Abstract
Lafora disease (LD, OMIM 254780, ORPHA501) is a fatal neurodegenerative disorder characterized by the presence of glycogen-like intracellular inclusions called Lafora bodies and caused, in the vast majority of cases, by mutations in either EPM2A or EPM2B genes, encoding respectively laforin and malin. In the last years, several reports have revealed molecular details of these two proteins and have identified several processes affected in LD, but the pathophysiology of the disease still remains largely unknown. Since autophagy impairment has been reported as a characteristic treat in both Lafora disease cell and animal models, and as there is a link between autophagy and mitochondrial performance, we sought to determine if mitochondrial function could be altered in those models. Using fibroblasts from LD patients, deficient in laforin or malin, we found mitochondrial alterations, oxidative stress and a deficiency in antioxidant enzymes involved in the detoxification of reactive oxygen species (ROS). Similar results were obtained in brain tissue samples from transgenic mice deficient in either the EPM2A or EPM2B genes. Furthermore, in a proteomic analysis of brain tissue obtained from Epm2b−/−mice, we observed an increase in a modified form of peroxiredoxin-6, an antioxidant enzyme involved in other neurological pathologies, thus corroborating an alteration of the redox condition. These data support that oxidative stress produced by an increase in ROS production and an impairment of the antioxidant enzyme response to this stress play an important role in development of LD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Progressive myoclonus epilepsy of the Lafora type (Lafora disease; LD, OMIM 254780, ORPHA501), first described in 1911 [1], is a rare and fatal autosomic recessive neurodegenerative disease that produces epileptic crises, myoclonus and absence seizures, ataxia and dementia. In the vast majority of patients, LD has been associated with mutations in two genes: EPM2A, encoding the glucan phosphatase laforin, and EPM2B, encoding the E3-ubiquitin ligase malin. However, the role of these two proteins in LD has remained elusive despite the accumulation of experimental data regarding the function of laforin and malin, the identification of alterations in various cell processes and the availability of several animal and cell models of the disease. Laforin and malin form a functional complex, which has been demonstrated to regulate the activity of proteins involved in glycogen synthesis [2–5], and laforin is the only human phosphatase able to dephosphorylate complex glucans [6, 7]. Therefore, for several years, LD has been linked to aberrant glycogen accumulation in neurons, in the so-called Lafora bodies (LBs). LBs are glycogen-like inclusions with a higher degree of phosphorylation and less branched than normal glycogen, hence LBs are largely insoluble. Since ubiquitinated proteins, chaperones and proteasome subunits have been detected in LBs [8], LD can also be linked to the group of neurodegenerative proteinopathies. Nevertheless, the cause–effect connection of LBs with the neurodegenerative processes that lead to the disease is not thoroughly understood. Recent work has demonstrated that neuronal death in LD could be influenced not only by the accumulation of LBs but also by impairment in mechanisms related to protein clearance. For instance, endoplasmic reticulum (ER) stress is significantly increased in mice models of LD [9–12] and molecular interactions between laforin, malin and the chaperone and proteasome systems have been described [8, 13]. These evidences point to a critical dysregulation of cellular protein homeostasis following the disruption of a functional laforin–malin complex that goes beyond alterations in glycogen metabolism regulation. In this sense, our previous work with malin knock-out mice (Epm2b−/−) [14] revealed a clear impairment of autophagy, previously observed in laforin-deficient cells [15] which correlated with neurological and behavioral abnormalities. Also defects in the ubiquitin–proteasome system have been found in Lafora disease models [16, 11].
Since both defects of autophagy and protein clearance have been related with an increase in reactive oxygen species (ROS) production and the consequent oxidative stress [17–20], we decided to analyze these parameters in LD cell and animal models. Oxidative stress is a process whose important role in the development of several neurological disorders is broadly documented [21–23] and production of ROS that oxidize proteins and lipids in the cells may lead to senescence and apoptosis. In many cases, ROS production is usually increased as a consequence of a previous mitochondrial malfunction, and a dysregulation of antioxidant cellular systems often contributes to the worsening of an already altered cell function. Thus, we started our approach examining mitochondrial function in Lafora disease cell models and found important alterations that correlated with significant increased levels of ROS and defects in the antioxidant response in these cells. These results were corroborated in brain tissue samples obtained from LD mice models. Finally, a proteomic approach in an Epm2b−/−mouse model identified the presence of a modified form of peroxiredoxin-6 (Prdx6), which further suggested an impairment of the antioxidant systems. Taken together, these results provide evidences of an alteration of the antioxidant responses in LD models.
Materials and Methods
Mammalian Cell Culture
Human fibroblasts were obtained from two patients with two different mutations, Y86X and R241X, in the laforin gene (EPM2A) [5] and from one patient with the P129H mutation in the malin gene (EPM2B) [(GM008935) from Coriell Institute for Medical Research (Camden, NJ, USA)], all of them with the clinical features of LD. For simplicity, we will name these cells as Epm2a and Epm2b, respectively. Control fibroblasts were matched by sex and age. The results on control and Epm2a fibroblasts correspond to the mean obtained from two independent healthy and two patient individuals (no significant differences were observed between samples from the two types of cells in the same group). Cells were cultured at 37 °C in a humidified 5 % CO2 atmosphere in MEM supplemented with 15 % fetal bovine serum, 1 % MEM amino acids, 0.5 % MEM non-essential amino acids, 1 % glutamine, 1 % vitamins, 100 units/ml penicillin and 100 μg/ml streptomycin (fibroblasts’ basal growth medium; low proteolysis), all from Invitrogen Life Technologies. For nutrient stress conditions (high proteolysis), cells were switched, after washing, from fibroblasts’ basal growth medium to Krebs–Henseleit medium (118.4 mM NaCl, 4.75 mM KCl, 1.19 mM KH2PO4, 2.54 mM MgSO4, 2.44 mM CaCl2 · 2H2O, 28.6 mM NaHCO3, 10 mM glucose) containing 10 mM Hepes, pH 7.4 and incubated for 2 h at 37 °C. Cell viability and growth curves were obtained by counting the cells with a hemocytometer chamber and using the trypan blue exclusion test. All experiments were performed at passage number 10–14 to avoid culture senescence effects.
Fluorescence Microscopy
Cells grown on coverslips were incubated with 75 nM MytoTracker Red for 15 min at 37 °C. Then, the cells were rinsed with phosphate buffered saline (PBS), fixed with 3.7 % formaldehyde/PBS for 15 min at 37 °C, washed again with PBS, mounted using FluorSave reagent (Calbiochem) and observed with an Apotome-equipped Axio Observer Z1 microscope (Carl Zeiss).
