Abstract
Autism spectrum disorder (ASD) is a neurodevelopmental disorder characterised by impaired social interaction and behavioural abnormalities. Growing evidence proved that impairment in mitochondrial functions could inhibit energy production and may contribute to the onset of ASD. Genetic variants in the genes of mitochondrial DNA (mtDNA) could interrupt the normal energy metabolism and production in the brain which lead to a wide range of structural and functional changes in the brain resulting in ASD. The present study aims to compare the activities of mitochondrial electron transport chain (ETC) complex I, pyruvate dehydrogenase (PDH) and specific mitochondrial DNA gene (MT-ND1 and MT-ND4) variants associated with ASD subjects in the Tamil Nadu population. Mutational analysis revealed that most mutations in ASD subjects showed synonymous type followed by missense in both the ND1 and ND4 genes. Interestingly, we found that the complex I and PDH dysfunctions may have a role in ASD compared to the controls (p ≤ 0.0001). Hence, the results of the present study suggest that mitochondrial dysfunction, specifically the complex I genes, may play a major role in the onset of ASD, concluding that further research on mitochondrial genes are mandatory to unravel the mechanism behind ASD pathogenesis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Autism spectrum disorder (ASD) is a neurodevelopmental condition characterised by difficulty in social interaction, problems with verbal and non-verbal communications and repetitive or restricted behaviour [1]. In India, almost around 1 in 100 children under 10 years of age has been affected with ASD [2]. Despite considerable efforts in the understanding of its etiological mechanism and biological basis, no reliable biomarker or distinct pathogenesis has yet been established for ASD. Initially, Lombard proposed that ASD is an impairment caused due to dysfunction in the mitochondria; therefore, it has become mandatory in analysing various aspects of the mitochondria in an ASD-affected patients [3], and few studies have also suggested that mitochondrial dysfunction and altered energy metabolism might be a major factor in the etiopathogenesis of ASD [4,5,6]. The mitochondria are solely responsible in the generation of energy via oxidative phosphorylation, a process that needs action of 5 electron transport chain (ETC) complexes (I to V). These ETCs in the mitochondria especially (complex I and III) are the main protagonist in the generation of free radicals. Also, it is important that the energy demand of a cell is directly proportional to the number of mitochondrial DNA (mtDNA) as it might vary depending on the presence of free radicals and oxidative stress [7, 8]. Remarkably, it has been observed that mutations in the mtDNA have been proposed as a biological biomarker to detect condition such as ASD [9]. Many researches have proved that genetic variants in the mtDNA genes could stimulate mitochondrial dysfunction which results into consecutive episodes of abnormal energy production for cell, irregular intracellular calcium buffering and increased production of ROS, which could contribute to ASD [10, 11]. Increasing evidence indicates that mutations in mtDNA variants especially in the complex I-related genes (ND1 and ND4) may affect the energy supply and key processes of the developing brain, triggering a cascade of structural and functional changes, thereby contributing to the aetiology of ASD [12, 13]. Increased copy numbers of mtDNA genes (ND1 and ND4) especially due to over-replication have been observed among ASD cases [14]. The dysfunction in the mitochondria can be easily described by analysing few biochemical markers like lactate, pyruvate, lactate to pyruvate (L:P) ratio, alanine aminotransferase (ALT) and aspartate aminotransferase (AST) [15,16,17]. An elevation or reduction in these biochemical markers is caused due to impaired mitochondrial aerobic respiration and reduction in TCA cycle functioning which could decrease the L:P ratio eventually affecting the ETC complexes and PDH activity in the mitochondria [18, 19]. These studies have established the proof that these biochemical markers such as AST, ALT, lactate, pyruvate and serotonin levels could impact the mitochondrial activity and end up in progression of ASD [15, 20].
