Abstract
Macroautophagy impairment plays a key role in sporadic Alzheimer’s disease (sAD) neurodegenerative process. Nevertheless, the mechanism(s) that lead to a deficiency in macroautophagy in AD remains elusive. In this work, we identify, for the first time that Beclin-1 acetylation status is implicated in the alterations in autophagy observed in AD neurodegeneration. We observed that Beclin-1 is deacetylated by sirtuin 1 (SIRT1) and acetylated by p300. In addition, Beclin-1 acetylation inhibits autophagosome maturation, leading to impairment in autophagic flux. We also analyzed some proteins known to be involved in the maturation of autophagosomes such as Rab7, which participates in the fusion step with lysosomes. We observed that increased expression of Rab7 might represent a response to boost the formation of large perinuclear lysosome clusters in accordance with an increase in lysosomal biogenesis determined by increase in LAMP-2A, LAMP-1, and cathepsin D expression in AD cells. Thus, our data provides strong evidences that Beclin-1 acetylation impairs the autophagic flux, and despite lysosomal biogenesis seems to be triggered as a compensatory response, autophagosome fusion with lysosomes is compromised contributing to AD neurodegeneration.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
There is an increasing body of evidence that supports an autophagic dysfunction in the neurodegenerative process of Alzheimer’s disease (AD). Accumulation of autophagic vesicles (AVs) in the neocortex from AD brains was the first evidence of altered autophagic activity in AD, where AV buildup, mostly in the form of autophagosomes, was observed in AD dystrophic neurites [1]. Even though AV detection is uncommon in healthy neurons, an accumulation of such vesicles was apparent in AD neuronal cells, pinpointing an impaired transport and maturation of AVs [2]. Additionally, many studies already reported that AVs accumulate in the brains of individuals with neurodegenerative diseases, such as Parkinson’s disease [3], strongly suggesting a defective transition from autophagosomes to autolysosomes [4, 5]. Also, depletion of key autophagy genes such as atg5 and atg7, which are essential for autophagosome maturation, leads to neurodegeneration in mice brains [6]. On the other hand, it was described that protein aggregate accumulation was a consequence of a flawed autophagic process, implying that one important role for autophagy is the turnover of cytosolic proteins and not the direct elimination of inclusion bodies [7].
Interestingly, it was established that dysfunctional autophagy influences the secretion of Aβ and may directly affect intracellular Aβ accumulation and extracellular Aβ plaque generation [8]. Autophagosomes that fuse with APP-containing endosomes and accumulate in the brain due to an inefficient traffic, lysosomal fusion, or decreased function of lysosomal enzymes are sites of Aβ formation. It was hypothesized that induction of autophagy before the development of AD-like pathology can reduce the levels of soluble Aβ, whereas induction after the formation of mature plaques and tangles has no beneficial effect on AD-like pathology [9]. Beclin-1 is a key protein in autophagosome formation, whose mRNA and protein levels were shown to be reduced in AD-affected brain areas [10]. Additionally, it was demonstrated a disruption of the autophagic pathway, increased neurodegeneration, and Aβ accumulation in Beclin-1 deficient mice [10]. The activation of autophagy by overexpressing Beclin-1 prevented neuronal cell death and improved the clearance of toxic protein aggregates [11]. Decreased levels of Beclin-1 seen in AD brain can be a consequence of impaired autophagosome–lysosomal fusion, which may activate a negative feedback loop, decreasing autophagy induction, thus preventing an uncontrolled activation of autophagy with potentially serious consequences for the cell [12].
Experimental data have also reported that Beclin-1 acetylation might regulate the maturation of the autophagosome. As such, it was clearly demonstrated that Beclin-1 acetylation by p300 inhibits the maturation of the autophagosome, while SIRT1-mediated deacetylation promotes the autophagic pathway [13].
Multiple evidences have proved that an upregulation or activation of SIRT1, a NAD+-dependent deacetylase, has a beneficial effect in AD models. Copious amount of data in AD transgenic mice models showed that an overexpression of SIRT1 not only reduced Aβ plaques accumulation and ameliorated learning and memory disabilities [14], but also increased lifespan of mice bearing brain-specific overexpression of SIRT1 [15]. Additionally, it has been shown that SIRT1 can deacetylate components of the autophagic machinery [16, 17]. One of such candidates is Beclin-1 and the variation in its acetylated levels may have an impact in the autophagic pathway. Therefore, we will tackle how the (de)acetylation of Beclin-1 influences the autophagic flux, as well as its impact of lysosomal function and localization in AD context.
Results
Effect of NAM and C646 on Cybrid Cells Viability
We determined cell viability using the MTT assay and established non-toxic concentrations of nicotinamide (NAM), an inhibitor of sirtuin 1 (SIRT1) [18], and C646, a relatively potent, selective, cell-permeable small inhibitor of p300 lysine acetyltransferase (p300) [19]. SIRT1, which deacetylate core histones and non-histone proteins, plays a relevant role in autophagy induction under starved conditions, regulating the deacetylation of multiple essential proteins involved in the autophagic pathway, such as LC3/Atg8 [20]. Acetyltransferase p300 is a critical regulator of eukaryotic gene expression, involved in several biological processes [21]. p300 is predominantly nuclear but can shuttle between the nucleus and cytoplasm and recently has been suggested to participate in the regulation of the autophagic pathway. Various ATG proteins, such as LC3/Atg8 and Atg5 are targeted by p300-mediated acetylation, leading to an inhibition of their autophagic activities [22].
Concentrations of 5 mM NAM (Fig. 1a) and 2.5 μM C646 (Fig. 1b) did not interfere with cybrid cells viability in CT, MCI, and sAD groups.
Effect of NAM and C646 on cell viability. a Cybrids cell lines were incubated with NAM at concentrations 0.5 mM, 1 mM, 5 mM, and 10 mM. b Cybrids cell lines were incubated with concentrations 1 μM, 2.5 μM, 5 μM, 10 μM, and 20 μM of C646. Data were analyzed by Student’s t test and are presented as mean ± SEM derived from three independent experiments
SIRT1 and p300 Role in the (de)Acetylation of Beclin-1
We evaluated the levels of phospho-SIRT1 (p-SIRT1), active form of SIRT1, in our cells. Our results point to a significant decrease of basal p-SIRT1 in sAD cybrids compared to MCI and CT cybrids (Fig. 2a), which is in accordance with previous reports [23, 24]. In fact, we observed that under basal conditions, p-SIRT1 levels are very similar between MCI and CT, suggesting that p-SIRT1 levels are of no value as an early AD marker. We did not see any significantly effect of NAM or C646 on p-SIRT1 levels in sAD cybrids. Because SIRT1 levels are decreased in the brains of AD patients [25] and also in our AD cells, we assessed the ratio between nuclear/cytosolic SIRT1, since its subcellular localization may account for the dual roles of SIRT1 in normal versus AD cells similarly to what observed for cancer cells [26]. We observed that this ratio was decreased in sAD cells, similarly to what was observed by Silva and coworkers [24] (Fig. 2b). Despite the decrease in the active SIRT1 form in AD cells, a cytosolic localization may promote Beclin-1 deacetylation at lysine 430 and lysine 437, potentiating autophagic degradation [13].