Flow Cytometry
For flow cytometry analysis, cells were detached by trypsinization and cells in suspension (5 × 105 cells/ml) were incubated with the different fluorophores. Mitochondrial membrane potential (ΔΨm) was measured by incubating the cells with 100 nM tetramethylrhodamine-methyl-ester (TMRM, Molecular Probes) for 15 min at 37 °C. TMRM was used in the redistribution mode to assess ΔΨm, and therefore, a reduction in TMRM fluorescence represents ΔΨm depolarization. As a ΔΨm control, 16 h before the experiment, the protonophore carbonyl cyanide m-chlorophenylhydrazone (CCCP, Sigma, St. Louis, MO, USA) was added at 10 μM. For measurement of ROS production, cells were incubated for 30 min al 37 °C with 2′,7′-dichlorofluorescein diacetate (2.5 μM, Sigma) and the emitted fluorescence for total ROS detection was measured. The emitted fluorescence was analyzed in 105 cells per sample by flow cytometry as described previously [24], using a Cytomics FC 500 flow cytometer (Beckman Coulter).
Measurements of Intracellular ATP and Superoxide Production
ATP was measured using the ATP bioluminescence assay kit HS II (Roche Applied Science). Briefly, human fibroblasts were grown in 12-well plates to 50 % confluence. Then cells were incubated in fresh medium for 3 h and intracellular ATP content was measured following the manufacturer’s instructions using a Spectra Max M5 microplate reader (Molecular Devices, Sunnyvale, CA, USA). Luminescence intensity was normalized to the protein present in each sample.
For measurements of superoxide production, cells were grown to confluence in six- chamber plates for 6 days. After two washes with prewarmed PBS medium, 2 μl/ml of diluted dihydroethidium (Sigma) was added to the plate. Cells were incubated at 37 °C for 20 min. After two additional washes with PBS, medium was replaced with fresh Dulbecco's modified eagle's medium (DMEM). The fluorescence was measured using a SpectraMAX GEMINIS microplate reader (Molecular Devices) at 530 nm of excitation and 610 nm of emission wavelengths. All samples were analyzed two times in three independent experiments.
Quantitative PCR
RNA Isolation and cDNA Synthesis
Total RNA was isolated from cells using the PARIS™ protein and RNA isolation system (Ambion, Austin, TX, USA) according to the manufacturer’s instructions. For reverse transcription (RT) reactions, 1 mg of the purified RNA was reverse transcribed using random hexamers with the high-capacity complementary DNA (cDNA) reverse transcription kit (Applied Biosystems, Foster City, CA, USA; P/N: 4322171) according to the manufacturer’s instructions. RT conditions comprised an initial incubation step at 25 °C for 10 min to allow annealing of random hexamers, followed by cDNA synthesis at 37 °C for 120 min and a final 5-min inactivation step at 95 °C.
Measurement of mRNA Levels
The messenger RNA (mRNA) levels were determined by quantitative real-time polymerase chain reaction (PCR) analysis using an ABI Prism 7900 HT fast real-time PCR system (Applied Biosystems). Gene-specific primer pairs and probes for SOD2 (MnSOD; Hs00167309_m1, Taqman® assays, Applied Biosystems) and CAT (catalase; Hs00989766_g1, Taqman® assays, Applied Biosystems) were used together with TaqManH universal PCR master mix (Applied Biosystems, P/N 4304437) and reverse transcribed sample RNA in 20 μl reaction volumes. PCR conditions were 10 min at 95 °C for enzyme activation, followed by 40 two-step cycles (15 s at 95 °C; 1 min at 60 °C). Expression levels of glyceraldehyde-3-phosphate dehydrogenase were measured in all samples to normalize differences in RNA input, RNA quality and reverse transcription efficiency. Each sample was analyzed in triplicate and the relative expression levels were calculated according to the 2−ΔΔCt method [25].
Western Blotting
Proteins in the sample were denatured using sample buffer (40 mM Tris, 15 mM EDTA, 0.01 % bromophenol blue, 40 % sucrose, 4 % SDS, 10 % β-mercaptoethanol) and heating to 95 °C for 5 min. The samples were subjected to SDS-PAGE (12 % acrylamide) and transferred onto nitrocellulose membranes (Whatman, Dassel, Germany). Membranes were blocked with 5 % milk in TBS-Tween for 1 h and slices of these membranes were incubated with the following specific antibodies: anti-catalase (1:1000, Sigma), anti-GPX1 (1:750, Abcam, MA, USA), anti-MnSOD (1:1000, Stressgen, Ann Arbor, MI, USA), anti-CuZnSOD (1:1000, Stressgen), anti-Prdx6 (1:1000, Abcam, UK) and, as loading control, antibody that recognizes β-actin (1:1000, Santa Cruz BioTech, USA). Thereafter, the blots were washed again with TBS-Tween and further incubated for 1 h with the corresponding secondary antibody conjugated with horseradish peroxidase. Finally, membranes were washed (3 × 5 min) with TBS-Tween and analyzed by chemiluminiscence (ECL Western blotting detection reagents, GE Healthcare, UK) using an image reader LAS-4000 (GE Healthcare). Densitometry of the bands was performed using Image-J.
Antioxidant Enzyme Activities
Antioxidant enzyme activities were performed as previously described [22]. For CuZnSOD and MnSOD, a superoxide dismutase assay kit (Cayman, Ann Arbor, MI, USA) was used, based on the reduction of the tetrazolium salt by superoxide to give rise to the formazan salt, which is measured at 460 nm. Catalase enzymatic activity was determined by a standard method based on the reaction of the enzyme with methanol in the presence of hydrogen peroxide to produce formaldehyde. Total glutathione peroxidase activity was measured with a glutathione peroxidase assay kit (Cayman) following the manufacturer’s instructions. All spectrophotometric measurements were carried out using a spectrophotometer SpectraMAX PLUS384 (Molecular Devices).
Measurement of Lipid Peroxide MDA
Lipid peroxides were determined by measuring the final product malondialdehyde (MDA). A 10-mg brain tissue was homogenized in 100 μl of 0.05 M potassium phosphate buffer, 1 mM EDTA pH 7.4. For plasma samples, 50 μl were used. Tissue homogenates and plasma samples were deproteinized using 2.5 ml of 20 % trichloroacetic acid. After centrifugation, 1 ml of 0.6 % thiobarbituric acid (TBA) was added to the supernatant and the mixture incubated at 100 °C during 30 min to form a MDA–TBA adduct. The derivatized samples were separated by high-performance liquid chromatography (HPLC) on a column of octadecyl silica gel to separate the MDA–TBA adduct from interfering chromogens. The adduct was eluted from the column with 50 mM phosphate buffer pH 6.8, 50 % methanol and was spectrophotometrically quantified at 532 nm. MDA levels were expressed as nmol/mg protein.
Generation of Epm2a and Epm2b Knock-Out Mice
A detailed description of the procedures that ended in the generation of the Epm2a−/−and Epm2b−/−mice (genetic background C57BL/6 J) is provided in Refs. [26] and [14], respectively. Mice were maintained at the Instituto de Biomedicina de Valencia (IBV-CSIC, Valencia) on a light/dark 12:12 h cycle under constant temperature (23 °C) with free access to water and regular chow diet (65 % carbohydrate, 11 % fat, and 24 % protein). Animal care and all procedures involving animals used in this study were in accordance with the Declaration of Helsinki principles and the guidelines from the Institutional Animal Care and Use Committee, and approved by the IBV-CSIC ethical review board.
