Abstract
For bladder cancer, a new diagnostic marker is needed to avoid painful cystoscopy. The aim of this study was to explore the efficacy of urinary miRNA-96 as molecular marker in bladder cancer diagnosis and its relation to bilharziasis. Urine cytology, serologic assessment of schistosomiasis and estimation of miRNA-96 by real-time PCR were carried out for 94 bladder cancer patients, 30 benign bladder lesions and 60 healthy individuals. Expression of miRNA-96 showed a significant difference among the three tested groups and also between benign and malignant bilharzial cases. Urinary miRNA-96 is a good noninvasive diagnostic biomarker for bladder cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The standard care for bladder cancer diagnosis and follow-up is through the combination of cystoscopic examination, histology and cytology [17]. However, these methods have a significant poor sensitivity for low-grade, well-differentiated lesions and high financial cost. They are also highly subjective testes and provide little about the molecular characteristics of cancers [12].
Recently, numerous urinary markers have been under study as noninvasive tests to reduce the frequency and cost of cystoscopy. An ideal test for the bladder tumors detection should have high specificity and sensitivity; moreover, it is necessary to be objective, rapid, accurate and easy to administer [1].
In the ongoing search for new markers to improve the bladder cancer diagnosis, microRNAs (miRNAs) may thus serve as biomarkers for early detection of bladder cancer. miRNAs are small endogenous noncoding RNAs that play crucial roles in multiple biological processes through regulating translational repression or cleavage of mRNAs. Recent studies have documented that miRNAs acted as tumor suppressors or oncogenes in a variety types of cancer, such as lung, hepatic, breast and pancreatic cancer [10, 11, 13, 16, 23, 24].
Several miRNAs properties make them attractive as potential biomarkers. miRNAs can be easily detected in small amount of samples using sensitive and specific real-time quantitative PCR (qPCR). miRNAs can be detectable in bodily fluids including serum, plasma, urine, saliva and tears and are stable against degradation [2, 9]. Furthermore, expression profiles of miRNAs would be changed in the serum and/or plasma of cancer patients and miRNAs have been shown to be released from tumor cells to the circulation. So circulating miRNAs will be a novel class of noninvasive biomarkers for cancer diagnosis and prognosis [19].
miR-96 have been found to be upregulated in various human cancers such as breast, lung, liver, colon, prostate, ovary, testis cancer and lymphoma [22]. These results implied that miR-96 is an onco-miRNA and might be a potential target of gene therapy of some human cancers. There are good results whereby expressions of miR-96 in urine was well correlated with tumor grade and stage; this miRNA is thus promising diagnostic tumor markers to distinguish BC patients from non-BC patients [21].
The aim of the present study was to evaluate miRNA-96 usefulness as a urine molecular marker for bladder cancer detection. Urinary miR-96 level was measured using real-time polymerase chain reaction (qPCR) in cellular pellets from a large number of voided urine samples, especially those with bilharziasis in comparison with urine cytology.
Materials and methods
Subjects
Approval of the study was conducted by ethical committee of faculty of medicine, Ain Shams University. This study included 184 Egyptian individuals: 124 inpatients were selected from Urology Department, Ain Shams University, Faculty of Medicine, Egypt, and 60 healthy normal volunteers were enrolled as a control group after obtaining informed consent.
Patients to be enrolled in the study must presented with chronic irritative voiding symptoms or hematuria and cystoscopic evidence confirmed by biopsy consistent with proven bladder carcinoma (in situ, low grade or high grade). Participants were excluded from the study if they had a past history of occupational exposure to known bladder carcinogens, bladder cancer or another urological malignancy within the past 5 years, received chemotherapy and radiation therapy.
According to histopathological examination of cystoscopy biopsies, 124 patients included in the study were classified into malignant and benign groups. The malignant group included 94 patients (mean age 58.8 ± 11.6 years; range 28–85 years) and benign group (n = 30, mean age 57.1 ± 17.4 years; range 27–82 years). From the laboratory staff, 60 healthy normal volunteers (mean age 34.3 ± 14.5 years; range 11–65 years) were recruited with matched age, sex and smoking status of patients. Of this malignant group, 52 were diagnosed by histopathology as transitional cell carcinoma (TCC), 33 cases as squamous cell carcinoma (SCC) and 9 as others types (Table 1). Tumor grading and staging were determined according to World Health Organization and TNM classification [4, 18].
