Abstract
This study is to analyze differentially expressed genes (DEGs) and mutation signatures of pancreatic head cancer and pancreatic body/tail cancer. Pancreatic Adenocarcinoma (PAAD) RNA-seq data, mutation data and clinical data were downloaded and collected from The Cancer Genome Atlas (TCGA), FireHose and CBioPortal. According to the anatomic location, the patients were divided into 146 cases of pancreatic head cancer and 28 cases of pancreatic body/tail cancer. Then survival analysis was performed by Kaplan–Meier and log-rank test. Furthermore, DEGs were screened by R package Deseq2. Gene Ontology (GO), Kyoto Encyclopedia of Genes and Genomes (KEGG) and protein–protein interaction (PPI) were then carried out by DAVID and String. Online tool TIMER was used to analyze the immune cells infiltration. R package maftools and GenVisR were applied to analyze frequently mutated genes and mutant-allele tumor heterogeneity (MATH) of PAAD. Survival of patients with pancreatic body/tail cancer was better than those with pancreatic head cancer (median survival, 24.05 vs 19.45 months, p = 0.048). And 496 significant DEGs (|log2 FoldChange| > 1.5,false discovery rate (FDR) < 0.05) were identified, including 253 downregulated genes and 243 upregulated genes. And there were 13 Go terms (4 biological processes, 6 cellular components and 3 molecular functions) and 3 KEGG pathways (Pancreatic secretion, Fat digestion and absorption, Protein digestion and absorption) (FDR < 0.05). B cells and CD4 + T cells infiltration were more significant in pancreatic head cancer. MATH scores of pancreatic body/tail cancer were higher than pancreatic head cancer, while χ2 test of top 10 frequently mutated genes showed little difference between them. There were prognostic and genetic differences between pancreatic head cancer and pancreatic body/tail cancer. PAAD originated from different location may have different biology natures and should not be treated with same strategy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pancreatic adenocarcinoma (PAAD) is one of the most malignant gastrointestinal tumors with difficulties in early diagnosis and treatment [1]. Despite of the development of diagnosis and treatment of PAAD, the 5-year survival rate is still as low as 8% [2]. Besides, since there is no obvious sign at early stage, the early diagnosis rate of PAAD is rather low. Treatment of PAAD includes surgery, chemotherapy, radiation therapy and palliative care, while surgery remains to be the only potentially curative option [3]. PAAD can occur in any part of the pancreas, but most of PAADs are found in the head of the pancreas, while pancreatic body/tail cancers account for only 20–25% [4]. Branching intraductal papillary mucinous tumors typically occur in the head of the pancreas, while mucinous cystic tumors are more common in the body or tail of pancreas [5]. Several studies have suggested that the anatomic location of PAADs plays an important role in survival. These studies indicate that there might be differences in clinical presentation, therapy and malignant potentials between pancreatic head cancer and pancreatic body/tail cancer [6,7,8]. Usually, pancreatic head cancer has a higher incidence and is easier to detect and has a better prognosis compared with pancreatic body/tail cancer [9]. Moreover, compared with patients with pancreatic head cancer, patients with early stage pancreatic body/tail cancer have a lower tumor recurrence rate after curative resection [10].
The effect of anatomic site on prognosis of PAAD has recently been studied extensively, yet the exact genetic differences between pancreatic head cancer and pancreatic body/tail cancer have not been fully elucidated [11]. Therefore, the main purpose of this study is to analyze the DEGs and mutation signatures of pancreatic head cancer and pancreatic body/tail cancer, and to find out the significant DEGs, GO functions, KEGG pathways and mutations which are related to PAAD sites, so as to contribute to the better diagnosis, treatment and prognosis of PAAD.
Methods
Data source
RNA-seq expression profiles, clinical data and mutation data were downloaded and collected from TCGA (https://xenabrowser.net/datapages/), CBioPortal (https://www.cbioportal.org/datasets) and FireHose, respectively. The RNA-seq expression profiles of PAAD were downloaded from TCGA, including 139 pancreatic head cancer cases and 28 pancreatic body/tail cancer cases (Fig. 1). Clinical data of PAAD was downloaded from CBioPortal, including age, gender, race, alcohol history, OS, DFS, and AJCC stage. Mutation data were downloaded from FireHose.
Clinical information analysis
To ensure the patients of pancreatic head cancer and pancreatic body/tail cancer were comparable, the χ2 test was performed on the age, sex, race, AJCC stage and alcohol history between the two groups. Then overall survival was calculated by R package survival, using the Kaplan–Meier method and log-rank test. The criterion for statistical significance was p < 0.05 on above analysis.