Two-Dimensional Differential In-Gel Electrophoresis (2D-DIGE)
Sample Preparation, Labelling and Protein Separation by 2-D Electrophoresis
Brain samples from control and Epm2b−/−mice were homogenized in 1 ml of lysis buffer containing 7 M urea, 2 M thiourea, 4 % CHAPS, 1 % dithiothreitol (DTT) (v/v) and 0.5 % Bio-Lyte (v/v) 3–10 ampholytes (GE Healthcare). Protein concentrations in the supernatants were measured with the Bradford assay kit (Bio-Rad) using bovine serum albumin (Sigma) as standard protein. Samples were desalted and concentrated with the cleanup kit (GE Healthcare) following the manufacturer’s recommendations and resuspended in 30 mM Tris, 7 M urea, 2 M thiourea and 4 % CHAPS (v/v).
Protein samples were labelled with CyDye® fluor minimal dyes in accordance with the manufacturer’s protocol (cyanine dye, Amersham Biosciences). Immobiline DryStrips pH 3–10 NL of 24 cm were passively rehydrated at 20 °C for 20 h. The Cy3- and Cy5-labelled samples to be separated on the same gel were pooled with an internal standard and mixed with an equal volume of 2× lysis buffer. Then an anodic cup-loading was used to load the samples in the strip for the first dimension. Isoelectic focusing (IEF) was run by applying 300 V for 4 h, 1,000 V for 6 h and 10,000 V for 3 h using an Amersham Ettan™ IPGphor unit (GE Healthcare). Focused strips were equilibrated in equilibration buffer [(50 mM Tris–HCl pH 7.5, 6 M Urea, 30 % glycerol (v/v), 2 % SDS (w/v)], first with 2 % DTT (w/v) and then with 2.5 % iodoacetamide. After equilibration, strips were loaded on 12.5 % SDS-PAGE gels in an Ettan Dalt-six separation unit (GE Healthcare) and run at 25 °C in two steps: 2 W/gel for 45 min and 17 W/gel for about 5 h. All electrophoresis procedures were performed in the dark. The protocol was repeated in three independent samples from control and Epm2b−/−mice.
Image Acquisition and Analysis
The gels were scanned with a Typhoon™ TRIO (GE Healthcare). Excitation and emission wavelengths were chosen specifically for each of the dyes according to the manufacturer’s recommendations. Image and statistical analyses were performed using DeCyder v7.0 (DIA and BVA module). Codetection and quantification of Cy2, Cy3 and Cy5 signals from each gel were determined in the differential in-gel analysis (DIA) module with an estimated number of spots set to 10,000 and an exclusion filter to omit spots with volume > 30,000. Data from the DIA module were exported to the biological variation analysis (BVA) module where gel-to-gel matching and two-way analysis of variance (ANOVA) statistical calculations were performed to determine the differentially expressed spots. Quantitative differences were only accepted at a fold change > 1.5. Statistical differences were calculated using Student’s t test and significance was accepted at p < 0.05.
LC-MS/MS Analysis (Q-TRAP 5500)
For each spot, a total volume of 2 μl of tryptic peptides was injected with a flow rate of 300 nl/min in a NanoLC Ultra1D plus system (Eksigent). A trap column Acclaim PepMap100 (100 μm × 2 cm; C18, 2 μm, 100 Å) and an analytical column Acclaim PepMap RSLC (75 μm × 15 cm, C18, 5 μm, 100 Å) from Dionex were used with the following gradient [Buffer A (0.1 % formic acid), Buffer B (100 % acetonitrile); only the amount of Buffer B is indicated]: 0–1 min (5 % B), 1–50 min (5–40 % B), 50–51 min (40–98 % B), 51–55 min (98 % B), 55–56 min (5 % B), 56–75 min (5 % B). MS analysis was performed on a Q-TRAP 5500 system (ABSciex) with a NanoSpray® III ion source (ABSciex) using rolling collision energy in positive mode. MS/MS data acquisition was performed using Analyst 1.5.2 (AB Sciex) and submitted to Phenyx™ Software (GeneBio, Geneva, Switzerland) against UniprotKB/Swiss-Prot database. Phenyx incorporates the true probabilistic and flexible scoring system OLAV developed at GeneProt Inc. [27]. The search was enzymatically constrained for trypsin and allowed for one missed cleavage site. Further search parameters were as follows: minimum significant scores for Phenyx were 5 and 6 for peptides and proteins, respectively; mass error tolerance was 50 ppm; there were no restrictions on molecular weight and isoelectric point; the fixed modification was carbamidomethylation of cysteine; and the variable modification was oxidation of methionine.
Statistical Analyses
For the statistical analysis of the results, the mean was taken as the measurement of the main tendency, while standard deviation was taken as the dispersion measurement. A one-way analysis of variance was used to determine the difference between groups, when analyzing superoxide levels and Western blotting densitometries and enzymatic activities of CuZnSOD, MnSOD, catalase and GPx1. When an interaction effect was found, multiple comparisons using the Student–Newman–Keuls method post hoc test were performed. The significance has been considered at * p < 0.05, ** p < 0.01 and *** p < 0.001, as indicated in each case. GraphPad Software v5.0 was used for statistical analysis and graphic representations.
Results
Human Fibroblasts from LD Patients Show Impaired Mitochondrial Function and Increased Oxidative Stress
To obtain further insights into the possible consequences of the previously described autophagy and protein clearance impairment in LD, we analyzed under low and high proteolysis conditions (see “Materials and Methods” section) a series of parameters of mitochondrial function in human fibroblasts from LD patients carrying mutations in the genes encoding either laforin (EPM2A) or malin (EPM2B) (for simplicity, we named these samples with altered laforin and malin as Epm2a and Epm2b, respectively; see “Materials and Methods” section). We started these studies analyzing the mitochondrial appearance with the fluorescence marker MitoTracker Red. We found that the tubular morphology of functional mitochondria observed in control fibroblasts was lost in both Epm2a and Epm2b fibroblasts (Fig. 1a). In fact, mitochondria in Epm2a and Epm2b fibroblasts looked aggregated (Fig. 1a, center and right), a morphology that also occurs under conditions of apoptosis [28]. Since mitochondrial dysfunction has been related with an elevated production of reactive oxygen species (ROS) [29, 30] and also with increased superoxide production [31, 22], we next analyzed the mitochondrial membrane potential by flow cytometry using TMRhodamine (TMRM) and found that it was decreased in LD human fibroblasts (Fig. 1b, the mitochondrial uncoupler CCCP was used as a positive control). The fact that morphology and membrane potential were altered in both groups of fibroblasts suggests an altered mitochondrial function. To corroborate this, we measured ATP production and also found decreased ATP levels in both groups of LD fibroblasts compared with control cells, a decrease that was also observed when cells were grown under nutrient stress conditions (high proteolysis medium; Fig. 1c). Thus, we conclude that mitochondrial function is affected in the fibroblasts from LD patients.
