Abstract
Variation in DNA repair genes is one of the mechanisms that may lead to variation in DNA repair capacity. Ku, a heterodimeric DNA-binding complex, is directly involved in repair of DNA double-strand breaks. Ku consists of two subunits, Ku70 and Ku80, which are encoded by the XRCC6 and XRCC5 genes, respectively. In the present study, we investigated whether common genetic variant in variable number of tandem repeats (VNTR) XRCC5 and T-991C XRCC6 was associated with an altered risk of breast cancer. The present study included 407 females with breast cancer and 395 age frequency-matched controls which were randomly selected from the healthy female blood donors. The XRCC5 and XRCC6 polymorphisms were determined using PCR-based methods. For XRCC5 polymorphism, in comparison with the 1R/1R genotype, the 0R/0R genotype increased breast cancer risk (OR 9.55, 95 %CI 1.19–76.64, P = 0.034). The 1R/3R genotype compared with 1R/1R genotype decreased the risk of breast cancer (Fisher’s exact test P = 0.015). There was no association between T-991C polymorphism of XRCC6 and breast cancer risk. Mean of age at diagnosis of breast cancer for 0, 1, 2, 3, and >4 repeat in XRCC5 were 39.2, 41.9, 44.3, 45.8, and 47.3 years, respectively. The Kaplan–Meier survival analysis revealed that the number of repeat was associated with age at diagnosis of breast cancer (log rank statistic = 13.90, df = 4, P = 0.008). The findings of the present study revealed that either breast cancer risk or age at diagnosis of breast cancer was associated with the VNTR polymorphism at promoter region of XRCC5.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
It is well established that DNA repair plays a crucial role in maintaining normal function and genetic material stability of mammalian [1]. After DNA damage caused by endogenous or exogenous factors, it may arise carcinogenesis or apoptosis [2]. Polymorphism in DNA repair genes is one of the mechanisms that may lead to variation in DNA repair capacity [3]. It has been suggested that polymorphisms in genes involved in the non-homologous end joining (NHEJ) pathway, influence DNA repair capacity [4]. Ku, a heterodimeric DNA-binding complex, is directly involved in repair of DNA double-strand breaks as a member of the NHEJ pathway. NHEJ is thought to proceed through three key steps: recognition of the breaks, DNA processing to remove non-ligatable ends or other forms of damage at the termini, and finally ligation of the DNA ends. Recognition of the double-strand breaks is carried out by the Ku heterodimer, which is also required for recruitment of DNA-PKcs, the XRCC4/DNA ligase IV complex and XLF, reflecting its essential role in NHEJ [5]. The Ku complex binds to the ends of double-stranded DNA in a cell cycle-dependent manner, being associated with chromosomes of interphase cells, followed by complete dissociation from the condensing chromosomes in early prophase [6]. Ku consists of two subunits, Ku70 and Ku80, which are encoded by the XRCC6 (X-ray repair cross-complementing 6; OMIM: 152690) and XRCC5 (X-ray repair cross-complementing 5; OMIM: 194364) genes, respectively [7].
Several polymorphisms in the XRCC5 have been reported. One of them is a variable number of tandem repeats of a 21 bp (VNTR, rs. 6147172) polymorphism in the promoter region of XRCC5. This polymorphism has four alleles: 3R, 2R, 1R and 0R [8]. Number copies of cis elements in the promoter region of XRCC5 regulate its expression [9]. It has been shown that the VNTR polymorphism of XRCC5 is associated with the risk of bladder cancer [10]. Single-nucleotide polymorphism of T-991C (rs. 5751129) in the promoter region of XRCC6 has been reported. This polymorphism is associated with the risk of gastric, oral and lung cancer and also hepatocellular and renal cell carcinomas [11–15].
To date, there is no study investigating the association between the above-mentioned polymorphisms and susceptibility to breast cancer. The aim of the present study is to investigate the association between XRCC5 and XRCC6 polymorphisms and susceptibility to breast cancer.
Materials and methods
For the current study, 407 female patients were recruited from the chemotherapy department of Nemazi hospital in Shiraz, south of Iran. Eligibility criteria for cases were patients with pathologically confirmed primary adenocarcinoma of the breast and mental competence to give written informed consent. During the same time period, 395 cancer-free female controls, recruited among randomly selected blood donors, frequency matched to the cases by age. Participants with any previous history of cancer or diagnosed psychiatric diseases were excluded from the control group. The mean age (SD; Min–Max) of the patients and controls was 45.3 (10.7; 22–80) and 43.9 (8.8; 24–72) years, respectively. Considering the high heterogeneity of Iranian populations [16, 17], the participants (patients and controls) were selected from Persian Muslims (Caucasians) living in Fars Province (southern Iran). Ethical approval for the current study was obtained from Shiraz University institutional review board.