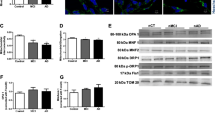
The role of SIRT1 and p300 in Beclin-1 acetylation. a Phospho-SIRT1 protein levels. b Nuclear and cytosolic SIRT1 levels. c Beclin-1 protein levels. d Acetylated-Beclin-1 levels and total lysine-acetylated levels. e NAM and C646 effect on acetylated-Beclin-1 levels and total lysine-acetylated levels. Data were analyzed by Student’s t test and are presented as mean ± SEM derived from three to five independent experiments. *p < 0.05; ***p < 0.001 significantly different from CT cells
Beclin-1 is an essential molecular anchor in autophagic pathway, regulating the initiation and nucleation of autophagosomes formation [27]. Studies have described that Beclin-1 expression is significantly decreased in samples from AD brains, when compared with age-matched controls [10, 28]. Moreover, Jaeger and colleagues described that a deficiency of Beclin-1 expression affects the degradation of autophagosomes in cultured cells [12]. In our study, Beclin-1 protein levels were reduced in basal sAD group, comparing with MCI and CT groups (Fig. 2c). Additionally, we observed a non-significative increase of Beclin-1 in sAD cybrids when SIRT1 was inhibited. We hypothesized that this result could be due to altered Beclin-1 acetylation status leading to an impairment of autophagy pathway in AD pathology [29]. To tackle this, we immunoprecipitated acetylated lysine residues and determined Beclin-1 acetylation levels. We found that basal levels of acetylated Beclin-1 were significantly increased in sAD cells, and showed a trend to increase in MCI when compared to CT cells. Intriguingly, total protein lysine acetylation levels decreased in sAD cell (Fig. 2d) suggesting the involvement of other deacetylases. To clarify the role of SIRT1 and p300 on Beclin-1 acetylation, we tested NAM and C646. We observed that in control cells, SIRT1 inhibition increased Beclin-1 acetylated levels, while p300 inhibition decreased these levels (Fig. 2e). The same profile was observed in AD cells, despite no effect was detected in MCI cells. Interestingly, NAM increased and C646 decreased total lysine acetylation levels in control cells (Fig. 2e).
Autophagic Flux—a Link to Pathophysiological Processes in AD
Since Beclin-1 is essential for the autophagic pathway and its acetylation levels may impact autophagy, we tackled the autophagic flux in our cells calculating the variation between the levels of LC3II—the cytosolic lipid conjugated form of LC3 in basal conditions and upon lysosomal inhibition with NH4Cl/Leupeptin. LC3 commands crucial steps in autophagy including autophagic membrane growth, cargo recognition, and the fusion between autophagosomes and lysosomes, therefore being widely used to monitor autophagic activity [16]. We observed that autophagic flux is significantly decreased in sAD cells and increased in MCI cells (Fig. 3a). This decrease in AD cells may represent a defective clearance of AVs, and the increase in MCI cells indicates an upregulation in the attempt to eliminate damaged organelles, such as mitochondria or protein aggregates [30]. Interestingly, SIRT1 inhibition decreases autophagic flow in control cells, probably due to a decrease in the deacetylated levels of Beclin-1 and LC3, both of which contribute to a failure in autophagy.
Autophagic flux. a Immunoblotting for LC3II protein levels for each separate cybrid group, untreated and treated with NH4CL/Leupeptin-NAM and NH4CL/Leupeptin-C646. b Immunoblotting for p62 protein levels for each separated cybrid group, untreated and treated with NH4CL/Leupeptin-NAM and NH4CL/Leupeptin-C646. Data were analyzed by Student’s t test and are presented as mean ± SEM derived from three to five independent experiments. *p < 0.05; ***p < 0.001 significantly different from CT cells. #p < 0.05, significantly different from AD untreated cells
Additionally, we also used p62, a multifunctional protein involved in the specific recognition of polyubiquitinated aggregates to be degraded by autophagy, as a marker for autophagic flux [31, 32]. We observed that the p62 flux was decreased in both MCI and AD cells (Fig. 3b). These results are in accordance with previous data showing that p62 strongly binds to neurofibrillary tangles (NFTs), most likely to target them for degradation and may indicate that p62 accumulation is due to inefficient autophagy [31, 33].
Recent data showed that Beclin-1 and LC3 could be acetylated by p300 and deacetylated by SIRT1 [13, 16, 34]. Our results suggest that SIRT1 inhibition significantly decreases autophagic flow, and p300 inhibition had no effect in CT cells (Fig. 3a). These observations may be in accordance with the hypothesis that SIRT1 deacetylates Beclin-1 allowing autophagosomes maturation [13]. Additionally, LC3 deacetylation by SIRT1 is necessary for its redistribution from the nucleus to cytoplasm and for LC3-Atg7 interaction [16]. In AD cells, we observed that p300 inhibition increased autophagic flow and the p62 flux (Fig. 3), indicating that Beclin-1 and LC3 acetylation inhibition may be in part beneficial in AD. Indeed, knockdown of p300 reduces acetylation of Atg8, and overexpressed p300 increases its acetylation [34]. Our data also suggests that acetylation of Beclin-1 promotes its interaction with Rubicon, that negatively regulates autophagosome and endosome maturation [35], leading to a decrease of autophagic flow observed in AD.
Lysosomal Response to Beclin-1 Acetylation Levels
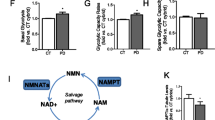
Lysosomal function and cellular localization are an essential cellular process, particularly important in neuronal cells due to their post-mitotic nature, asymmetry, and the long distance between the cell body and dendritic arborization [35]. Multiple evidences suggest that changes or mutations in components of the lysosome-positioning machinery may cause neurological disorders [36]. Indeed, AD progression is accompanied by complex pathological changes, such as massive accumulation of lysosomes, autolysosomes, and autophagosomes [37]. In order to infer lysosomal biogenesis in our experimental conditions, we determined the levels of LAMP-2A, a protein associated to the lysosomal membrane, and cathepsin D (CatD), a soluble lumen lysosomal aspartic endopeptidase [38, 39]. We observed an increase of basal LAMP-2A levels in sAD group and a non-significant increase in MCI cells compared with CT cells (Fig. 4a). This result is supported by the fact that LAMP-2A is significantly increased in the CSF from AD patients [40]. Additionally, these results may indicate that an accumulation of both oligomeric Aβ1–42 and tau aggregates signal the cell to increase its expression. It was described that tau protein can associate with hsc70, the cytosolic chaperone responsible for targeting substrates through the chaperon-mediated autophagy (CMA) to lysosomes [41]. An increase in LAMP-2A levels may indicate an increase in lysosomal biogenesis or an upregulation to boost CMA [42]. We also observed a significantly increase of CatD levels in untreated sAD cells and a slight increase in MCI cells (Fig. 4b). These data reflect previous knowledge that CatD might be involved in the clearance of Aβ and tau protein through the autophagy–lysosomal system [43, 44]. Also, higher levels of CatD and increased gene expression were reported in the hippocampus of MCI individuals [44]. Unexpectedly, we did not observe an increase in LAMP-2A or CatD levels in CT cells after NH4Cl/Leupeptin treatment, which induces lysosomal inhibition (Fig. 4). Upon SIRT1 inhibition with NAM, we did not detect any alterations in LAMP-2A and CatD levels, which may indicate that SIRT1 inhibition do not potentiate the alteration already seen in MCI and AD cells (Fig. 4). However, when p300 was inhibited with C646, we noticed a decrease of LAMP-2A and CatD levels in the AD group (Fig. 4).