Analysis of RNA-seq data
Firstly, RNA-seq with count per million (CPM) > 1 in more than 10% of the PAAD samples were retained for differential expression genes analysis. Then R package DESeq2 (https://github.com/mikelove/DESeq2) was applied to screen DEGs of pancreatic head cancer and pancreatic body/tail cancer. We performed multiple comparisons using the Benjamini and Hochberg approach to acquire the false discovery rate (FDR). DEGs were defined with the thresholds of FDR < 0.05 and |log2 FoldChange| > 1.5. To analyze the function and the potential pathway of DEGs, the online tool DAVID was used to conduct the functional annotation GO and KEGG enrichment analysis. FDR < 0.05 was defined as the criteria of statistical significance. Moreover, the top 300 genes were used to build PPI network by online tool String and Cytoscape software. We used nodes to represent the proteins and edges to represent interactions between two proteins. The nodes and edges indicate proteins and interactions between two proteins, respectively. The minimum required interaction score was set up as 0.400.
Tumor immune estimation resource (TIMER)
To analyze the immune infiltration of the tumor microenvironment of PAAD, we used the TIMER, a web-based tool which was validated in several cancers (https://cistrome.shinyapps.io/timer/). Six types of tumor-infiltrating cell populations (CD4 + T cells, CD8 + T cells, B cells, Macrophages, Neutrophils and Dendritic cells) were included in the analysis based on the PAAD data from TCGA database.
Analysis of somatic mutation data
Mutation annotation format (MAF) files for somatic mutation were downloaded from FireHose for the analysis. A total of 140 patients were available for somatic mutation analysis, including 20 pancreatic head cancer and 106 pancreatic body/tail cancer. Then, the R packages maftools (https://github.com/PoisonAlien/maftools) and GenVisR (https://bioconductor.org/packages/GenVisR/) were used to analyze the top 10 frequently mutated genes of PDAC. To compare the top 10 frequently mutated genes of pancreatic head cancer and pancreatic body/tail cancer, Chi-square test was performed on them. Furthermore, to evaluate the degree of intra-tumor heterogeneity, MATH was calculated for each patient by R package maftools. Student’s t test was used to compare the MATH score between two groups.
Result
Clinicopathological characteristics
There was no significant difference in the Chi-square test of age (p = 0.89982), sex (p = 0.99982), race (p = 0.14), AJCC stage (p = 0.61) and alcohol history (p = 0.16) of pancreatic head cancer and pancreatic body/tail cancer (Table 1). The Kaplan–Meier survival analysis showed that the anatomic location of PAAD was closely related to the prognosis of patients with PAAD. The survival of patients with pancreatic body/tail cancer was superior to those with pancreatic head cancer (p = 0.048) (Fig. 2).
Analysis of DEGs
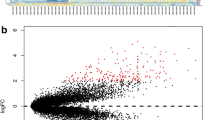
There were 496 significant DEGs (|log2 FoldChange| > 1.5, p < 0.05), of which 243 genes were upregulated and 253 genes were downregulated (Fig. 3a, b; Supplementary Table S1). The GO enrichment analysis showed that there were 13 significant functions/structures (FDR < 0.05), 4 biological processes, 6 cellular components and 3 molecular functions, including digestion, chemical synaptic transmission, regulation of insulin secretion, glucose homeostasis, extracellular space, extracellular region, plasma membrane, secretory granule, integral component of plasma membrane, voltage-gated potassium channel complex neuropeptide hormone activity, serine-type endopeptidase activity and heparin binding terms (Fig. 3c). The KEGG demonstrated that there were 3 significant pathways (FDR < 0.05), including pancreatic secretion, fat digestion and absorption and protein digestion and absorption (Fig. 3d).The PPI network of top 300 genes consisted of 278 nodes and 684 edges (Fig. 3e, Supplementary Table S2). Genes CCR7, PENK, PPY, SSTR2, CXCL13, GNG4, ADCY1, CCR4, CPR18 and CNR2 were at the core of interaction network (Fig. 3f).
