Abstract
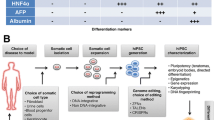
The generation of human induced pluripotent stem cells (hiPSCs) from an individual patient provides a unique tool for disease modeling, drug discovery, and cell replacement therapies. Patient-specific pluripotent stem cells can be expanded in vitro and are thus suitable for genetic manipulations. To date, several genetic liver disorders have been modeled using patient-specific hiPSCs. Here, we present the generation of corrected hepatocyte-like cells (HLCs) from hiPSCs of a familial hypercholesterolemia (FH) patient with a homozygous mutation in the low-density lipoprotein receptor (LDLR) gene. We generated hiPSCs from a patient with FH with the mutated gene encoding a truncated non-functional receptor. In order to deliver normal LDLR to the defective cells, we used a plasmid vector carrying the normal receptor ORF to genetically transform the hiPSCs. The transformed cells were expanded and directed toward HLCs. Undifferentiated defective hiPSCs and HLCs differentiated from the defective hiPSCs did not have the ability to uptake labeled low-density lipoprotein (LDL) particles. The differentiated transformed hiPSCs showed LDL-uptake ability and the correction of disease phenotype as well as expressions of hepatocyte-specific markers. The functionality of differentiated cells was also confirmed by indo-cyanine green (ICG) uptake assay, PAS staining, inducible cyp450 activity, and oil red staining. These data suggest that hiPSC technology can be used for generation of disease-corrected, patient-specific HLCs with potential value for disease modeling and drug discovery as well as cell therapy applications in future.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Familial hypercholesterolemia (FH) is primarily caused by inherited mutations in the low-density lipoprotein receptor (LDLR) encoding gene and is characterized by elevated blood cholesterol level. The deposition of cholesterol can result in xanthoma in the tendons of feet and hands, cutaneous planar, corneal arcus, and atheroma of the aortic root and valves, which can lead to fatal myocardial infarctions before the age of 30 years [1–3]. Currently, FH management primarily involves lifestyle modifications and drug (statins) treatments [3, 4], so having a disease model based on iPSC technology in vitro might facilitate the process of finding new treatments and studying the impacts of specific allelic variations on control of low-density lipoprotein cholesterol. Recently, patient-specific iPSC-derived hepatocytes are shown to be a valuable tool for studying the functional contribution of various genomic loci in regulating lipid metabolism and recapitulating the complex pathophysiology of FH in culture [5].
Orthotopic liver transplantation (OLT) in homozygous patients is the best way to correct the disease phenotype [6], but this aggressive procedure carries considerable risks. Therefore, there is a need for development of alternative treatments. An ideal treatment for FH requires the genetic correction of patients’ liver cells in order to restore normal phenotype. Successful ex vivo gene delivery experiments on rabbits and non-human primates [6–8] have demonstrated the feasibility of gene therapy for FH, which was subsequently performed in humans [9, 10]. In these first clinical experiments [9, 10], patients’ primary hepatocytes were manipulated in vitro with retroviral vectors that carried the normal LDLR encoding gene and then injected back into the same patients. These treatments resulted in a stable therapeutic decline in blood LDL/HDL ratio. However, the invasiveness of the surgical intervention necessary for obtaining patients’ hepatocytes and difficulties in culturing and maintenance of primary hepatocytes during the genetic manipulation procedure were the major drawbacks of this treatment protocol. In order to develop alternative therapies, we need to achieve a comprehensive insight about FH by in vitro evaluation of potential cell sources applicable for ex vivo genetic manipulation.
The generation of human induced pluripotent stem cells (hiPSCs) from dermal fibroblasts by epigenetic reprogramming [11] holds great promise for advancements in regenerative medicine and disease modeling. iPSCs can be generated from most mammalian somatic tissues [12] using different approaches [13] and can be differentiated into functional specialized cell lineages of all three embryonic germ layers. This wide differentiation potential provides fascinating possibilities and tools for drug discovery, developmental and pathogenesis studies, and investigation of possible treatment for inherited disorders (reviewed in [14]). To date, several diseases have been modeled [15–25] or corrected in vitro [26–29] using patients’ derived iPSCs. A number of disease-specific cell lines have been generated from patients with genetic liver disorders such as α1 anti-trypsin deficiency, glycogen storage disease types 1a and 1b, Crigler–Najjar syndrome, tyrosinemia type 1, progressive familial hereditary cholestasis, and FH [30, 31]. These liver disease-specific iPSC lines have been differentiated toward hepatocyte-like cells (HLCs) and some of them were used for disease modeling; however, in none of the above cases genetic manipulation was examined as a means for correction of the disease phenotype. For the first time in this study, we generated disease-corrected HLCs from manipulated hiPSCs that were derived from the dermal fibroblasts of a homozygous FH patient.