Mitochondrial morphology, membrane potential and cellular ATP levels in human fibroblasts from LD patients. a Representative fluorescence microscopy images of human fibroblasts from LD patients (Epm2a and Epm2b; see “Materials and Methods” section) and controls incubated with MitoTracker Red for 15 min. b The same groups of cells were incubated with TMRM for 15 min and analyzed by flow cytometry as described in “Materials and Methods” section. CCCP treatment (16 h, 10 μM) was used as control of membrane potential dysfunction. Results are represented as arbitrary units (a.u.) and expressed in percentage relative to the control fibroblasts. c Intracellular ATP levels from LD and control human fibroblasts grown in either basal (low proteolysis) or nutrient stress (high proteolysis) medium were measured as described in “Materials and Methods” section. Error bars represent SD of three independent measurements with duplicate samples. Statistical significance refers to the value of control samples in each condition (* for low proteolysis conditions; # for high proteolysis conditions; ** p < 0.01 and ***/### p < 0.001)
Next, we measured ROS levels (Fig. 2a) and superoxide production (Fig. 2b) in LD human fibroblasts and observed a significant raise of these parameters in both groups of LD human fibroblasts. These increases were more evident under basal conditions, probably because under these conditions, ROS levels and superoxide production are low in control cells. When these levels were enhanced by growing the cells under nutrient stress conditions (high proteolysis), only a slight increase in superoxide production was noticed, whereas no increase in the levels of ROS occurred. Therefore, we conclude that oxidative stress is increased in the fibroblasts from LD patients.
Analysis of oxidative stress parameters in LD human fibroblasts. Human fibroblasts grown in either basal (low proteolysis) or nutrient stress (high proteolysis) medium were incubated 30 min with 2′,7′-dichlorofluorescein diacetate (a) for ROS levels or with diluted dihydroethidium (b) for mitochondrial superoxide levels, as described in “Materials and Methods” section. Results are shown as arbitrary units (a.u.), normalized to control samples. Bars represent the mean ± SD from three independent experiments with duplicate samples and statistical significance refers to the value of control samples in each condition (* for low proteolysis conditions; # for high proteolysis conditions; **/## p < 0.01 and ***/### p < 0.001)
Antioxidant Enzymatic Activities Are Altered in LD Fibroblasts
We next analyzed a group of antioxidant enzymes in LD fibroblasts to characterize the antioxidant profile in these cells. First, we determined by Western blotting the levels of the superoxide dismutases MnSOD and CuZnSOD, the main enzymes involved in the detoxification of superoxide. A significant decrease in the protein levels of MnSOD, the mitochondrial variant of SOD, was evident in Epm2a and Epm2b cells both in low and high proteolysis conditions, a decrease that could be explained by a reduction of the levels of mRNA (Fig. 3a and Online Resource 1), whereas the protein levels of cytosolic CuZnSOD were not significantly altered in LD fibroblasts when compared to their controls (Fig. 4a). In accordance with these results, the enzymatic activity of MnSOD decreased in both Epm2a and Epm2b fibroblasts (Fig. 3b), confirming the inability of MnSOD to respond to oxidative stress. On the other hand, we detected an increase in the activity of the cytosolic form of SOD (CuZnSOD), but only in Epm2b cells growing in low proteolysis conditions (Fig. 4b). This may reflect either differences in the severity of the mutations present on LD fibroblasts or independent functions of laforin and malin in the cell. However, there was a clear decrease in this activity when both Epm2a and Epm2b cells were grown under high proteolysis conditions (Fig. 4b). At any rate, all these results clearly suggest an altered response of LD cells against superoxide.
Analysis of mitochondrial superoxide dismutase in LD human fibroblasts. a Protein extracts (20 μg protein) from cultured LD and control fibroblasts grown in either basal (low proteolysis) or nutrient stress (high proteolysis) medium were loaded in SDS-PAGE gels and analyzed by Western blotting using antibodies anti-MnSOD and anti-actin as loading control. Histograms show the quantification of various blots by densitometry with the values normalized to the corresponding actin levels and referred to the value of control samples in low proteolysis conditions. b MnSOD activity was measured in extracts from LD and control fibroblasts. Results represent the mean ± SD from three independent experiments and statistical significance refers to the value of control samples in each condition (* for low proteolysis conditions; # for high proteolysis conditions; * p < 0.05 and ***/### p < 0.001)
Analysis of cytosolic superoxide dismutase in LD human fibroblasts. a Protein extracts (20 μg protein) from cultured LD and control fibroblasts grown in either basal (low proteolysis) or nutrient stress (high proteolysis) medium were loaded in SDS-PAGE gels and analyzed by Western blotting using anti-CuZnSOD and anti-actin as loading control. Histograms show the quantification of various blots by densitometry with the values normalized to the corresponding actin levels and referred to the value of control samples in low proteolysis conditions. b CuZnSOD activity was measured in extracts from LD and control fibroblasts. Results represent the mean ± SD from three independent experiments and statistical significance refers to the value of control samples in each condition (* for low proteolysis conditions; # for high proteolysis conditions; ** p < 0.01 and ### p < 0.001)
In addition, the protein levels (Fig. 5a) and the enzymatic activity (Fig. 5b) of catalase, an enzyme involved in the detoxification of hydrogen peroxide, clearly decreased in Epm2a and Epm2b cells in both low and high proteolysis conditions (the decrease in protein amount was also corroborated by analyzing the mRNA levels, see Online Resource 1). However, in the latter conditions, the differences in enzymatic activity between control and LD cells were largely attenuated, probably due to a clear decrease of the activity of the control cells. Since hydrogen peroxide can also be detoxified by glutathione peroxidases (GPx), we analyzed GPx activity. We found that there were no differences between human LD and control fibroblasts either in the protein levels of the major isoform of glutathione peroxidase (GPx1; Fig. 6a), or in the total GPx activity, in either low or high proteolysis conditions (Fig. 6b). In summary, these results suggest that Epm2a and Epm2b fibroblasts have a partially defective antioxidant enzymatic defense that specifically affects some of their components.