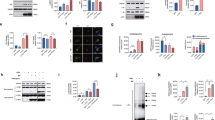
Lysosomes localization and function. a Immunoblotting for LAMP-2A protein levels for each separate cybrid group, untreated and treated with NH4Cl/Leupeptin-NAM and NH4CL/Leupeptin-C646. b Immunoblotting for CatD protein levels for each separated cybrid group, untreated and treated with NH4Cl/Leupeptin-NAM and NH4CL/Leupeptin-C646. Data were analyzed by Student’s t test and are presented as mean ± SEM derived from three to five independent experiments. *p < 0.05; ***p < 0.001 significantly different from CT cells. #p < 0.05, significantly different from AD untreated cells
Lysosomal function also depends on their cellular localization. Recently, it was described that the balance between Rab7 and Arl8b determines their localization. Moreover, peripheral lysosomes are less acidic than juxtanuclear ones [45]. We found an increase in Rab7 levels in the sAD group, accordingly, to the observed upregulation of Rab7 levels within frontal cortex and hippocampus in MCI and AD, which also correlated with Braak staging [46]. Rab7 is a Ras small GTPase that controls aggregation and fusion of late endocytic structures/lysosomes, essential for maintenance of the perinuclear lysosome localization [47]. Upon p300 inhibition, we observed a significant decrease in Rab7 levels in AD cells (Fig. 5a). These results may infer that Beclin-1 acetylation may affect not only autophagy–lysosomal pathway but also the endocytosis pathway, compromising the degradation of toxic proteins, affecting neuronal cells, and ultimately leading to neurodegenerative processes. We determined, in our cells, lysosomal dispersion and localization by detecting LAMP-1, a glycoprotein present in luminal side of lysosomes that serves as a barrier to prevent inadvertent release of soluble cathepsins and other hydrolases. Similarly to what was described for the cerebral cortex of AD patients [48], MCI and AD cells have an increase in LAMP-1 fluorescence (Fig. 5b). Moreover, we observed that LAMP-1 positive lysosomes are dispersed in AD cells, which could explain the increase in Rab7 expression to promote lysosomal clustering and perinuclear localization. Also, an inhibition of p300 increases the formation of juxtanuclear clusters in sAD groups. Indeed, dispersed, more peripheral, and less acidic lysosomes may promote Aβ deposition in AD brains [49]. Although peripheral lysosomes have decreased levels of Rab7, we saw that AD cells try to respond to the presence of less efficient lysosomes by increasing Rab7 expression. We also observed that p300 inhibition decreases Rab7 levels and increases lysosomal clustering (Fig. 5), which indicates that Beclin-1 acetylation and its downstream consequences lead to a global impairment of autophagy–lysosomal and endocytic pathways.
Endo-lysosomal fusion: an essential key on autophagic flux. a Immunoblotting for Rab7 protein levels for each separate cybrid group, treated with NAM and C646. b. Immunofluorescence for LAMP-1 protein levels for each separated cybrid group, treated with NAM and C646. Data were analyzed by Student’s t test and are presented as mean ± SEM derived from three to five independent experiments. *p < 0.05; **p < 0.01 significantly different from CT cells. #p < 0.05, ##p < 0.01 significantly different from AD untreated cells
Discussion
Autophagy is an ubiquitous process essential for the maintenance of cell homeostasis, on which neurons rely for correct function. In AD patients, deleterious effects in autophagy system have been extensively reported, accompanied by an increase in cellular accumulation of Aβ plaques [50]. Along with these observations, dysfunctional autophagy also induces abnormalities in AD neurons such as swelled and dystrophic neurites, due to the impairment of the lysosomal degradative process and consequent buildup of AVs and dysfunctional mitochondria [51]. Moreover, AD cybrid cells, extensively studied by our group that contain sporadic AD patients’ mitochondria, show reduced cytochrome oxidase activity, elevated ROS, and reduced ATP levels. Our data further confirm that AD- and MCI-driven cells have increased Tau phosphorylation levels and accumulation of autophagosomes and Aβ oligomers [30]. Indeed, AD and MCI cybrids with higher ADP/ATP and lower NAD+/NADH ratios [24] could have impaired SIRT1 activation due to a decrease in ATP to activate SIRT1 to pSIRT1 and due to a decrease in SIRT1 cofactor, NAD+.
Nowadays, it is accepted that post-translational modifications (PTM) are pivotal to the autophagic process. For instance, many proteins included in the autophagic machinery such as mTOR, AMPK, and ATG proteins are tightly orchestrated by numerous PTM with relevant impact in the endolysosomal process [52]. Acetylation is emerging as an important metabolic regulatory mechanism in autophagy, since it was demonstrated that p300 acetyltransferase activity was involved in regulating several ATG proteins [34]. Strikingly, Marzi and coworkers elegantly determined that late-onset AD patients’ acetylome is in fact completely altered [53].
SIRT1 regulation has proven successful in AD pathogenesis and, therefore, has received much attention as a potential therapeutic target in neurodegenerative disorders [54]. Interestingly, increased expression of SIRT1 in mammalian neurons prevents Aβ production by promoting the anti-amyloidogenic cleavage of APP by α-secretase, involving ROCK1 inhibition [55]. Additionally, some evidences on the autophagic machinery demonstrate that SIRT1 coimmunoprecipitated with Atg5, Atg7, and Atg8/LC3. All these proteins can be deacetylated by SIRT1 in a NAD+-dependent manner in vitro [56].
Our goal was to determine how SIRT1 and p300 modulates autophagy through Beclin-1 in AD cells. SIRT1 active form (p-SIRT1) was decreased in our AD cells as well as its nuclear localization, which could possibly lead to impairment in autophagic activity, as well as, an accumulation of harmful protein aggregates or damaged organelles. Since SIRT1 is predominantly nuclear in the nervous system [57], our results imply an apparent deregulation of its downstream pathway. Indeed, SIRT1 deacetylates LC3 at K49 and K51 in the nucleus, which is required for Atg8/LC3 interaction with Atg7 and so for the initiation of autophagy [16]. Additionally, SIRT1 may deacetylate Beclin-1 and favor autophagosomes and endosome maturation [13]. Our results showed that total acetylated lysine residues were decreased in sAD cells, despite the increase observed for the acetylated form of Beclin-1. Not only our results corroborate previous findings regarding the expression of both proteins, but we also clearly show that Beclin-1 acetylation is increased. Therefore, the inactivated form seems to be favored in this case, which can be explained by a possible decreased function of SIRT1 and the consequent disruption in the SIRT1/p300 orchestration of Beclin-1 acetylation. As many ATG proteins are deacetylated by SIRT1 in order to exert their function [17], presented data in sAD cells allow us to infer that AD acetylome might be aberrant with severe consequences in autophagy. Indeed, we detected an effective decrease in the autophagic flux of AD cells, together with a decrease in AV flow in control cells exposed to NAM. Since SIRT1 induction improves autophagic flux in neurons [17], it seems plausible that defective clearance of AVs observed in our AD cells can be, at least, partially modulated by SIRT1. Interesting data revealed that in Beclin-1 knockdown cells, rapamycin was not able to induce autophagosome maturation and lysosomal biogenesis determined by the increase of LC3 or LAMP-1 and cathepsin B levels [58]. Nevertheless, in an AD context with decreased levels of Beclin-1, lysosomal/endosomal proteins such as LAMP-2A, LAMP-1, Rab3, and Rab7 are found to be significantly increased in CSF from AD patients compared with age-matched controls [59]. In our cellular model, we also observed higher levels of LAMP-2A, LAMP-1, and CatD in untreated sAD cybrids. Indeed, increased and altered intracellular distribution of lysosomal hydrolases was described in many degenerated neurons, particularly in brain areas known to become affected in AD [60]. Accordingly, CatD has been associated with important factors of AD pathogenesis, such as APP, APOE, and Tau protein [39], and an abnormal immunoreactivity of CatD in senile plaques of AD patients was reported [43, 61]. CatD polymorphisms have impact in intracellular routing and maturation of pro-enzymes [62] and influence the progression of AD [63]. Moreover, an impairment of lysosomal pathway has been demonstrated to occur early in AD pathology before the robust accumulation of NFTs and senile plaques [64]. Our data shows that CatD levels in untreated sAD cells are significantly increased comparing with both CT and MCI cells. Although, when p300 is inhibited, CatD levels decrease in sAD treated cells. These results suggest that acetylated Beclin-1 may compromise the lysosome organelle and affect CatD enzyme correct functioning, consequently, leading to an increase of expression levels. Therefore, an upregulation of CatD and LAMP-2A expression in AD condition may represent a compensatory protective mechanism to counteract the autophagic–lysosomal deficit of the neurons in eliminating toxic protein aggregates.