Materials and Methods
Generation of hiPSCs from Patient’s Fibroblasts
Dermal fibroblasts were harvested from a 14-year-old female patient with homozygous FH [32]. The ethical principles of the Helsinki Declaration were followed and the human dermal biopsy sampling was approved by the local Royan Institutional Review Board and Ethical Committee. There were nine independent patient-specific hiPSC clones (FH-hiPSCs) generated from the patient’s dermal fibroblasts by transduction of retroviral vectors carrying OCT4, SOX2, c-MYC, and KLF4, as described earlier in serum and feeder-free culture conditions [33].
Karyotype Analysis and Bisulfite Sequencing
Karyotype analysis and bisulfite sequencing were performed as described previously [34]. The promoter regions of human OCT4 and NANOG gene were amplified with PCR and the products were subcloned into the InsTAclone PCR Cloning Kit (Fermentas). The cloned fragments were sequenced with M13 universal primers and analyzed with BIQ Analyzer software.
RT-PCR and Quantitative RT-PCR Analysis
Total RNA was isolated using the RNeasy Kit (Fermentas). Two micrograms of total RNA was used for reverse transcription reaction with the RevertAid First Strand cDNA Synthesis Kit (Fermentas) and random hexamer primer (Fermentas), according to the manufacturer’s instructions. The first strand cDNA was used for PCR reaction and the products were examined by electrophoresis on 2–3 % agarose gel in TAE buffer.
Quantitative real-time PCR (qRT-PCR) reactions were set up in experimental duplicate for the three biological repeats with the Power SYBR Green Master Mix (Applied Biosystems) and analyzed with a 7500 RT-PCR system (Applied Biosystems). Expression values were normalized to the average expression of housekeeping gene [ActB by the comparative CT method (2−ΔΔct)]. The primer sequences have been previously presented [31, 33].
Genetic Manipulation of FH-hiPSCs
The pENTR/D-TOPO and pLenti6/UbC/V5-DEST vectors (Invitrogen, K2400-20 and V499-10) were used for cloning LDLR cDNA (Variant number 1, accession number: NM_000527.4). The pLenti6/UbC/V5-DEST derived construct that carried the normal ORF was subsequently used for lipofection of FH-hiPSCs with Fugene6 reagent (Roche, 05061377001). Transfection was performed according to the manufacturer’s instructions, 24 h after plating 200,000 single cells per well in a 6-well plate.
Direct Differentiation of hiPSCs into HLCs
Human induced pluripotent stem cells differentiation toward the hepatic lineage was performed using Basma et al. [35] protocol with some modifications. Briefly, embryoid bodies (EBs) were generated by plating collagenase/dispase-treated cells at a density of 1–5 × 104 cells/cm2 on bacterial petri dishes for 48 h in DMEM/F12 supplemented with 20 % KOSR, 1 mM nonessential amino acids, and 2 mM l-glutamine. The EBs were then plated on Matrigel-coated plates in DMEM/F12 supplemented with activin A (100 ng/mL, R&D, 338-AC) for 3 days to induce a definitive endoderm lineage. The concentration of KOSR was started from 0 % for the first 24 h, changed to 0.2 % for the second 24 h and 2.0 % for the final 24 h. Cells were then grown for 8 days in DMEM/F12 that contained 2.0 % KOSR, 1 mM nonessential amino acids, 2 mM l-glutamine, 1 % dimethyl sulfoxide (Sigma-Aldrich), and 100 ng/mL HGF (R&D Systems, 294-HG). The culture was continued for an additional 5 days in hepatocyte culture medium (HCM) (Lonza, Inc., CC-1399) that consisted of 2 % KOSR, 1 mM nonessential amino acids, 2 mM l-glutamine, and 0.1 μM dexamethasone (Sigma-Aldrich, D-2915).
Alkaline Phosphatase Staining, Immunofluorescence Staining, and Flow Cytometry
Alkaline phosphatase (ALP) staining was conducted based on the manufacturer’s recommendations (Sigma, 86R).
Immunofluorescence staining for pluripotency and hepatic markers in addition to flow cytometry analysis for ALB and asialoglycoprotein receptor 1 (ASGPR1) were performed as described previously [31, 34].
Low-Density Lipoprotein (LDL) Uptake Assay
LDL DylighTM 549 kit was used (Cayman Chemical, 10011125) and the assay was performed according to the manufacturer’s instructions. Cells were visualized using a fluorescence microscope (IX71, Olympus, Japan).
Cytochrome P450 Activity and Inducibility
Cytochrome P450-dependent pentoxyresorufin O-dealkylase activity (PROD) was evaluated using pentoxyresorufin substrates. Pentoxyresorufin is O-dealkylated by CYP and changes a nonfluorescent compound into resorufin, a fluorescent compound [36]. To evaluate the inducibility of cytochrome P450, the differentiated cells were exposed to sodium phenobarbital for three days and subsequently washed. Then an incubation mixture containing 7-pentoxyresorufin substrate (Sigma-Aldrich) and dicumarol (Sigma-Aldrich) in Hank’s balanced salt solution (HBSS) was added and plates were incubated at 37 °C in a 5 % CO2 incubator for 30 min. Cell supernatant medium was collected after 30 min for three biological repeats and examined for the quantity of resorufin produced using a fluorospectrophotometer (Cary-Eclipse, Varian) at 530 nm excitation/590 nm emission against resorufin standards. To give a comparative view, the results were normalized to the number of cells.