a Volcano plots of DEGs between pancreatic head cancer and pancreatic body/tail cancer. X-axis indicates the fold change (log scaled), whereas the Y-axis shows the p values (log scaled). Each symbol represents a different gene, and the red/blue color of the symbols categorize the upregulated/downregulated genes falling under different criteria (p value and fold change threshold). p value < 0.05 is considered as statistically significant, whereas fold change = 1.5 is set as the threshold. b Heatmaps of the top 20 and last 20 DEGs between pancreatic head cancer and pancreatic body/tail cancer in TCGA. c GO analyses of the DEGs according to their biological process, cellular component and molecular function. X-axis indicates the gene number, whereas the Y-axis shows the most enriched GO terms. d Bubble plot of significant KEGG pathways. X-axis indicates the gene ratio, whereas the Y-axis shows the most enriched KEGG pathways. The size of bubble presents genes number, and the color of bubble presents the p value. e PPI network of the top 300 DEGs. And the red/blue color of the symbols categorize the upregulated/downregulated genes. f Ten hub genes were identified by Cytoscape Cytohubba plug-in
Mutation signatures
Among the 186 patients with PAAD, 140 had both mutation and tumor site information, including 118 pancreatic head cancer and 22 pancreatic body/tail cancer. By analyzing the MAF files of PAAD. The top 10 frequently mutated genes of PAAD were respectively KRAS (90.7%), TP53 (69.0%), TNN (2.9%), MUC16 (7.8%), CDKN2A (14.3%), SMAD4 (25.0%), FLG (7.1%), GNAS (7.1%), RYR (7.1%), and OBSCN (7.1%). Among them, the KRAS gene mutation rate was the highest and 127 out of 140 patients with PAAD had KRAS gene mutations. Moreover, the results of Chi-square test showed no significant difference in KRAS (p = 0.2213), TP53 (p = 0.2558), TNN (p = 1.000), MUC16 (p = 0.379), CDKN2A (p = 1.000), SMAD4 (p = 0.283), FLG (p = 0.949), GNAS (p = 0.362), RYR (p = 0.193), and OBSCN (p = 0.363) (Table 2). Then, to evaluate the degree of intra-tumor heterogeneity, MATH was calculated. The result showed the median MATH score of pancreatic body/tail cancer (91.5) was significantly higher than pancreatic head cancer (78.1) (p = 0.009) (Fig. 4b).
Infiltration level of immune cells in pancreatic head cancer and pancreatic body/tail cancer
With the estimation of TIMER, the infiltration levels of CD4+ T cells, CD8+ T cells, B cells, Macrophages, Neutrophils and DCs in two groups were retrieved. The infiltration levels of CD4+ T cells and B cells were higher in pancreatic head cancers than those in the pancreatic body/tail cancers (Fig. 5, both p < 0.01). There is no significant difference of CD8+ T cells (p = 0.450), Macrophages (p = 0.450), Neutrophils (p = 0.051) and DCs (p = 0.067) between two groups.
Discussion
Pancreatic head cancer and pancreatic body/tail cancer may not be the same type of tumor. Several studies have demonstrated that there are differences in clinical manifestations, treatment, prognosis and recurrence [10, 12]. As for clinical presentation, most patients with PAAD are asymptomatic until the disease progresses to an advanced stage [13]. Patients with PAAD usually presents jaundice, indigestion, pain and weight loss. Usually, pancreatic head cancer presents jaundice early due to the obstruction of the common bile duct, while pancreatic body/tail cancer presents with weight loss and pain, symptoms more in keeping with advanced disease [14]. As for treatment, tumor location is an important factor in treatment of PAAD. Operation plan is different between pancreatic head cancer and pancreatic body/tail cancer. Pancreatic head cancer requires a pancreaticoduodenectomy (Whipple operation), whereas pancreatic tail cancer requires a distal pancreatectomy with an en bloc splenectomy. Pancreatic neck/body cancer may require a pancreaticoduodenectomy, distal pancreatectomy or, rarely, a total pancreatectomy [15, 16]. The lower frequency of pancreatic body cancer resection may be explained by the fact that pancreatic body lesions may be the most challenging to manage due to the co-involvement of major vessels and the late diagnosis [17].
The anatomic location of PAAD also plays an important role in prognosis. Our study based on 174 PAAD cases from TCGA cohort indicates that the prognosis of patients with pancreatic body/tail cancer is better than those with pancreatic head cancer. However, the prognosis of pancreatic head cancer and pancreatic body/tail cancer is still controversial. Some studies support that patients with pancreatic body/tail cancer have an increased risk of death compared to those with pancreatic head cancer [6, 18]. This may result from late diagnosis of pancreatic body/tail cancer, increased frequency of metastasis and lower resecting rate [6]. However, some studies have shown that patients with pancreatic head cancer have a poorer survival than those with pancreatic body/tail cancer. Although patients with pancreatic head cancer have higher resection rates and utilization of adjuvant therapy, those with pancreatic body/tail cancer experience better overall survival [19]. Besides, study shows that the earlier stage at diagnosis has better survival than later stages, especially for pancreatic body/tail cancer [8]. Study has demonstrated that the patients with early stage pancreatic body/tail cancer have a lower tumor recurrence rate after curative resection [10].