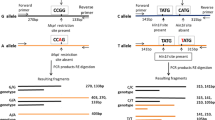
Genomic DNA was extracted from EDTA-treated blood samples. Genotyping for the XRCC5 VNTR polymorphism was carried out using high-resolution melting analysis (HRMA), following a pre-amplification step by Rotor-Gene 6000 instrument (Corbett Life Science). The reaction employed SYBR Premix Ex Taq II PCR master mix (Takara Bio Inc.) and previously described primers, 5′-AGG CGG CTC AAA CAC CAC AC-3′ (forward), and 5′-CAA GCG GCA GAT AGC GGA AAG-3′ (reverse). The PCR was performed in a total volume of 20 μl containing 100 ng genomic DNA, 10 μl SYBR Premix Ex Taq II, 0.8 μl (0.4 μM) of each forward and reverse primers, and the total reaction volume was brought to 20 μl with dH2O. Real-time PCR conditions were as follows: one cycle at 95 °C for 1 min followed by 40 cycles of denaturation at 95 °C for 10 s, annealing at 67 °C for 20 s and extension at 72 °C for 20 s. Melting curve data were collected from 85 to 95 °C at a ramping rate of 0.1 °C per second. Melting curve analysis was performed using SYBR green I channel, employing Rotor-Gene 6000 software (version 1.7) [18]. The T-991C polymorphism of XRCC6 was determined using the PCR–RFLP method with the primers as described previously [19]. The primers used were forward 5′-AAC TCA TGG ACC CAC GGT TGT GA-3′ and backward 5′-CAA CTT AAA TAC AGG AAT GTC TTG-3′. The cycling condition for the Ku70 promoter C-991T polymorphism was set as follows: one cycle at 94 °C for 8 min, 30 cycles at 94 °C for 30 s, 55 °C for 30 s, and 72 °C for 30 s, and one final cycle of extension at 72 °C for 10 min. The resultant 301 bp PCR product was mixed with 2 U of DpnII. The restriction site was located at −991 with a C/T polymorphism, and the C form PCR products could be further digested, while the T form could not. Two fragments measuring 101 and 200 bp were present if the product was digestible (C). The reaction was incubated for 2 h at 37 °C. Then, 10 μl of product was loaded into a 3 % agarose gel containing ethidium bromide for electrophoresis. The polymorphism was categorized as either (1) a C/C homozygote (digested), (2) T/T homozygote (undigested), or (3) C/T heterozygote.
A chi-square test was performed for the polymorphisms to determine if the control sample demonstrated Hardy–Weinberg equilibrium. The risk of breast cancer associated with the XRCC5 and XRCC6 polymorphisms was estimated. The relative associations between the genotypes and breast cancer risk were assessed by calculating crude odds ratios (ORs) and 95 % confidence intervals (CIs). To determine the effect(s) of XRCC5 and XRCC6 polymorphisms on age at diagnosis of breast cancer, the Kaplan–Meier survival analysis and Cox proportional hazards regression model were used. In analysis, breast cancer was defined as event, and age at diagnosis was included in the analysis as time period to event. Data analysis was performed using SPSS software version 11.5. All statistical tests are two-sided.
Results and discussion
Table 1 shows the genotype distribution of the studied polymorphisms in breast cancer cases and healthy controls. Control subjects were at Hardy–Weinberg equilibrium for both polymorphisms (For VNTR XRCC5 polymorphism: χ2 = 3.065, df = 6, P = 0.800; For T-991C XRCC6 polymorphism: χ2 = 1.767, df = 1, P = 0.183).
For the VNTR XRCC5 polymorphism, in comparison with the 1R/1R genotype, the 0R/0R genotype significantly increased the risk of breast cancer (OR 9.55, 95 % CI 1.19–76.64, P = 0.034). The 1R/3R genotype compared with 1R/1R genotype decreased the risk of breast cancer (Fisher’s exact test P = 0.015). There was no significant association between T-991C polymorphism of XRCC6 and risk of breast cancer.
To determine the effect(s) of XRCC5 and XRCC6 polymorphisms on age at diagnosis of breast cancer, the Kaplan–Meier survival analysis and Cox proportional hazards regression model were used. Mean age at diagnosis of breast cancer for 0, 1, 2, 3, and >4 repeat in XRCC5 were 39.2, 41.9, 44.3, 45.8, and 47.3 years, respectively. The Kaplan–Meier survival analysis revealed that number of repeat was associated with age at diagnosis of breast cancer (log rank statistic = 13.90, df = 4, P = 0.008; Fig. 1). This means that more number of VNTR is associated with higher age at diagnosis of breast cancer. However, the XRCC6 polymorphism was not associated with age at diagnosis of breast cancer (log rank statistic = 0.584, df = 2, P = 0.747).