Periodic Acid-Schiff (PAS) Staining
Glycogen storage of hiPSC-derived HLCs was evaluated by PAS staining at day 23. The cells were fixed with 4 % paraformaldehyde, oxidized in 1 % periodic acid for 5 min, then washed and treated with Schiff’s reagent for 15 min, with subsequent color development in dH2O for 5–10 min and assessed under light microscope (BX71, Olympus).
ICG Uptake and Release
The day 23 differentiated cells were incubated with ICG (Cardiogreen, Sigma-Aldrich) in basal medium for 1 h at 37 °C. Uptake of ICG was detected by light microscopy (IX51, Olympus). ICG elimination from the positive cells was verified after 6 h.
Secretion Analysis
On day 23, conditioned media with fully differentiated hiPSCs were collected and stored at −20 °C until assayed. The media were assayed for alpha-fetoprotein (AFP) secretion using a chemiluminescence immunoassay kit (Pishtaz-Teb); for ALB secretion with an ALB ELISA kit (Bethyl); and for urea secretion with a colorimetric assay kit (Pars Azmun) according to the manufacturers’ instructions. Secretion was normalized to the total number of cells.
Oil Red Staining
The storage of lipid vesicles in cells was assessed by oil red staining. Differentiated cells were fixed with 4 % paraformaldehyde, and incubated for 1 h with oil red. The stained cells were washed and analyzed with an inverted light microscope (IX51, Olympus).
Statistical Analysis
mRNA expression levels and other experiments were measured for three independent biological repeats and expressed as the mean ± standard deviation (SD). Statistical analysis was performed using the Student’s t test for independent samples. The mean difference was considered significant at the p < 0.05 level.
Results
Generation and Characterization of FH-hiPSCs
Dermal fibroblasts were harvested from the skin biopsy of a 14-year-old homozygous FH female patient. The patient had tendon xanthoma, xanthelasma, and a total blood cholesterol level of 801 mg/dL. The LDLR encoding gene of the harvested cells was previously shown to have a single base pair insertion in position 2411 in exon 17 [32]. The abnormal phenotype is the result of this frame-shift in the LDLR coding sequence and the subsequent production of a truncated non-functional receptor. Next, these somatic cells were reprogrammed to iPSCs under feeder and serum-free conditions by the 4-factor approach as described before [33]. We obtained nine hiPSC colonies (FH-hiPSC3) out of around 200,000 plated human fibroblast cells. Of these, one hiPSC colony was picked and expanded for further characterization. The cell line had the normal karyotype (Fig. 1a) and was used for further analyses. The cell line exhibited the characteristic human embryonic stem cell (hESC) morphology (colonies of round compact cells with high nucleus to cytoplasm ratio) and strong ALP activity. FH-hiPSCs also expressed pluripotency markers including OCT4, SSEA4, TRA-1-60, and TRA-1-81 (Fig. 1a). The pluripotency of the FH-hiPSCs was further assessed by RT-PCR analysis of lineage specific markers after spontaneous differentiation (Fig. 1b). The methylation status of the OCT4 and NANOG promoters was also monitored with bisulfite sequencing, which revealed that these regions were highly unmethylated in hESCs and established hiPSCs, whereas the CpG dinucleotides of the regions were highly methylated in the parental HDFs (Fig. 1c). The qRT-PCR analysis, with primers specific for retroviral transcripts of OCT4, SOX2, c-MYC, and KLK4, indicated complete silencing of the four transgenes (data not shown). The line maintained a normal karyotype in a continuous culture for several months by weekly passaging with a split ratio of 1:3 to 1:6. Collectively, these results demonstrated that the hiPSC line established in this study was fully reprogrammed.
Generation of iPSCs from patient’s fibroblasts. a Normal karyotype, morphology, and expression of different pluripotency and surface markers of FH-hiPSC cell line are presented. The lines are characterized after at least ten passages. Nuclei were stained with DAPI (blue). b RT-PCR analyses of various differentiation markers for the three germ layers in spontaneously differentiated “D” and undifferentiated “U” cells. c Bisulfite genomic sequencing of the promoter regions of OCT4 and NANOG. Open and closed circles indicate unmethylated and methylated CpGs, respectively. Human ESCs (Royan H6) is the control (Color figure online)
Genetic Transformation of FH-hiPSCs
A normal human LDLR complete ORF was PCR amplified from the normal HDFs’ cDNA and cloned into the pLenti6/UbC/V5-DEST vector under the control of the ubiquitin (UbC) constitutive promoter. DNA sequencing revealed the proper cloning of the LDLR ORF in the vector, without any mutations. Next, the plasmid was used to transform feeder-free cultured FH-hiPSCs. Blasticidin (10 μg/mL) was added into medium from day 3. This dose of Blasticidin was observed to kill all non-transfected FH-hiPSCs in 3 days. After 3 weeks, a single genetic-transformed FH-hiPSC (GT-FH-hiPSC) colony was generated from the starting population of around 1,200,000 iPSCs. The resistant colony was picked up, expanded and differentiated toward hepatocytes according to the protocol described previously [35].