To consider the above statements, we investigated the activities of mitochondrial ETC complex I and pyruvate dehydrogenase (PDH) in ASD subjects. To the best of our knowledge, this is the first kind of study in Indian population, where we had explored the clinical spectrum of children with ASD and mitochondrial dysfunction. Hence, the aim of the present study is to find out the mitochondrial activities of ETC complexes and other biochemical parameters like AST, ALT, lactate and pyruvate which helps in mitochondrial dysfunction among the ASD subjects. Further, we analysed the mtDNA complex I gene (ND1and ND4) variants as whether they have any role in the onset of ASD due to mitochondrial dysfunction.
Methodology
Subject Recruitment
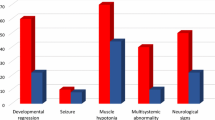
The subjects aged 3 to 18 years were recruited from various hospitals and rehabilitation centres in the study area (Tamil Nadu, India) during 2017 to 2020. Control samples who were age and sex matched with ASD cases (± 2) were also recruited. Environmental, lifestyle, reproductive, maternal lifestyle and detailed demographic characters were collected from the study subjects using a detailed filled questionnaire. The Institutional Human Ethics Committee (IHEC) informed consent was obtained from the participants. The study protocol follows the ethical guidelines of the Declaration of Helsinki (2002). The ASD patients were confirmed via clinical examinations using the Diagnostic and Statistical Manual of Mental Disorders (Fifth Edition) (DSM-V) and Childhood Autism Rating Scale (CARS) to detect the severity of symptoms (Fig. 1).
Sample Size and Details
Totally, 174 samples were recruited for the present study and were grouped according to the severity of the disorder: group 1, mild ASD (n = 57); group 2, moderate ASD (n = 98); and group 3. severe ASD (n = 19). The inclusion criteria for the present study were children aged from 3 to 18 years and who had been diagnosed to have ASD according to DSM-V from a certified paediatrician and neurologist. The exclusion criteria were children with a comorbid diagnosis of other disorders thought to be associated with ASD (fragile X syndrome, Asperger’s syndrome, cerebral palsy, tuberous sclerosis and neurofibromatosis) and children who had other physical illnesses.
Blood Sample Collection
Blood samples were obtained from ASD and control subjects by venipuncture in EDTA and heparin tubes.
Analysis of Biochemical Parameters
A 300 μl blood was taken and centrifuged at 1500 × g, 4 °C for 15 min. The detection reagents were prepared according to the manufacturer’s instructions. For AST and ALT, 10 μl of the samples were transferred to the cuvette, and the samples were diluted with 40 μl of saline. A 300 μl of detection reagent was added, and the values read at 340 nm. For lactate, 4 μl of plasma was added and diluted with 200 μl of saline. A 400 μl of detection reagent was added, and the values were measured at 340 nm.
Assay for Activities of Mitochondrial ETC Complex I
The activity of mitochondrial ETC complex I was analysed using microplate assay kits. All the tests were done in duplicate. The absorbance was read by spectrophotometer microplate reader. For the activity assays of complex I, detergent (1/10 volume) was added to the sample to extract transmembrane proteins into the solution. To select the optimal amount of the sample for the activity assays, a dose–effect curve was plotted for each assay.
Assay for Mitochondrial PDH Enzyme Activity
The PDH enzyme activity was analysed using microplate assay kit. Intact and functional PDH enzymes were extracted after adding detergent (1/19 volume) to the sample, and 100 mg proteins from each sample were used for this assay. PDH activity was determined by following the reduction of NADþ to NADH, coupled to the reduction of a reporter dye to yield a yellow-coloured reaction product whose concentration was monitored by measuring the increase in absorbance at 450 nm. The mitochondrial citrate synthase is an important marker to check the mitochondrial activity. Hence, its level was determined by using extraction buffer, solubilised and determined using immunocapture-based method.
