Abstract
Amyotrophic lateral sclerosis (ALS) and frontotemporal dementia (FTD) primarily affect the motor and frontotemporal areas of the brain, respectively. These disorders share clinical, genetic, and pathological similarities, and approximately 10–15% of ALS-FTD cases are considered to be multisystemic. ALS-FTD overlaps have been linked to families carrying an expansion in the intron of C9orf72 along with inclusions of TDP-43 in the brain. Other overlapping genes (VCP, FUS, SQSTM1, TBK1, CHCHD10) are also involved in similar functions that include RNA processing, autophagy, proteasome response, protein aggregation, and intracellular trafficking. Recent advances in genome sequencing have identified new genes that are involved in these disorders (TBK1, CCNF, GLT8D1, KIF5A, NEK1, C21orf2, TBP, CTSF, MFSD8, DNAJC7). Additional risk factors and modifiers have been also identified in genome-wide association studies and array-based studies. However, the newly identified genes show higher disease frequencies in combination with known genes that are implicated in pathogenesis, thus indicating probable digenetic/polygenic inheritance models, along with epistatic interactions. Studies suggest that these genes play a pleiotropic effect on ALS-FTD and other diseases such as Alzheimer’s disease, Ataxia, and Parkinsonism. Besides, there have been numerous improvements in the genotype–phenotype correlations as well as clinical trials on stem cell and gene-based therapies. This review discusses the possible genetic models of ALS and FTD, the latest therapeutics, and signaling pathways involved in ALS-FTD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Amyotrophic lateral sclerosis (ALS) and frontotemporal dementia (FTD) are debilitating, progressive adult-onset brain disorders [1]. Clinically, ALS and FTD are considered as movement and cognitive disorders [2]. Most commonly, ALS affects the upper motor neurons (UMNs) and lower motor neurons (LMNs) as well as the brain stem and the corticospinal tracts of the brain, while FTD causes a gradual degeneration of both the temporal and frontal cortex [3]. ALS and FTD are clinically heterogeneous entities, although some phenotypes of ALS, such as cognitive impairment, overlap with that of FTD [4]. An estimated 50% of ALS patients demonstrate subtle impairments in behavior, cognition, and dementia-like features, while up to 15% of ALS patients also suffer from FTD; these individuals are called ALS-FTD or FTD-ALS patients [4]. Most of these ALS-FTD patients do not have a family history and are considered sporadic (sALS/sFTD). Nevertheless, ~ 10% of ALS cases are inherited in an autosomal dominant manner (fALSs) [5], while ~ 10–30% of FTD cases also show autosomal dominant inheritance or familial FTDs (fFTDs), which increases to 40% if a history of neurodegenerative disease is considered [6]. Moreover, both sALS and sFTD lack a family history that could be attributed to alternative modes of inheritance, such as oligogenic/polygenic inheritance or, most likely, reduced penetrance [7]. Additionally, pleiotropic mechanisms have been hypothesized at the genetic and environmental levels, possibly explaining individual differences due to the involvement of distinct disease pathways [8]. Clinically, most fALS cases are indistinguishable from sALS and vice versa. For example, TAR DNA-binding protein-43 (TARDBP) mutations are rare (~ 1%) in both ALS and FTD patients [1]. However, neuropathological studies have shown the presence of TDP-43 aggregates in ~ 97% of ALS cases and up to 50% of FTD cases [1, 9]. These studies suggest there is a definitive pathological link between familial and sporadic ALS and FTD. Nevertheless, several well-known genes have been identified in a larger population of ALS/FTD patients [4]. Such examples include those of chromosome 9 open reading frame 72 (C9orf72), TARDBP, fused in sarcoma (FUS), sequestrome-1 (SQSTM1), etc. [10]. With the advancement of sequencing technologies and their ease of use, scientists have been able to produce larger datasets for international projects and consortia such as Project MinE (https://www.projectmine.com/) and GENFI3 (https://www.genfi.org/) which have further identified many novel genes and risk factors, along with genetic modifiers of ALS and FTD, respectively [11]. ALS-FTD is related to over 30 genetic loci, > 100 genes, and numerous mutations, and new genes are continually being discovered [12, 13]. A number of GWASs have revealed the role of several modifier genes in ALS/FTD pathogenesis [14, 15]. Nevertheless, these low penetrating variants, which appear to contribute substantially to disease pathology, must be validated across different populations. There is still uncertainty about whether these genes play a role in disease penetrance or whether environmental factors trigger the disease. It is well established that ALS and FTD are not two separate diseases, but instead, constitute a syndrome or a disease spectrum [16]. These diseases illustrate the presence of oligogenic or polygenic effects, or even pleiotropy, where more than one gene or numerous mutations are present in a variety of genes, resulting in a wide variety of phenotypes [3, 4, 17]. Besides, phenotypic variability might be triggered by other genetic phenomena such as epistatic interactions. For example, multiple genes involved in ALS may interact epistatically, explaining the substantial differences in phenotypic behavior observed between individuals who carry the same mutation(s); thus, this may also explain the incomplete penetrance of the disease within some families [18]. Therefore, this leads to newer challenges for appropriate diagnostics and prognostics of ALS and FTD. New genetic testing, patient education and counseling, and gene-based clinical trials are equally important. Interestingly, gene-specific therapies such as superoxide dismutase 1 (SOD1) and granulin (GRN)/C9orf72 have shown promise in treating ALS and FTD [19,20,21]. Hence, this review discusses the latest genetic findings related to ALS and FTD in addition to hotspot/modifier genes like SOD1, GRN, microtubule-associated protein tau (MAPT), TARDBP, and C9orf72. These genes are not only congregated in corresponding pathways, but they also function in DNA repair, axonal transport, RNA metabolism, and formation of stress granules [22]. These insights also spurred mechanistic studies that uncovered a common pathway between ALS and FTD [4]. Despite these discoveries, understanding why mutations in these genes cause ALS, FTD, or both remains obscure.
Novel Genetic Landscape of ALS and FTD
ALS and FTD share many clinical, genetic, and pathological features as evidenced in Table 2. Numerous known and novel genes have also been detected in each group (Tables 1 and 2 and Fig. 1). Mutations in genes known to cause ALS (SOD1, C9orf72, TARDBP, VCP, FUS, PFN1, etc.) and FTD (PGRN, MAPT, C9orf72, CHMP2B, TBP, CHCHD10, etc.) have explained only ~ 60–70% of fALS/fFTD and ~ 10% of sALS/sFTD cases [23]. Furthermore, the hexanucleotide expansion mutations in C9orf72 have been a major causal factor for both ALS and FTD [24, 25]. Despite the identification of several known causal factors, there are still many genetic forms of the disease that are yet to be discovered. Recent advances in sequencing capabilities, including whole-genome sequencing (WGS), and whole-exome sequencing (WES), have made identification of causal alleles and additional genes easier. The WGS focuses to cover all regions in the genome, while WES targets mostly the coding region of the genome. In comparison to WGS, WES is much cheaper and cost-effective and takes lesser time to analyze raw data. With the advent of next-generation sequencing technology, greater in-depth discoveries have been made possible in the genomics of ALS-FTD. Genome-wide association studies (GWAS) is another powerful tool that can be used to discover novel genetic loci involved in ALS-FTD pathogenesis. GWAS involves rapid scanning of markers scattered throughout the genome or examining complete DNA sets from many individuals to trace the genetic variation(s) associated with a particular trait or disease. GWAS generally identifies common variants, while rare variants conferring disease risk remain undetected. Despite this, identifying novel genes or newer defects involved in disease pathogenesis is extremely crucial, and thus, scientists are sequencing more samples to identify rare variants and genes that significantly contribute to both ALS and FTD. Recently, several genes that carry these rare genetic variants have been identified [10]. This includes two new FTD loci, six ALS genes, and two ALS-FTD genes discovered in the last 7 years (Table 1, Fig. 1, and Fig. 2). There is a higher penetrance for these genes and co-occurrence of multiple mutations in combination with C9orf72 which are based upon familial, case–control, and candidate-based studies [4]. Among them are TBK1, CCNF, KIF5A, GLT8D1, TIA1, NEK1, C21orf2, DNAJC7, TBP, CTSF, and MFSD8 (see Table 1, Fig. 1, and Fig. 2). We have described each of these genes in detail, and this includes gene descriptions, along with their pathological mechanisms.
(A) Flowchart indicating numerous factors that trigger the initiation of complex phenotypes observed in FTD along with the disease progression. (B) Depiction of different pathways involved in FTD pathogenesis, that can be affected by the cellular stress and, cellular microenvironment as well as external factors
Hotspot Genes Known in ALS and FTD: a Brief Overview
Superoxide Dismutase 1 (SOD1)
SOD1 was first linked to ALS through genetic linkage analysis in 1993 [26]. Later, the first transgenic mouse model was developed using the Gly93Ala mutation in SOD1 [27]. The most prevalent mutations in SOD1 are Asp90Ala (aspartic acid to alanine) and Ala4Val (alanine to valine). It has been observed that Asp90Ala substitutions may be inherited both as dominant and recessive forms of the disorder; however, the majority of the cases are recessive [28]. To date, more than 200 genetic mutations (insertions, deletions, point mutations, and truncation mutations) have been documented in the SOD1 (http://www.hgmd.cf.ac.uk/ac/gene.php?gene=SOD1 and Supplementary information). Different mutations of SOD1 lead to destabilization and reduced protein affinity, as well as the formation of protein inclusion and aggregation [29]. Therefore, a variety of mutations in SOD1 can either increase or decrease its function or lead to complete loss of function (LoF) [30]. Genetically and clinically, SOD1 mutations are extremely heterogeneous. Symptoms are often very similar to sALS and fALS. SOD1 has been linked to intra-familial variations in clinical and genetic severity as well as in progression of the disease [31]. Even though the exact pathological process is unknown, it might be that the LoF of proteins, or aggregation of misfolded proteins, or prion formation may create the molecular basis of disease pathogenesis [32]. SOD1 encodes CuZn superoxide dismutase (32 kDa), which is ubiquitously expressed in the brain [33]. The main function of SOD1 is to remove the free radicals such as the toxic superoxide anion O–2 to O2 and H2O2, thus reducing the oxidative stress within the cell [34]. During, normal oxygen metabolism, oxygen radicals such as superoxide and hydrogen peroxide are produced, and these are involved in normal signaling within the cell. However, their excess amounts increase cellular stress and harm a wide range of biological processes [35]. Consequently, these superoxide and hydrogen peroxide molecules in the cytoplasm must be converted to H2O by catalase, glutathione peroxidases, or peroxiredoxins [36]. SOD1 has been reported to protect cells against such molecules in the intermembrane space of the cell as well as in the cytoplasm [37,38,39,40]. Furthermore, in the case of SOD1 mutation or altered SOD1 functionality, the scavenging systems might not be efficient enough to maintain the homeostasis of normal reactive oxygen species (ROS). Consequently, this further increases both cellular stress and the elevated levels of H2O2 that might ultimately lead to mitochondrial damage [41]. Furthermore, SOD1 is localized in distinct places within the cell, which includes the cytoplasm, nucleus, mitochondria, and lysosomes [27, 33]. Transgenic mice were created harboring these mutations and were seen to generate motor neuron syndrome [42]. In order to develop gene therapy approaches against mutant SOD1, one needs to understand the detailed molecular pathogenesis of each and every mutation identified so far [43].
SOD1 Pathogenesis
Mutations in the SOD1 cause exclusively ALS and most likely a toxic gain-of-function protein [30]. So far, SOD1 mutations have caused protein misfolding, nuclear and cytoplasmic aggregation, and cytotoxicity, among others. ALS is a very heterogeneous disease involving genetic, environmental, and other interrelated factors including smoking, aging, etc. (Fig. 3). ALS research was revolutionized by Charcot’s description of the disease as well as the discovery of the SOD1 gene [44]. Despite extensive research, the actual and realistic functions of the SOD1 remains obscure; however, both toxic gain and LoF are implicated. Regarding the pathogenesis of the SOD1, several hypotheses have been proposed, including protein aggregation and protein cytotoxicity [30]. Mutations in the SOD1 create toxic protein aggregation and RNA toxicity, leading to pathogenesis in multiple parts of the brain, which has been demonstrated by PET studies [45]. ALS autopsy samples and transgenic mice models have demonstrated ubiquitinated SOD1 protein and intracellular inclusions in both neuronal and astrocyte cells [46]. The disorder occurs in both UMN and LMN phenotypes in ALS and in other regions such as the posterior column, thalamus, Clarke’s nucleus, and spinocerebellar tracts [47]. There is still uncertainty about the consequences and contributing effects of these inclusion bodies. However, individuals with mutations in the SOD1 gene are more likely to be vulnerable to stress and cellular injuries in their lifetimes [48]. In this case, the alpha motor neurons (largest post-mitotic neurons) are responsible for intracellular transport, and energy metabolism, and have extensive neurofilament networks [48]. Currently, it is unknown whether a toxic gain of function or a LoF of SOD1 may be involved in the pathology of motor neuron disease [30].