Rab7 is commonly viewed as a late endosome marker protein and involved in multiple processes including endosome maturation and transport, lysosome biogenesis, and vesicle fusion [65], being relevant for the progression of autophagy in mammalian cells [66]. Remarkably, an expression of a constitutively active form of Rab7 resulted in the formation of large perinuclear lysosome clusters, while dominant-negative Rab7 cells displayed dispersed lysosomes throughout the cytosol [47]. Additionally, Rab7 protein expression was found to be increased in CSF of AD patients [59] and in basal forebrain, frontal cortex, and hippocampus of MCI and AD patients; Rab7 gene expression and protein levels have been reported to be upregulated [67].
LAMP-1 is a glycoprotein associated with lysosomes, whose expression was shown to be increased in AD patients’ brains and other tauopathies [68], despite a negative correlation in immunoreactivity with NFTs [48]. Moreover, CatD immunoreactivity was found to be reduced in the neocortex of adult LAMP-1-deficient mice [69], which demonstrates the intrinsic function of LAMP-1 in mediating lysosomal function. We observed an increase of Rab7 basal levels in sAD cybrids, which could be reversed by inhibiting p300 following C646 treatment. Additionally, despite having detected an increase in LAMP-1 basal expression in sAD cybrids, we curiously noticed a decrease in large-sized clusters containing LAMP-1 immunoreactivity and a subsequent shift from large to small clusters in untreated sAD cells. Although Rab7 expression increases, we do observe an increase in lysosomal dispersion in sAD cybrids, which allows us to infer that Rab7 might not be successfully integrated in the clusters to address them correctly. Additionally, we also observed that LAMP-1 total levels increased in AD cells, despite observing a significant decrease of large clusters, which allow us to infer that a feedback mechanism might be promoting the upregulation of these proteins in order to facilitate lysosome clusters positioning and autophagosome clearance. Interestingly, the decreased total cluster numbers observed in C646-treated sAD cells indicate that Beclin-1 acetylation may be implicated in autophagic–lysosomal dysfunction that characterizes AD pathology.
In a concluding remark, it is conceived that activation of autophagy promotes neuronal survival in pathological conditions and pharmacological studies have already demonstrated that autophagy induction by SIRT1 can benefit neuronal fitness and ameliorate AD pathology in numerous disease models. We identify for the first time in this study that regulation of Beclin-1 acetylation is deeply relevant in AD, since we clearly demonstrated that Beclin-1 deacetylation can improve several processes related with the autophagic pathway such as the autophagic flux and the maturation of autophagosomes. Moreover, we also established that Beclin-1 acetylation participates in later lysosome confinement, which is crucial to the proper clearance of toxic aggregates.
Although we consider that further research is needed to clarify in detail the mechanisms behind autophagy–lysosomal pathway, we believe that regulation of Beclin-1 acetylation status by modulating SIRT1 and p300 function may be a feasible approach to overcome some defects observed in AD patients.
Material and Methods
Reagents
NAM, Leupeptin, and MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide) were purchased from Sigma Chemical Co., (St. Louis, MO, USA).
Ammonium Chloride (NH4Cl) was obtained from Merck KGaA (Darmstadt, Germany), and C646 was purchased from Calbiochem, Merck KGaA (Darmstadt, Germany). For immunoblotting analysis, the following secondary antibodies were used and the working dilutions are given in brackets: goat anti-mouse IgG alkaline phosphatase conjugated antibody (1:10,000) from Thermo Fisher Scientific, Inc. (Rockford, IL, USA) and goat anti-rabbit IgG alkaline phosphatase conjugated antibody (1:20,000) was from GE Healthcare (Bio-Sciences, Uppsala, Sweden).
The following primary antibodies were used, and the working dilutions are given in brackets: mouse monoclonal anti-cathepsin D (C-5) antibody (1:1000) was from Santa Cruz Biotechnology, Inc. (Santa Cruz, CA, USA); rabbit polyclonal anti-SIRT1 (D739) antibody (1:1000), rabbit polyclonal anti-phospho-SIRT1 (Ser47) antibody (1:2000), rabbit polyclonal anti-Beclin-1 antibody (1:1000), rabbit polyclonal anti-Rab7 antibody (1:1000), rabbit polyclonal anti-LC3B antibody, mouse monoclonal anti-α-tubulin (11H10) antibody (1:1000), rabbit polyclonal anti-acetylated-lysine antibody (1:1000) all were from Cell Signaling Technology, Inc. (Danvers, MA, USA); and rabbit polyclonal anti-p62 antibody (1:1000), mouse monoclonal anti-β-actin antibody (1:5000), all from Sigma (St. Louis, MO, USA). LAMP-2A (1:100) from Santa Cruz Biotechnology, Inc. (Santa Cruz, CA, USA). For immunoprecipitation, acetylated-lysine antibody (1:500) was used from Cell Signaling Technology, Inc. (Danvers, MA, USA), and for immunofluorescence, LAMP-1 antibody (1:100) was used from Hybridoma Bank, USA.
Human Subjects
Subject participation was approved by the Kansas University Medical Center’s Institutional Review Board. For this study, subjects were recruited from the University of Kansas Alzheimer’s Disease Center (KU ADC). Additionally, each subject was determined, based on cognitive testing and by a memory disorder subspecialist clinician, to meet criteria for normal cognition–control status (CT), mild cognitive impairment (MCI), or sporadic AD (sAD).
After providing informed consent, sAD (n = 8), MCI (n = 7), and CT (n = 7) subjects underwent a 10 mL phlebotomy using tubes containing acid-citrate-dextrose as an anticoagulant. The age of the sAD subject platelet donors was 71.5 ± 9.7 years; the age of the MCI platelet donors was 72.3 ± 6.6, and the age of the CT subject platelet donors was 73.9 ± 7.7 [70].
Creation of Cybrid Cell Lines
Cytoplasmic hybrid (cybrid) technique was first described by King and Attardi, that consists in the transfer of mitochondrial (mtDNA) exogenous to mtDNA-depleted recipient cells (Rho 0 line cells), generating cybrids. To generate the cybrid cell lines used in this study, we used platelets from human subjects, as already referred, CT, MCI, and sAD, which were mixed with SH-SY5Y cells previously depleted of endogenous mtDNA (Rho 0 cells) as previously described [70].
During the overall cybrid generation procedure, several different types of media were used. Dulbecco’s modified Eagle’s medium: Nutrient Mixture F-12 (DMEM/ F12) was obtained from Gibco-Invitrogen (Life Technologies Ltd., UK), while non-dialyzed FBS was obtained from Sigma (St. Louis, MO, USA). SH-SY5Y Rho 0 cell growth medium consisted of DMEM/F12 supplemented with 10% non-dialyzed FBS, 200 μg/mL sodium pyruvate obtained from Sigma (St. Louis, MO, USA), 150 μg/mL uridine obtained from Sigma (St. Louis, MO, USA), and 1% penicillin–streptomycin (pen-strep) solution. The SH-SY5Y cybrid selection medium consisted of DMEM/F12 supplemented with 10% dialyzed FBS and 1% pen-strep solution.