Mitochondrial DNA Extraction from Blood
A 2 ml of the whole blood was treated with low salt buffer (10 mM Tris–HCl pH 7.6, 10 Mm KCl, 10 mM MgCl2 and 2 mM EDTA) to which 125 µl Triton X 100 was added. The contents were centrifuged at 2200 rpm for 10 min, and the supernatant was collected. Further centrifugation was carried out at 15,000 rpm for 30 min. Repeated washing with low salt buffer was carried out till white pellets were obtained which was collected and treated with high salt buffer (10 mM Tris–HCl pH 7.6, 10 Mm KCl, 10 mM MgCl2, 2 mM EDTA and 0.4 M NaCl) to which 75 µl of 10% SDS was added and incubated at 55 °C. A 200 µl 6 M NaCl was added and centrifuged at 11,300 rpm for 20 min. The supernatant was collected and treated with twice the volume of 100% ethanol for precipitation of mtDNA. Contents were centrifuged at 15,000 rpm for 10 min, and pellets were collected and washed with 70% ethanol. The mtDNA obtained was stored in 50 µl TE buffer.
mtDNA Amplification and Sequencing
In order to screen the mutations of MT-ND1and MT-ND4, polymerise chain reaction (PCR) amplification was carried out using specific primers. Further, the PCR products were electrophoresed through 1.5% agarose gel, and the nucleotide sequences of the specific amplicons were determined. The obtained mtDNA sequences were aligned with a multiple sequence alignment interface of Finch TV software and DNA baser and aligned by BLAST. Identified variations were confirmed by repeated analysis of both strands.
Statistical Analysis
Summary statistics are expressed as mean (95% confidence intervals [CIs]), whereas individual data are expressed as mean (SD). The data were evaluated using the correlation in each independent feature, a 95% confidence interval (CI). Mean and SD levels were calculated along with chi square test. All values were considered to be significant at p < 0.05 level throughout the study.
Results
In demographic characteristics, age, sex, passive smoking, stress and family history play a major role in the onset of ASD (significant at p < 0.05 level) compared to other factors; these results have been explained elaborately in Table 1. Table 2 depicts the mutational changes in the ASD patients (mild, moderate and severe) compared to the control subjects. The mitochondrial genes ND1 and ND4 show mutational changes, where most of them were synonymous mutation followed by missense mutation. The values were found to be significant (p < 0.05) for the following 10,895 (ND4), 10,819 (ND4), 11,176 (ND4), 11,150 (ND4), 3622 (ND1) and 4216 (ND1) nucleotide positions in ASD subjects.
The nucleotide coverage of the ND1 gene, its alleles, class and mutation type, amino acid residues and its coordinates has been depicted in Table 3, while in Table 4, we have depicted the total single-nucleotide polymorphisms observed in ASD subjects for ND4 gene. The nucleotide coverage of the ND4 gene, its alleles, class and mutation type, amino acid residues and its coordinates where most of them were of synonymous type followed by missense type.
When lactate levels were measured in autistic children and controls, a higher number of children exhibit hyperlactaemia especially among mild ASD cases than compared to controls. The L:P ratio (lactate:pyruvate ratio) was also found to be reduced among the ASD cases than controls, especially in the moderate ASD cases. Children with ASD also exhibited significantly higher levels of ALT (p < 0.0005) and AST (p < 0.0002) as compared to the controls. Furthermore, hyperserotonaemia was observed in the autistic group where all the three groups of ASD children showed higher serotonin levels as compared to the controls. Interestingly, the chi square test also revealed that all the biochemical parameters showed statistically significant levels when compared with controls except for the lactate levels (p = 0.5). All the control subjects have normal range of values for the respective biochemical parameters. The detailed analysis of the biochemical parameters has been explained in Table 5.
The activities of mitochondrial ETC complex I and PDH normalised to the activity of citrate synthase of ASD and control subjects are represented in Table 6. Our results showed that the activity of complex I was significantly reduced by p ≤ 0.0001) in ASD cases (X2 = 29.8) compared to control group. Further, PDH activity was also significantly reduced by p ≤ 0.0001 in ASD cases (X2 = 34.1) compared to controls.
