., Miller M., Wang N., Pang H. et al. 2000 Agrobacterium mediated sorghum transformation. Plant Mol. Biol. 44, 789–798.
Author information
Authors and Affiliations
Corresponding author
Additional information
[Yellisetty V., Reddy L. A. and Mandapaka M. 2015 In planta transformation of sorghum (Sorghum bicolor (L.) Moench) using TPS1 gene for enhancing tolerance to abiotic stresses. J. Genet. 94, xx–xx]
Rights and permissions
About this article
Cite this article
YELLISETTY, V., REDDY, L.A. & MANDAPAKA, M. In planta transformation of sorghum (Sorghum bicolor (L.) Moench) using TPS1 gene for enhancing tolerance to abiotic stresses. J Genet 94, 425–434 (2015). https://doi.org/10.1007/s12041-015-0540-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12041-015-0540-y