Using Cox proportional hazards regression model, after adjustment for smoking habit, marital status, and the XRCC6 genotypes, there were significant associations between the number of repeats of XRCC5 and age at diagnosis of breast cancer. The 4 repeats (Hazard ratio, HR 0.229, 95 % CI 0.09–0.56, P = 0.002), 3 repeats (HR 0.216, 95 % CI 0.09–0.49, P < 0.001), 2 repeats (HR 0.231, 95 % CI 0.10–0.52, P < 0.001), and 1 repeat (HR 0.205, 95 % CI 0.08–0.47, P < 0.001) versus to the 0 repeat showed higher age at diagnosis of breast cancer.
There is strong evidence that some transcription factors can bind VNTR sequences [20]. The VNTR polymorphism of XRCC5 can alter the number of cis elements. The alleles with more tandem repeats include more Sp1 binding sites and could increase the affinity of Sp1 to the promoter of XRCC5 [10]. Our previous study indicated that the increase in the overall number of tandem repeats in the promoter region of XRCC5 down-regulates the gene expression [5].
The present study showed that 0R/0R genotype verses 1R/1R genotype is associated with increased risk of breast cancer. Also, we found that the 1R/3R vs 1R/1R genotypes decreased the risk of breast cancer (Table 1). It should be noted that based on the present study, the 2R/2R and 1R/2R genotypes did not alter the risk of breast cancer compared with the 1R/1R genotype (Table 1). It is self-evident that when 0R/0R and 1R/3R genotypes increased and decreased the risk of breast cancer compared with the 1R/1R genotype, respectively, it means that intermediate genotypes (such as 2R/2R and 1R/2R) may not alter the breast cancer risk. This finding is consistent with a report, indicating that fewer repeats of XRCC5 VNTR polymorphism is associated with increased risk of bladder cancer [10]. It should be mentioned that the over-expression of XRCC5 protein in several types of cancers have been reported previously [21–23]. Taken together, it might be suggested that in fewer tandem repeats, over-expression of XRCC5 leads to excess DNA repair, which interfere with normal apoptosis, thus increasing the likelihood for the development of breast cancer.
The strength of the results is tempered by small sample size, and this finding needs further replication in a larger sample. Considering the fact that ethnicity may influence the observed associations in multifactorial disease [24, 25], replication of this study in other countries is recommended.
References
Hakem R. DNA-damage repair; the good, the bad, and the ugly. EMBO J. 2008;27:589–605.
Hoeijmakers JH. DNA damage, aging, and cancer. N Engl J Med. 2009;361:1475–85.
Mohrenweiser HW, Jones IM. Variation in DNA repair is a factor in cancer susceptibility: a paradigm for the promises and perils of individual and population risk estimation? Mutat Res. 1998;400:15–24.
Fu YP, Yu JC, Cheng TC, Lou MA, Hsu GC, Wu CY, Chen ST, Wu HS, Wu PE, Shen CY. Breast cancer risk associated with genotypic polymorphism of the nonhomologous end-joining genes: a multigenic study on cancer susceptibility. Cancer Res. 2003;63:2440–6.
Dobbs TA, Tainer JA, Less-Miller SP. A structural model for regulation of NHEJ by DNA-PKcs autophosphorylation. DNA Repair. 2010;10:1307–14.
Park SJ, Ciccone SL, Freie B, Kurimasa A, Chen DJ, Li GC, Clapp DW, Lee SH. A positive role for the Ku complex in DNA replication following strand break damage in mammals. J Biol Chem. 2004;279:6046–55.
Kuschel B, Auranen A, McBride S, Novik KL, Antoniou A, Lipscombe JM, Day NE, Easton DF, Ponder BA, Pharoah PD, Dunning A. Variants in DNA double-strand break repair genes and breast cancer susceptibility. Hum Mol Genet. 2002;11:1399–407.
Rajaei M, Saadat I, Saadat M. Introducing a novel allele for the polymorphism of variable number of tandem repeats in the promoter region of XRCC5. Biochem Biophys Res Commun. 2012;427:503–5.
Rajaei M, Saadat I, Saadat M. The novel allele (3R) of the VNTR polymorphism in the XRCC5 promoter region dramatically decreases the gene expression. Biochem Biophys Res Commun. 2013;430:640–1.