Collection of samples
Voided urine (30–60 mL) and sera (1 mL) samples were obtained from all individuals before any surgical or other therapeutic intervention. Urine samples were collected using the urine collection cup that measures volume, sealed immediately and placed on ice and then centrifuged at 2,500–4,000×g for 15–20 min. The urinary pellets were washed twice with phosphate-buffered saline (PBS) at pH 7.0. A portion of the pellet was used for microscopic and cytological examination [15], and the other portion was treated with 0.15 ml of RNA later, RNA Stabilization Reagent (Qiagen, Valencia, CA, USA), and stored at −80 °C for further processing to extract miRNAs.
Schistosomiasis antibodies detection in serum
Assessment of bilharzial infestation was done by schistosomiasis antibodies detection in sera via indirect haemagglutination test, using the Cellognost Schistosomiasis H kit (Dade Behring Marburg GmbH, Marburg, Germany) [5].
Extraction of miRNA from urine pellet samples
Total miRNAs were extracted from all urine sediments using miRNeasy Mini Kit (Qiagen, Germany, Cat. no. 217004). The miRNeasy Mini Kit combines phenol-/guanidine-based lysis of samples with silica-membrane-based purification of total miRNAs according to the manufacturer’s instructions. Then, total miRNAs were treated with RNase inhibitors and kept at −80 °C until its use in the reverse transcription quantitative polymerase chain reaction (RT-qPCR) for detection of miRNA-96 expression. The concentrations of RNA were determined spectrophotometrically.
Detection of miRNA-96 by reverse transcription quantitative polymerase chain reaction (RT-qPCR)
First, the cDNA was synthesized from the total miRNAs of the urine pellets using Miscript II RT kit in accordance with the manufacturer’s recommended protocol (Qiagen, Germany). The resultant cDNA was subjected to real-time quantitative polymerase chain reaction (qPCR) using miScript SYBR® Green PCR Kit with miScript Primer assays (Qiagen, Germany). This kit includes QuantiTect SYBR Green PCR Master Mix and the miScript Universal Primer (reverse primer that use to detect miRNAs) in combination with a miScript Primer Assay (Cat. no. MS00003360) that specifically recognizes the targeted miRNA. The real-time qPCR was performed on a Step One Plus™ System (Applied Biosystems Inc, Foster, CA). The PCR conditions were as follows: 95 °C for 15 min, then 94 °C for 15 s, 55 °C for 30 s and 70 °C for 30 s for 40 cycles. The data were normalized using the endogenous RNU U6 as reference control. The threshold cycle (Ct) value of each sample was calculated with Step One Plus™ software v2.2.2 (Applied Biosystems), and the 2−ΔΔCT method was used in the analysis of PCR data for relative quantification of miRNA-96.
Statistical methods
Data analyses were performed using Chi-square and nonparametric tests, and the level of significance was determined to be less than 0.05. The threshold value for optimal sensitivity and specificity of miRNA-96 was determined by receiver operating characteristics (ROC) curve. All analyses were performed using Statistical Package for the Social Sciences software (SPSS Inc, Chicago, IL, USA).
Results
The present study included 184 subjects. Ninety-four bladder cancer: 35 of them were bilharzial and 59 were non-bilharzial. Thirty patients with benign urological diseases: 12 of them with bilharzial lesions and 18 cases without. All 60 normal volunteers were without bilharziasis (Table 1).
Urinary miRNA-96 level in investigated groups
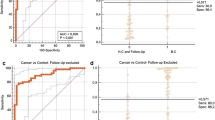
In urine sediment cells, miRNA-96 expression was measured using real-time qPCR and normalized to RNU 6 as reference control. As shown in Table 2, the expression level of miR-96 was significantly upregulated in the bladder cancer group (mean rank = 121.49), in comparison with the normal and benign groups (mean rank = 55.02 and 76.63, respectively, P < 0.001) using the RQ values. The best cutoff point for miRNA-96 using the ROC curve was 1.63 (Fig. 1). Using this cutoff value, 68 out of 94 (72.3 %) malignant patients, 8 out of 30 (26.7 %) benign patients and 2 out of 60 (3.3 %) normal individuals were miRNA-96 positive (P < 0.001), as shown in Table 2.
ROC curve analysis for miRNA-96 to calculate the best cutoff point that discriminates between malignant and non-malignant groups. Area under the curve = 0.822 and standard error = 0.033. The best cutoff point of miRNA-96 was 1.63, 95 % confidence limits range = 0.758–0.887, sensitivity = 72.3 % and specificity = 88.9 % and P < 0.001
Positive urine cytology results were reported in 6.7 % of benign cases and in 34 % of malignant cases (Table 2). Bilharziasis was found in 40 % of benign cases and 37.2 % of malignant cases (Table 2). There was a significant difference in urinary miRNA-96 expression between benign and malignant bilharzial cases (Table 3).