The selection process lasted 6 weeks. After cell line selection was completed, each line was continuously maintained in a cybrid growth medium containing DMEM/F12 supplemented with 10% non-dialyzed FBS and 1% pen-strep solution for over 2 months prior to biochemical and molecular assays [70].
Cell Media and Treatments
Cells were grown in 75 cm2 tissue culture flasks maintained in a humidified incubator at 37 °C and 5% CO2. For the MTT assay, cells were plated in 24-well plates at a density of 0.2 × 106 cells/mL.
For Western blot analysis and immunoprecipitation assay, cells were plated in 6-well plates at a density of 0.25 × 106 cells/mL or in petri dishes (10 cm). For immunocytochemistry analysis, cybrid cell lines were grown on coverslips in 12-well plates at a density of 0.1 × 106 cells/mL.
Prior to experiments, cell lines were maintained in the cybrid growth medium, cybrids containing DMEM supplemented with 10% non-dialyzed FBS, and 1% pen–strep solution.
Cybrids cell lines were incubated for 6 h with 5 mM NAM, a SIRT1 inhibitor, or with 2.5 μM C646, a p300 inhibitor. Autophagy modulation consisted of the treatment of the respective cell line with 20 mM NH4Cl and 10 mM Leupeptin for 4 h. For all experimental procedures, controls were performed in the absence of those agents.
MTT Reduction Test
Cell viability was determined by the MTT reduction test. In viable cells, the enzyme succinate dehydrogenase metabolizes MTT into a formazan that absorbs light at 570 nm. Following the cell treatment protocol, the medium was aspirated and 0.5 mL MTT (0.5 mg/mL) was added to each well. The plate was then incubated at 37 °C for 1 h 30 m protected from light. At the end of the incubation period, the formazan precipitates were solubilized with 0.5 mL of acidic isopropanol (0.04 M HCl/isopropanol). The absorbance was measured at 570 nm [71]. Cell reduction ability was expressed as a percentage of the CT cybrids.
Preparation of Whole Cellular Extracts
Individual cell lines were washed with phosphate-buffered saline 1 × (PBS), lysed on ice in a hypotonic lysis buffer containing 1% Triton X-100, 25 mM 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES), pH 7.5, 2 mM magnesium chloride (MgCl2), 1 mM ethylenediaminetetraacetic acid (EDTA), and 1 mM ethylene glycol-bis (β-aminoethyl ether)-N,N,N′,N′-tetraacetic acid (EGTA) and supplemented with 2 mM dithiothreitol (DTT) and protease inhibitors (0.1 mM phenylmethanesulfonyl fluoride [PMSF], and a 1:1000 dilution of a protease inhibitor cocktail). Cells were then scrapped on ice. Subsequently, cell suspensions were frozen three times in liquid nitrogen and centrifuged at 20,000×g for 10 min.
The resultant supernatant was removed and stored at − 80 °C. Protein content was determined using Bradford Protein Assay (Bio-Rad, Hercules, CA, USA).
Preparation of Cytosolic and Nuclear Cellular Extracts
Cell lines were washed with PBS (1×), lysed on ice in a first buffer, buffer I containing 10 mM HEPES, 10 mM NaCl, 3 mM MgCl2, 1 mM EGTA, and pH 7.5, supplemented with 0.1% Triton X-100, with 1 mM DTT, with protease inhibitors (1 mM DTT, 1 mM PMSF, and 1:1000 dilution of a protease inhibitor cocktail), and with phosphatase inhibitors (2 mM sodium orthovanadate [Na3VO4] and 50 mM of sodium fluoride [NAF]). Cells were then scraped on ice.
Afterwards, cell suspensions were centrifuged at 2300×g for 10 min at 4 °C. The resulting supernatant was the cytosolic fraction. Then, the resulting pellet was resuspended in a second buffer, buffer II containing 20 mM HEPES, 300 mM NaCl, 5 mM MgCl2, 0.2 mM EGTA, 20% glycerol, and pH 7.5, supplemented with 1 mM DTT, 1 mM PMSF, and 1:1000 dilution of a protease inhibitor cocktail. Cell suspensions were centrifuged at 12,000×g for 20 min at 4 °C. The resulting supernatant was the nuclear fraction. Both fractions were stored at − 80 °C. Protein content was determined using Bradford protein assay (Bio-Rad, Hercules, CA, USA).
Western Blot Analyses
For Western blot analysis samples were resuspended in 6 × sample buffer (4 × Tris–Cl/SDS, pH 6.8, 30% glycerol, 10% SDS, 0.6 M DTT, 0.012% bromophenol blue) under reducing conditions. Depending on the protein molecular weight, samples containing 30 μg of protein were loaded onto accordingly percent SDS-polycrylamide (SDS-PAGE) gels. Specifically, for the analysis of LC3, LAMP-2A, and cathepsin D, samples were separated by electrophoresis on a 15% gel, and all the other protein samples were separated by electrophoresis on a 10% gel.
After electrophoresis, gels were transferred to polyvinylidene difluoride (PVDF) membranes (Millipore, Billerica, MA, USA). Subsequently, non-specific binding was blocked by gently agitating the membranes in 3% bovine serum albumin (BSA) for 1 h at room temperature. The blots were subsequently incubated with the respective primary antibodies overnight at 4 °C with gentle agitation. The next day, membranes were washed in Tris-buffered solution (TBS) contain 0.1% Tween, three times, each time for 5 min, and then incubated with the appropriate horseradish peroxidase-conjugated secondary antibody for 2 h at room temperature with gentle agitation. After three washes, specific bands of interest were detected by developing with an alkaline phosphatase enhanced chemical fluorescence reagent (ECF from GE Healthcare). Fluorescence signals were detected using a Bio-Rad VersaDoc Imager.
Immunoprecipitation Assay
Cells were scraped in buffer containing 20 mM Tris, 100 mM NaCl, 2 mM EDTA, 2 mM EGTA (pH 7.0), and protease inhibitors (100 mM PMSF and a commercial protease inhibitor cocktail). CT, MCI, and sAD cybrid cell suspensions were centrifuged at 20,000×g for 10 min at 4 °C.
Supernatants were collected and stored at − 80 °C. Protein content was determined using Bradford Protein Assay (Bio-Rad, Hercules, CA, USA), and 500 μg of cell lysates were incubated with the acetylated-lysine (1:500) primary antibody overnight at 4 °C, and with gentle agitation. Lysates were then incubated with 100 μL of protein-A beads for 2 h at 4 °C and with gentle agitation. After completing this incubation, lysate tubes were centrifuged at 65×g for 5 min at 4 °C; the supernatant was removed, and the beads were washed buffer five times (each time centrifuging at 4 °C and removing the supernatant). For the two first washes, the buffer was supplemented with 1% Triton X-100. For the next two washes, the buffer was supplemented with 1% Triton X-100 and 5 mM NaCl. The two final washes were performed with unsupplemented buffer. Finally, the last supernatant was removed, and 25 μL of sample buffer (2×) was added.
The sample was boiled at 95–100 °C for 5 min to denature the protein and to separate it from the protein-A beads. The boiled proteins were centrifuged at 20,000×g for 5 min at room temperature, and the supernatants were collected. The resulting immunoprecipitated samples were separated by SDS-PAGE and subjected to Western blot analysis using anti-Beclin-1 antibody and anti-acetylated-lysine antibody, as previously described.