Protein aggregation may cause misfolding; however, their monomerization and subsequent clump formation may cause cytotoxicity. The SOD1 protein forms aggregates and microaggregates of denatured monomers as a result of various genetic mutations. It has been documented that these are common factors driving the formation of numerous microaggregates that disrupt cellular transport, mitochondrial function, proteasomal function, and homeostasis [49]. The latest SOD1 (G93A) mouse model has shown that morphological and functional changes (dendritic regression and glutamatergic excitation) are related to cortical neuronal hyperexcitability as an early feature of the ALS pathogenesis [49,50,51]. Motor neuron cells derived from induced pluripotent stem cells (iPSCs) have also displayed similar patterns of neuronal hyperexcitability, and inhibition of this process has been shown to be neuroprotective [52]. Several animal studies support this notion, suggesting that hyperexcitability in the cortex contributes to ALS [53, 54]. Therefore, SOD1 pathogenesis is integral to ALS, and more research is necessary to understand SOD1 pathogenesis.
Granulin (GRN)
GRN gene mutations were first identified in families with FTD (5–20%), mainly as autosomal dominant forms associated with chromosome 17 [55]. Likewise, these mutations in GRN make up about 5–20% of fFTD and ~ 1–5% of sFTD in this study [55]. There are more than 240 genetic variations known to date (http://www.hgmd.cf.ac.uk/ac/gene.php?gene=GRN and Supplementary information), and most of the pathogenic mutations result in null alleles or haploinsufficiency [56]. The fact that heterozygous mutations in GRN cause FTD (FTD-GRN), whereas homozygous mutations cause neuronal ceroid lipofuscinosis, a lysosomal storage disorder, suggests the role of progranulin (PGRN) in lysosomal biogenesis and homeostasis [57]. This gene encodes a PGRN, which is secreted into the extracellular space and absorbed by the cells. Granulin is produced when progranulin is cleaved to form proinflammatory granulin subunits, and the catabolism of progranulin was performed in the lysosomes using cathepsins [58, 59]. Further, PGRN is mainly found in the lysosome where it may have an impact on lysosomal functions such as enzymatic activity and acidification [59]. This gene is strongly expressed in neurons, including microglia, and astrocytes in the central nervous system [60]. FTD affects both the frontal and temporal lobes, leading to behavioral, linguistic, and executive dysfunction. Therefore, variations in clinical presentation among individuals within a family have been observed. Mutations in the GRN lead to lipid droplet accumulation in microglia [61], a phenomenon that is also seen with aging. Neurons and microglia derived from patients with heterozygous GRN mutations show increased lipofuscinosis and lysosomal storage materials, supporting the notion that FTD-GRN may be caused by lysosomal dysfunction [62]. The other molecular pathologies are deficient autophagy, inflammation, and nuclear depletion of TDP-43 or accumulation of pathogenic forms of TDP-43 [60, 63] (Fig. 4). FTD-GRN patients have been found to have increased reactive microglia, proinflammatory cytokines, and microglial dystrophy [64]. Complete loss of PGRN in Grn–/– mice is known to increase complementary pathways and induce GABAergic synaptic pruning, leading to hyperexcitability in the thalamus [61]. Mice models exhibiting PGRN deficiency show increased NF-kB-TNFα signaling, leading to hyperexcitability in the striatum [65]. The pathogenesis of FTD-related neurodegeneration associated with PGRN deficiency remains a focus of intense investigation [61]. FTD also exhibits neuropathological protein aggregation, glial hyperproliferation, inflammation, and lysosome dysfunction [65] (Fig. 4).
Microtubule-Associated Protein Tau (MAPT)
The MAPT gene was identified in 1975 on chromosome 17q21 [66]. MAPT encodes tau protein which is abundantly expressed in neurons along with low expression in glial cells [67]. Abundant MAPT has also been seen in various areas of the brain including the cerebral cortex, subcortical, white matter, and brain stem, as well as the spinal cord, which are vulnerable to tau deposition [68]. In 1998, the first mutations were discovered in exons 9, 10, and 13, as well as in intron 10 [69,70,71]. Mutations in MAPT occur primarily in the repeats and result in tau proteins that have less affinity to microtubules [72]. The MAPT gene variant is inherited in an autosomal dominant manner, and more than 100 mutations have been reported in MAPT (http://www.hgmd.cf.ac.uk/ac/gene.php?gene=MAPT and Supplementary information). MAPT mutations are associated with tau-positive neuropathology in familial frontotemporal lobar degeneration (FTLD), and their frequencies are variable across populations, ranging from 5 to 50% [73]. MAPT alleles exhibit complete linkage disequilibrium with several polymorphisms, resulting in two distinct haplotypes H1 and H2 [74]. According to most studies, the predominant haplotype, H1, is associated with sporadic tauopathies and a weaker association with FTLD [74]. MAPT mutations are associated with an autosomal dominant form of FTD and parkinsonism (FTDP-17 T), affecting approximately 10% of people with familial FTD (http://www.molgen.ua.ac.be/FTDMutations/). The pathology is characterized by hyperphosphorylated tau inclusions in cortical and subcortical gray and white matter. It shares similarities with sporadic tauopathies in terms of anatomical distribution, inclusion morphology, and biochemistry. It is likely that genetic mutations have some correlation with clinical and pathological features among families [68]. The clinical features linked to MAPT mutations are behavior, language, motor, and memory function. Also, the clinical phenotypes often overlap with other symptoms including Alzheimer’s disease (AD), progressive supranuclear palsy (PSP), corticobasal degeneration (CBD), Pick disease, and primary progressive aphasia [75]. Tau plays an important role in microtubule assembly, which is crucial for axoplasmic transport and neuronal integrity [76]. The abnormal intracellular accumulation of hyperphosphorylated tau is thought to contribute to a number of neurodegenerative disorders (tauopathies), including FTD (FTLD-tau) [76]. Based on neuropathological findings, FTLD is characterized by hyperproliferation of glial cells and abnormal protein inclusions in the cytoplasm and nuclei of neurons [77]. It has been shown that FTLD-tau and TDP-43 protein aggregates (~ 45% of FTLD-tau and ~ 50% of FTLD-TDP) have been observed in the brains of FTLD patients [78]. There are heterogeneous genetic features in FTLD, reflecting its complex clinical and pathological nature. Despite enormous advances in FTLD genetics over the past 20 years, it’s clear that molecular mechanisms remain the major obstacle in unraveling FTLD pathogenesis to date [79].
Chromosome 9 Open Reading Frame 72 (C9orf72)
The massive GGGGCC repeat expansion (∼700–1600 copies) in the first intron of C9orf72 has been identified in 2011 with familial and sporadic forms of both FTD and ALS [24, 80]. Since then, it has become a hallmark not only of ALS and FTD but also of AD, PD, PSP, ataxia, CJD, and Huntington’s disease (HD) [24, 25]. A total of 29 genetic variants have been linked to C9orf72 (http://www.hgmd.cf.ac.uk/ac/gene.php?gene=C9orf72 and Supplementary information), including missense, splice site, and repeat variations. C9orf72 is responsible for up to 80% of familial ALS-FTD, 20–50% of familial ALS, 5–20% of sporadic ALS, and 10–30% of FTD cases [81]. Critical clinical phenotypes include behavioral variant of FTD, primary progressive aphasia, movement issues, memory problems, and psychosis [82]. Patients with the C9orf72 repeat expansion have been shown to have higher rates of bulbar onset ALS, cognitive impairment with an earlier onset of disease, and accelerated disease progression [83]. The actual function of C9orf72 is yet to be defined, but accumulating evidence suggests that it functions (directly or indirectly) in coordinating autophagic flux or endosomal trafficking [84]. The genetic, clinical, and neurological heterogeneity of C9orf72 is also evident in families with GRN mutations. Different neuropathological phenotypes also demonstrate overlapping inclusions of abnormal proteins, including FTLD-TDP and typical ALS-TDP-43 inclusions in LMN of different patients [77, 82, 85]. Brain pathologies of patients have also shown overlapping phenotypic categories [77, 82]. Apart from TDP-43 pathology, ubiquitin–proteasome system (UPS) dysfunction, repeat-associated non-AUG (RAN) translation, dipeptide repeat (DPR) protein synthesis, and intracellular RNA accumulation can also be detected as a resultant of C9orf72 mutations [24, 84]. In order to determine the role of C9orf72 in cellular physiology, studies are being aimed at targeted reduction of mutant C9orf72 expression, to deplete the formation of toxic nuclear foci. Further, lysosome–autophagosome connections and proteasomal activities may also be associated with C9orf72-mediated pathways [1].
TAR DNA-Binding Protein 43 (TDP-43)
TDP-43 was initially cloned as a repressor protein involved in HIV-1 transcription [86]. It binds to a trans-active response element within viral DNA sequences and is essential for the regulation of viral gene expression [86]. The TDP-43 (1p36.22) encodes 414 amino acids and functions as an hnRNP. TDP-43 has been implicated in both ALS and FTD [87]. TDP-43 is ubiquitously expressed in almost all tissues, including the brain [87]. It contains two RNA recognition motifs (RRM1and RRM2) and a glycine-rich C-terminal sequence, which is a “hot spot” for mutations in TDP43 [87]. A total of 70 different genetic variants have been reported so far in TARDBP (http://www.hgmd.cf.ac.uk/ac/gene.php?gene=TARDBP and Supplementary information). One of the major pathways perturbed in TDP-43 pathogenesis might be dysregulation of RNA splicing, which could result in functional defects in vital proteins. However, further research is needed to elucidate the role of TDP-43 in normal cellular function, as well as in other mechanistic processes affected by mutations in TARDBP. The most important domain of TDBP-43 is its glycine-rich domain, which has been implicated in exon skipping, splicing inhibition, and biogenesis of mRNA [88]. Recently, it has been demonstrated that TDP-43 regulates spermatogenesis and acts as a scaffold for nuclear bodies called Gemini of coiled bodies (GEMs) and interacts with the survival motor neuron protein [89, 90]. Although its biological functions are not fully understood, TDP-43 has been shown to function by binding to single-stranded DNA, RNA, and/or proteins [88, 91]. Likewise, the anatomical distribution of DPR pathology does not correlate with the patterns of neurodegeneration and pathology related to TDP-43 [92] Currently, it is debated whether TDP-43 or DPR pathology modulates the pathogenesis caused by C9orf72 mutations [92].
Novel Genes Identified in ALS and FTD
TANK-Binding Kinase (TBK1)
In 2015, mutations in the TBK1 gene were reported to be associated with ALS [93, 94], FTD [95], and ALS-FTD [17]. Similarly, a large-scale GWAS study identified TBK1, which is located on chromosome 12q14.2, as a risk locus in ALS [96]. There are many LoF mutation in TBK1 resulting because of frameshifts, nonsense mutations, splice site alterations, and possibly pathogenic read-through mutations [97]. Many of these missense mutations in TBK1 are clustered in the kinase and ubiquitin-like domains [98]. Missense mutations in the functional domains as well as those that impair TBK1 phosphorylation and binding of target proteins to TBK1 may result in disease pathogenesis [98]. This was supported by the difference in the odd ratio between cases and controls observed in this study (11.78 and 1.62, respectively) [99]. Furthermore, patients with ALS, FTD, and ALS-FTD are estimated to have a mutation frequency between 0.4 and 1.7% in TBK1 [95, 100]. A recent meta-analysis found that the two mutation combinations (LoF mutations at 1% and missense mutations at 1.8%) are more prevalent in European populations than in Asian populations suffering from ALS and/or FTD [99]. Probably, this was because the numbers of Caucasian cohorts studied were more than that of Asian cohorts [1]. TBK1 mutations were also found to be the second most common cause of ALS-FTD pathogenesis, after C9orf72 mutations [101]. Mutations in TBK1 are related to bulbar onset in ~ 15% of ALS patients and progressive behavioral changes in FTD patients [94]. It has been shown that both ALS and FTD patients exhibit increased inclusions of TDP43 and p62 in their affected brain regions [97]. The knockout of TBK1 mutations in rodent models led to similar symptoms of cognitive impairment and motor dysfunction as those seen in ALS/FTD patients [102]. Further, a germline TBK1 deletion has been associated with embryonic lethality, suggesting that it plays an important role in development. Moreover, T2K-deficient mice showed embryonic lethality and apoptotic liver degeneration, along with altered NF-kB gene expression [103].
TBK1 belongs to the IkB kinase gene family, and it is essential for binding and phosphorylating several proteins involved in autophagy [104], mitophagy [105], and innate immunity [106]. TBK1 is highly expressed in neurons of the hippocampal formation, the cerebral cortex, and the lateral ventricles [107]. TBK1 interacts with optineurin (OPTN) and SQSTM1 to form an autophagic adaptor complex [108]. Mutations in these two genes as well as TBK1 have been detected in patients with ALS-FTD [17, 101, 109]. Furthermore, TBK1 mutations have been found along with C9orf72 expansion [95, 100], TARDBP [94, 98], FUS [109], or FUS and DCTN1 [110]. Mutations harboring TBK1 with TARDBP [98] or TBK1 with FUS [94] have a more rapid onset of disease pathogenesis than those harboring TBK1 alone [94, 111]. Mutations in TBK1 and repeat expansion in C9orf72 have also been associated with alterations in disease onset and penetrance [95]. The clinical phenotypes of patients with TBK1 mutations showed variable onsets, progressions, and survival rates [95, 98, 100, 111]. TBK1 mutations have also been detected in other cohorts, including patients with motor neuron disease alone or with FTD [17, 93,94,95]. TBK1 mutations have been linked to a variety of overlapping diseases, including behavioral FTD, prominent parkinsonism, and semantic paraphasias [95, 97]. Future studies should investigate clinical, phenotypic, and clinicopathological correlations to confirm or disprove the distinct genetic identification and treatment associated with TBK1.