Analysis of catalase in LD human fibroblasts. a Protein extracts (20 μg protein) from LD and control fibroblasts grown in either basal (low proteolysis) or nutrient stress (high proteolysis) medium were loaded in SDS-PAGE gels and analyzed by Western blotting using anti-catalase and anti-actin as a loading control. Histograms show the quantification of various blots by densitometry with the values normalized to the corresponding actin levels and referred to the value of control samples in low proteolysis conditions. b Catalase activity was analyzed as described in “Materials and Methods” section. Results represent the mean ± SD from three independent experiments and statistical significance refers to the value of control samples in each condition (* for low proteolysis conditions; # for high proteolysis conditions; **/## p < 0.01 and *** p < 0.001)
Analysis of glutathione peroxidase in LD human fibroblasts. a Protein extracts (20 μg protein) from cultured LD and control fibroblasts grown in either basal (low proteolysis) or nutrient stress (high proteolysis) medium were loaded on SDS-PAGE gels and analyzed by Western blotting using anti-GPx1 and anti-actin as a loading control. Densitometry of the bands (histograms) was normalized to the actin levels and referred to the value of control samples in low proteolysis conditions. b Total GPx activity was analyzed in total extracts obtained from LD and control fibroblasts, as described in “Materials and Methods” section. Results represent the mean ± SD from three independent experiments
Defective Antioxidant Response Elements in LD Animal Models
To get a more physiological overview of the oxidative stress conditions in LD, we analyzed two different models of transgenic mice deficient in laforin or malin (hereafter Epm2a−/−and Epm2b−/−, respectively) [14, 26]. Total brain homogenates from three control and three transgenic Epm2a−/−and Epm2b−/−mice of 7 months of age were lysed and mitochondrial and cytosolic superoxide dismutase activities were measured as described in the previous section. Although in this case we did not observe changes in protein levels of MnSOD or CuZnSOD (Fig. 7a), we detected a significant decrease in enzymatic activity for both proteins in laforin- and malin-deficient mice (Fig. 7b), suggesting that the superoxide detoxification process was affected in LD mice models. We next analyzed the levels of catalase and GPx (Fig. 8) and their corresponding enzymatic activities, but no apparent changes were obtained in these enzymes besides a small increase on catalase activity observed only in malin-deficient mice (Fig. 8b).
Analysis of mitochondrial and cytosolic superoxide dismutases in brain extracts from LD mice models. a Protein extracts (20 μg protein) obtained from brain homogenates of three independent wild type (WT); Epm2a−/−and Epm2b−/−mice were loaded in SDS-PAGE gels and analyzed by Western blotting using anti-MnSOD (left), anti-CuZnSOD (right) and anti-actin as loading control. Histograms show the quantification of various blots by densitometry with the values normalized to the corresponding actin levels and referred to the value of WT samples. b MnSOD (left) and CuZnSOD (right) activities were measured in extracts from brain homogenates of wild type (WT), Epm2a−/−and Epm2b−/−mice. Results represent the mean ± SD from three independent samples in each case (** p < 0.01)
Analysis of catalase and glutathione peroxidase in brain extracts from LD mice models. a Protein extracts (20 μg protein) obtained from brain homogenates of three independent wild type (WT), Epm2a−/−and Epm2b−/−mice were loaded in SDS-PAGE gels and analyzed by Western blotting using anti-catalase (left), anti-GPx1 (right) and anti-actin as a loading control. Histograms show the densitometry of the bands normalized to the actin levels and referred to the value of WT samples. b Catalase and GPx activities were analyzed as described in “Materials and Methods” section. Results represent the mean ± SD from three independent samples in each case (* p < 0.05)
Finally, we addressed the oxidative stress condition in these animals by measuring its effect on lipid peroxidation. Lipid hydroperoxides lead to a wide variety of by-products, one of which is malondialdehyde (MDA), considered as a reliable marker of chronic oxidative stress [32]. Data presented in Fig. 9 show high levels of MDA in plasma for Epm2b−/−mice, clearly indicating oxidative stress. On the other hand, only a tendency of increasing MDA levels was detected in brain tissue samples for both Epm2a−/−and Epm2b−/−mice models.
Analysis of MDA levels in plasma and brain from LD mice models. Plasma samples (a) and brain homogenates (b) from three independent wild type (WT), Epm2a−/−and Epm2b−/−mice were analyzed as described in “Materials and Methods” section, and MDA levels were calculated normalizing to the total protein amount in the samples. Results are expressed as mean ± SD from three independent samples in each case (* p < 0.05)
2D-DIGE Analysis of Malin-Deficient Mice
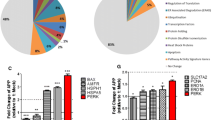
To further investigate the effects of the alterations in antioxidant response elements observed in animal models of LD, we chose a proteomic approach consisting in a two-dimensional differential in-gel electrophoresis analysis (2D-DIGE) using brain tissue homogenates. Animals of 7 months of age were used and samples from Epm2b−/−and control mice (three mice per group) were processed as described in the “Materials and Methods” section. Spots with at least ±1.5-fold differential change were selected for analysis and, under these conditions, only three spots were significantly changed (Fig. 10a). The first one (with a 2.02-fold increase in Epm2b−/−samples) was identified as voltage-dependent anion-selective channel protein 2 (VDAC2), a mitochondrial transporter implicated in diverse epileptogenic and neurodegenerative processes [33–35]. The two other spots (one with a 5.53-fold increase and the second with a 1.70-fold decrease in Epm2b−/−samples) corresponded to two different isoelectric locations of the same protein, peroxiredoxin-6 (Prdx6). Prdx6 is a bifunctional enzyme with peroxidase and phospholipase activities involved in the detoxification of hydrogen peroxides and lipids [36].
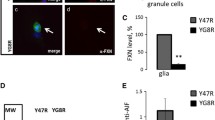
Proteomic analysis of brain extracts from Epm2b−/−mice. a Summary of 2D-DIGE results. Graphical view of the relative expression of each spot in samples from control or Epm2b−/−mice, as measured by the DeCyder differential analysis software. The correspondence between the number of each spot, the average intensity ratio of Epm2b−/−to control mice (Av. Ratio), the statistical t test values and their identity, as determined after mass spectrometry, are summarized in the right panel (see “Materials and Methods” section for details). b Analysis of Prdx6 electrophoretic mobility in Epm2b−/−mice brain samples. Total brain homogenates (10 μg protein) from mice of 6 months of age were subjected to conventional 2D electrophoresis and analyzed by Western blotting using anti-Prdx6 antibody. The calculated pI for each spot is indicated
Since the change in the levels of modified forms of Prdx6 were more important than those related to VDAC2, we sought to confirm the observations derived from the 2D-DIGE analysis by Western blotting using anti-Prdx6 antibody, in brain homogenates from three different control and Epm2b−/−mice, of 6 months of age. Although no clear differences in the total levels of Prdx6 were found (not shown), we observed a change in electrophoretic mobility when the samples were subjected to two-dimensional electrophoresis. In contrast to the brain homogenates from control mice, in which only one Prdx6 spot with a pI of about 6.14 was evident, an additional spot with a pI of approximately 5.75 was found in the samples from Epm2b−/−mice (Fig. 10b). These data are in agreement with other results indicating the presence of similar post-translational modifications of Prdx6 due to oxidative stress conditions [37–39]. In addition, this observation confirms the data obtained in the 2D-DIGE analysis and supports the existence of an oxidative stress in brain tissue of Epm2b−/−mice.
In all, the results obtained for the oxidative stress parameters analyzed in LD mice models are consistent with the previous study of LD fibroblasts and corroborates that the major antioxidant response elements are altered as a consequence of laforin or malin depletion.