Discussion
Autism spectrum disorder is a complex neurodevelopmental disorder with multifactorial disorder which has varied aetiologies like genetic, environment, as well as high level of oxidative stress [21, 22]. Based on the previous review article, it was reported that in India among the group of monogenic diseases, ASD has been commonly diagnosed in the Indian populations [23, 24]. The focus into unravelling the relations between mitochondrial dysfunction and ASD was initiated because there has been evidence that mutations in mtDNA have a crucial impact on neurodevelopment of the foetus. Many studies have supported the hypothesis that mtDNA has a pivotal role in the aetiopathogenesis of ASD. Therefore, we aimed to undo the potential as well as possible role of ETC complex I and its related genes in causing the onset of ASD in the present study population. In the present study, we found that defective mitochondria were observed among the ASD cases, where we successfully found low activity of PDH which was allied with low L:P ratio and ETC complex I activity when compared with the controls. Interestingly, even the scatter plot also revealed that the complex I activity and PDH activity have a positive correlation, as the data points are making a straight line among the two variables.
The complex I of the mitochondria is one of the largest complex among the rest four complexes of ETC, also is has the major role in oxidative phosphorylation in the cell, even though its complete mechanism is yet to be discovered, mainly because of its sub-units and their interactions [25]. Our present study was in accordance with Giulivi et al. [14] where the authors found that the activity of complex I was reduced up to 60% among autism cases than controls. In accordance to our study, another author has also stated that either dysfunction in mitochondrial ETC enzyme or mutations in them might have a role in the onset of autism in their study [15, 26]. In the present study, we found that the ASD cases had reduced levels of L:P ratio which suggest that there might be a deficiency in PDH activity among the ASD cases. It has been reported that reduced levels of L:P ratio would remove insufficient amount of pyruvate and lactate resulting into decreased energy production in to brain dysfunction in autism [27]. Studies have reported that mitochondrial dysfunction allied with elevated levels of lactate and alanine in autism cases [28, 29]. Our study is in accordance with previous study which suggested that damage or dysfunction in PDH activity could be a reason for the abnormal levels of other biochemical parameters or metabolites in ASD cases [30].
Numerous biochemical anomalies have been found in individuals with autism. Tests pertaining to lactate revealed elevations in the study group varied from the controls. This was also observed by other researchers as well [19, 30]. Nonetheless, a decrease in these levels was found in a study conducted [30]. As no specific control was maintained on the diet of autistic children and control, these discrepancies can be attributed to this as well. When AST and ALT levels were measured, it was significantly elevated in the children with autism in accordance with previous studies [15, 31, 32]. Similarly, the thyroid profile was also significantly altered in the autism group as compared to their controls. Serotonin has been considered a biomarker for ASD, and hyperserotonaemia was observed in about 25% of the individuals affected with ASD [33]. Similar to prior reports, the present study showed elevated serotonin levels in the autistic group. Despite tremendous research in this realm, the association between hyperserotonaemia and ASD still remains in its infancy. A deeper understanding of the underlying pathology may help in providing novel therapeutic strategies for ASD-like symptoms.
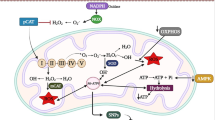
An increase in the production of these free radicals and OXPHOS due to dysfunction in the mitochondria may result in increased mtDNA replication or repair. In our study, we found that most of the mutations were synonymous followed by missense mutation. We also found that the MT-ND1and MT-ND4 gene variants were significant for our study (ND4), 10,819 (ND4), 11,176 (ND4), 11,150 (ND4), 3622 (ND1) and 4216 (ND1) nucleotide positions. Our study was in accordance with a previous study where mtDNA deletion in ND4 gene was almost 44% among the ASD cases than controls [30]. There are reports which also suggest that variations in the nucleotide sequences of mtDNA genes involved in metabolic processes could possibly be a part in provoking ASD in the offspring [10, 34]. Chen et al. [35] observed the expression of mtDNA genes like MT-ND1 and MT-ND4 copy number variations in ASD cases compared to healthy controls. This high prevalence of mtDNA and its metabolites dysfunction among the ASD cases observed in our study could be either primary or secondary, as this could be assessed with further rigorous research work. The present study has few limitations. The major reason could be the sample size to assess the mitochondrial dysfunction as we have divided our ASD cases into three groups. Nevertheless, caution should be exercised with regard to the generalisation of findings in a larger population.