Generation of Disease-Free HLCs
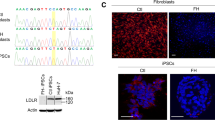
Morphological analysis of the cells revealed that during the differentiation process, the hiPSCs gradually formed polygonal and highly granulated HLCs with less density, and a lower nucleus to cytoplasm ratio (Fig. 2a). Functional correction of the GT-FH-HLCs was demonstrated by LDL-uptake assays (Fig. 2a). Immunofluorescence staining of LDLR in HLCs using a polyclonal antibody that binds both normal LDLR and the truncated non-functional receptor of the defective cells demonstrated the expression of this receptor in the cells (Fig. 2a). However, the defective FH-HLCs did not show the ability to uptake fluorescent LDL particles from the medium and this function was restored in GT-FH-HLCs. The morphology, receptor staining, and LDL-uptake of HLCs derived from hESCs (Royan H6) [37] were used as positive controls for the experiment. RT-PCR expression analysis of hiPSCs as well as FH-hiPSC- and GT-FH-hiPSC-derived HLCs (FH-HLCs and GT-FH-HLCs), with primers specific for the exogenous LDLR encoding mRNA, also revealed expression of exogenous receptor mRNA only in the GT cell lines (Fig. 2b). qRT-PCR expression analysis of the endogenous LDLR mRNA showed up-regulation of the receptor in FH-hiPSCs and its down-regulation in GT-FH-HLCs upon differentiation (Fig. 2c).
Functional correction in GT-FH-HLCs. a FH-HLCs and GT-FH-HLCs were shown to have polygonal and granulated morphological characteristics of mature hepatocytes. LDLR staining with a polyclonal antibody that binds both normal and mutated receptors showed the presence of LDLR on FH-HLCs and GT-FH-HLCs membrane. However, the LDLRs of the FH-HLCs were not functional and unable to uptake LDL particles. The LDL-uptake assay showed the successful gain of function upon genetic transformation of FH-hiPSCs and their differentiation. hESC (Royan H6)-derived HLCs were used as positive controls. Nuclei were stained with DAPI (blue). Scale bar 100 μm. b Exogenous LDLR gene was only expressed in genetic-transformed cells. c Endogenous LDLR gene was down-regulated in HLCs derived from genetic-transformed cells relative to those derived from defective cells (Color figure online)
To further assess the transition from hiPSCs to HLCs in direct differentiation, qRT-PCR analysis was performed on hiPSCs and hiPSC-HLCs for expression of hepatic lineage cells (AFP, ALB, HNF4α, and CYP3A4) and the undifferentiated hiPSCs marker (OCT4). The results showed that while OCT4 mRNA level decreased significantly upon hepatic induction; HLCs-specific mRNAs were increased (Fig. 3a). The expression differences between FH-HLCs and GT-FH-HLCs were not statistically significant (Fig. 3a). Immunofluorescence staining for ALB, AAT, and CYP1A1 verified hepatic-specific gene expression at the protein level (Fig. 3b). Flow cytometry was performed to determine the percentage of ALB and ASGPR1 expression, which is a definitive feature of hepatocytes. The percent of positive cells in the populations of FH-HLCs and GT-FH-HLCs were 22 and 25 % for ASGPR1 and 61 and 62 % for ALB, respectively (Fig. 3c).
mRNA and protein expression in differentiated cells. Directed differentiation of hiPSCs toward HLCs, and characterization of FH-HLCs and GT-FH-HLCs. a Gene expression analysis. The mRNA fold change of hepatic-specific genes (AFP, ALB, HNF4A, and CYP3A4) and a pluripotency gene (OCT4) have been examined for corrected and non-corrected hiPSC–HLCs relative to their undifferentiated stage. Expressions of endoderm and hepatocyte markers increased during the time span of differentiation whereas OCT4 expression decreased. b Protein expression analysis by immunofluorescence staining. HLCs were positive for hepatocyte-specific markers (ALB, AAT, and CYP1A) at the end of the differentiation period. The data proves efficient induction and maturation of FH-HLCS and GT-FH-HLCs. Nuclei were stained with DAPI (blue). Scale bar 100 μm. c Efficient differentiation was further confirmed by flow cytometry quantification of ALB and ASGPR1 positive cells (Color figure online)
To evaluate the other functionalities of FH-HLCs and GT-FH-HLCs, cells were examined for hepatic-specific functions in vitro. The cells’ ability to store lipids was demonstrated by oil red staining (Fig. 4a) and their glycogen storage ability was proved by PAS staining (Fig. 4b) and the presence of specific membrane transporters by ICG uptake (Fig. 4c). Hepatocytes-like cells had also high secretion activity and were able to secret ALB, AFP, and urea to the medium (Fig. 4d–f). The PROD test was performed to assess human hepatocyte-specific cytochrome P450 activity before and after induction with phenobarbital and the results showed an approximate 1.5-fold increased activity of CYP upon induction (p < 0.05, Student's t test, Fig. 4g). All of the above assays were performed for three biological repeats and the differences between FH-HLCs and GT-FH-HLCs were not statistically significant.