Discussion
The ethiopathology of Lafora disease has remained largely elusive, in part due to the high variety of its symptoms. The initial hypothesis that an abnormal glycogen metabolism, due to genetic alterations of the genes encoding the phosphatase laforin or the E3 ubiquitin ligase malin, has been enriched by novel data pointing out to an accumulative process in which an ER stress caused by impairment in the unfolded protein response, as well as alterations in protein degradation mechanisms, could be also relevant [10–12]. In the present work, we provide evidence that the antioxidant defense systems are severely and specifically affected in both cell and animal models of LD, probably as a consequence of the previously described impairments.
The brain is considered a highly sensitive tissue to oxidative damage because it contains a large amount of oxidizable fatty acids and its antioxidant defenses are scarce [40]. Oxidative stress has been previously shown in other mice models of myoclonus epilepsy [41], hence it is regarded as a possible event contributing to the pathogenesis of epilepsy [42]. Interestingly, high levels of 8-hydroxydeoxyguanosine, one of the by-products of DNA oxidation, have been found in brain autopsies from LD patients with mutations in EPM2B [43]. Although mitochondrial dysfunction and oxidative stress-related mechanisms remain obscure in epileptogenic syndromes, recent hypotheses have established these mechanisms as contributing factors involved in epileptogenesis and seizures [21], probably by altering homeostasis of intracellular calcium, affecting neuronal excitability and synaptic transmission [44] and contributing to an increased susceptibility to epileptic seizures [45]. In fact, it has been shown in animal models in which epilepsy is induced that mitochondrial electron transport and ATP synthesis is seriously compromised [46]. Our results obtained in human fibroblasts from LD patients, in which mitochondrial dysfunction was suspected on the basis of an altered mitochondrial morphology, a decreased membrane potential and a low production of ATP, fit also with these observations. Since mitochondrial dysfunction generates high superoxide production and ROS accumulation, this could be the cause of the oxidative stress observed in these cells. Following this line of evidence, we found in laforin- and malin-deficient fibroblasts an alteration in important antioxidant enzymes involved in the detoxification of ROS: the superoxide dismutase isoforms MnSOD, CuZnSOD and catalase. The levels of hydrogen peroxide in cells can also be detoxified by glutathione peroxidases, but we did not detect any significant changes either in protein amount or in enzymatic activity of these proteins. Moreover, some of these alterations were only observed, or were clearly enhanced, after growing cells under a nutrient stress condition that triggers proteolytic processes, suggesting a relationship between the increase in oxidative stress and the impairment of protein clearance processes previously defined for these experimental models. In a recent work, we described in laforin-deficient fibroblasts a deficiency in thioredoxin 1 (Trx1), one of the most important antioxidant enzymes that participates in the reduction of oxidized cysteines in proteins and also an alteration in its subcellular localization [16]. Those results also corroborate the present data. Defects in the specific antioxidant enzymes MnSOD and CuZnSOD were also found when analyzing brain tissue samples from transgenic mice deficient in laforin or malin: we observed decreased MnSOD and CuZnSOD activities in brain tissue of LD mice models. In addition, levels of MDA (a marker of lipid peroxidation) were increased in Epm2b−/−plasma samples, suggesting high chronic oxidative stress in this particular mouse model. These results further suggest that oxidative stress plays a role in LD and corroborates the pattern observed in LD fibroblasts.
Since antioxidant defense systems are composed of a heterogeneous set of proteins and its regulation varies significantly between cell types and tissues, we performed a high-throughput 2D-DIGE proteomic analysis to gain a more general view of the proteomic alterations in malin-deficient mice. This approach revealed accumulation of a modified form of Prdx6, a bifunctional enzyme with peroxidase and phospholipase activities involved in the cellular detoxification of hydrogen peroxide [36]. Increase in Prdx6 expression has been documented in mice models of Huntington’s disease and, more recently, Prdx6 overexpression has been shown to accelerate the development of Alzheimer’s disease in mice [47, 48]. In malin-deficient mice, we found no increase in the total amount of Prdx6 but a difference in the relative distribution of its post-translationally modified forms. Similar modifications of Prdx6 have been described in the literature being due to oxidative stress-induced alterations [37–39]. Therefore, our results suggest the presence of oxidative stress conditions in the brain samples of Epm2b−/−mice. Proteomic analysis also revealed an increase in VDAC2, an outer mitochondrial membrane channel associated to different kinds of epilepsy [34, 33]. This is consistent with an alteration in mitochondrial function and in oxidative stress, two processes that are tightly interrelated. As Epm2b−/−mice have impaired autophagy [14], our data support the hypothesis that a defective mitophagy in these animals may lead to the accumulation of damaged and dysfunctional mitochondria and a corresponding increase in intracellular ROS levels, as described in other pathological conditions [49].
In summary, we propose that increase of ROS production due to mitochondrial dysfunction, together with a partially decreased antioxidant defense, may be important events in the progression of the pathophysiology of LD in humans. Our work clearly shows an implication of various components of the enzymatic antioxidant response in LD, which must be taken into account to build up a complete landscape of the pathophysiology of LD. Since there is evidence that antioxidant therapy may reduce lesions induced by oxidative stress and mitochondrial dysfunction, these results also open the possibility for such treatments.