Cyclin F (CCNF)
Genome-wide linkage analysis led to the identification of CCNF in 2016 in an Australian family with British ancestry afflicted with ALS-FTD [112]. The CCNF mutation frequency ranged from 0.6 to 3.3% among Australian, American, European, and Asian populations with fALS-FTD [112, 113]. CCNF encodes the cyclin F protein (part of the E3 ubiquitin–protein ligase complex), also known as the Skp1-Cul1-F-box E3 ubiquitin ligase complex [114]. CCNF catalyzes and transfers ubiquitin molecules to the specified proteins, which are then degraded by the UPS [115]. The mutation Ser621Gly in CCNF inhibits this degradation by disrupting ubiquitination, resulting in the accumulation of ubiquitinated proteins, such as TDP-43 and ribonucleoside-diphosphate reductase subunit M2 [112]. Interestingly, CCNF also acts on SQSTM1 like TBK1 and triggers ubiquitinated protein degradation by autophagous receptor binding [116]. CCNF is expressed in neuronal cells, and mutations in CCNF are associated with defects in ubiquitination processing and autophagosome–lysosome fusion [112, 116]. Recently, a zebrafish model of the p.Ser621Gly mutation has been used to demonstrate its effects on axonal growth [116, 117]. CCNF has been shown to interact with VCP and increase its ATPase activity, leading to the aggregation of TDP-43 [118]. The genetic and functional findings obtained in cellular as well as model organisms indicate that mutant CCNF leads to impaired UPS/autophagy pathways resulting in impaired axonal growth. Hence, this gene has been implicated in the ALS-FTD spectrum and is also known as locus FTDALS5 (Table 1).
Kinesin Family Member 5A (KIF5A)
For the first time, KIF5A was identified using WES followed by rare variant analysis in fALS patients (case = 426, control = 6137) [119]. Splice site mutations were reported to be clustered in the C-terminal region of KIF5A [119]. Further, functional analyses of lymphoblast cell lines demonstrated defective splicing and haploinsufficiency, resulting in disease pathogenesis [5, 119]. This study was initiated as there have been previous findings on the involvement of KIF5A in other diseases, such as monogenic spastic paraplegia and Charcot–Marie–Tooth disease type 2 (CMT2) [120]. In 2018, similar findings were reported using two analogous methods: (1) the GWAS in 20, 806 ALS cases, and 59,804 controls and (2) the rare variant burden in 1,138 fALS cases and 19,494 controls [120]. Consequently, a missense heterozygous variant (p.Pro986Leu, rs113247976) and other LoF variants (within amino acids 996–999) were identified, suggesting the need for additional cohort validation [119]. The variant (p.Pro986Leu, rs113247976) was checked in additional cohorts of ALS patients (cases = 4159, controls = 18,650) and confirmed the original GWAS findings [119]. Further validation of this observation was done by meta-analysis and replication studies (cases = 24,965, controls = 78,454) that generated a highly significant p value (7.09 × 10 − 13) [120]. An independent study using rare variant burden analysis showed that this variant (rs113247976) was associated with fALS [120]. Three additional carriers with LoF variants in KIF5A were also found [120]. In their research, the authors suggested that the variant p.Pro986Leu in KIF5A posed a lower penetrance with a relatively common occurrence in fALS [119, 120]. There is also evidence that LoF variants in KIF5A are rare and have a higher penetrance and are at higher risk factors for ALS [119, 120].
KIF5A belongs to a family of motor proteins, called kinesins (KIF-5 major isoform). It plays a role in the transport of organelles within the cells, such as axonal transport [119, 120]. It has also been reported that mutant KIF5 plays a role in the pathogenesis of motor neuron degeneration [121, 122]. Nevertheless, KIF5 transports granules in axons and dendrites that interact with RNA and RNA-related molecules. [123]. Most of these granules/cargos are related to other genes associated with ALS, for example, FUS, VAPB, and hnRNPA1 [124, 125]. The knockout mice of KIF5A demonstrated abnormal neurofilament transport [126] and pathological accumulation of these neurofilaments in ALS [127]. Similarly, KIF5A−/− mice also show defects in axonal transport and survival [128]. ALS patients also have impaired axonal transport and mitochondrial activity [124, 129]. In addition to the KIF5 family members, other receptors including the α-amino-3-hydroxyl-5-methyl-4-isoxazole-propionate (AMPA) and γ-Aminobutyric acid type A receptors (GABAA) types are also affected [130, 131]. Furthermore, other ALS genes (NEK1 and PFN1) modulate KIF5A expression [132, 133], and this impacts the formation of neurite-like membrane protrusions [125]. In contrast, the majority of the LoF variants have been identified in the C-terminal domain of KIF5A, and haploinsufficiency has been observed which may disrupt the binding affinity of KIF5A with other cargo proteins [119, 120].
Furthermore, the zebrafish models with a truncated C-terminus of KIF5A led to mitochondrial translocation disruption [134]. It has been suggested that lack of KIF5A expression can lead to accumulations of downstream proteins, such as phosphorylated neurofilaments and amyloid precursor proteins, which are associated with neurodegeneration in patients with multiple sclerosis (MS) [135]. Recently, KIF5A has emerged as a promising and predictive biomarker in MS [135]. KIF5A mutations show genetic heterogeneity that can result in a variety of distinct phenotypes depending on their location within the gene [119, 135]. For example, KIF5A mutations can lead to classical ALS phenotypes, myoclonus in infancy, CMT2, and hereditary spastic paralysis, etc., suggesting a pleiotropic role of this gene in various neurological conditions [119, 135].
Glycosyltransferase 8 Domain Containing 1 (GLT8D1)
GLT8D1 was discovered by WES in a family with autosomal dominant ALS that harbored variant p.Arg92Cys within exon 4 [136]. Among the affected individuals, the heterozygous p.Arg92Cys mutation in GLT8D1 co-segregated in an autosomal dominant manner [136]. The findings were confirmed by Sanger sequencing [136]. Within the same exon, additional rare deleterious mutations within GLT8D1 have been linked to ALS [136]. The mutations p.Arg92Cys and p.Gly78Trp are located at the substrate-binding site and alter the enzyme activity of GLT8D1 [136]. Further, in vitro cytotoxicity analysis using zebrafish revealed p.Arg92Cys and p.Gly78Trp variants in GLT8D1 cause insufficient motor activity resembling ALS, suggesting that the glycosyltransferase activity is lost in these mutants [136]. The possible roles of GLT8D1 are related to synthesizing signaling molecules, such as gangliosides, which are important in motor neuron functions [137]. A mice model of Ugt8a (the orthologue of UGT8) has shown the involvement of different brain-gangliosides in dysfunction of motor coordination [138, 139]. It is interesting to note that GLT8D1 deficiency has been linked to a number of other neurodegenerative conditions, including AD, PD, HD, and schizophrenia [140, 141].
TIA1 Cytotoxic Granule-Associated RNA-Binding Protein (TIA1
The TIA1 gene was first discovered by WES [142] in a European family with ALS-FTD. Heterozygous missense mutation (p.Pro362Leu) in TIA1 segregated in the affected family members [142]. Five additional mutations in six different patients were identified in the low complexity sequence domain (LCD) [142]. TIA1 encodes an RNA-binding protein with a C-terminal LCD similar to that of TDP-43 or FUS [142]. TIA1 is also found in membrane-less organelles like stress granules and can modify liquid–liquid phase separation (LLPS) using LCD [142]. Due to the TIA1 mutation, the LLPS undergoes changes in their biophysical properties, which lead to changes in the formation of stress granules [142]. A genetic mutation occurring in this domain causes Welander distal myopathy and is also associated with TDP-43 and p62 aggregation [143,144,145]. Mutations in TIA1 affect stress granule dynamics and result in TDP-43 accumulation [142]. In a case–control study (cases = 1039, controls = 3036), additional six nonsynonymous variants were present in the LCD domain of TIA1 in affected members [146]. A significant value with ~ 2% with fALS and < 0.5% of sALS patients were observed [142]. Further, neuropathological examination of autopsied brains and spinal cords of TIA1 carriers revealed TDP-43 pathology [147]. Additionally, inclusions of hyaline-like cytoplasmic Lewy bodies were also observed in LMNs [142, 147]. There may be a link between the inclusions present in the LMNs and the TIA1 mutation, which is characteristic of ALS patients [148].
NIMA-Related Kinase 1 (NEK1) and Chromosome 21 Open Reading Frame 2 (C21orf2)
NEK1 was first identified as a hit in WES study of > 2,000 ALS patients [4, 93, 149]. The rare variants in NEK1 were significantly enriched in ALS patients compared to that of controls (cases = 2303, controls = 1059) [150]. Approximately 3–4% of ALS patients of European descent had the presence of rare variants in NEK1, including heterozygous LoF variants in 1% of patients [4, 93, 149, 150]. Mutations in this gene were also reported in another WES study that examined 827 controls and 265 index ALS patients [150].
NEK1 is a member of the conserved NIMA-related serine/threonine kinases family, and it plays an important role in cell cycle, DNA damage repair, mitochondrial membrane regulation, and ciliogenesis [151,152,153]. A heterozygous mutation (p.Arg812Xaa) in NEK1 has been demonstrated in patient-derived neuronal cells with an increased DNA repair response to irradiation [152]. NEK1 is involved in the maintenance of the cytoskeleton system in neurons [154] and interacts with other ALS genes like tubulin alpha 4a (TUBA4A) [155] and PFN1 [133]. Along with other ALS proteins, such as VAPB and ALS2 (alsin), NEK1 binds to C21orf2 in response to DNA damage [93, 156].
C21orf2 has recently been identified as a novel ALS gene containing nonsynonymous and LoF variants [96]. Additionally, in vitro cell-based assays have shown that knocking down NEK1 expression results in cellular death, resulting in the LoF mechanism [157]. The function of C21orf2 is unknown, but it is believed that it interacts with NEK1 and participates in primary cilia formation [158,159,160]. Mutations in C21orf2 and NEK1, like those in TBK1 and GLT8D1, are associated with other diseases, including axial spondylopathies and retinal dystrophies [159].
DnaJ Heat Shock Protein Family (Hsp40) Member C7 (DNAJC7)
DNAJC7 has been recently identified in a large WES-based case–control study (cases = 3864 and controls = 7839) [161]. Eight individuals were found to have six protein-truncating variants (PTVs) (cases = 5095, controls = 28,911). A higher rate of significant PTVs has been observed in cases with constrained genes. Gene-based analyses have shown the presence of PTVs in the well-known ALS genes like SOD1, FUS, and NEK1. In GWAS, DNAJC7 showed a greater occurrence of PTVs, such as variant p.Arg156Ter [162]. Western blot analysis of patient-derived fibroblasts revealed a depletion of DNAJC7 protein (p.Arg156Ter) [162]. Thus, it was concluded that DNAJC7 loss was responsible for neurodegeneration, since it encodes a chaperone and contributes to protein folding and clearance of mature proteins, along with protein homeostasis. The study was limited to a single population, and therefore, further research is needed both in ALS and FTD cohorts from other populations. DNAJC7 encodes mainly a heat shock protein (HSP40), which facilitates the homeostasis of HSP70 by modulating folding, misfolding, and clearance of other proteins and peptides [162].
TATA-Box Binding Protein (TBP)
Recently, TBP has been identified in an Irish family that contributed to partial frontal dysfunction and cerebellar atrophy [163]. The proband had a 43 repeat expansion in TBP that co-segregated with other family members [163]. TBP encodes a transcription initiation factor that may have a role in RNA metabolism [164]. The mutations in TBP have been associated with spinocerebellar ataxia 17, suggesting that dementia and ataxia share overlapping mechanisms [163]. Future studies in TBP warrant better clinical profiling and genetic testing.
Cathepsin F (CTSF) and Major Facilitator Superfamily Domain 8 (MFSD8)
CTSF and MFSD8 have recently been identified in FTD [165, 166]. Two unrelated FTD patients in a Belgian cohort carried the heterozygous p.Arg245His mutation in CTSF [165]. Nevertheless, case–control, genetic association, and rare burden analyses showed a higher rate of risk variants in MFSD8, which has been linked to FTD [166]. It is interesting to note that these two genes are involved in the lysosomal–autosomal pathway that leads to ALS/FTD. The CTSF encodes cathepsin F, which is an enzyme involved in lysosomal cysteine protease activity [167, 168]. Recently, another missense variant (p.Leu465Trp) was identified in CTSF of an individual affected with FTD [169]. The MFSD8 encodes a ubiquitous integral membrane protein that is found in lysosomes [170]. According to this study, mutations in MFSD8 cause FTD by increasing the concentration of other proteins such as LAMP2 and CTSD and, concurrently, by decreasing the rate of protein degradation [166]. The presence of homozygous LoF in these two genes can result in neuronal ceroid lipofuscinoses [171, 172], suggesting that lysosomal dysfunction may exacerbate the disease. Further studies are needed to confirm the role of these genes in FTD pathogenesis.