Conclusion
In conclusion, the present study was the first to report the activities of mitochondrial ETC complex I, PDH as well as mtDNA gene (MT-ND1 and MT-ND4) variants associated with ASD subjects in Tamil Nadu population. Previous studies and the present study results strongly suggest that dysfunction in mitochondrial processes could be a potential pathogenetic criteria for the onset of ASD. Therefore, abnormalities observed in the mtDNA or ETC functions along with oxidative stress may be responsible for the onset of ASD. Even though more research is recommended in the same stream to understand the clear molecular background behind the mitochondrial dysfunction, studies are still in its infancy, and additional research are needed to understand how it triggers the cascade of structural and functional changes, leading to the autistic phenotype if mitochondrial functions are impaired and the energy need of the brain is not fulfilled.
Availability of Data and Material
Not applicable.
Code Availability
Not applicable.
Abbreviations
- ALT:
-
Aminotransferase
- ASD:
-
Autism spectrum disorder
- AST:
-
Aspartate aminotransferase
- CARS:
-
Childhood Autism Rating Scale
- DSM-V:
-
Diagnostic and Statistical Manual of Mental Disorders (Fifth Edition)
- EDTA:
-
Ethylenediaminetetraacetic acid
- ETC:
-
Electron transport chain
- HCl:
-
Hydrogen chloride
- IHEC:
-
Institutional Human Ethics Committee
- KCl:
-
Potassium chloride
- L:P:
-
Lactate to pyruvate
- MgCL2:
-
Magnesium dichloride
- mtDNA:
-
Mitochondrial DNA
- NaCl:
-
Sodium chloride
- NADH:
-
Nicotinamide adenine dinucleotide
- ND1:
-
NADH dehydrogenase 1
- ND4:
-
NADH dehydrogenase 4
- OXPHOS:
-
Oxidative phosphorylation
- PDH:
-
Pyruvate dehydrogenase
- SDS:
-
Sodium dodecyl sulfate
- CI:
-
Confidence interval
- X 2 :
-
Chi square
References
Ganesan H, Balasubramanian V, Iyer M, Venugopal A, Subramaniam MD, Cho SG, Vellingiri B (2019) mTOR signalling pathway - a root cause for idiopathic autism? BMB Rep 52(7):424–433. https://doi.org/10.5483/BMBRep.2019.52.7.137
Chauhan A, Sahu JK, Jaiswal N, Kumar K, Agarwal A, Kaur J, Singh S et al (2019) Prevalence of autism spectrum disorder in Indian children: a systematic review and meta-analysis. Neurol India 67(1):100–104. https://doi.org/10.4103/0028-3886.253970
Lombard J (1998) Autism: a mitochondrial disorder? Med Hypotheses 50:497–500. https://doi.org/10.1016/s0306-9877(98)90270-5
Filipek PA, Juranek J, Smith M et al (2003) Mitochondrial dysfunction in autistic patients with 15q inverted duplication. Ann Neurol 53(6):801–804. https://doi.org/10.1002/ana.10596
Fillano JJ, Goldenthal MJ, Rhodes CH, Marín-García J (2002) Mitochondrial dysfunction in patients with hypotonia, epilepsy, autism, and developmental delay: HEADD syndrome. J Child Neurol 17(6):435–439. https://doi.org/10.1177/088307380201700607
Oliveira G, Diogo L, Grazina M et al (2005) Mitochondrial dysfunction in autism spectrum disorders: a population- based study. Dev Med Child Neurol 47(3):185–189. https://doi.org/10.1017/s0012162205000332
Liu CS, Tsai CS, Kuo CL et al (2003) Oxidative stress-related alteration of the copy number of mitochondrial DNA in human leukocytes. Free Radic Res 37:1307–1317. https://doi.org/10.1080/10715760310001621342
Lee HC, Wei YH (2005) Mitochondrial biogenesis and mitochondrial DNA maintenance of mammalian cells under oxidative stress. Int J Biochem Cell Biol 37:822–834. https://doi.org/10.1016/j.biocel.2004.09.010