Abbreviations
- CCCP :
-
carbonyl cyanide m-chlorophenylhydrazone
- ER :
-
endoplasmic reticulum
- LBs :
-
Lafora bodies
- LD :
-
Lafora disease
- 2D-DIGE :
-
two-dimensional differential in gel electrophoresis
- Prdx6 :
-
peroxiredoxin-6
- ROS :
-
reactive oxygen species
- VDAC2 :
-
voltage-dependent anion-selective channel protein 2
References
Lafora GR, Glueck BZ (1911) Beitrag zur Histopathologie der myoklonischen Epilepsie. Ges Neurol Psychiat 6:1–14
Lohi H, Ianzano L, Zhao XC, Chan EM, Turnbull J, Scherer SW, Ackerley CA, Minassian BA (2005) Novel glycogen synthase kinase 3 and ubiquitination pathways in progressive myoclonus epilepsy. Hum Mol Genet 14:2727–2736. doi:10.1093/hmg/ddi306
Rubio-Villena C, García-Gimeno MA, Sanz P (2013) Glycogenic activity of R6, a protein phosphatase 1 regulatory subunit, is modulated by the laforin–malin complex. Int J Biochem Cell Biol 45:1479–1488. doi:10.1016/j.biocel.2013.04.019
Worby CA, Gentry MS, Dixon JE (2008) Malin decreases glycogen accumulation by promoting the degradation of protein targeting to glycogen (PTG). J Biol Chem 283:4069–4076. doi:10.1074/jbc.M708712200
Solaz-Fuster MC, Gimeno-Alcaniz JV, Ros S, Fernandez-Sanchez ME, Garcia-Fojeda B, Criado Garcia O, Vilchez D, Dominguez J, Garcia-Rocha M, Sanchez-Piris M, Aguado C, Knecht E, Serratosa J, Guinovart JJ, Sanz P, Rodriguez de Cordoba S (2008) Regulation of glycogen synthesis by the laforin–malin complex is modulated by the AMP-activated protein kinase pathway. Hum Mol Genet 17:667–678. doi:10.1093/hmg/ddm339
Wang J, Stuckey JA, Wishart MJ, Dixon JE (2002) A unique carbohydrate binding domain targets the Lafora disease phosphatase to glycogen. J Biol Chem 277:2377–2380. doi:10.1074/jbc.C100686200
Worby CA, Gentry MS, Dixon JE (2006) Laforin, a dual specificity phosphatase that dephosphorylates complex carbohydrates. J Biol Chem 281:30412–30418. doi:10.1074/jbc.M606117200
Rao SN, Maity R, Sharma J, Dey P, Shankar SK, Satishchandra P, Jana NR (2010) Sequestration of chaperones and proteasome into Lafora bodies and proteasomal dysfunction induced by Lafora disease-associated mutations of malin. Hum Mol Genet 19:4726–4734. doi:10.1093/hmg/ddq407
Garyali P, Siwach P, Singh PK, Puri R, Mittal S, Sengupta S, Parihar R, Ganesh S (2009) The malin–laforin complex suppresses the cellular toxicity of misfolded proteins by promoting their degradation through the ubiquitin–proteasome system. Hum Mol Genet 18:688–700. doi:10.1093/hmg/ddn398
Liu Y, Wang Y, Wu C, Zheng P (2009) Deletions and missense mutations of EPM2A exacerbate unfolded protein response and apoptosis of neuronal cells induced by endoplasm reticulum stress. Hum Mol Genet 18:2622–2631. doi:10.1093/hmg/ddp196
Vernia S, Rubio T, Heredia M, Rodríguez de Córdoba S, Sanz P (2009) Increased endoplasmic reticulum stress and decreased proteasomal function in Lafora disease models lacking the phosphatase laforin. PLoS ONE 4:e5907. doi:10.1371/journal.pone.0005907
Zeng L, Wang Y, Baba O, Zheng P, Liu Y (2012) Laforin is required for the functional activation of malin in endoplasmic reticulum stress resistance in neuronal cells. FEBS J 279:2467–2478. doi:10.1111/j.1742-4658.2012.08627.x
Rao SN, Sharma J, Maity R, Jana NR (2010) Co-chaperone CHIP stabilizes aggregate-prone malin, a ubiquitin ligase mutated in Lafora disease. J Biol Chem 285:1404–1413. doi:10.1074/jbc.M109.006312
Criado O, Aguado C, Gayarre J, Duran-Trio L, Garcia-Cabrero AM, Vernia S, San Millan B, Heredia M, Romá-Mateo C, Mouron S, Juana-Lopez L, Dominguez M, Navarro C, Serratosa JM, Sanchez M, Sanz P, Bovolenta P, Knecht E, Rodriguez de Cordoba S (2012) Lafora bodies and neurological defects in malin-deficient mice correlate with impaired autophagy. Hum Mol Genet 21:1521–1533. doi:10.1093/hmg/ddr590
Aguado C, Sarkar S, Korolchuk VI, Criado O, Vernia S, Boya P, Sanz P, Rodríguez de Córdoba S, Knecht E, Rubinsztein DC (2010) Laforin, the most common protein mutated in Lafora disease, regulates autophagy. Hum Mol Genet 19:2867–2876. doi:10.1093/hmg/ddq190
García-Giménez JL, Seco-Cervera M, Aguado C, Romá-Mateo C, Dasi F, Priego S, Markovic J, Knecht E, Sanz P, Pallardo FV (2013) Lafora disease fibroblasts exemplify the molecular interdependence between thioredoxin 1 and the proteasome in mammalian cells. Free Radic Biol Med 65:347–359. doi:10.1016/j.freeradbiomed.2013.07.001
Dutta D, Xu J, Kim JS, Dunn WA Jr, Leeuwenburgh C (2013) Upregulated autophagy protects cardiomyocytes from oxidative stress-induced toxicity. Autophagy 9:328–344. doi:10.4161/auto.22971
Kiffin R, Bandyopadhyay U, Cuervo AM (2006) Oxidative stress and autophagy. Antioxid Redox Signal 8:152–162. doi:10.1089/ars.2006.8.152
Lee J, Giordano S, Zhang J (2012) Autophagy, mitochondria and oxidative stress: cross-talk and redox signalling. Biochem J 441:523–540. doi:10.1042/BJ20111451
Navarro-Yepes J, Burns M, Anandhan A, Khalimonchuk O, Del Razo LM, Quintanilla-Vega B, Pappa A, Panayiotidis MI, Franco R (2014) Oxidative stress, redox signaling, and autophagy: cell death versus survival. Antioxid Redox Signal. doi:10.1089/ars.2014.5837
Aguiar CC, Almeida AB, Araujo PV, de Abreu RN, Chaves EM, do Vale OC, Macedo DS, Woods DJ, Fonteles MM, Vasconcelos SM (2012) Oxidative stress and epilepsy: literature review. Oxid Med Cell Longev 2012:795259. doi:10.1155/2012/795259
Garcia-Gimenez JL, Gimeno A, Gonzalez-Cabo P, Dasi F, Bolinches-Amoros A, Molla B, Palau F, Pallardo FV (2011) Differential expression of PGC-1alpha and metabolic sensors suggest age-dependent induction of mitochondrial biogenesis in Friedreich ataxia fibroblasts. PLoS ONE 6:e20666. doi:10.1371/journal.pone.0020666
Viña J, Lloret A, Valles SL, Borras C, Badia MC, Pallardo FV, Sastre J, Alonso MD (2007) Mitochondrial oxidant signalling in Alzheimer’s disease. J Alzheimers Dis 11:175–181
Esteve JM, Armengod ME, Knecht E (2010) BRCA1 negatively regulates formation of autophagic vacuoles in MCF-7 breast cancer cells. Exp Cell Res 316:2618–2629. doi:10.1016/j.yexcr.2010.06.019
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) method. Methods 25:402–408. doi:10.1006/meth.2001.1262