In our study, digestion, chemical synaptic transmission, regulation of insulin secretion, glucose homeostasis, extracellular space, extracellular region, plasma membrane, secretory granule, integral component of plasma membrane, voltage-gated potassium channel complex neuropeptide hormone activity, serine-type endopeptidase activity and heparin binding terms are the most enriched GO terms. KEGG is mainly enriched in three pathways, including Pancreatic secretion, Fat digestion and absorption and Protein digestion and absorption. We can find that the most enriched GO terms and KEGG pathways are intensely correlated to digestion and pancreatic excretion. The main functions of the pancreas include digestion and excretion, while the head of pancreas and body/tail of pancreas have different roles in digestion and secretion. Study has confirmed that the concentration of insulin-positive endocrine cells in the tail of the pancreas is higher than that in the head [20]. Therefore, different clinical features of the cancers from head and tail could be caused by the different histological components and different biological functions.
PPIs plays an important role in gene expression, cell growth, proliferation and apoptosis [21]. Numerous studies have shown that PPIs is the basis of a variety of aggregation-related diseases, especially related to the occurrence and progression of cancer [22, 23]. In our study, genes CCR7, PENK, PPY, SSTR2, CXCL13, GNG4, ADCY1, CCR4, CPR18 and CNR2 are at the core of PPI network. However, CCR7 is the most significant gene of PPI network. As one of chemokine receptors, CCR7 expression is highly associated with lymph node metastasis in PAAD [24]. Although CCR7 expression may have no power to predict the recurrence of PAAD, CCR7 positivity is closely associated with a high incidence of metastasis [24]. Study has identified that tumor metastasis is highly related to cancer mortality, especially PAAD [25]. Therefore, we can conclude that the expression of CCR7 may affect the survival and prognosis of patients with PAAD. KRAS, TP53, TTN, SMAD4, CDKN2A, MUC16, FLG, GNAS, RYR1 and OBSCN are top 10 frequently mutated genes of PAAD. However, the results of our study show little difference of the top 10 frequently mutated genes between pancreatic head cancer and pancreatic body/tail cancer. Study shows heterogeneity may have a significant impact on tumor invasiveness, disease prognosis, and response to treatment, and thus may be a major obstacle to effective treatment of cancer and individualized medicine [26]. MATH has been proven to be a simple, quantitative and universally applicable method to evaluate intra-tumor heterogeneity [27]. In our study, the MATH score of pancreatic body/tail cancer is higher than pancreatic head cancer, which means that pancreatic body/tail cancer might be more heterogeneous than pancreatic head cancer. Tumor microenvironment analysis regarding the immune cells infiltration showed that the levels of B cells and CD4 + cells are higher in the pancreatic head cancer. RNA-seq data above suggests that CXCL13 is highly expressed in the pancreatic head which is an important recruitment factor for B cells [28]. B cells have a cancer-promoting effect which might play a role in the worse outcome of pancreatic head cancer. Tregs (T regulatory cells) are important subpopulation of CD4 + cells and can promote the pancreatic cancer progression [29].
The main advantage of this study is the reliable data source, which is downloaded from TCGA. The TCGA is a public funded project which aims to catalog and discover major cancer-causing genome alterations in large cohorts of over 30 human tumors through large-scale genome sequencing and integrated multi-dimensional analyses. In addition, TCGA can provide comprehensive analysis of cancer genome profiles, which is constantly updated [30]. So, the RNA-seq expression profiles, clinical data and genes mutation data of our study are believable and comprehensive. However, there are some limitations to our study. Firstly, the sample size in the confirmation by qRT-PCR is small and large numbers of samples of pancreatic head cancer and pancreatic body/tail cancer are needed for further research. Secondly, the DEGs obtained in our study are from TCGA cohort, which lacks the validation of other cohorts. Thirdly, the analyses of the current study are mainly based on the gene expression and mutation of the PAADs in TCGA cohort. To fully explore genetic differences of pancreatic head cancer and pancreatic body/tail cancer, vascular stability and immune response genes, in vitro and in vivo experimental approaches are needed.