Golzio C, Katsanis N (2016) Mitochondrial copy number as a biomarker for autism? Pediatrics 137(4):e20160049. https://doi.org/10.1542/peds.2016-0049
Patowary A, Nesbitt R, Archer M, Bernier R, Brkanac Z (2017) Next generation sequencing mitochondrial DNA analysis in autism spectrum disorder. Autism Res 10:1338–1343. https://doi.org/10.1002/aur.179210:1338-1343
Cruz A, Ferrasa A, Muotri AR, Herai RH (2019) Frequency and association of mitochondrial genetic variants with neurological disorders. Mitochondrion 46:345–360. https://doi.org/10.1016/j.mito.2018.09.005
Chauhan A, Gu F, Chauhan V (2012) Mitochondrial respiratory chain defects in autism and other neurodevelopmental disorders. J Pediatr Biochem 2:213–223. https://doi.org/10.3233/JPB-120063
Palmieri L, Persico AM (2010) Mitochondrial dysfunction in autism spectrum disorders: cause or effect? Biochim Biophys Acta 1797:1130–1137. https://doi.org/10.1016/j.bbabio.2010.04.018
Giulivi C, Zhang YF, Omanska-Klusek A et al (2010) Mitochondrial dysfunction in autism. JAMA 304(21):2389–2396. https://doi.org/10.1001/jama.2010.1706
Weissman JR, Kelley RI, Bauman ML et al (2008) Mitochondrial disease in autism spectrum disorder patients: a cohort analysis. PLoS One 3:e3815. https://doi.org/10.1371/journal.pone.0003815
Haas RH, Parikh S, Falk MJ, Saneto RP, Wolf NI, Darin N, Cohen BH (2007) Mitochondrial disease: a practical approach for primary care physicians. Pediatrics 120(6):1326–1333. https://doi.org/10.1542/peds.2007-0391
Frye RE (2009) 15q11.2-13 duplication, mitochondrial dysfunction, and developmental disorders. J Child Neurol 24(10):1316–1320. https://doi.org/10.1177/0883073809333531
Zeviani M, Bertagnolio B, Uziel G (1996) Neurological presentations of mitochondrial diseases. J Inherit Metab Dis 19(4):504–520. https://doi.org/10.1007/BF01799111
Rossignol D, Frye R (2012) Mitochondrial dysfunction in autism spectrum disorders: a systematic review and meta-analysis. Mol Psychiatry 17:290–314. https://doi.org/10.1038/mp.2010.136
László A, Horváth E, Eck E, Fekete M (1994) Serum serotonin, lactate and pyruvate levels in infantile autistic children. Clin Chim Acta 229:205–207. https://doi.org/10.1016/0009-8981(94)90243-7
Geetha B, Sukumar C, Dhivyadeepa E, Reddy JK, Balachandar V (2019) Autism in India: a case-control study to understand the association between socio-economic and environmental risk factors. Acta Neurol Belg 119(3):393–401. https://doi.org/10.1007/s13760-018-01057-4
Bharathi G, Jayaramayya K, Balasubramanian V, Vellingiri B (2019) The potential role of rhythmic entrainment and music therapy intervention for individuals with autism spectrum disorders. J Exerc Rehabil 15(2):180–186. https://doi.org/10.12965/jer.1836578.289
Bharathi G, Venugopal A, Vellingiri B (2019) Music therapy as a therapeutic tool in improving the social skills of autistic children. Egypt J Neurol Psychiatry Neurosurg 55:44. https://doi.org/10.1186/s41983-019-0091-x
Venugopal A, Chandran M, Eruppakotte N, Kizhakkillach S, Breezevilla SC, Vellingiri B (2018) Monogenic diseases in India. Mutat Res 776:23–31. https://doi.org/10.1016/j.mrrev.2018.03.003
Pollard AK, Craig EL, Chakrabarti L (2016) Mitochondrial complex 1 activity measured by spectrophotometry is reduced across all brain regions in ageing and more specifically in neurodegeneration. PLoS One 11(6):e0157405. https://doi.org/10.1371/journal.pone.0157405
Balachandar V, Kamarajan R, Kaavya J, Madesh J, Mahalaxmi I (2021) Mitochondrial dysfunction: a hidden trigger of autism? Genes Dis 8:629–639. https://doi.org/10.1016/j.gendis.2020.07.002