Involvement of Common Interlinked Pathways in ALS-FTD Pathogenesis
Numerous genes are known to play a vital role in different pathways linked to ALS and FTD [4]. Some genes are exclusively associated with ALS/FTD. These genes include SOD1, MAPT, and GRN [80]. In SOD1, the disease has been classified as a pure case of ALS, while in MAPT and GRN, it has been classified as a pure case of FTD. The development of the disease is also influenced by related pathways, for example, genes that regulate autophagy and proteasome damage (C9orf72, OPTN, VCP, UBQLN2, SQSTM1, CCNF, FUS, TARDBP, TBK1, CHMP2B), RNA metabolism (C9orf72, UBQLN2, SETX, ANG, ATXN2, hnRNPA1, MATR3, FUS, TARDBP), mitochondrial dysfunction (TBK1, CHCHD10, VCP), and the cytoskeleton dynamics (TUBA4A, SQSTM1, CCNF, SPG11, KIF5A, PFN1) [4] (Fig. 2). However, other pathways such as lysosomal–endosomal involvement (NEK1, FIG1, ALS2), unfolded protein response (VAPB, SIGMAR1), stress granule formation (TAF15, FUS, ATXN2, TIA1, EWSR1, hnRNPA1), excitotoxicity, oxidative stress, neuronal survival, neurite outgrowth, and neuroinflammation are other important emerging mechanisms [4, 163] (Fig. 1). A systemic pathway that plays a role in ALS, FTD, and ALS-FTD pathogenesis is illustrated in Figs. 3, 4, and 5, respectively. ALS, FTD, and ALS-FTD appear to be characterized by a complex interaction between genetic and environmental factors, leading to dysfunction of critical molecular pathways and neurodegeneration and neuronal cell death (Figs. 3, 4, and 5). It still remains unclear as to how these steps or factors work. A combination of genetic, environmental, and epigenetic factors appears to contribute to the disease pathogenesis (Fig. 3, 4, and 5). Neurophysiological tests have shown that the hyperexcitability of the cortex may play an important role in ALS and FTD [3]. It is shown that both ALS and FTD are associated with similar pathogenic mechanisms that involve RNA and protein imbalances, as well as disturbed cellular homeostasis that promotes disease progression (Fig. 3, 4, and 5).
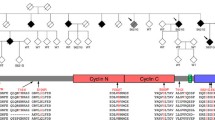
Genetic spectrum of ALS, FTD, and ALS-FTD. The timelines and coordinates of the genes discovered in ALS are shown in (A), those discovered in FTD are shown in (B), and overlapping genes implicated in both ALS as well as FTD patients are shown in (C). Additionally, TDP-43 inclusions were detected in both ALS-FTD patients. The diameter of each gene indicates its relative frequency
New Genetic Risk Loci Identified in ALS and FTD
ALS-FTD, ALS, and FTD genes can be classified into three major pathways, namely, autophagy/proteasome impairment (C9orf72, SQSTM1, TBK1, OPTN, VCP, CHMP2B, UBQLN2, CCNF, FUS, TARDBP), RNA processing (C9orf72, FUS, TARDBP, UBQLN2, SPG11, KIF5A, MATR3, SETX, hnRNPA, PFN1), and cytoskeleton dysfunction (CHCHD10, TBK1, ALS2, FIG1, NEK1, VCP, TUBA4A, CCNF, SQSTM1) [13, 173] (Fig. 3). Risk factors or modifier genes are equally important for understanding the molecular basis of the disease, along with the onset of disease, as well as subtle modulations in the disease phenotype [5]. Recent GWAS studies have revealed numerous novel genes related to ALS and FTD [174]. For example, p.A152T in MAPT increases the risk for FTD, while loss of EPHA4 variants is associated with a higher survival rate in patients with ALS [3, 175]. GWAS analysis identified UNC13A on chromosome 19p13.3, which has been shown to play a role in susceptibility to ALS [6, 176]. Recently, UNC13A was found to be a risk factor in FTD patients along with deposition of TDP-43 [6, 176]. Studies have previously linked C21orf2 (explained above) with SCFD1, MOBP, and TNIP1 to ALS [96, 177]. Currently, gene expression analysis and patient stratification are being conducted in ALS, and this could lead to the development of personalized therapy [178]. GWAS and DNA methylation studies have recently identified a number of genes (DLG1, METTL7A, KIAA1147, PCNX, UBTD2, WDR35, ELP2/SLC39A6), which harbor mutations and perhaps have connections to differential DNA methylation or epigenetic changes leading to the involvement of common pathways [174]. Therefore, these genes indirectly contribute to neurodegeneration in ALS, FTD, and ALS-FTD [174]. Recently, another study has shown that known genetic risk loci for AD/FTD have a nominal causal effect on immune-related traits [115]. It was found that a higher rate of fold change in immune-related genes in FTD was seen when compared with other diseases such as rheumatoid arthritis, ulcerative colitis, and celiac disease [115]. Moreover, a number of other genes comprising of ATXN2, TMEM106B, UNC13A/KCNN1, and ZNF512B have also been identified as modifiers or risks for ALS/FTD with distinctive stages of signs [13, 173]. Thus, these genes/variants with low penetrance have been found in a subgroup of ALS/FTD patients, suggesting that many genetic elements contribute to the ALS/FTD spectrum. Although these risk factors and modifiers are not directly linked to ALS/FTD, they increase the likelihood of developing the disease [13, 115]. Further, these observations indicate that environmental triggers may be also involved, thus leading to oligogenic/polygenic models of ALS/FTD, which have been described in details below. Several recent studies have suggested that ALS/FTD patients may carry more than one genetic variant, which may have additive effect on the disease pathogenesis [3, 17]. However, more studies are needed to identify risk, modifier, and protective factors in ALS/FTD biology to cover the spectrum of these diseases [4, 17].
Oligogenic/Polygenic or Pleiotropy Models in ALS and FTD
As discussed above, ALS is closely linked to FTD at various stages. There can be more than one gene or variant(s) that causes both ALS and FTD; for example, mutations in TARDBP, C9orf72, or TBK1 may occur even within the same family [23, 179]. However, phenotypic pleiotropy is speculated among a multitude of disorders beyond ALS or FTD [180]. In fact, most C9orf72 mutations have also been reported in AD, Alzheimer’s dementia, and PD [85, 181]. Additionally, recently identified TBK1, which has been observed with LoF leading to haploinsufficiency, also could be a cause for cerebellar ataxia and progressive supranuclear palsy [182]. Intriguingly, KIF5A has also been associated with a variety of other neurodegenerative disorders, including hereditary spastic paraplegia and hereditary neuropathy (CMT2A) and neurodevelopmental disorder NEIMY (see above under KIF5A section). Thus, continuous high-throughput sequencing of patients will reveal new genes which will help in dissection of phenotypic nuances.
Oligogenic Architecture of ALS/FTD
Several years ago, it was assumed that most genetic causes of ALS and FTD could be explained by pathogenic mutations that do not co-segregate in carriers or unaffected family members, such as SOD1 in ALS [26]. A similar phenomenon has also been observed in FTD patients [24, 25]. Together with other studies, these observations have addressed genetic penetrance, expressivity, epigenetics, and environmental influences [183, 184]. Moreover, new discoveries in this era of ALS/FTD genetics point at the oligogenic/polygenic or reduced penetrance models for these diseases [4]. Past and ongoing studies indicate a significantly higher number of patients with two or more mutations, which is more than expected, suggesting an additive or synergistic impact of oligogenic diseases on disease penetrance and progression [3, 4, 7, 15, 17]. There is a similar pattern of oligogenic disease onset, penetrance, and progression associated with C9orf72 repeat expansion in ALS [185]. This may be explained by the higher prevalence, incomplete penetrance, and the co-occurrence of numerous other mutations with the C9orf72 repeat expansion in carriers with both ALS-FTD and ALS [184,185,186]. Further, the loss and gain of C9orf72 function in ALS may also allow identification of a common therapeutic pathway in the near future [24, 187]. A recent study has shown that activating the autophagy pathway can ameliorate TDP-43 proteinopathies, in mice models, suggesting the possibility of a potential treatment for neurodegenerative disorders involving TDP-43 inclusions [188]. New therapeutic approaches, including stereopure oligonucleotides, may have some striking correlations with future genetic and model-based research [189]. Numerous other known and novel genes, as well as their inheritance patterns, have resulted in a wide spectrum of ALS-FTD disease presentations [22]. The discovery of ALS and FTD gene mutations has created new ways to diagnose/treat as well as identify disease penetrance. In recent months, advances in sequencing techniques have made it possible to analyze gene panels and to document multiple gene mutations simultaneously in ALS and FTD [179, 190]. An analysis of one such study found that cases with more than one mutation tend to be more penetrant than those with single-gene mutations [179]. According to another study, patients with mutations in more than one gene had a lower survival rate than those with mutations in only one gene. Thus, mutations in multiple genes within one individual lead to a more severe and progressive disease onset (imitating oligogenic and polygenic disease models) [2, 109].
Advances in Novel Gene-Based Therapies
In 1995, riluzole, an antiglutamatergic agent, was identified as the only effective treatment for ALS with a minimum survival time of 3 months [191]. Edaravone (Radicava, a neuroprotective agent) has been approved by the US Food and Drug Administration for the treatment of ALS [192], but no treatment has yet been approved for FTD (https://clinicaltrials.gov/ct2/show/NCT04220021). Nevertheless, various treatment strategies are being used in different drug classes, such as antipsychotics, antidepressants, cholinesterase inhibitors, memantine, riluzole, and serotonin inhibitors, depending on the phenotype [85]. For example, specific serotonin inhibitors are prescribed to treat behavioral problems, and riluzole is prescribed to treat conditions related to motor function or parkinsonism [193]. Despite this, these treatments are effective for up to 6 or 8 months after they are administered to patients with a milder form of the disease [85, 193].
ALS and FTD, however, do not have any other beneficial and effective treatments that can extend life expectancy significantly; hence, it is crucial to continuously develop better therapies, treatments, and patient care [85]. Until now, advances in genetics and molecular understanding of these diseases have led to numerous novel gene identifications and subsequently newer therapeutic approaches [85]. Antisense oligonucleotides (ASOs) and stereopure oligonucleotides, for instance, target majority of the ALS genes [189]. In ongoing clinical trials, it has been shown that ASOs function directly against SOD1 mutations in the CSF of patients without causing any adverse effects [194]. Different phases of clinical trials are currently being conducted with the latest generation of ASOs (NCT02623699). Further, iPSCs generated from fALS patients carrying SOD1 mutations with ASOs have shown decreased apoptotic markers, which indicates improved survival rates [195]. The p.Gly93Ala mutation in SOD1 transgenic mice was shown to delay the disease onset and improved survival when combined with a microRNA linked to niR-155 (miR-155) [196, 197]. Interestingly, copper-based therapeutic diacetylbis(N(4)-methylthiosemicarbazonato) copper II (CuATSM) pet analysis rendered controlled activity in mice with p.Gly93Ala mutation in SOD1 [198]. CuATSM is delivered into the CNS with a metal cofactor (copper), and without this copper, these proteins become misfolded and toxic, potentially leading to motor neuron degeneration [198]. In addition, ASO inhibited the formation of RNA foci linked to C9orf72 repeat expansion and thus prevented dipeptide repeat pathogenesis [19, 199]. In a mouse model, stereotactic ASO injection reduced RNA foci formation by 60–70% in the brain and spinal cord [200]. ASO has been used for ALS and FTD modifiers as well as targeting causal genes. Moreover, ATXN2 has been demonstrated to be associated with ALS pathogenesis and acts primarily as a modifier of TDP-43 toxicity in cellular and animal models [201]. ASO has been used to silence ATXN2, and this has also been validated in knockout mice, demonstrating that the reduced pathology associated with TDP-43 is due to silenced ATXN2 [201].
Other tools, including CRISPR/Cas9 technology and adeno-associated viral vectors, have been used to develop mutant and wild-type alleles for personalized and precision therapy [194, 200]. Systematic review and meta-analysis of human transcriptomics also revealed neuroinflammation [20]. Each technology has its limitations, but an ongoing effort to develop and decipher efficient therapeutics is the basis of any study. The study of epigenetic changes in these genes, such as hypermethylation, is an emerging field that poses new challenges and requires new approaches [174]. In both ALS and FTD research, CRISPR/Cas9 technology is being used to create new animal models for testing genes and their penetrance in patients. This will, in turn, open up new therapeutic options such as personalized, permissive, and precise therapies [187].
Conclusions
Genetic research on ALS and FTD continues to reveal genotype–phenotype correlations based on monogenic/oligogenic gene(s), genetic risk factor(s), and disease modifier(s). Further studies are needed to examine numerous patients and control samples, and this can help in better understanding of the data and increases the statistical power to make new discoveries and validate findings. Our predictions are that new gene discovery will continue, and within a few years, this may lead to more effective treatment options for overlapping conditions. However, progress is being made in modeling oligogenic risks and conquering non-coding regions of the human genome [2, 3]. Genetic findings have shed light on individualized medicine and gene-specific treatments over the past 10–20 years of study [85]. This serves as a proof-of-principle for the treatment of patients using novel techniques such as ASO-based splice modifiers in young patients with spinal muscular atrophy. Patients with SOD1 mutations/C9ORF72 mutations are in clinical trials [19,20,21]. ALS and FTD have similar pathological mechanisms, such as protein aggregation, RNA metabolism, autophagy–lysosomal dysfunction, cytoskeleton and mitochondrial dysfunction, and stress granules, suggesting a causal relationship between the two disorders [4, 19, 177]. The genetic basis of SOD1 in ALS, MAPT, and GRN in FTD is still debated as distinct pathologies, so it is argued that pure ALS and pure FTD cases should be treated separately [57]. It has been suggested that diagnosis and prognosis biomarkers should be developed based on their distinct neuropathological findings. ALS/FTD genetics has been studied using a variety of advanced sequencing tools (e.g., WES, WGS, and targeted sequencing) to discover novel and rare genetic variants. This adds not only to the genetics of ALS/FTD, but also to the genetics of other diseases [143, 169]. Further, by including interaction between genes/epistasis, other aspects of genetics have been dissected, such as reduced penetrance, variable expression, and oligogenic/polygenic/pleiotropic inheritance models [180]. Numerous studies have shown the potential for personalized medicine and future therapies for ALS/FTD in the near future [85, 181].