Immunofluorescence and Confocal Microscopy
Cybrid cells were washed with PBS (1×) and fixed for 30 min at room temperature using 4% paraformaldehyde. Fixed cells were then washed again with PBS (1×), permeabilized with methanol at − 20 °C for 30 min, and blocked with 3% BSA for 30 min. Afterwards, cells were incubated with primary antibodies (1:100 anti-LAMP-1 (H4A30) from the Developmental Studies Hybridoma Bank (University of Iowa, Iowa City, IA, USA) overnight in a wet chamber at 4 °C. Later, cells were incubated 1 h with the appropriate secondary antibody (1:250 Alexa Fluor 488 or 594 from Molecular Probes, Eugene, OR, USA). Finally, cells were washed in PBS (1×), incubated for 5 min with Hoechst (15 μg/μL) in the dark. Cells were then washed twice in PBS (1×), and the coverslips were immobilized on a glass slide with mounting medium DakoCytomation (Dako, Glostrup, Denmark). Confocal images were obtained using a Plan-Apochromat/1.4NA 63× lens on an Axio Observer.Z1 confocal microscope (Zeiss Microscopy, Germany) with Zeiss LSM 710 software.
LAMP1-positive clusters per cell were quantified using the “analyze particle” function of the ImageJ, as previously described by Bandyopadhyay and colleagues [72].
Punctate lysosomes were selected with a size ≤ 0.2 μM, whereas lysosome clusters were selected with a size between 0.2 and 5 μM. LAMP-1 fluorescence intensity was quantified using ImageJ v1.39 k (National Institute of Health, USA) program.
Data Acquisition and Analysis
All data result was expressed as mean ± standard error of the mean (SEM) in at least three independent experiments. Analyses and graphical presentation were performed with the GraphPad Prism software version 5 (GraphPad Software, Inc., San Diego, CA, USA). Differences between two data sets were evaluated by unpaired Student’s t test. Statistical tests between multiple data sets and conditions were carried out using a one way analysis of variance (ANOVA) with pairwise multiple comparison procedures using the post hoc Bonferroni’s test to determine statistical significance, as appropriate. A p value < 0.05 was considered statistically significant.
References
Nah J (2015) Autophagy in neurodegenerative diseases: from mechanism to therapeutic approach. Mol Cells 38(5):381–389. https://doi.org/10.14348/molcells.2015.0034
Nixon RA, Wegiel J, Kumar A, Yu WH, Peterhoff C, Cataldo A, Cuervo AM (2005) Extensive involvement of autophagy in Alzheimer disease: an immuno-electron microscopy study. J Neuropathol Exp Neurol 64(2):113–122
Esteves AR, Arduino DM, Silva DF, Viana SD, Pereira FC, Cardoso SM (2018) Mitochondrial metabolism regulates microtubule acetylome and autophagy trough sirtuin-2: impact for Parkinson’s disease. Mol Neurobiol 55(2):1440–1462. https://doi.org/10.1007/s12035-017-0420-y
Guo F, Liu X, Cai H, Le W (2018) Autophagy in neurodegenerative diseases: pathogenesis and therapy. Brain Pathol 28(1):3–13. https://doi.org/10.1111/bpa.12545
Arduino DM, Esteves AR, Cortes L, Silva DF, Patel B, Grazina M, Swerdlow RH, Oliveira CR et al (2012) Mitochondrial metabolism in Parkinson’s disease impairs quality control autophagy by hampering microtubule-dependent traffic. Hum Mol Genet 21(21):4680–4702. https://doi.org/10.1093/hmg/dds309
Komatsu M, Waguri S, Chiba T, Murata S, Iwata J, Tanida I, Ueno T, Koike M et al (2006) Loss of autophagy in the central nervous system causes neurodegeneration in mice. Nature 441(7095):880–884. https://doi.org/10.1038/nature04723
Hara T, Nakamura K, Matsui M, Yamamoto A, Nakahara Y, Suzuki-Migishima R, Yokoyama M, Mishima K et al (2006) Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 441(7095):885–889. https://doi.org/10.1038/nature04724
Nilsson P, Loganathan K, Sekiguchi M, Matsuba Y, Hui K, Tsubuki S, Tanaka M, Iwata N et al (2013) Abeta secretion and plaque formation depend on autophagy. Cell Rep 5(1):61–69. https://doi.org/10.1016/j.celrep.2013.08.042
Majumder S, Richardson A, Strong R, Oddo S (2011) Inducing autophagy by rapamycin before, but not after, the formation of plaques and tangles ameliorates cognitive deficits. PLoS One 6(9):e25416. https://doi.org/10.1371/journal.pone.0025416
Pickford F (2008) The autophagy-related protein beclin 1 shows reduced expression in early Alzheimer disease and regulates amyloid β accumulation in mice. J Clin Invest 118(6):2190–2199. https://doi.org/10.1172/jci33585
Lonskaya I, Hebron ML, Desforges NM, Franjie A, Moussa CE (2013) Tyrosine kinase inhibition increases functional parkin-Beclin-1 interaction and enhances amyloid clearance and cognitive performance. EMBO Mol Med 5(8):1247–1262. https://doi.org/10.1002/emmm.201302771
Jaeger PA, Pickford F, Sun CH, Lucin KM, Masliah E, Wyss-Coray T (2010) Regulation of amyloid precursor protein processing by the Beclin 1 complex. PLoS One 5(6):e11102. https://doi.org/10.1371/journal.pone.0011102
Sun T, Li X, Zhang P, Chen WD, Zhang HL, Li DD, Deng R, Qian XJ et al (2015) Acetylation of Beclin 1 inhibits autophagosome maturation and promotes tumour growth. Nat Commun 6:7215. https://doi.org/10.1038/ncomms8215
Kim D, Nguyen MD, Dobbin MM, Fischer A, Sananbenesi F, Rodgers JT, Delalle I, Baur JA et al (2007) SIRT1 deacetylase protects against neurodegeneration in models for Alzheimer’s disease and amyotrophic lateral sclerosis. EMBO J 26(13):3169–3179. https://doi.org/10.1038/sj.emboj.7601758
Satoh A, Brace CS, Rensing N, Clifton P, Wozniak DF, Herzog ED, Yamada KA, Imai S (2013) Sirt1 extends life span and delays aging in mice through the regulation of Nk2 homeobox 1 in the DMH and LH. Cell Metab 18(3):416–430. https://doi.org/10.1016/j.cmet.2013.07.013
Huang R, Xu Y, Wan W, Shou X, Qian J, You Z, Liu B, Chang C et al (2015) Deacetylation of nuclear LC3 drives autophagy initiation under starvation. Mol Cell 57(3):456–466. https://doi.org/10.1016/j.molcel.2014.12.013
Lee IH, Cao L, Mostoslavsky R, Lombard DB, Liu J, Bruns NE, Tsokos M, Alt FW et al (2008) A role for the NAD-dependent deacetylase Sirt1 in the regulation of autophagy. Proc Natl Acad Sci U S A 105(9):3374–3379. https://doi.org/10.1073/pnas.0712145105
Wang T, Cui H, Ma N, Jiang Y (2013) Nicotinamide-mediated inhibition of SIRT1 deacetylase is associated with the viability of cancer cells exposed to antitumor agents and apoptosis. Oncol Lett 6(2):600–604. https://doi.org/10.3892/ol.2013.1400