Hepatocyte-specific functional assays. Qualitative functions of the differentiated cells derived from both cell lines were examined. Cells were able to a store lipid (oil red staining) and b glycogen, as well as (c) uptake anionic ICG from culture media. Scale bar 100 μm. Quantitative functions of the differentiated cells from both cell lines were examined. d ALB, e AFP, and f urea secretion during the time span of differentiation indicate the functionality of the cells in vitro. g Cyp450 activity and its inducibility upon phenobarbital administration. PROD activity increased significantly in response to phenobarbital induction (p < 0.05, Student's t test). The differences between FH-HLCs and GT-FH-HLCs were not statistically significant (Color figure online)
Discussion
Hepatocytes are responsible for about 70 % of body LDL-uptake. Genetic defects that interfere with this hepatic function will lead to the lethal disorder of FH. To date, OLT has been the only available option for curing this severe genetic liver disorder. Due to the major limitations of OLT (e.g., the availability of suitable donors and the risk of an immune response), the search for a replacement therapy has been an attractive issue for many scientists over the past decade [9, 10, 38].
In this study, we represent the differentiation of hiPSCs into HLCs expressed various liver markers and liver-specific functions, such as albumin secretion, urea production, ICG glycogen storage, and inducible cytochrome P450 activity. Considering the gene expression and functional analysis, these cells showed characteristics of immature fetal hepatocytes as reported previously [39, 40].
Further, we have previously showed that patient-specific hiPSCs can be generated in serum and feeder-free culture conditions from patients suffering from various genetic liver disorders [31]. Several recent papers have reported that regardless of the origin of the reprogrammed somatic cell, the generated hiPSCs can be directed to hepatocytes in the same manner as hESCs [30, 31, 41–44]. These cell lines can serve as tools for modeling disorders, drug screening, and therapeutic applications [45–47].
We observed that the hiPSCs–HLCs had hepatic-specific functions in vitro; however, due to a mutation in the LDL receptor gene which is responsible for most of the liver LDL-uptake, differentiated cells from the defective patient-specific iPSCs were not able to uptake LDL. On the other hand, the genetically transformed hiPSC–HLCs were shown to uptake LDL from the media, which has proven successful return of the lost function. Exogenous LDLR mRNA was expressed under control of a housekeeping promoter (UbC) in genetically transformed cells. This promoter expresses the receptor in physiological amounts and the presence of LDL in differentiating media results in the in vitro incorporation of these particles in functionally corrected cells. This incorporation activates the negative feed-back loop [48, 49] in GT-FH-HLCs and results in a decrease in endogenous LDLR mRNA.
These disease-corrected HLCs from patient-specific hiPSCs might provide an accessible therapeutic source (Fig. 5), as the principle for therapeutic application of iPSC-HLCs has been proved by treating diseases in animal models [50]. It has been reported that iPSC-HLCs have both functional and proliferative potential for liver regeneration after transplantation in an acute liver failure model or after partial hepatectomy in mice with fumarylacetoacetate hydrolase deficiency [51] as well as their potential as a source for disease modeling and evaluation of the pathophysiology of the metabolic diseases such as FH [5]. Therefore, the next step to evaluate the therapeutic potential of GT-hiPSC–HLCs is to examine them in vivo in animal models of FH, such as LDLR-knockout mice [52] and Watanabe heritable hyperlipidemic (WHHL) rabbits [53].
Taken together, our results suggest the possible application of iPS technology and gene correction methods for personalized medicine and bring new hopes for treatment of metabolic liver patients. However, before clinical application of these genetic manipulated hiPSC–HLCs, more detailed studies are needed to address safety issues concerning iPSCs production [54], immunologic response [55], tumor formation [56], and genetic manipulations. In addition, further studies are required to show the functionality of cell in vivo. There is also a need for development of more efficient, chemically defined differentiation protocols. Recently, it has been shown that functional HLCs can also be generated directly from mouse fibroblasts [57]. This strategy can be an alternative way for producing disease-specific HLCs applicable for genetic modifications and gene therapy. Further studies, however, are needed to examine this approach on human fibroblasts. Until then, FH-HLCs and GT-FH-HLCs can serve as tools for the basic investigation of the LDL-uptake pathway and in vitro disease models for studying FH and its potential treatments.
References
Yamamoto, A., Kamiya, T., Yamamura, T., Yokoyama, S., Horiguchi, Y., Funahashi, T., et al. (1989). Clinical features of familial hypercholesterolemia. Arteriosclerosis, 9, I66–I74.