Summary
Even after years of research in the field of ALS-FTD, it is still not clear as to what extent the two diseases overlap. Several new genes have been identified in ALS-FTD, ALS, and FTD over the past few years (Table 1). There are also phenotypic characteristics common to the ALS and FTD phenotype, such as language and behavioral difficulties (Table 2). The overlap between ALS and FTD is also incomplete; for example, SOD1 is included in ALS cases, while GRN and MAPT are included in FTD cases. It is still unclear what causes their distinct pathologies. Intriguingly, much of the pathophysiology in ALS/FTD shows a massive overlap, which is still a matter of debate among scientists. However, clinical and genetic heterogeneity provides clues beyond these overlays, for example, the C9orf72 repeat expansion, which includes intra- and inter-family variations in disease onset, incidence, penetrance, and progression, as well as cognitive and motor impairments. [24].
It is due to a combination of genes and environmental factors that the disease develops and progresses. Environmental factors can include toxic compounds, heavy metals, and electromagnetic frequencies, and many of these factors have been associated with neurological disorders in humans [202]. A person’s habits and lifestyle are equally important, and studies indicate that metabolic changes such as eating habits, weight loss, insulin resistance, and body cholesterol levels may have an effect on neurodegenerative diseases including ALS and FTD [174] (Fig. 3, 4, and 5). A study conducted with > 700 phenotypic traits and ALS GWAS data (cases = 20,806; controls = 59,804) revealed a shared polygenic risk for this disease [203]. These include smoking, educational attainment, physical activity, anxiety/agitation, and an elevated level of low-density lipoprotein, suggesting more controlled trials are needed to prove causality [203]. Environmental factors do interact with many gene sets and subsets, and this can be related to genes with dissimilar functions. Recent studies on the gene TET2 (which promotes demethylation and is considered an environmental interacting gene) have confirmed the presence of coding and non-coding variants in individuals with neurodegenerative diseases including ALS and FTD patients [204]. TET2 is essential for methylation conversion to 5-hydroxymethylation, a process essential for aging, learning, and memory [204]. Therefore, a combined approach to studying non-coding parts of the genome is also important. In the future, it is crucial that we focus on genetic epidemiology studies, cohort studies based on population data, and clinical phenotyping, which considers epigenetic features, genetics, lifestyle, and environmental factors.
Data Availability
Not applicable.
Code Availability
Not applicable.
Abbreviations
- fALS:
-
Familial amyotrophic lateral sclerosis
- fFTD:
-
Familial frontotemporal dementia
- sALS:
-
Sporadic amyotrophic lateral sclerosis
- sFTD:
-
Sporadic frontotemporal dementia
- TARDBP :
-
TAR DNA-binding protein-43
- TDP-43 :
-
TAR DNA-binding protein 43
- HIV-1:
-
Human immunodeficiency virus
- c9orf72:
-
Chromosome 9 open reading frame 72
- FUS :
-
Fused in sarcoma
- SQSTM1 :
-
Sequestrome-1
- SOD1 :
-
Superoxide dismutase 1
- GRN :
-
Granulin
- MAPT :
-
Microtubule-associated protein tau
- VCP :
-
Valosin-containing protein
- PFN1 :
-
Profilin1
- TBK1 :
-
TANK-binding kinase
- CCNF :
-
Cyclin F
- KIF5A :
-
Kinesin family member 5A
- GLT8D1 :
-
Glycosyltransferase 8 domain containing 1
- NEK1 :
-
NIMA-related kinase 1
- CHMP2B:
-
Charged multivesicular body protein 2B
- CHCHD10:
-
Coiled-coil-helix-coiled-coil-helix domain containing 10
- WGS:
-
Whole-genome sequencing
- WES:
-
Whole-exome sequencing
- GWAS:
-
Genome-wide association studies
- RAN:
-
Repeat-associated non-AUG
- DPR:
-
Dipeptide repeat
- OPTN :
-
Optineurin
- C21orf2 :
-
Chromosome 21 open reading frame 2
- DNAJC7:
-
DnaJ heat shock protein family (Hsp40) member C7
- TBP:
-
TATA-box binding protein
- CTSF:
-
Cathepsin F
- MFSD8:
-
Major facilitator superfamily domain 8
- PGRN:
-
Glycoprotein called progranulin
- TDP-43:
-
TAR DNA-binding protein-43
- hnRNP:
-
Heterogeneous nuclear ribonucleoprotein
- DPRs:
-
Dipeptide repeat proteins
- TBK1:
-
TANK-binding kinase 1
- UMN:
-
Upper motor neuron
- LMN:
-
Lower motor neuron
- FTLD:
-
Frontotemporal lobar degeneration
- FTDP-17T:
-
Frontotemporal dementia with parkinsonism-17 T
- PSP:
-
Progressive supranuclear palsy
- CBD:
-
Corticobasal degeneration
- NFTs:
-
Neurofibrillary tangles
- IHC:
-
Immunohistochemistry
- AD:
-
Alzheimer’s disease
- PD:
-
Parkinson’s disease
- CJD:
-
Creutzfeldt-Jakob disease
- UPS:
-
Ubiquitin–proteasome system
- RRM2:
-
Ribonucleoside-diphosphate reductase subunit M2
- VCP:
-
Valosin-containing protein
- VAPB:
-
VAMP-associated protein B And C
- HNRNPA1:
-
Heterogeneous nuclear ribonucleoprotein A1
- AMPA:
-
α-Amino-3-hydroxyl-5-methyl-4-isoxazole-propionate
- GABAA:
-
γ-Aminobutyric acid type A receptors
- LLPS:
-
Liquid–liquid phase separation
- LCD:
-
Low complexity domain
- TUBA4A:
-
Tubulin Alpha 4a
- ALS2:
-
Alsin
- LAMP-2:
-
Lysosomal associated membrane protein-2
- CTSD:
-
Cathepsin D
- CSF:
-
Cerebrospinal fluid
- ASOs:
-
Antisense oligonucleotides
- CuATSM:
-
Diacetylbis(N(4)-methylthiosemicarbazonato) copper II
- iPSCs:
-
Induced pluripotent stem cells
- CSF:
-
Cerebrospinal fluid
References
Abramzon YA, Fratta P, Traynor BJ, Chia R (2020) The overlapping genetics of amyotrophic lateral sclerosis and frontotemporal dementia. Front Neurosci 14:42
Ji AL, Zhang X, Chen WW, Huang WJ (2017) Genetics insight into the amyotrophic lateral sclerosis/frontotemporal dementia spectrum. J Med Genet 54. https://doi.org/10.1136/jmedgenet-2016-104271
Lattante S, Ciura S, Rouleau GA, Kabashi E (2015) Defining the genetic connection linking amyotrophic lateral sclerosis (ALS) with frontotemporal dementia (FTD). Trends Genet 31:263–73
Nguyen HP, Van Broeckhoven C, van der Zee J (2018) ALS genes in the genomic era and their implications for FTD. Trends Genet 34
Brenner D, Weishaupt JH (2019) Update on amyotrophic lateral sclerosis genetics. CurrOpin Neurol 32
Sirkis DW, Geier EG, Bonham LW, et al (2019) Recent advances in the genetics of frontotemporal dementia. Curr Genet Med Rep 7. https://doi.org/10.1007/s40142-019-0160-6
Keogh MJ, Wei W, Aryaman J, et al (2018) Oligogenic genetic variation of neurodegenerative disease genes in 980 postmortem human brains. J NeurolNeurosurg Psychiatry 89. https://doi.org/10.1136/jnnp-2017-317234
Bradley WG, Andrew AS, Traynor BJ, et al (2019) Gene-environment-time interactions in neurodegenerative diseases: hypotheses and research approaches. Ann Neurosci 25. https://doi.org/10.1159/000495321
Chou CC, Zhang Y, Umoh ME, et al (2018) TDP-43 pathology disrupts nuclear pore complexes and nucleocytoplasmic transport in ALS/FTD. Nat Neurosci 21. https://doi.org/10.1038/s41593-017-0047-3
Parobkova E, Matej R (2021) Amyotrophic lateral sclerosis and frontotemporal lobar degenerations: similarities in genetic background. Diagnostics 11
Conforti FL, Renton AE, Houlden H (2021) Editorial: multifaceted genes in amyotrophic lateral sclerosis-frontotemporal dementia. Front Neurosci 15
Chia R, Chiò A, Traynor BJ (2018) Novel genes associated with amyotrophic lateral sclerosis: diagnostic and clinical implications. Lancet Neurol 17
Guerreiro R, Gibbons E, Tábuas-Pereira M, et al (2020) Genetic architecture of common non-Alzheimer’s disease dementias. Neurobiol Dis 142
Ciani M, Benussi L, Bonvicini C, Ghidoni R (2019) Genome wide association study and next generation sequencing: a glimmer of light toward new possible horizons in frontotemporal dementia research. Front Neurosci 13. https://doi.org/10.3389/fnins.2019.00506
Zhang S, Cooper-Knock J, Weimer AK, et al (2021) Genome-wide identification of the genetic basis of amyotrophic lateral sclerosis. SSRN Electron J. https://doi.org/10.2139/ssrn.3744427
Seelaar H, Rohrer JD, Pijnenburg YAL, et al (2011) Clinical, genetic and pathological heterogeneity of frontotemporal dementia: a review. J Neurol Neurosurg Psychiatry 82
Pottier C, Bieniek KF, Finch NC, et al (2015) Whole-genome sequencing reveals important role for TBK1 and OPTN mutations in frontotemporal lobar degeneration without motor neuron disease. Acta Neuropathol 130. https://doi.org/10.1007/s00401-015-1436-x
Marangi G, Traynor BJ (2015) Genetic causes of amyotrophic lateral sclerosis: new genetic analysis methodologies entailing new opportunities and challenges. Brain Res 1607
Lagier-Tourenne C, Baughn M, Rigo F, et al (2013) Targeted degradation of sense and antisense C9orf72 RNA foci as therapy for ALS and frontotemporal degeneration. Proc Natl Acad Sci U S A 110. https://doi.org/10.1073/pnas.1318835110
Martier R, Liefhebber JM, García-Osta A, et al (2019) Targeting RNA-mediated toxicity in C9orf72 ALS and/or FTD by RNAi-based gene therapy. Mol Ther - Nucleic Acids 16. https://doi.org/10.1016/j.omtn.2019.02.001
Berry JD, Cudkowicz ME, Windebank AJ, et al (2019) NurOwn, phase 2, randomized, clinical trial in patients with ALS: safety, clinical, and biomarker results. Neurology 93.https://doi.org/10.1212/WNL.0000000000008620
Noori A, Mezlini AM, Hyman BT, et al (2021) Systematic review and meta-analysis of human transcriptomics reveals neuroinflammation, deficient energy metabolism, and proteostasis failure across neurodegeneration. Neurobiol Dis 149. https://doi.org/10.1016/j.nbd.2020.105225
Renton AE, Chiò A, Traynor BJ (2014) State of play in amyotrophic lateral sclerosis genetics. Nat Neurosci 17
DeJesus-Hernandez M, Mackenzie IR, Boeve BF, et al (2011) Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron 72. https://doi.org/10.1016/j.neuron.2011.09.011
Renton AE, Majounie E, Waite A, et al (2011) A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron 72.https://doi.org/10.1016/j.neuron.2011.09.010
Rosen DR, Siddique T, Patterson D, et al (1993) Mutations in Cu/Zn superoxide dismutase gene are associated with familial amyotrophic lateral sclerosis. Nature 362.https://doi.org/10.1038/362059a0
Gurney ME, Pu H, Chiu AY, et al (1994) Motor neuron degeneration in mice that express a human Cu Zn superoxide dismutase mutation. Science (80- ) 264 https://doi.org/10.1126/science.8209258
Andersen PM (2006) Amyotrophic lateral sclerosis associated with mutations in the CuZn superoxide dismutase gene. Curr Neurol Neurosci Rep 6
de AraújoBrasil A, de Carvalho MDC, Gerhardt E, et al (2019) Characterization of the activity, aggregation, and toxicity of heterodimers of WT and ALS-associated mutant Sod1. Proc Natl Acad Sci U S A 116. https://doi.org/10.1073/pnas.1902483116
Sau D, De Biasi S, Vitellaro-Zuccarello L, et al (2007) Mutation of SOD1 in ALS: a gain of a loss of function. Hum Mol Genet 16. https://doi.org/10.1093/hmg/ddm110
Perrone B, Conforti FL (2020) Common mutations of interest in the diagnosis of amyotrophic lateral sclerosis: how common are common mutations in ALS genes? Expert Rev Mol Diagn 20
Sibilla C, Bertolotti A (2017) Prion properties of SOD1 in amyotrophic lateral sclerosis and potential therapy. Cold Spring Harb Perspect Biol 9. https://doi.org/10.1101/cshperspect.a024141
Zelko IN, Mariani TJ, Folz RJ (2002) Superoxide dismutase multigene family: a comparison of the CuZn-SOD (SOD1), Mn-SOD (SOD2), and EC-SOD (SOD3) gene structures, evolution, and expression. Free Radic Biol Med 33