Wang S, Wang M, Yin S, Fu G, Li C, Chen R, Li A, Zhou J, Zhang Z, Liu Q. A novel variable number of tandem repeats (VNTR) polymorphism containing Sp1 binding elements in the promoter of XRCC5 is a risk factor for human bladder cancer. Mutat Res. 2008;638:26–36.
Bau DT, Tseng HC, Wang CH, Chiu CF, Hua CH, Wu CN, Liang SY, Wang CL, Tsai CW, Tsai MH. Oral cancer and genetic polymorphism of DNA double strand break gene Ku70 in Taiwan. Oral Oncol. 2008;44:1047–51.
Yang MD, Wang HC, Chang WS, Tsai CW, Bau DT. Genetic polymorphisms of DNA double strand break gene Ku70 and gastric cancer in Taiwan. BMC Cancer. 2011;11:174–8.
Hsia TC, Liu CJ, Chu CC, Hang LW, Chang WS, Tsai CW, Wu CI, Lien CS, Liao WL, Ho CY, Bau DT. Association of DNA double-strand break gene XRCC6 genotypes and lung cancer in Taiwan. Anticancer Res. 2012;32:1015–20.
Chang WS, Ke HL, Tsai CW, Lien CS, Liao WL, Lin HH, Lee MH, Wu HC, Chang CH, Chen CC, Lee HZ, Bau DT. The role of XRCC6 T-991C functional polymorphism in renal cell carcinoma. Anticancer Res. 2012;32:3855–60.
Hsu CM, Yang MD, Chang WS, Jeng LB, Lee MH, Lu MC, Chang SC, Tsai CW, Tsai Y, Tsai FJ, Bau DT. The contribution of XRCC6/Ku70 to hepatocellular carcinoma in Taiwan. Anticancer Res. 2013;33:529–35.
Mohamadynejad P, Saadat M. Genetic polymorphisms of XRCC1 (at codons 194 and 399) in Shiraz population (Fars province, southern Iran). Mol Biol Rep. 2008;35:669–72.
Rafiee L, Saadat I, Saadat M. Glutathione S-transferase genetic polymorphisms (GSTM1, GSTT1 and GSTO2) in three Iranian populations. Mol Biol Rep. 2010;37:155–8.
Rajaei M, Saadat I, Saadat M. High resolution melting analysis for detection of variable number of tandem repeats polymorphism of XRCC5. Biochem Biophys Res Commun. 2012;425:398–400.
Tsai YY, Bau DT, Chiang CC, Cheng YW, Tseng SH, Tsai FJ. Pterygium and genetic polymorphism of DNA double strand break repair gene Ku70. Molecular Vis. 2007;13:1436–40.
Nakamura Y, Koyama K, Matsushima M. VNTR (variable number of tandem repeat) sequences as transcriptional, translational, or functional regulators. J Hum Genet. 1998;43:149–52.
Pucci S, Mazzarelli P, Rabitti C, Giai M, Gallucci M, Flammia G, Alcini A, Altomare V, Fazio VM. Tumor specific modulation of KU70/80 DNA binding activity in breast and bladder human tumor biopsies. Oncogene. 2001;20:739–47.
Lim JW, Kim H, Kim KH. Expression of Ku70 and Ku80 mediated by NF-kappa B and cyclooxygenase-2 is related to proliferation of human gastric cancer cells. J Biol Chem. 2002;277:46093–100.
Hosoi Y, Watanabe T, Nakagawa K, Matsumoto Y, Enomoto A, Morita A, Nagawa H, Suzuki N. Up-regulation of DNA-dependent protein kinase activity and Sp1 in colorectal cancer. Int J Oncol. 2004;25:461–8.
Saadat M, Ansari-Lari M. Polymorphism of XRCC1 (at codon 399) and susceptibility to breast cancer, a meta-analysis of the literatures. Breast Cancer Res Treat. 2009;115:137–44.
Saadat M. Haplotype analysis of XRCC1 (at codons 194 and 399) and susceptibility to breast cancer, a meta-analysis of the literatures. Breast Cancer Res Treat. 2010;124:785–91.
Acknowledgments
The authors are indebted to the participants for their close cooperation. The authors are indebted to Dr. Maryam Ansari-Lari for critical reading of the manuscript and for her contribution in discussion. This study was supported by Shiraz University.
Conflict of interest
No conflicts of interest exist.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rajaei, M., Saadat, I., Omidvari, S. et al. Association between polymorphisms at promoters of XRCC5 and XRCC6 genes and risk of breast cancer. Med Oncol 31, 885 (2014). https://doi.org/10.1007/s12032-014-0885-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-014-0885-8