Kwiterovich, P. O., Jr, Bachorik, P. S., Smith, H. H., McKusick, V. A., Connor, W. E., Teng, B., et al. (1981). Hyperapobetalipoproteinaemia in two families with xanthomas and phytosterolaemia. Lancet, 1, 466–469.
Gordon, M. Y., Levicar, N., Pai, M., Bachellier, P., Dimarakis, I., Al-Allaf, F., et al. (2006). Characterization and clinical application of human CD34+ stem/progenitor cell populations mobilized into the blood by granulocyte colony-stimulating factor. Stem Cells, 24, 1822–1830.
Beigel, R., & Beigel, Y. (2009). Homozygous familial hypercholesterolemia: long term clinical course and plasma exchange therapy for two individual patients and review of the literature. Journal of Clinical Apheresis, 24, 219–224.
Cayo, M. A., Cai, J., Delaforest, A., Noto, F. K., Nagaoka, M., Clark, B. S., et al. (2012). JD induced pluripotent stem cell-derived hepatocytes faithfully recapitulate the pathophysiology of familial hypercholesterolemia. Hepatology, 56(6), 2163–2171.
Hoeg, J. M., Starzl, T. E., & Brewer, H. B., Jr. (1987). Liver transplantation for treatment of cardiovascular disease: comparison with medication and plasma exchange in homozygous familial hypercholesterolemia. American Journal of Cardiology, 59, 705–707.
Grossman, M., Raper, S. E., & Wilson, J. M. (1992). Transplantation of genetically modified autologous hepatocytes into nonhuman primates: feasibility and short-term toxicity. Human Gene Therapy, 3, 501–510.
Chowdhury, J. R., Grossman, M., Gupta, S., Chowdhury, N. R., Baker, J. R., Jr, & Wilson, J. M. (1991). Long-term improvement of hypercholesterolemia after ex vivo gene therapy in LDLR-deficient rabbits. Science, 254, 1802–1805.
Grossman, M., Raper, S. E., Kozarsky, K., Stein, E. A., Engelhardt, J. F., Muller, D., et al. (1994). Successful ex vivo gene therapy directed to liver in a patient with familial hypercholesterolaemia. Nature Genetics, 6, 335–341.
Grossman, M., Rader, D. J., Muller, D. W., Kolansky, D. M., Kozarsky, K., Clark, B. J, 3rd, et al. (1995). A pilot study of ex vivo gene therapy for homozygous familial hypercholesterolaemia. Nature Medicine, 1(11), 1148–1154.
Takahashi, K., Tanabe, K., Ohnuki, M., Narita, M., Ichisaka, T., Tomoda, K., et al. (2007). Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell, 131, 861–872.
Stadtfeld, M., & Hochedlinger, K. (2010). Induced pluripotency: history, mechanisms, and applications. Genes and Development, 24, 2239–2263.
Gonzalez, F., Boue, S., & Belmonte, J. C. (2011). Methods for making induced pluripotent stem cells: Reprogramming a la carte. Nature Reviews Genetics, 12, 231–242.
Nishikawa, S., Goldstein, R. A., & Nierras, C. R. (2008). The promise of human induced pluripotent stem cells for research and therapy. Nature Reviews Molecular Cell Biology, 9, 725–729.
Ye, Z., Zhan, H., Mali, P., Dowey, S., Williams, D. M., Jang, Y. Y., et al. (2009). Human-induced pluripotent stem cells from blood cells of healthy donors and patients with acquired blood disorders. Blood, 114, 5473–5480.
Park, I. H., Arora, N., Huo, H., Maherali, N., Ahfeldt, T., Shimamura, A., et al. (2008). Disease-specific induced pluripotent stem cells. Cell, 134, 877–886.
Lee, G., Papapetrou, E. P., Kim, H., Chambers, S. M., Tomishima, M. J., Fasano, C. A., et al. (2009). Modelling pathogenesis and treatment of familial dysautonomia using patient-specific iPSCs. Nature, 461, 402–406.
Ebert, A. D., Yu, J., Rose, F. F., Jr, Mattis, V. B., Lorson, C. L., Thomson, J. A., et al. (2009). Induced pluripotent stem cells from a spinal muscular atrophy patient. Nature, 457, 277–280.
Dimos, J. T., Rodolfa, K. T., Niakan, K. K., Weisenthal, L. M., Mitsumoto, H., Chung, W., et al. (2008). Induced pluripotent stem cells generated from patients with ALS can be differentiated into motor neurons. Science, 321, 1218–1221.
Carvajal-Vergara, X., Sevilla, A., D’Souza, S. L., Ang, Y. S., Schaniel, C., Lee, D. F., et al. (2010). Patient-specific induced pluripotent stem-cell-derived models of LEOPARD syndrome. Nature, 465, 808–812.