McCord JM, Fridovich I (1969) Superoxide dismutase. An enzymic function for erythrocuprein (hemocuprein). J Biol Chem 244
Dröge W (2002) Free radicals in the physiological control of cell function. Physiol Rev 82
Rhee SG, Yang KS, Kang SW, et al (2005) Controlled elimination of intracellular H2O2: regulation of peroxiredoxin, catalase, and glutathione peroxidase via post-translational modification. Antioxidants Redox Signal 7
O’Brien EM, Dirmeier R, Engle M, Poyton RO (2004) Mitochondrial protein oxidation in yeast mutants lacking manganese- (MnSOD) or copper- and zinc-containing superoxide dismutase (CuZnSOD): evidence that mnsod and cuznsod have both unique and overlapping functions in protecting mitochondrial proteins from. J BiolChem 279. https://doi.org/10.1074/jbc.M405958200
Aquilano K, Vigilanza P, Rotilio G, et al (2006) Mitochondrial damage due to SOD1 deficiency in SH‐SY5Y neuroblastoma cells: a rationale for the redundancy of SOD1. FASEB J 20. https://doi.org/10.1096/fj.05-5225fje
Klöppel C, Michels C, Zimmer J, et al (2010) In yeast redistribution of Sod1 to the mitochondrial intermembrane space provides protection against respiration derived oxidative stress. Biochem Biophys Res Commun 403. https://doi.org/10.1016/j.bbrc.2010.10.129
Fischer LR, Igoudjil A, Magrané J, et al (2011) SOD1 targeted to the mitochondrial intermembrane space prevents motor neuropathy in the Sod1 knockout mouse. Brain 134.https://doi.org/10.1093/brain/awq314
Vehviläinen P, Koistinaho J, Goldsteins G (2014) Mechanisms of mutant SOD1 induced mitochondrial toxicity in amyotrophic lateral sclerosis. Front Cell Neurosci 8
Valentine JS, Hart PJ (2003) Misfolded CuZnSOD and amyotrophic lateral sclerosis. Proc Natl Acad Sci U S A 100
Pansarasa O, Bordoni M, Diamanti L, et al 2018 Sod1 in amyotrophic lateral sclerosis: “ambivalent” behavior connected to the disease Int J Mol Sci 19
Zarei S, Carr K, Reiley L, et al (2015) A comprehensive review of amyotrophic lateral sclerosis. Surg Neurol Int 6
Brownell AL, Kuruppu D, Kil KE, et al (2015) PET imaging studies show enhanced expression of mGluR5 and inflammatory response during progressive degeneration in ALS mouse model expressing SOD1-G93A gene. J Neuroinflammation 12.https://doi.org/10.1186/s12974-015-0439-9
Benkler C, O’Neil AL, Slepian S, et al (2018) Aggregated SOD1 causes selective death of cultured human motor neurons. Sci Rep 8. https://doi.org/10.1038/s41598-018-34759-z
Grad LI, Rouleau GA, Ravits J, Cashman NR (2017) Clinical spectrum of amyotrophic lateral sclerosis (ALS). Cold Spring Harb Perspect Med 7
Saccon RA, Bunton-Stasyshyn RKA, Fisher EMC, Fratta P (2013) Is SOD1 loss of function involved in amyotrophic lateral sclerosis? Brain 136
Saba L, Viscomi MT, Caioli S, et al (2016) Altered functionality, morphology, and vesicular glutamate transporter expression of cortical motor neurons from a presymptomatic mouse model of amyotrophic lateral sclerosis. Cereb Cortex 26. https://doi.org/10.1093/cercor/bhu317
van den Bos MAJ, Geevasinga N, Higashihara M, et al (2019) Pathophysiology and diagnosis of ALS: insights from advances in neurophysiological techniques. Int J Mol Sci 20
Özdinler PH, Benn S, Yamamoto TH, et al (2011) Corticospinal motor neurons and related subcerebral projection neurons undergo early and specific neurodegeneration in hSOD1G93A transgenic ALS mice. J Neurosci 31. https://doi.org/10.1523/JNEUROSCI.4184-10.2011
Wainger BJ, Kiskinis E, Mellin C, et al (2014) Intrinsic membrane hyperexcitability of amyotrophic lateral sclerosis patient-derived motor neurons. Cell Rep 7. https://doi.org/10.1016/j.celrep.2014.03.019
Saxena S, Roselli F, Singh K, et al (2013) Neuroprotection through excitability and mTOR required in ALS motoneurons to delay disease and extend survival. Neuron 80. https://doi.org/10.1016/j.neuron.2013.07.027
Leroy F, Lamotted’Incamps B, Imhoff-Manuel RD, Zytnicki D (2014) Early intrinsic hyperexcitability does not contribute to motoneuron degeneration in amyotrophic lateral sclerosis. Elife 3. https://doi.org/10.7554/eLife.04046
Baker M, Mackenzie IR, Pickering-Brown SM, et al (2006) Mutations in progranulin cause tau-negative frontotemporal dementia linked to chromosome 17. Nature 442.https://doi.org/10.1038/nature05016
Gass J, Prudencio M, Stetler C, Petrucelli L (2012) Progranulin: an emerging target for FTLD therapies. Brain Res 1462
Ferrari R, Hernandez DG, Nalls MA, et al (2014) Frontotemporal dementia and its subtypes: a genome-wide association study. Lancet Neurol 13.https://doi.org/10.1016/S1474-4422(14)70065-1
De Muynck L, Van Damme P (2011) Cellular effects of progranulin in health and disease. In J Mole Neurosci
Holler CJ, Taylor G, Deng Q, Kukar T (2017) Intracellular proteolysis of progranulin generates stable, lysosomal granulins that are haploinsufficient in patients with frontotemporal dementia caused by GRN mutations. eNeuro 4 https://doi.org/10.1523/ENEURO.0100-17.2017
Chang MC, Srinivasan K, Friedman BA, et al (2017) Progranulin deficiency causes impairment of autophagy and TDP-43 accumulation. J Exp Med 214.https://doi.org/10.1084/jem.20160999
Amin S, Carling G, Gan L (2022) New insights and therapeutic opportunities for progranulin-deficient frontotemporal dementia. Curr Opin Neurobiol 72
Valdez C, Wong YC, Schwake M, et al (2017) Progranulin-mediated deficiency of cathepsin D results in FTD and NCL-like phenotypes in neurons derived from FTD patients. Hum Mol Genet 26. https://doi.org/10.1093/hmg/ddx364
Ward ME, Taubes A, Chen R, et al (2014) Early retinal neurodegeneration and impaired Ran-mediated nuclear import of TDP-43 in progranulin-deficient FTLD. J Exp Med 211.https://doi.org/10.1084/jem.20140214
Azam S, Haque ME, Kim IS, Choi DK (2021) Microglial turnover in ageing-related neurodegeneration: therapeutic avenue to intervene in disease progression. Cells 10
Krabbe G, Minami SS, Etchegaray JI, et al (2017) Microglial NFκB-TNFα hyperactivation induces obsessive-compulsive behavior in mouse models of progranulin-deficient frontotemporal dementia. Proc Natl Acad Sci U S A 114. https://doi.org/10.1073/pnas.1700477114
Weingarten MD, Lockwood AH, Hwo SY, Kirschner MW (1975) A protein factor essential for microtubule assembly. ProcNatl Acad Sci U S A 72. https://doi.org/10.1073/pnas.72.5.1858
Ghetti B, Oblak AL, Boeve BF, et al (2015) Invited review: frontotemporal dementia caused by microtubule-associated protein tau gene (MAPT) mutations: a chameleon for neuropathology and neuroimaging. Neuropathol Appl Neurobiol 41 115
Spillantini MG, Bird TD, Ghetti B (1998) Frontotemporal dementia and parkinsonism linked to chromosome 17: a new group of tauopathies. In Brain Pathology
Poorkaj P, Bird TD, Wijsman E, et al (1998) Tau is a candidate gene for chromosome 17 frontotemporal dementia. Ann Neurol 43. https://doi.org/10.1002/ana.410430617
Hutton M, Lendon CL, Rizzu P, et al (1998) Association of missense and 5’-splice-site mutations in tau with the inherited dementia FTDP-17. Nature 393.https://doi.org/10.1038/31508
Spillantini MG, Murrell JR, Goedert M, et al (1998) Mutation in the tau gene in familial multiple system tauopathy with presenile dementia. Proc Natl Acad Sci U S A 95. https://doi.org/10.1073/pnas.95.13.7737
Lee Virginia MY, Zhukareva V, Vogelsberg-Ragaglia V, et al (1998) Mutation-specific functional impairments in distinct tau isoforms of hereditary FTDP-17. Science (80- ) 282 https://doi.org/10.1126/science.282.5395.1914
Poorkaj P, Grossman M, Steinbart E, et al (2001) Frequency of tau gene mutations in familial and sporadic cases of non-Alzheimer dementia. Arch Neurol 58. https://doi.org/10.1001/archneur.58.3.383
Pittman AM, Myers AJ, Abou-Sleiman P, et al (2005) Linkage disequilibrium fine mapping and haplotype association analysis of the tau gene in progressive supranuclear palsy and corticobasal degeneration. J Med Genet 42.https://doi.org/10.1136/jmg.2005.031377
Lillo P, Hodges JR (2009) Frontotemporal dementia and motor neurone disease: overlapping clinic-pathological disorders. J Clin Neurosci 16
Lee VMY, Goedert M, Trojanowski JQ (2001) Neurodegenerative tauopathies. Annu Rev Neurosci 24
Mackenzie IRA, Neumann M (2016) Molecular neuropathology of frontotemporal dementia: insights into disease mechanisms from postmortem studies. J Neurochem
Halliday G, Bigio EH, Cairns NJ, et al (2012) Mechanisms of disease in frontotemporal lobar degeneration: gain of function versus loss of function effects. Acta Neuropathol 124
Raffaele F, Claudia M, John H (2019) Genetics and molecular mechanisms of frontotemporal lobar degeneration: an update and future avenues. Neurobiol Aging 78
Bennion Callister J, Pickering-Brown SM (2014) Pathogenesis/genetics of frontotemporal dementia and how it relates to ALS. Exp Neurol 262
Van Blitterswijk M, Dejesus-Hernandez M, Rademakers R (2012) How do C9ORF72 repeat expansions cause amyotrophic lateral sclerosis and frontotemporal dementia: can we learn from other noncoding repeat expansion disorders? Curr Opin Neurol 25
Hsiung GYR, Dejesus-Hernandez M, Feldman HH, et al (2012) Clinical and pathological features of familial frontotemporal dementia caused by C9ORF72 mutation on chromosome 9p. Brain 135. https://doi.org/10.1093/brain/awr354
Byrne S, Elamin M, Bede P, et al (2012) Cognitive and clinical characteristics of patients with amyotrophic lateral sclerosis carrying a C9orf72 repeat expansion: A population-based cohort study. Lancet Neurol 11.https://doi.org/10.1016/S1474-4422(12)70014-5
Zhu Q, Jiang J, Gendron TF, et al (2020) Reduced C9ORF72 function exacerbates gain of toxicity from ALS/FTD-causing repeat expansion in C9orf72. Nat Neurosci 23. https://doi.org/10.1038/s41593-020-0619-5
Ferrari R, Kapogiannis D, D. Huey E, Momeni P (2011) FTD and ALS: a tale of two diseases. Curr Alzheimer Res 8.https://doi.org/10.2174/156720511795563700
Ou SH, Wu F, Harrich D, et al (1995) Cloning and characterization of a novel cellular protein, TDP-43, that binds to human immunodeficiency virus type 1 TAR DNA sequence motifs. J Virol 69. https://doi.org/10.1128/jvi.69.6.3584-3596.1995
Liscic RM, Grinberg LT, Zidar J, et al (2008) ALS and FTLD: two faces of TDP-43 proteinopathy. Eur J Neurol 15
Prasad A, Bharathi V, Sivalingam V, et al (2019) Molecular mechanisms of TDP-43 misfolding and pathology in amyotrophic lateral sclerosis Front Mol Neurosci 12
Geser F, Martinez-Lage M, Kwong LK et al (2009) Amyotrophic lateral sclerosis frontotemporal dementia and beyond:The TDP-43 diseases. J Neurol 256:1205
Forman MS, Trojanowski JQ, Lee VMY (2007) TDP-43: a novel neurodegenerative proteinopathy. Curr Opin Neurobiol 17:548–55
Charif SE, Vassallu MF, Salvañal L, Igaz LM (2022) Protein synthesis modulation as a therapeutic approach for amyotrophic lateral sclerosis and frontotemporal dementia. Neural Regen Res 17:1423–1430. https://doi.org/10.4103/1673-5374.330593
Schludi MH, May S, Grässer FA, et al (2015) Distribution of dipeptide repeat proteins in cellular models and C9orf72 mutation cases suggests link to transcriptional silencing. Acta Neuropathol 130. https://doi.org/10.1007/s00401-015-1450-z
Cirulli ET, Lasseigne BN, Petrovski S et al (2015) Exome sequencing in amyotrophic lateral sclerosis identifies risk genes and pathways. Science 347:1436–41. https://doi.org/10.1126/science.aaa3650