Shrimp JH, Sorum AW, Garlick JM, Guasch L, Nicklaus MC, Meier JL (2016) Characterizing the covalent targets of a small molecule inhibitor of the lysine acetyltransferase P300. ACS Med Chem Lett 7(2):151–155. https://doi.org/10.1021/acsmedchemlett.5b00385
Ou X, Lee MR, Huang X, Messina-Graham S, Broxmeyer HE (2014) SIRT1 positively regulates autophagy and mitochondria function in embryonic stem cells under oxidative stress. Stem Cells 32(5):1183–1194. https://doi.org/10.1002/stem.1641
Ghosh AK, Varga J (2007) The transcriptional coactivator and acetyltransferase p300 in fibroblast biology and fibrosis. J Cell Physiol 213(3):663–671. https://doi.org/10.1002/jcp.21162
Wan W, You Z, Xu Y, Zhou L, Guan Z, Peng C, Wong CCL, Su H et al (2017) mTORC1 phosphorylates acetyltransferase p300 to regulate autophagy and lipogenesis. Mol Cell 68(2):323–335.e326. https://doi.org/10.1016/j.molcel.2017.09.020
Kumar R, Chaterjee P, Sharma PK, Singh AK, Gupta A, Gill K, Tripathi M, Dey AB et al (2013) Sirtuin1: A promising serum protein marker for early detection of Alzheimer’s disease. PLoS One 8(4):e61560. https://doi.org/10.1371/journal.pone.0061560
Silva DF, Selfridge JE, Lu J, E L, Roy N, Hutfles L, Burns JM, Michaelis EK et al (2013) Bioenergetic flux, mitochondrial mass and mitochondrial morphology dynamics in AD and MCI cybrid cell lines. Hum Mol Genet 22(19):3931–3946. https://doi.org/10.1093/hmg/ddt247
Julien C, Tremblay C, Emond V, Lebbadi M, Salem N Jr, Bennett DA, Calon F (2009) Sirtuin 1 reduction parallels the accumulation of tau in Alzheimer disease. J Neuropathol Exp Neurol 68(1):48–58. https://doi.org/10.1097/NEN.0b013e3181922348
Bai W, Zhang X (2016) Nucleus or cytoplasm? The mysterious case of SIRT1’s subcellular localization. Cell Cycle 15(24):3337–3338. https://doi.org/10.1080/15384101.2016.1237170
McKnight NC, Zhenyu Y (2013) Beclin 1, an essential component and master regulator of PI3K-III in health and disease. Curr Pathobiol Rep 1(4):231–238. https://doi.org/10.1007/s40139-013-0028-5
Jaeger PA, Wyss-Coray T (2010) Beclin 1 complex in autophagy and Alzheimer disease. Arch Neurol 67(10):1181–1184. https://doi.org/10.1001/archneurol.2010.258
Uddin MS, Stachowiak A, Mamun AA, Tzvetkov NT, Takeda S, Atanasov AG, Bergantin LB, Abdel-Daim MM et al (2018) Autophagy and Alzheimer’s disease: from molecular mechanisms to therapeutic implications. Front Aging Neurosci 10:04. https://doi.org/10.3389/fnagi.2018.00004
Silva DF, Esteves AR, Oliveira CR, Cardoso SM (2017) Mitochondrial metabolism power SIRT2-dependent deficient traffic causing Alzheimer’s-disease related pathology. Mol Neurobiol 54(6):4021–4040. https://doi.org/10.1007/s12035-016-9951-x
Liu WJ, Ye L, Huang WF, Guo LJ, Xu ZG, Wu HL, Yang C, Liu HF (2016) p62 links the autophagy pathway and the ubiqutin–proteasome system upon ubiquitinated protein degradation. Cell Mol Biol Lett 21:29. https://doi.org/10.1186/s11658-016-0031-z
Caccamo A, Ferreira E, Branca C, Oddo S (2017) p62 improves AD-like pathology by increasing autophagy. Mol Psychiatry 22(6):865–873. https://doi.org/10.1038/mp.2016.139
Yue Z (2007) Regulation of neuronal autophagy in axon: Implication of autophagy in axonal function and dysfunction/degeneration. Autophagy 3(2):139–141
Lee IH, Finkel T (2009) Regulation of autophagy by the p300 acetyltransferase. J Biol Chem 284(10):6322–6328. https://doi.org/10.1074/jbc.M807135200
Matsunaga K, Saitoh T, Tabata K, Omori H, Satoh T, Kurotori N, Maejima I, Shirahama-Noda K et al (2009) Two Beclin 1-binding proteins, Atg14L and Rubicon, reciprocally regulate autophagy at different stages. Nat Cell Biol 11(4):385–396. https://doi.org/10.1038/ncb1846
Ballabio A (2016) The awesome lysosome. EMBO Mol Med 8(2):73–76. https://doi.org/10.15252/emmm.201505966
Nixon RA (2016) The lysosome in aging-related neurodegenerative diseases. In: Lysosomes: Biology, Diseases, and Therapeutics. doi:https://doi.org/10.1002/9781118978320.ch9
Wartosch L, Bright NA, Luzio JP (2015) Lysosomes. Curr Biol CB 25(8):R315–R316. https://doi.org/10.1016/j.cub.2015.02.027
Di Domenico F, Tramutola A, Perluigi M (2016) Cathepsin D as a therapeutic target in Alzheimer’s disease. Expert Opin Ther Targets 20(12):1393–1395. https://doi.org/10.1080/14728222.2016.1252334
Sjödin S, Öhrfelt A, Brinkmalm G, Zetterberg H, Blennow K, Brinkmalm A (2016) Targeting LAMP2 in human cerebrospinal fluid with a combination of immunopurification and high resolution parallel reaction monitoring mass spectrometry. Clin Proteomics 13:4. https://doi.org/10.1186/s12014-016-9104-2
Wang Y, Martinez-Vicente M, Kruger U, Kaushik S, Wong E, Mandelkow EM, Cuervo AM, Mandelkow E (2009) Tau fragmentation, aggregation and clearance: the dual role of lysosomal processing. Hum Mol Genet 18(21):4153–4170. https://doi.org/10.1093/hmg/ddp367
Patel B, Cuervo AM (2015) Methods to study chaperone-mediated autophagy. Methods 75:133–140. https://doi.org/10.1016/j.ymeth.2015.01.003
Cataldo AM, Nixon RA (1990) Enzymatically active lysosomal proteases are associated with amyloid deposits in Alzheimer brain. Proc Natl Acad Sci U S A 87(10):3861–3865
Perez SE, He B, Nadeem M, Wuu J, Ginsberg SD, Ikonomovic MD, Mufson EJ (2015) Hippocampal endosomal, lysosomal, and autophagic dysregulation in mild cognitive impairment: correlation with abeta and tau pathology. J Neuropathol Exp Neurol 74(4):345–358. https://doi.org/10.1097/nen.0000000000000179
Johnson DE, Ostrowski P, Jaumouillé V, Grinstein S (2016) The position of lysosomes within the cell determines their luminal pH. J Cell Biol 212(6):677–692. https://doi.org/10.1083/jcb.201507112
Ginsberg SD, Alldred MJ, Counts SE, Cataldo AM, Neve RL, Jiang Y, Wuu J, Chao MV et al (2010) Microarray analysis of hippocampal CA1 neurons implicates early endosomal dysfunction during Alzheimer’s disease progression. Biol Psychiatry 68(10):885–893. https://doi.org/10.1016/j.biopsych.2010.05.030
Bucci C, Thomsen P, Nicoziani P, McCarthy J, van Deurs B (2000) Rab7: a key to lysosome biogenesis. Mol Biol Cell 11(2):467–480. https://doi.org/10.1091/mbc.11.2.467
Barrachina M, Maes T, Buesa C, Ferrer I (2006) Lysosome-associated membrane protein 1 (LAMP-1) in Alzheimer’s disease. Neuropathol Appl Neurobiol 32(5):505–516. https://doi.org/10.1111/j.1365-2990.2006.00756.x