Ohnishi, T., Saito, K., Maeda, S., Matsumoto, K., Sakuda, M., & Inoki, R. (1990). Intracerebroventricular treatment of mice with pertussis toxin induces hyperalgesia and enhances 3H-nitrendipine binding to synaptic membranes: Similarity with morphine tolerance. Naunyn Schmiedebergs Archives of Pharmacology, 341, 123–127.
Brennand, K. J., Simone, A., Jou, J., Gelboin-Burkhart, C., Tran, N., Sangar, S., et al. (2011). Modelling schizophrenia using human induced pluripotent stem cells. Nature, 473(7346), 221–225.
Baek, K. H., Zaslavsky, A., Lynch, R. C., Britt, C., Okada, Y., Siarey, R. J., et al. (2009). Down’s syndrome suppression of tumour growth and the role of the calcineurin inhibitor DSCR1. Nature, 459, 1126–1130.
Soldner, F., Hockemeyer, D., Beard, C., Gao, Q., Bell, G. W., Cook, E. G., et al. (2009). Parkinson’s disease patient-derived induced pluripotent stem cells free of viral reprogramming factors. Cell, 136, 964–977.
Marchetto, M. C., Carromeu, C., Acab, A., Yu, D., Yeo, G. W., Mu, Y., et al. (2010). A model for neural development and treatment of Rett syndrome using human induced pluripotent stem cells. Cell, 143, 527–539.
Raya, A., Rodriguez-Piza, I., Guenechea, G., Vassena, R., Navarro, S., Barrero, M. J., et al. (2009). Disease-corrected haematopoietic progenitors from Fanconi anaemia induced pluripotent stem cells. Nature, 460, 53–59.
Kazuki, Y., Hiratsuka, M., Takiguchi, M., Osaki, M., Kajitani, N., Hoshiya, H., et al. (2010). Complete genetic correction of ips cells from Duchenne muscular dystrophy. Molecular Therapy, 18, 386–393.
Meyer, J. S., Howden, S. E., Wallace, K. A., Verhoeven, A. D., Wright, L. S., Capowski, E. E., et al. (2011). Optic vesicle-like structures derived from human pluripotent stem cells facilitate a customized approach to retinal disease treatment. Stem Cells, 29, 1206–1218.
Zagoura, D. S., Roubelakis, M. G., Bitsika, V., Trohatou, O., Pappa, K. I., Kapelouzou, A., et al. (2011). Therapeutic potential of a distinct population of human amniotic fluid mesenchymal stem cells and their secreted molecules in mice with acute hepatic failure. Gut, 61(6), 894–906.
Rashid, S. T., Corbineau, S., Hannan, N., Marciniak, S. J., Miranda, E., Alexander, G., et al. (2010). Modeling inherited metabolic disorders of the liver using human induced pluripotent stem cells. The Journal of Clinical Investigation, 120, 3127–3136.
Ghodsizadeh, A., Taei, A., Totonchi, M., Seifinejad, A., Gourabi, H., Pournasr, B., et al. (2010). Generation of liver disease-specific induced pluripotent stem cells along with efficient differentiation to functional hepatocyte-like cells. Stem Cell Reviews, 6, 622–632.
Mohamadnejad, M., Pournasr, B., Bagheri, M., Aghdami, N., Shahsavani, M., Hosseini, L. A., et al. (2010). Transplantation of allogeneic bone marrow mesenchymal stromal cell-derived hepatocyte-like cells in homozygous familial hypercholesterolemia. Cytotherapy, 12(4), 566–568.
Totonchi, M., Taei, A., Seifinejad, A., Tabebordbar, M., Rassouli, H., Farrokhi, A., et al. (2010). Feeder- and serum-free establishment and expansion of human induced pluripotent stem cells. International Journal of Developmental Biology, 54, 877–886.
Mollamohammadi, S., Taei, A., Pakzad, M., Totonchi, M., Seifinejad, A., Masoudi, N., et al. (2009). A simple and efficient cryopreservation method for feeder-free dissociated human induced pluripotent stem cells and human embryonic stem cells. Human Reproduction, 24, 2468–2476.
Basma, H., Soto-Gutierrez, A., Yannam, G. R., Liu, L., Ito, R., Yamamoto, T., et al. (2009). Differentiation and transplantation of human embryonic stem cell-derived hepatocytes. Gastroenterology, 136, 990–999.
Tzanakakis, E. S., Hsiao, C. C., Matsushita, T., Remmel, R. P., & Hu, W. S. (2001). Probing enhanced cytochrome P450 2B1/2 activity in rat hepatocyte spheroids through confocal laser scanning microscopy. Cell Transplantation, 10, 329–342.
Baharvand, H., Ashtiani, S. K., Taee, A., Massumi, M., Valojerdi, M. R., Yazdi, P. E., et al. (2006). Generation of new human embryonic stem cell lines with diploid and triploid karyotypes. Development, Growth & Differentiation, 48, 117–128.
Asgari, S., Pournasr, B., Salekdeh, G. H., Ghodsizadeh, A., Ott, M., & Baharvand, H. (2010). Induced pluripotent stem cells: a new era for hepatology. Journal of Hepatology, 53, 738–751.