Freischmidt A, Wieland T, Richter B, et al (2015) Haploinsufficiency of TBK1 causes familial ALS and fronto-temporal dementia. Nat Neurosci 18. https://doi.org/10.1038/nn.4000
Gijselinck I, Van Mossevelde S, Van Der Zee J, et al (2015) Loss of TBK1 is a frequent cause of frontotemporal dementia in a Belgian cohort. Neurology 85. https://doi.org/10.1212/WNL.0000000000002220
Van Rheenen W, Shatunov A, Dekker AM, et al (2016) Genome-wide association analyses identify new risk variants and the genetic architecture of amyotrophic lateral sclerosis. Nat Genet 48. https://doi.org/10.1038/ng.3622
Van Mossevelde S, Van Der Zee J, Gijselinck I, et al (2016) Clinical features of TBK1 carriers compared with C9orf72, GRN and non-mutation carriers in a Belgian cohort. Brain 139.https://doi.org/10.1093/brain/awv358
de Majo M, Topp SD, Smith BN, et al (2018) ALS-associated missense and nonsense TBK1 mutations can both cause loss of kinase function. Neurobiol Aging 71. https://doi.org/10.1016/j.neurobiolaging.2018.06.015
Cui R, Tuo M, Li P, Zhou C (2018) Association between TBK1 mutations and risk of amyotrophic lateral sclerosis/frontotemporal dementia spectrum: a meta-analysis. Neurol Sci 39:811–820
van der Zee J, Gijselinck I, Van Mossevelde S, et al (2017) TBK1 mutation spectrum in an extended european patient cohort with frontotemporal dementia and amyotrophic lateral sclerosis. Hum Mutat 38. https://doi.org/10.1002/humu.23161
Dols-Icardo O, García-Redondo A, Rojas-García R, et al (2018) Analysis of known amyotrophic lateral sclerosis and frontotemporal dementia genes reveals a substantial genetic burden in patients manifesting both diseases not carrying the C9orf72 expansion mutation. J Neurol Neurosurg Psychiatry 89. https://doi.org/10.1136/jnnp-2017-316820
Duan W, Guo M, Yi L et al (2019) Deletion of Tbk1 disrupts autophagy and reproduces behavioral and locomotor symptoms of FTD-ALS in mice. Aging (Albany NY) 11:2457–2476. https://doi.org/10.18632/aging.101936
Bonnard M (2000) Deficiency of T2K leads to apoptotic liver degeneration and impaired NF-kappaB-dependent gene transcription. EMBO J 19. https://doi.org/10.1093/emboj/19.18.4976
Korac J, Schaeffer V, Kovacevic I, et al (2013) Ubiquitin-independent function of optineurin in autophagic clearance of protein aggregates. J Cell Sci 126. https://doi.org/10.1242/jcs.114926
Heo JM, Ordureau A, Paulo JA, et al (2015) The PINK1-PARKIN mitochondrial ubiquitylation pathway drives a program of OPTN/NDP52 recruitment and TBK1 activation to promote mitophagy. Mol Cell 60. https://doi.org/10.1016/j.molcel.2015.08.016
Weidberg H, Elazar Z (2011) TBK1 mediates crosstalk between the innate immune response and autophagy. Sci Signal 4:pe39
Uhlen M, Uhlén M, Fagerberg L et al (2015) Proteomics Tissue-based map of the human proteome. Science 347:1260419
Li X, Zhang Q, Ding Y, et al (2016) Methyltransferase Dnmt3a upregulates HDAC9 to deacetylate the kinase TBK1 for activation of antiviral innate immunity. Nat Immunol 17. https://doi.org/10.1038/ni.3464
Lattante S, Doronzio PN, Marangi G, et al (2019) Coexistence of variants in TBK1 and in other ALS-related genes elucidates an oligogenic model of pathogenesis in sporadic ALS. Neurobiol Aging 84. https://doi.org/10.1016/j.neurobiolaging.2019.03.010
Müller K, Brenner D, Weydt P, et al (2018) Comprehensive analysis of the mutation spectrum in 301 German ALS families. J NeurolNeurosurg Psychiatry 89. https://doi.org/10.1136/jnnp-2017-317611
Pozzi L, Valenza F, Mosca L, et al (2017) TBK1 mutations in Italian patients with amyotrophic lateral sclerosis: genetic and functional characterization. J Neurol Neurosurg Psychiatry 88. https://doi.org/10.1136/jnnp-2017-316174
Williams KL, Topp S, Yang S, et al (2016) CCNF mutations in amyotrophic lateral sclerosis and frontotemporal dementia. Nat Commun 7. https://doi.org/10.1038/ncomms11253
Pan C, Jiao B, Xiao T, et al (2017) Mutations of CCNF gene is rare in patients with amyotrophic lateral sclerosis and frontotemporal dementia from Mainland China. Amyotroph Lateral Scler Front Degener 18.https://doi.org/10.1080/21678421.2017.1293111
Galper J, Rayner SL, Hogan AL et al (2017) Cyclin F: a component of an E3 ubiquitin ligase complex with roles in neurodegeneration and cancer. Int J Biochem Cell Biol 89:216–220
Li QS, Tian C, Hinds D, Seabrook GR (2020) The association of clinical phenotypes to known AD/FTD genetic risk loci and their interrelationship. PLoS One 15.https://doi.org/10.1371/journal.pone.0241552
Lee A, Rayner SL, Gwee SSL, et al (2018) Pathogenic mutation in the ALS/FTD gene, CCNF, causes elevated Lys48-linked ubiquitylation and defective autophagy. Cell Mol Life Sci 75.https://doi.org/10.1007/s00018-017-2632-8
Hogan AL, Don EK, Rayner SL, et al (2017) Expression of ALS/FTD-linked mutant CCNF in zebrafish leads to increased cell death in the spinal cord and an aberrant motor phenotype. Hum Mol Genet 26. https://doi.org/10.1093/hmg/ddx136
Yu Y, Nakagawa T, Morohoshi A, et al (2019) Pathogenic mutations in the ALS gene CCNF cause cytoplasmic mislocalization of cyclin F and elevated VCP ATPase activity. Hum Mol Genet 28. https://doi.org/10.1093/hmg/ddz119
Brenner D, Yilmaz R, Müller K, et al (2018) Hot-spot KIF5A mutations cause familial ALS. Brain 141.https://doi.org/10.1093/brain/awx370
Nicolas A, Kenna K, Renton AE, et al (2018) Genome-wide analyses identify KIF5A as a novel ALS gene. Neuron 97.https://doi.org/10.1016/j.neuron.2018.02.027
Hirokawa N, Niwa S, Tanaka Y (2010) Molecular motors in neurons: transport mechanisms and roles in brain function development and disease. Neuron 68:610–38
Millecamps S, Julien JP (2013) Axonal transport deficits and neurodegenerative diseases. Nat Rev Neurosci 14:161–76
Kanai Y, Dohmae N, Hirokawa N (2004) Kinesin transports RNA: isolation and characterization of an RNA-transporting granule. Neuron 43.https://doi.org/10.1016/j.neuron.2004.07.022
Guo W, Naujock M, Fumagalli L, et al (2017) HDAC6 inhibition reverses axonal transport defects in motor neurons derived from FUS-ALS patients. Nat Commun 8. https://doi.org/10.1038/s41467-017-00911-y
Matsuzaki F, Shirane M, Matsumoto M, Nakayama KI (2011) Protrudin serves as an adaptor molecule that connects KIF5 and its cargoes in vesicular transport during process formation. Mol Biol Cell 22. https://doi.org/10.1091/mbc.E11-01-0068
Xia CH, Roberts EA, Her LS, et al (2003) Abnormalneurofilament transport caused by targeted disruption of neuronal kinesin heavy chain KIF5A. J Cell Biol 161. https://doi.org/10.1083/jcb.200301026
Al-Chalabi A, Andersen PM, Nilsson P, et al (1999) Deletions of the heavy neurofilament subunit tail in amyotrophic lateral sclerosis. Hum Mol Genet 8. https://doi.org/10.1093/hmg/8.2.157
Karle KN, Möckel D, Reid E, Schöls L (2012) Axonal transport deficit in a KIF5A -/- mouse model. Neurogenetics 13.https://doi.org/10.1007/s10048-012-0324-y
Smith EF, Shaw PJ, De Vos KJ (2019) The role of mitochondria in amyotrophic lateral sclerosis. Neurosci Lett 710:132933
Heisler FF, Lee HK, Gromova K V., et al (2014) GRIP1 interlinks N-cadherin and AMPA receptors at vesicles to promote combined cargo transport into dendrites. Proc Natl Acad Sci U S A 111. https://doi.org/10.1073/pnas.1304301111
Nakajima K, Yin X, Takei Y, et al (2012) Molecular motor KIF5A is essential for GABAA receptor transport, and KIF5A deletion causes epilepsy. Neuron 76.https://doi.org/10.1016/j.neuron.2012.10.012
Thiel C, Kessler K, Giessl A, et al (2011) NEK1 mutations cause short-rib polydactyly syndrome type majewski. Am J Hum Genet 88. https://doi.org/10.1016/j.ajhg.2010.12.004
Wu CH, Fallini C, Ticozzi N, et al (2012) Mutations in the profilin 1 gene cause familial amyotrophic lateral sclerosis. Nature 488.https://doi.org/10.1038/nature11280
Campbell PD, Shen K, Sapio MR, et al (2014) Unique function of kinesin Kif5A in localization of mitochondria in axons. J Neurosci 34.https://doi.org/10.1523/JNEUROSCI.2770-14.2014
Hares K, Kemp K, Loveless S, et al (2021) KIF5A and the contribution of susceptibility genotypes as a predictive biomarker for multiple sclerosis. J Neurol 268.https://doi.org/10.1007/s00415-020-10373-w
Cooper-Knock J, Moll T, Ramesh T, et al (2019) Mutations in the glycosyltransferase domain of GLT8D1 are associated with familial amyotrophic lateral sclerosis. Cell Rep 26. https://doi.org/10.1016/j.celrep.2019.02.006
Harschnitz O, Jongbloed BA, Franssen H, et al (2014) MMN: from immunological cross-talk to conduction block. J Clin Immunol 34. https://doi.org/10.1007/s10875-014-0026-3
Yu RK, Tsai YT, Ariga T, Yanagisawa M (2011) Structures, biosynthesis, and functions of gangliosides-an overview. J Oleo Sci 60:537–44
Moll T, Shaw PJ, Cooper-Knock J (2020) Disrupted glycosylation of lipids and proteins is a cause of neurodegeneration. Brain 143.https://doi.org/10.1093/brain/awz358
Sasayama D, Hori H, Yamamoto N, et al (2014) ITIH3 polymorphism may confer susceptibility to psychiatric disorders by altering the expression levels of GLT8D1. J Psychiatr Res 50. https://doi.org/10.1016/j.jpsychires.2013.12.002
Yang CP, Li X, Wu Y, et al (2018) Comprehensive integrative analyses identify GLT8D1 and CSNK2B as schizophrenia risk genes. Nat Commun 9. https://doi.org/10.1038/s41467-018-03247-3
Mackenzie IR, Nicholson AM, Sarkar M, et al (2017) TIA1 mutations in amyotrophic lateral sclerosis and frontotemporal dementia promote phase separation and alter stress granule dynamics. Neuron 95.https://doi.org/10.1016/j.neuron.2017.07.025
Brand P, Dyck PJB, Liu J, et al (2016) Distal myopathy with coexisting heterozygous TIA1 and MYH7 Variants. NeuromusculDisord 26. https://doi.org/10.1016/j.nmd.2016.05.012
Hackman P, Sarparanta J, Lehtinen S, et al (2013) Welander distal myopathy is caused by a mutation in the RNA-binding protein TIA1. Ann Neurol 73. https://doi.org/10.1002/ana.23831
Klar J, Sobol M, Melberg A, et al (2013) Welander distal myopathy caused by an ancient founder mutation in TIA1 associated with perturbed splicing. Hum Mutat 34. https://doi.org/10.1002/humu.22282
van der Spek RA, van Rheenen W, Pulit SL et al (2018) Reconsidering the causality of TIA1 mutations in ALS. Amyotroph. Lateral Scler Front Degener 19:1–3
Hirsch-Reinshagen V, Nicholson A, Pottier C, et al (2018) Clinical and neuropathological features of ALS/FTD with TIA1 mutations . Can J NeurolSci / J Can des Sci Neurol 45. https://doi.org/10.1017/cjn.2018.48
Hirsch-Reinshagen V, Pottier C, Nicholson AM, et al (2017) Clinical and neuropathological features of ALS/FTD with TIA1 mutations. ActaNeuropathol Commun 5. https://doi.org/10.1186/s40478-017-0493-x
Brenner D, Müller K, Wieland T et al (2016) NEK1 mutations in familial amyotrophic lateral sclerosis. Brain 139:e28
Kenna KP, Van Doormaal PTC, Dekker AM, et al (2016) NEK1 variants confer susceptibility to amyotrophic lateral sclerosis. Nat Genet 48. https://doi.org/10.1038/ng.3626
Chen Y, Chen CF, Riley DJ, Chen PL (2011) Nek1 kinase functions in DNA damage response and checkpoint control through a pathway independent of ATM and ATR. Cell Cycle 10.https://doi.org/10.4161/cc.10.4.14814
Higelin J, Catanese A, Semelink-Sedlacek LL, et al (2018) NEK1 loss-of-function mutation induces DNA damage accumulation in ALS patient-derived motoneurons. Stem Cell Res 30. https://doi.org/10.1016/j.scr.2018.06.005
Shalom O, Shalva N, Altschuler Y, Motro B (2008) The mammalian Nek1 kinase is involved in primary cilium formation. FEBS Lett 582.https://doi.org/10.1016/j.febslet.2008.03.036