In conclusion, by comparing DEGs and mutation signatures of pancreatic head cancer and pancreatic body/tail cancer from TCGA cohort, our study identifies that there are prognostic and genetic differences between pancreatic head cancer and pancreatic body/tail cancer, which can provide the basis for the individualized treatment and prognosis assessment of pancreatic head cancer and pancreatic body/tail cancer.
References
Ilic M, Ilic I. Epidemiology of pancreatic cancer. World J Gastroenterol. 2016;22(44):9694–705. https://doi.org/10.3748/wjg.v22.i44.9694.
Siegel RLMK, Jemal A. Cancer statistics, 2018. CA Cancer J Clin. 2018;68:7–30. https://doi.org/10.3322/caac.21442.
Neoptolemos JPPD, Ghaneh P, et al. Comparison of adjuvant gemcitabine and capecitabine with gemcitabine monotherapy in patients with resected pancreatic cancer (ESPAC-4): a multicentre, open-label, randomised, phase 3 trial. Lancet. 2017;389:1011–24.
Modolell IGL, Malagelada JR. Vagaries of clinical presentation of pancreatic and biliary tract cancer. Ann Oncol. 1999;10(Suppl 4):82–4. https://doi.org/10.1093/annonc/10.
Pitman MB, Lewandrowski K, Shen J, Sahani D, Brugge W, Fernandez-del CC. Pancreatic cysts. 2010;118(1):1–13. https://doi.org/10.1002/cncy.20059.
Artinyan A, Soriano PA, Prendergast C, Low T, Ellenhorn JD, Kim J. The anatomic location of pancreatic cancer is a prognostic factor for survival. HPB (Oxford). 2008;10(5):371–6. https://doi.org/10.1080/13651820802291233.
Watanabe I, Sasaki S, Konishi M, Nakagohri T, Inoue K, Oda T, et al. Onset Symptoms and Tumor Locations as Prognostic Factors of Pancreatic Cancer. 2004;28(2):160–5.
Lau MK, Davila JA, Shaib YH. Incidence and Survival of Pancreatic Head and Body and Tail Cancers: A Population-Based Study in the United States. 2010;39(4):458–62. https://doi.org/10.1097/MPA.0b013e3181bd6489.
Ling Q, Xu X, Zheng S-S, Kalthoff H. The diversity between pancreatic head and body/tail cancers: clinical parameters and in vitro models. Hepatobiliary Pancreatic Dis Int. 2013;12(5):480–7.
Ling Q, Xu X, Ye P, Xie H, Gao F, Hu Q, et al. The prognostic relevance of primary tumor location in patients undergoing resection for pancreatic ductal adenocarcinoma. Oncotarget. 2017;8(9):15159–677. https://doi.org/10.18632/oncotarget.14768.
Bailey P, Chang DK, Nones K, Johns AL, Patch A-M, Gingras M-C, et al. Genomic analyses identify molecular subtypes of pancreatic cancer. Nature. 2016;531(7592):47–52.
Birnbaum DJ, Bertucci F, Finetti P, Birnbaum D, Mamessier E. Head and body/tail pancreatic carcinomas are not the same tumors. Cancers (Basel). 2019. https://doi.org/10.3390/cancers11040497.
Kamisawa T, Wood LD, Itoi T, Takaori K. Pancreatic cancer. Lancet. 2016;388(10039):73–85. https://doi.org/10.1016/s0140-6736(16)00141-0.
Walter FM, Mills K, Mendonça SC, Abel GA, Basu B, Carroll N, et al. Symptoms and patient factors associated with diagnostic intervals for pancreatic cancer (SYMPTOM pancreatic study): a prospective cohort study. Lancet Gastroenterol Hepatol. 2016;1(4):298–306. https://doi.org/10.1016/S2468-1253(16)30079-6.
Winter JM, Cameron JL, Campbell KA, Arnold MA, Chang DC, Coleman J, et al. 1423 Pancreaticoduodenectomies for pancreatic cancer: a single-institution experience. J. Gastroint. Surg. 2006;10(9):1199–211. https://doi.org/10.1016/j.gassur.2006.08.018.
Nathan H, Wolfgang CL, Edil BH, Choti MA, Herman JM, Schulick RD, et al. Peri-operative mortality and long-term survival after total pancreatectomy for pancreatic adenocarcinoma: a population-based perspective. J Surg Oncol. 2009;99(2):87–92. https://doi.org/10.1002/jso.21189.
Hartwig W, Werner J, Jäger D, Debus J, Büchler MW. Improvement of surgical results for pancreatic cancer. Lancet Oncol. 2013;14(11):e476–e485485. https://doi.org/10.1016/S1470-2045(13)70172-4.