Mathews PM, Guerra CB, Jiang Y, Grbovic OM, Kao BH, Schmidt SD, Dinakar R, Mercken M et al (2002) Alzheimer’s disease-related overexpression of the cation-dependent mannose 6-phosphate receptor increases Abeta secretion: role for altered lysosomal hydrolase distribution in beta-amyloidogenesis. J Biol Chem 277(7):5299–5307. https://doi.org/10.1074/jbc.M108161200
Funderburk SF, Marcellino BK, Yue Z (2010) Cell “self-eating” (autophagy) mechanism in Alzheimer’s disease. Mt Sinai J Med 77(1):59–68. https://doi.org/10.1002/msj.20161
Nixon RA, Yang DS (2011) Autophagy failure in Alzheimer’s disease—locating the primary defect. Neurobiol Dis 43(1):38–45. https://doi.org/10.1016/j.nbd.2011.01.021
Xie Y, Kang R, Sun X, Zhong M, Huang J, Klionsky DJ, Tang D (2015) Posttranslational modification of autophagy-related proteins in macroautophagy. Autophagy 11(1):28–45. https://doi.org/10.4161/15548627.2014.984267
Marzi SJ, Leung SK, Ribarska T, Hannon E, Smith AR, Pishva E, Poschmann J, Moore K, Troakes C, Al-Sarraj S, Beck S, Newman S, Lunnon K, Schalkwyk L, Mill J (2018) A histone acetylome-wide association study of Alzheimer's disease identifies disease-associated H3K27ac differences in the entorhinal cortex. Nat Neurosci 21(11):1618–1627
Donmez G, Outeiro TF (2013) SIRT1 and SIRT2: emerging targets in neurodegeneration. EMBO Mol Med 5(3):344–352. https://doi.org/10.1002/emmm.201302451
Gan L, Mucke L (2008) Paths of convergence: sirtuins in aging and neurodegeneration. Neuron 58(1):10–14. https://doi.org/10.1016/j.neuron.2008.03.015
Ng F, Tang BL (2013) Sirtuins’ modulation of autophagy. J Cell Physiol 228(12):2262–2270. https://doi.org/10.1002/jcp.24399
Zakhary SM, Ayubcha D, Dileo JN, Jose R, Leheste JR, Horowitz JM, Torres G (2010) Distribution analysis of deacetylase SIRT1 in rodent and human nervous systems. Anat Rec (Hoboken) 293(6):1024–1032. https://doi.org/10.1002/ar.21116
Fekadu J, Rami A (2016) Beclin-1 deficiency alters autophagosome formation, lysosome biogenesis and enhances neuronal vulnerability of HT22 hippocampal cells. Mol Neurobiol 53(8):5500–5509. https://doi.org/10.1007/s12035-015-9453-2
Armstrong A, Mattsson N, Appelqvist H, Janefjord C, Sandin L, Agholme L, Olsson B, Svensson S et al (2014) Lysosomal network proteins as potential novel CSF biomarkers for Alzheimer’s disease. NeuroMolecular Med 16(1):150–160. https://doi.org/10.1007/s12017-013-8269-3
Cataldo AM, Paskevich PA, Kominami E, Nixon RA (1991) Lysosomal hydrolases of different classes are abnormally distributed in brains of patients with Alzheimer disease. Proc Natl Acad Sci U S A 88(24):10998–11002
Cataldo AM, Hamilton DJ, Nixon RA (1994) Lysosomal abnormalities in degenerating neurons link neuronal compromise to senile plaque development in Alzheimer disease. Brain Res 640(1–2):68–80
Touitou I, Capony F, Brouillet JP, Rochefort H (1994) Missense polymorphism (C/T224) in the human cathepsin D pro-fragment determined by polymerase chain reaction—single strand conformational polymorphism analysis and possible consequences in cancer cells. Eur J Cancer 30a(3):390–394
Ntais C, Polycarpou A, Ioannidis JP (2004) Meta-analysis of the association of the cathepsin D Ala224Val gene polymorphism with the risk of Alzheimer’s disease: a HuGE gene-disease association review. Am J Epidemiol 159(6):527–536
Cataldo AM, Hamilton DJ, Barnett JL, Paskevich PA, Nixon RA (1996) Properties of the endosomal-lysosomal system in the human central nervous system: disturbances mark most neurons in populations at risk to degenerate in Alzheimer’s disease. J Neurosci 16(1):186–199
Wen H, Zhan L, Chen S, Long L, Xu E (2017) Rab7 may be a novel therapeutic target for neurologic diseases as a key regulator in autophagy. J Neurosci Res 95(10):1993–2004. https://doi.org/10.1002/jnr.24034
Gutierrez MG, Munafo DB, Beron W, Colombo MI (2004) Rab7 is required for the normal progression of the autophagic pathway in mammalian cells. J Cell Sci 117(Pt 13):2687–2697. https://doi.org/10.1242/jcs.01114
Ginsberg SD, Mufson EJ, Counts SE, Wuu J, Alldred MJ, Nixon RA, Che S (2010) Regional selectivity of rab5 and rab7 protein up regulation in mild cognitive impairment and Alzheimer’s disease. J Alzheimers Dis 22(2):631–639. https://doi.org/10.3233/jad-2010-101080
Piras A, Collin L, Gruninger F, Graff C, Ronnback A (2016) Autophagic and lysosomal defects in human tauopathies: analysis of post-mortem brain from patients with familial Alzheimer disease, corticobasal degeneration and progressive supranuclear palsy. Acta Neuropathol Commun 4:22. https://doi.org/10.1186/s40478-016-0292-9
Andrejewski N, Punnonen EL, Guhde G, Tanaka Y, Lullmann-Rauch R, Hartmann D, von Figura K, Saftig P (1999) Normal lysosomal morphology and function in LAMP-1-deficient mice. J Biol Chem 274(18):12692–12701
Silva DF, Santana I, Esteves AR, Baldeiras I, Arduino DM, Oliveira CR, Cardoso SM (2013) Prodromal metabolic phenotype in MCI cybrids: implications for Alzheimer’s disease. Curr Alzheimer Res 10(2):180–190
Esteves AR, Arduino DM, Swerdlow RH, Oliveira CR, Cardoso SM (2010) Dysfunctional mitochondria uphold calpain activation: contribution to Parkinson’s disease pathology. Neurobiol Dis 37(3):723–730. https://doi.org/10.1016/j.nbd.2009.12.011
Bandyopadhyay D (2014) Lysosome transport as a function of lysosome diameter. PLoS One 9(1):e86847. https://doi.org/10.1371/journal.pone.0086847
Funding
Work in our laboratory was supported by Fundação para a Ciência e a Tecnologia (FCT) and by EU-FEDER funding through the Operational Competitiveness Program—COMPETE grant UID/NEU/04539/2013, by the European Regional Development Fund, Centro 2020 Regional Operational Program (CENTRO-01-0145-FEDER-000012-HealthyAging2020), by Prémio Santa Casa Neurociências Mantero Belard MB-40-2016, and by Project POCI-01-0145-FEDER-030712. AR Esteves and DF Silva are supported by post-doctoral fellowship from FCT-MCTES, Portugal. JD Magalhães is supported by a MB-40-2016 fellowship.
Author information
Authors and Affiliations
Contributions
FF, ARE, and DFS acquired the data; ARE and SMC discussed the results, and JDM and SMC wrote the article.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Esteves, A.R., Filipe, F., Magalhães, J.D. et al. The Role of Beclin-1 Acetylation on Autophagic Flux in Alzheimer’s Disease. Mol Neurobiol 56, 5654–5670 (2019). https://doi.org/10.1007/s12035-019-1483-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-019-1483-8