Chang J, Baloh RH, Milbrandt J (2009) The NIMA-family kinase Nek3 regulates microtubule acetylation in neurons. J Cell Sci 122.https://doi.org/10.1242/jcs.048975
Smith BN, Ticozzi N, Fallini C, et al (2014) Exome-wide rare variant analysis identifies TUBA4A mutations associated with familial ALS. Neuron 84.https://doi.org/10.1016/j.neuron.2014.09.027
Fang X, Lin H, Wang X, et al (2015) The NEK1 interactor, C21ORF2, is required for efficient DNA damage repair. ActaBiochim Biophys Sin (Shanghai) 47. https://doi.org/10.1093/abbs/gmv076
Amin P, Florez M, Najafov A, et al (2018) Regulation of a distinct activated RIPK1 intermediate bridging complex I and complex II in TNFα-mediated apoptosis. ProcNatl Acad Sci U S A 115. https://doi.org/10.1073/pnas.1806973115
Khan AO, Eisenberger T, Nagel-Wolfrum K, et al (2015) C21orf2 is mutated in recessive earlyonset retinal dystrophy with macular staphyloma and encodes a protein that localises to the photoreceptor primary cilium. Br J Ophthalmol 99.https://doi.org/10.1136/bjophthalmol-2015-307277
Wang Z, Horemuzova E, Iida A, et al (2017) Axial spondylometaphyseal dysplasia is also caused by NEK1 mutations. J Hum Genet 62.https://doi.org/10.1038/jhg.2016.157
Wheway G, Schmidts M, Mans DA, et al (2015) An siRNA-based functional genomics screen for the identification of regulators of ciliogenesis and ciliopathy genes. Nat Cell Biol 17. https://doi.org/10.1038/ncb3201
Farhan SMK, Howrigan DP, Abbott LE, et al (2019) Exome sequencing in amyotrophic lateral sclerosis implicates a novel gene, DNAJC7, encoding a heat-shock protein. Nat Neurosci 22. https://doi.org/10.1038/s41593-019-0530-0
Farhan SMK, Howrigan DP, Abbott LE et al (2019) Exome sequencing in amyotrophic lateral sclerosis implicates a novel gene, DNAJC7, encoding a heat-shock protein. Nat Neurosci 22:1966–1974. https://doi.org/10.1038/s41593-019-0530-0
Olszewska DA, Fallon EM, Pastores GM, et al (2019) Autosomal dominant gene negative frontotemporal dementia-think of SCA17. Cerebellum 18. https://doi.org/10.1007/s12311-018-0998-2
Ranganathan R, Haque S, Coley K et al (2020) Multifaceted genes in amyotrophic lateral sclerosis-frontotemporal dementia. Front Neurosci 14:684
Van Der Zee J, Mariën P, Crols R, et al (2016) Mutated CTSF in adult-onset neuronal ceroidlipofuscinosis and FTD. Neurol Genet 2. https://doi.org/10.1212/NXG.0000000000000102
Geier EG, Bourdenx M, Storm NJ, et al (2019) Rare variants in the neuronal ceroid lipofuscinosis gene MFSD8 are candidate risk factors for frontotemporal dementia. Acta Neuropathol 137. https://doi.org/10.1007/s00401-018-1925-9
Tang C-H, Lee J-W, Galvez MG, et al (2006) Murine cathepsin F deficiency causes neuronal lipofuscinosis and late-onset neurological disease. Mol Cell Biol 26.https://doi.org/10.1128/mcb.26.6.2309-2316.2006
Wang B, Shi GP, Yao PM et al (1999) Human cathepsin F Molecular cloning functional expression, tissue localization, and enzymatic characterization. J Biol Chem 273:3200. https://doi.org/10.1074/jbc.273.48.32000
Blauwendraat C, Wilke C, Simón-Sánchez J, et al (2018) The wide genetic landscape of clinical frontotemporal dementia: systematic combined sequencing of 121 consecutive subjects. Genet Med 20. https://doi.org/10.1038/gim.2017.102
Siintola E, Topcu M, Aula N, et al (2007) The novel neuronal ceroid lipofuscinosis gene MFSD8 encodes a putative lysosomal transporter. Am J Hum Genet 81. https://doi.org/10.1086/518902
Aiello C, Terracciano A, Simonati A, et al (2009) Mutations in MFSD8/CLN7 are a frequent cause of variant-late infantile neuronal ceroidlipofuscinosis. Hum Mutat 30. https://doi.org/10.1002/humu.20975
Smith KR, Dahl HHM, Canafoglia L, et al (2013) Cathepsin F mutations cause type B Kufs disease, an adult-onset neuronal ceroidlipofuscinosis. Hum Mol Genet 22. https://doi.org/10.1093/hmg/dds558
Guerreiro R, Brás J, Hardy J (2015) SnapShot: genetics of ALS and FTD. Cell 160:798
Taskesen E, Mishra A, van der Sluis S, et al (2017) Susceptible genes and disease mechanisms identified in frontotemporal dementia and frontotemporal dementia with amyotrophic lateral sclerosis by DNA-methylation and GWAS. Sci Rep 7. https://doi.org/10.1038/s41598-017-09320-z
Zhao J, Cooper LT, Boyd AW, Bartlett PF (2018) Decreased signalling of EphA4 improves functional performance and motor neuron survival in the SOD1G93A ALS mouse model. Sci Rep 8. https://doi.org/10.1038/s41598-018-29845-1
Karch CM, Wen N, Fan CC, et al (2018) Selective genetic overlap between amyotrophic lateral sclerosis and diseases of the frontotemporal dementia spectrum. JAMA Neurol 75. https://doi.org/10.1001/jamaneurol.2018.0372
Benyamin B, He J, Zhao Q, et al (2017) Cross-ethnic meta-analysis identifies association of the GPX3-TNIP1 locus with amyotrophic lateral sclerosis. Nat Commun 8. https://doi.org/10.1038/s41467-017-00471-1
Dash BP, Naumann M, Sterneckert J, Hermann A (2020) Genome wide analysis points towards subtype-specific diseases in different genetic forms of amyotrophic lateral sclerosis. Int J MolSci 21. https://doi.org/10.3390/ijms21186938
Giannoccaro MP, Bartoletti-Stella A, Piras S, et al (2017) Multiple variants in families with amyotrophic lateral sclerosis and frontotemporal dementia related to C9orf72 repeat expansion: further observations on their oligogenic nature. J Neurol 264. https://doi.org/10.1007/s00415-017-8540-x
Cady J, Allred P, Bali T, et al (2015) Amyotrophic lateral sclerosis onset is influenced by the burden of rare variants in known amyotrophic lateral sclerosis genes. Ann Neurol 77. https://doi.org/10.1002/ana.24306
Majounie E, Abramzon Y, Renton AE, et al (2012) Repeat expansion in C9ORF72 in Alzheimer’s disease . N Engl J Med 366. https://doi.org/10.1056/nejmc1113592
Goldman JS, Quinzii C, Dunning-Broadbent J, et al (2014) Multiple system atrophy and amyotrophic lateral sclerosis in a family with hexanucleotide repeat expansions in C9orf72. JAMA Neurol 71.https://doi.org/10.1001/jamaneurol.2013.5762
Cooper DN, Krawczak M, Polychronakos C et al (2013) Where genotype is not predictive of phenotype: towards an understanding of the molecular basis of reduced penetrance in human inherited disease. Hum Genet 132:1077–130
Volk AE, Weishaupt JH, Andersen PM et al (2018) Current knowledge and recent insights into the genetic basis of amyotrophic lateral sclerosis. Med Genet 30:252–258
Van Blitterswijk M, Van Es MA, Hennekam EAM, et al (2012) Evidence for an oligogenic basis of amyotrophic lateral sclerosis. Hum Mol Genet 21. https://doi.org/10.1093/hmg/dds199
Murphy NA, Arthur KC, Tienari PJ, et al (2017) Age-related penetrance of the C9orf72 repeat expansion. Sci Rep 7. https://doi.org/10.1038/s41598-017-02364-1
Mis MSC, Brajkovic S, Tafuri F et al (2017) Development of therapeutics for C9ORF72 ALS/FTD-related disorders. Mol Neurobiol 54:4466–4476
Kumar S, Phaneuf D, Cordeau P, et al (2021) Induction of autophagy mitigates TDP-43 pathology and translational repression of neurofilament mRNAs in mouse models of ALS/FTD. MolNeurodegener 16. https://doi.org/10.1186/s13024-020-00420-5
Liu Y, Dodart JC, Tran H, et al (2021) Variant-selective stereopure oligonucleotides protect against pathologies associated with C9orf72-repeat expansion in preclinical models. Nat Commun 12. https://doi.org/10.1038/s41467-021-21112-8
Van Giau V, An SS, Kim S, Bagyinszky E (2016) TD-P-002: genetic diagnosis of neurodegenerative disorders based on gene panels and primers by next-generation sequencing. Alzheimer’s Dement 12. https://doi.org/10.1016/j.jalz.2016.06.248
Lacomblez L, Bensimon G, Leigh PN, et al (1996) Dose-ranging study of riluzole in amyotrophic lateral sclerosis. Lancet 347. https://doi.org/10.1016/S0140-6736(96)91680-3
Abe K, Aoki M, Tsuji S, et al (2017) Safety and efficacy of edaravone in well defined patients with amyotrophic lateral sclerosis: a randomised, double-blind, placebo-controlled trial. Lancet Neurol 16.https://doi.org/10.1016/S1474-4422(17)30115-1
Tsai RM, Boxer AL (2014) Treatment of frontotemporal dementia. Curr Treat Options Neurol 16:319
Abati E, Bresolin N, Comi G, Corti S (2020) Silence superoxide dismutase 1 (SOD1): a promising therapeutic target for amyotrophic lateral sclerosis (ALS). Expert Opin Ther Targets 24:295–310
Li Y, Balasubramanian U, Cohen D, et al (2015) A comprehensive library of familial human amyotrophic lateral sclerosis induced pluripotent stem cells. PLoS One 10.https://doi.org/10.1371/journal.pone.0118266
Miller TM, Pestronk A, David W, et al (2013) An antisense oligonucleotide against SOD1 delivered intrathecally for patients with SOD1 familial amyotrophic lateral sclerosis: a phase 1, randomised, first-in-man study. Lancet Neurol 12.https://doi.org/10.1016/S1474-4422(13)70061-9
Butovsky O, Jedrychowski MP, Cialic R, et al (2015) Targeting miR-155 restores abnormal microglia and attenuates disease in SOD1 mice. Ann Neurol 77. https://doi.org/10.1002/ana.24304
Vieira FG, Hatzipetros T, Thompson K, et al (2017) CuATSM efficacy is independently replicated in a SOD1 mouse model of ALS while unmetallated ATSM therapy fails to reveal benefits. IBRO Reports 2. https://doi.org/10.1016/j.ibror.2017.03.001
Balendra R, Isaacs AM (2018) C9orf72-mediated ALS and FTD: multiple pathways to disease. Nat Rev Neurol 14:544–558
Biferi MG, Cohen-Tannoudji M, Cappelletto A, et al (2017) A new AAV10-U7-mediated gene therapy prolongs survival and restores function in an ALS mouse model. Mol Ther 25. https://doi.org/10.1016/j.ymthe.2017.05.017
Scoles DR, Pulst SM (2018) Oligonucleotide therapeutics in neurodegenerative diseases. RNA Biol 15:707–714
Pritchard C, Silk A, Hansen L (2019) Are rises in electro-magnetic field in the human environment, interacting with multiple environmental pollutions, the tripping point for increases in neurological deaths in the Western World? Med Hypotheses 127.https://doi.org/10.1016/j.mehy.2019.03.018
Bandres-Ciga S, Noyce AJ, Hemani G, et al (2019) Shared polygenic risk and causal inferences in amyotrophic lateral sclerosis. Ann Neurol 85. https://doi.org/10.1002/ana.25431
Cochran JN, Geier EG, Bonham LW, et al (2020) Non-coding and loss-of-function coding variants in TET2 are associated with multiple neurodegenerative diseases. Am J Hum Genet 106. https://doi.org/10.1016/j.ajhg.2020.03.010
Funding
LK is supported by Malaviya Postdoctoral Fellowship by Institute of Eminence (IoE), Banaras Hindu University (BHU) from Ministry of Education (MoE), Government of India (GOI). Funding from IoE, BHU to MM and AM is duly acknowledged. MM is supported by SERB-Power Fellowship by Science and Engineering Research Board, GOI.
Author information
Authors and Affiliations
Contributions
L. K. and M. M. generated the theme for this article, and L. K. executed the literature search, tabulations, figures, data collection, and analysis. The final draft was edited and modified by L. K., A. M., and M. M.
Corresponding author
Ethics declarations
Ethics Approval
Not applicable.
Consent to Participate
Not applicable.
Consent for Publication
All authors have given their consent to publish this manuscript in Molecular Neurobiology.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kirola, L., Mukherjee, A. & Mutsuddi, M. Recent Updates on the Genetics of Amyotrophic Lateral Sclerosis and Frontotemporal Dementia. Mol Neurobiol 59, 5673–5694 (2022). https://doi.org/10.1007/s12035-022-02934-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-022-02934-z