Behbahan, I. S., Duan, Y., Lam, A., Khoobyari, S., Ma, X., Ahuja, T. P., et al. (2011). New approaches in the differentiation of human embryonic stem cells and induced pluripotent stem cells toward hepatocytes. Stem Cell Reviews, 7, 748–759.
Chistiakov, D. A., & Chistiakov, P. A. (2012). Strategies to produce hepatocytes and hepatocyte-like cells from pluripotent stem cells. Hepatology Research, 42(2), 111–119.
Sullivan, G. J., Hay, D. C., Park, I. H., Fletcher, J., Hannoun, Z., Payne, C. M., et al. (2010). Generation of functional human hepatic endoderm from human induced pluripotent stem cells. Hepatology, 51, 329–335.
Song, Z., Cai, J., Liu, Y., Zhao, D., Yong, J., Duo, S., et al. (2009). Efficient generation of hepatocyte-like cells from human induced pluripotent stem cells. Cell Research, 19, 1233–1242.
Si-Tayeb, K., Noto, F. K., Nagaoka, M., Li, J., Battle, M. A., Duris, C., et al. (2010). Highly efficient generation of human hepatocyte-like cells from induced pluripotent stem cells. Hepatology, 51, 297–305.
Liu, H., Ye, Z., Kim, Y., Sharkis, S., & Jang, Y. Y. (2010). Generation of endoderm-derived human induced pluripotent stem cells from primary hepatocytes. Hepatology, 51, 1810–1819.
Zhu, H., Lensch, M. W., Cahan, P., & Daley, G. Q. (2011). Investigating monogenic and complex diseases with pluripotent stem cells. Nature Reviews Genetics, 12, 266–275.
Inoue, H., & Yamanaka, S. (2011). The use of induced pluripotent stem cells in drug development. Clinical Pharmacology and Therapeutics, 89, 655–661.
Sadelain, M. (2010). The need for genetically engineering therapeutic pluripotent stem cells. Molecular Therapy, 18, 2039.
Brown, M. S., & Goldstein, J. L. (1999). A proteolytic pathway that controls the cholesterol content of membranes, cells, and blood. Proceedings of the National Academy of Sciences of the United States of America, 96, 11041–11048.
Brown, M. S., & Goldstein, J. L. (1997). The SREBP pathway: regulation of cholesterol metabolism by proteolysis of a membrane-bound transcription factor. Cell, 89, 331–340.
Liu, H., Kim, Y., Sharkis, S., Marchionni, L., & Jang, Y. Y. (2011). In vivo liver regeneration potential of human induced pluripotent stem cells from diverse origins. Science Translational Medicine, 3(82), 82ra39.
Espejel, S., Roll, G. R., McLaughlin, K. J., Lee, A. Y., Zhang, J. Y., Laird, D. J., et al. (2010). Induced pluripotent stem cell-derived hepatocytes have the functional and proliferative capabilities needed for liver regeneration in mice. The Journal of Clinical Investigation, 120, 3120–3126.
Ishibashi, S., Goldstein, J. L., Brown, M. S., Herz, J., & Burns, D. K. (1994). Massive xanthomatosis and atherosclerosis in cholesterol-fed low density lipoprotein receptor-negative mice. The Journal of Clinical Investigation, 93, 1885–1893.
Watanabe, Y., Ito, T., & Shiomi, M. (1985). The effect of selective breeding on the development of coronary atherosclerosis in WHHL rabbits. An animal model for familial hypercholesterolemia. Atherosclerosis, 56, 71–79.
Seifinejad, A., Tabebordbar, M., Baharvand, H., Boyer, L. A., & Salekdeh, G. H. (2010). Progress and promise towards safe induced pluripotent stem cells for therapy. Stem Cell Reviews, 6, 297–306.
Zhao, T., Zhang, Z. N., Rong, Z., & Xu, Y. (2011). Immunogenicity of induced pluripotent stem cells. Nature, 474, 212–215.
Ben-David, U., & Benvenisty, N. (2011). The tumorigenicity of human embryonic and induced pluripotent stem cells. Nature Reviews Cancer, 11, 268–277.
Huang, P., He, Z., Ji, S., Sun, H., Xiang, D., Liu, C., et al. (2011). Induction of functional hepatocyte-like cells from mouse fibroblasts by defined factors. Nature, 475, 386–389.
Acknowledgments
This study was funded by a grant provided from Royan Institute and the Iranian Council of Stem Cell Research and Technology.
Conflict of interest
We hereby confirm that any and all potential conflicts of interest have been fully and properly disclosed in the manuscript as outlined.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Fattahi, F., Asgari, S., Pournasr, B. et al. Disease-Corrected Hepatocyte-Like Cells from Familial Hypercholesterolemia-Induced Pluripotent Stem Cells. Mol Biotechnol 54, 863–873 (2013). https://doi.org/10.1007/s12033-012-9635-3
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-012-9635-3