Bilici A. Prognostic factors related with survival in patients with pancreatic adenocarcinoma. World J Gastroenterol. 2014;20(31):10802–12. https://doi.org/10.3748/wjg.v20.i31.10802.
Winer LK, Dhar VK, Wima K, Morris MC, Lee TC, Shah SA, et al. The impact of tumor location on resection and survival for pancreatic ductal adenocarcinoma. J Surg Res. 2019;239:60–6. https://doi.org/10.1016/j.jss.2019.01.061.
Gidekel Friedlander SY, Chu GC, Snyder EL, Girnius N, Dibelius G, Crowley D, et al. Context-dependent transformation of adult pancreatic cells by oncogenic K-Ras. Cancer Cell. 2009;16(5):379–89. https://doi.org/10.1016/j.ccr.2009.09.027.
Hu Y, Zhang Y, Ren J, Wang Y, Wang Z, Zhang J. Statistical Approaches for the Construction and Interpretation of Human Protein-Protein Interaction Network. BioMed Res Int. 2016;2016:5313050. https://doi.org/10.1155/2016/5313050.
Huang Z, Duan H, Li H. Identification of gene expression pattern related to breast cancer survival using integrated tcga datasets and genomic tools. BioMed Res Int. 2015;2015:878546. https://doi.org/10.1155/2015/878546.
Vishnubalaji R, Hamam R, Abdulla MH, Mohammed MAV, Kassem M, Al-Obeed O, et al. Genome-wide mRNA and miRNA expression profiling reveal multiple regulatory networks in colorectal cancer. Cell Death Dis. 2015;6(1):e16–e1414. https://doi.org/10.1038/cddis.2014.556.
Nakata B, Fukunaga S, Noda E, Amano R, Yamada N, Hirakawa K. Chemokine receptor CCR7 expression correlates with lymph node metastasis in pancreatic cancer. Oncology. 2008;74(1–2):69–75. https://doi.org/10.1159/000139126.
Zhang L, Wang D, Li Y, Liu Y, Xie X, Wu Y, et al. CCL21/CCR7 axis contributed to CD133+ pancreatic cancer stem-like cell metastasis via EMT and Erk/NF-kappaB pathway. PLoS ONE. 2016;11(8):e0158529. https://doi.org/10.1371/journal.pone.0158529.
Almendro V, Marusyk A, Polyak K. Cellular Heterogeneity and Molecular Evolution in Cancer. 2013;8(1):277–302. https://doi.org/10.1146/annurev-pathol-020712-163923.
Mroz EA, Rocco JW. MATH, a novel measure of intratumor genetic heterogeneity, is high in poor-outcome classes of head and neck squamous cell carcinoma. Oral Oncol. 2013;49(3):211–5. https://doi.org/10.1016/j.oraloncology.2012.09.007.
Pylayeva-Gupta Y, Das S, Handler JS, Hajdu CH, Coffre M, Koralov SB, et al. IL35-producing B cells promote the development of pancreatic neoplasia. Cancer Discov. 2016;6(3):247–55.
Liu L, Zhao G, Wu W, Rong Y, Jin D, Wang D, et al. Low intratumoral regulatory T cells and high peritumoral CD8+ T cells relate to long-term survival in patients with pancreatic ductal adenocarcinoma after pancreatectomy. Cancer Immunol Immunother. 2016;65(1):73–82.
Tomczak K, Czerwinska P, Wiznerowicz M. The Cancer Genome Atlas (TCGA): an immeasurable source of knowledge. Contemp Oncol. (Pozn). 2015;19(1A):A68–77. https://doi.org/10.5114/wo.2014.47136.
Acknowledgements
This work was supported by the grants from the National Natural Science Foundation of China (81672449, 81871980), The Project of Invigorating Health Care through Science, Technology and Education, Jiangsu Provincial Medical Outstanding Talent (to Yi Miao, JCRCA2016009), Innovation Capability Development Project of Jiangsu Province (No. BM2015004), Jiangsu Key Medical Discipline (General Surgery) (ZDXKA2016005) and Wu Jiepin Medical Foundation (No. 320.2710.1858).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
All authors have declared no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yin, L., Xiao, L., Gao, Y. et al. Comparative bioinformatical analysis of pancreatic head cancer and pancreatic body/tail cancer. Med Oncol 37, 46 (2020). https://doi.org/10.1007/s12032-020-01370-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-020-01370-0