Ganesh S, Delgado-Escueta AV, Sakamoto T, Avila MR, Machado-Salas J, Hoshii Y, Akagi T, Gomi H, Suzuki T, Amano K, Agarwala KL, Hasegawa Y, Bai DS, Ishihara T, Hashikawa T, Itohara S, Cornford EM, Niki H, Yamakawa K (2002) Targeted disruption of the Epm2a gene causes formation of Lafora inclusion bodies, neurodegeneration, ataxia, myoclonus epilepsy and impaired behavioral response in mice. Hum Mol Genet 11:1251–1262. doi:10.1093/hmg/11.11.1263
Colinge J, Masselot A, Giron M, Dessingy T, Magnin J (2003) OLAV: towards high-throughput tandem mass spectrometry data identification. Proteomics 3:1454–1463. doi:10.1002/pmic.200300485
Fransson A, Ruusala A, Aspenstrom P (2003) Atypical Rho GTPases have roles in mitochondrial homeostasis and apoptosis. J Biol Chem 278:6495–6502. doi:10.1074/jbc.M208609200
Kato T, Kapczinski F, Berk M (2010) Mitochondrial dysfunction and oxidative stress. In: Bipolar disorder. Wiley, p 244–254. doi:10.1002/9780470661277.ch18
Lin MT, Beal MF (2006) Mitochondrial dysfunction and oxidative stress in neurodegenerative diseases. Nature 443:787–795. doi:10.1038/nature05292
Du G, Mouithys-Mickalad A, Sluse FE (1998) Generation of superoxide anion by mitochondria and impairment of their functions during anoxia and reoxygenation in vitro. Free Radical Biol Med 25:1066–1074. doi:10.1016/S0891-5849(98)00148-8
Ohkawa H, Ohishi N, Yagi K (1979) Assay for lipid peroxides in animal tissues by thiobarbituric acid reaction. Anal Biochem 95:351–358
Liu XY, Yang JL, Chen LJ, Zhang Y, Yang ML, Wu YY, Li FQ, Tang MH, Liang SF, Wei YQ (2008) Comparative proteomics and correlated signaling network of rat hippocampus in the pilocarpine model of temporal lobe epilepsy. Proteomics 8:582–603. doi:10.1002/pmic.200700514
Jiang W, Du B, Chi Z, Ma L, Wang S, Zhang X, Wu W, Wang X, Xu G, Guo C (2007) Preliminary explorations of the role of mitochondrial proteins in refractory epilepsy: some findings from comparative proteomics. J Neurosci Res 85:3160–3170. doi:10.1002/jnr.21384
Cheng EH, Sheiko TV, Fisher JK, Craigen WJ, Korsmeyer SJ (2003) VDAC2 inhibits BAK activation and mitochondrial apoptosis. Science 301:513–517. doi:10.1126/science.1083995
Fisher AB (2011) Peroxiredoxin 6: a bifunctional enzyme with glutathione peroxidase and phospholipase A(2) activities. Antioxid Redox Signal 15:831–844. doi:10.1089/ars.2010.3412
Baraibar MA, Hyzewicz J, Rogowska-Wrzesinska A, Ladouce R, Roepstorff P, Mouly V, Friguet B (2011) Oxidative stress-induced proteome alterations target different cellular pathways in human myoblasts. Free Radic Biol Med 51:1522–1532. doi:10.1016/j.freeradbiomed.2011.06.032
Chevallet M, Wagner E, Luche S, van Dorsselaer A, Leize-Wagner E, Rabilloud T (2003) Regeneration of peroxiredoxins during recovery after oxidative stress: only some overoxidized peroxiredoxins can be reduced during recovery after oxidative stress. J Biol Chem 278:37146–37153. doi:10.1074/jbc.M305161200
Jeong J, Kim Y, Kyung Seong J, Lee KJ (2012) Comprehensive identification of novel post-translational modifications in cellular peroxiredoxin 6. Proteomics 12:1452–1462. doi:10.1002/pmic.201100558
Henchcliffe C, Beal MF (2008) Mitochondrial biology and oxidative stress in Parkinson disease pathogenesis. Nat Clin Pract Neurol 4:600–609. doi:10.1038/ncpneuro0924
Lehtinen MK, Tegelberg S, Schipper H, Su H, Zukor H, Manninen O, Kopra O, Joensuu T, Hakala P, Bonni A, Lehesjoki AE (2009) Cystatin B deficiency sensitizes neurons to oxidative stress in progressive myoclonus epilepsy, EPM1. J Neurosci 29:5910–5915. doi:10.1523/JNEUROSCI.0682-09.2009
Chang SJ, Yu BC (2010) Mitochondrial matters of the brain: mitochondrial dysfunction and oxidative status in epilepsy. J Bioenerg Biomembr 42:457–459. doi:10.1007/s10863-010-9317-4
Singh S, Sethi I, Francheschetti S, Riggio C, Avanzini G, Yamakawa K, Delgado-Escueta AV, Ganesh S (2006) Novel NHLRC1 mutations and genotype-phenotype correlations in patients with Lafora’s progressive myoclonic epilepsy. J Med Genet 43:e48. doi:10.1136/jmg.2005.039479
Patel M (2004) Mitochondrial dysfunction and oxidative stress: cause and consequence of epileptic seizures. Free Radic Biol Med 37:1951–1962. doi:10.1016/j.freeradbiomed.2004.08.021
Kudin AP, Zsurka G, Elger CE, Kunz WS (2009) Mitochondrial involvement in temporal lobe epilepsy. Exp Neurol 218:326–332. doi:10.1016/j.expneurol.2009.02.014
Liang LP, Ho YS, Patel M (2000) Mitochondrial superoxide production in kainate-induced hippocampal damage. Neuroscience 101:563–570
Yata K, Oikawa S, Sasaki R, Shindo A, Yang R, Murata M, Kanamaru K, Tomimoto H (2011) Astrocytic neuroprotection through induction of cytoprotective molecules; a proteomic analysis of mutant P301S tau-transgenic mouse. Brain Res 1410:12–23. doi:10.1016/j.brainres.2011.06.064
Yun HM, Jin P, Han JY, Lee MS, Han SB, Oh KW, Hong SH, Jung EY, Hong JT (2013) Acceleration of the development of Alzheimer’s disease in amyloid beta-infused peroxiredoxin 6 overexpression transgenic mice. Mol Neurobiol 48:941–951. doi:10.1007/s12035-013-8479-6
Wu JJ, Quijano C, Chen E, Liu H, Cao L, Fergusson MM, Rovira II, Gutkind S, Daniels MP, Komatsu M, Finkel T (2009) Mitochondrial dysfunction and oxidative stress mediate the physiological impairment induced by the disruption of autophagy. Aging 1:425–437
Acknowledgments
This work was supported by grants from the Spanish Ministry of Education and Science (SAF2011-27442, BFU2011-22630), Fundació La Marató de TV3 (ref. 100130), EU-Funded FRAILOMIC-HEALTH.2012.2.1.1, TREAT-CMT IRDiRC consortium Research fellowship, Generalitat Valenciana (Prometeo 2009/051, Prometeo 2012/061) and an ACCI2012 action from CIBERER, an initiative of the Instituto de Salud Carlos III.
Author information
Authors and Affiliations
Corresponding author
Additional information
Carlos Romá-Mateo Carmen Aguado and José Luis García-Giménez contributed equally to this work.
Rights and permissions
About this article
Cite this article
Romá-Mateo, C., Aguado, C., García-Giménez, J.L. et al. Increased Oxidative Stress and Impaired Antioxidant Response in Lafora Disease. Mol Neurobiol 51, 932–946 (2015). https://doi.org/10.1007/s12035-014-8747-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-014-8747-0