De Meirleir L, Lissens W, Denis R et al (1993) Pyruvate dehydrogenase deficiency. Pediatr Neurol 9(3):216–220. https://doi.org/10.1016/0887-8994(93)90088-t
Haas RH (2010) Autism and mitochondrial disease. Dev Disabil Res Rev 16:144–153. https://doi.org/10.1002/ddrr.112
Gargus JJ, Imtiaz F (2008) Mitochondrial energy-deficient endophenotype in autism. Am J Biochem Biotech 4:198–207. https://doi.org/10.3844/ajbbsp.2008.198.207
Gu F, Chauhan V, Kaur K et al (2013) Alterations in mitochondrial DNA copy number and the activities of electron transport chain complexes and pyruvate dehydrogenase in the frontal cortex from subjects with autism. Transl Psychiatry 3(9):e299. https://doi.org/10.1038/tp.2013.68
Karim M, Begum S, Shahzadi S (2015) Serum lactate, AST, ALT in male autistic children in Bangladesh. J Bangladesh Soc Physiol 10:56–60. https://doi.org/10.3329/jbsp.v10i2.27165
Shahjadi S, Khan AS, Ahmed MU (2017) Mitochondrial dysfunction in early diagnosed autism spectrum disorder children. J Dhaka Med Coll 26:43–47. https://doi.org/10.3329/jdmc.v26i1.34000
Muller CL, Anacker AM, Veenstra-VanderWeele J (2016) The serotonin system in autism spectrum disorder: from biomarker to animal models. Neuroscience 321:24–41. https://doi.org/10.1016/j.neuroscience.2015.11.010
Varga NÁ, Pentelényi K, Balicza P, Gézsi A, Reményi V, Hársfalvi V, Molnár MJ (2018) Mitochondrial dysfunction and autism: comprehensive genetic analyses of children with autism and mtDNA deletion. Behav Brain Funct 14:4. https://doi.org/10.1186/s12993-018-0135-x
Chen S, Li Z, He Y et al (2015) Elevated mitochondrial DNA copy number in peripheral blood cells is associated with childhood autism. BMC Psychiatry 15:50. https://doi.org/10.1186/s12888-015-0432-y
Acknowledgements
The author Dr. VB would like to thank the Bharathiar University for providing the necessary infrastructure facility and the Science and Engineering Research Board (SERB) (ECR/2016/001688), Government of India, New Delhi, for providing necessary help in carrying out the manuscript. We acknowledge the Department of Human Genetics and Molecular Biology, Bharathiar University, for providing necessary infrastructure facilities, ethical approval, and technical assistance to conduct this article.
Funding
This study was funded by Science and Engineering Research Board (SERB) Early Career Research (ECR) Award funded by the Government of India, New Delhi (Grant No. ECR/2016/ 001688).
Author information
Authors and Affiliations
Contributions
Concept and design, VB; literature search, IM and MDS; experimental studies, VB, IM and MDS; data analysis and statistical analysis, VB, IM and MDS; manuscript preparation, IM and MDS; manuscript editing and manuscript review, VB and AVG.
Corresponding author
Ethics declarations
Ethics Approval
Institutional Human Ethics had been obtained from the Bharathiar University.
Consent to Participate
Informed consent was obtained from all study participants.
Consent for Publication
All authors have approved for publication.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Mahalaxmi, I., Subramaniam, M.D., Gopalakrishnan, A.V. et al. Dysfunction in Mitochondrial Electron Transport Chain Complex I, Pyruvate Dehydrogenase Activity, and Mutations in ND1 and ND4 Gene in Autism Spectrum Disorder Subjects from Tamil Nadu Population, India. Mol Neurobiol 58, 5303–5311 (2021). https://doi.org/10.1007/s12035-021-02492-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-021-02492-w