Relation between urinary miRNA-96 and different clinicopathological factors in the malignant group
No significant difference was detected between miRNA-96 expression and any of the studied clinicopathological factors in the malignant group (P > 0.05) as shown in Table 4.
Overall sensitivity, specificity, PPV, NPV and accuracy of urine miRNA-96
When miRNA-96 was tested independently using real-time qPCR, it showed the highest sensitivity and specificity (72.3, 88.9 %) even in low-grade, early-stage or bilharzial bladder cancer (Table 5). Moreover, the sensitivity of urine cytology (34 %) when combined with miRNA-96 was improved to 79.8 %.
Discussion
For bladder cancer, a new diagnostic marker has been under study in order to reduce the cost and the frequency of cystoscopy or replace them by noninvasive tests during the initial diagnosis and follow-up period. In recent years, the aberrant expression of miRNAs in bladder cancer has been studied. Some miRNAs have been reported to be upregulated in tissues of bladder cancer. For example, miR-129 was the most commonly upregulated and its upregulation was associated with poor outcome [3].
In the ongoing search for new markers to improve the bladder cancer diagnosis, the efficacy of urinary miRNA-96 in diagnosis of bladder cancer and its relation to bilharziasis was performed in this study.
In the current study, real-time qPCR was used to detect the expression level of miRNA-96 in 184 voided urine samples collected from patients with different types of bladder cancer (n = 94), benign bladder lesions (n = 30) and normal volunteers (n = 60). The expression level of miR-96 was significantly upregulated in bladder cancer group (mean rank = 121.49), in comparison with the normal and benign groups (mean rank = 55.02 and 76.63, respectively, P < 0.001) using the RQ values. Upregulation of miR-96 in TCC tumorigenesis is one of the mechanisms of repression of transcription factors (FOXO) of Forkhead Box O subfamily, which is a tumor-suppressor gene-causing G1 cell cycle arrest and cell death ([6, 14]). Also, hsa-miR-96 by upregulating MAP4K1 and insulin receptor substrate 1 (IRS1) levels may affect the growth of bladder cancer cells [20].
Yamada et al. [21] and Han et al. [7] identified a great number of miRNAs that were significantly upregulated in bladder cancer group using miRNA qRT-PCR and microarray profiling. Yamada et al. [21] observed that miR-96 was significantly higher expressed in urine of 100 bladder cancer group than in healthy controls (miR-96, P = 0.0059) and significantly correlated with tumor grade and stage. Han et al. [7] also revealed that miR-96 was the most significantly upregulated miRNA in bladder cancer group. This miRNA, therefore, can be regarded as a promising diagnostic marker in bladder cancer. In addition, expression of this miRNA decreased significantly after radical surgery, suggesting that it can be used also as a prognostic molecular marker of cancer recurrence [21].
Yan et al. [22] and Wang et al. [20] observed that miR-96 expression was higher in bladder carcinoma compared with normal bladder tissues using northern blot analysis and quantitative real-time PCR (qRT-PCR). Also, miR-96 expression in the superficial bladder tumors was lower than in invasive tumors and significantly related to the clinical stages of bladder carcinoma and the pathological types. These results revealed that miR-96 maybe plays a role in the process of development, occurrence and infiltration of bladder carcinomas [20].
This is the first study to investigate miRNA-96 expression in bilharzial bladder cancer. Interestingly, 27 out of 35 bilharzial malignant group showed positive miRNA-96 with no statistical significance between them (P > 0.05). While in bilharzial benign group, 4 out of 12 showed positive miRNA-96 with also no statistical significance between them (P > 0.05). This study is among the first to investigate urinary miRNA-96 in bilharzial bladder cancer. There was significant difference between malignant and benign bilharzial cases regarding urinary miRNA-96, indicating a potential role for bilharziasis in the aberrant expression of urinary miRNA-96 underlying carcinogenesis.
An ideal urine biomarker for bladder cancer diagnosis should have high positive predictive value (PPV) and sensitivity [8]. In this study, miRNA-96 showed high sensitivity and specificity even in low-grade, early-stage or bilharzial bladder cancer than that of cytology (Table 4). Accordingly, urinary miRNA-96 was superior to urine cytology for bladder cancer diagnosis. Moreover, the sensitivity of urine cytology was improved when combined with miRNA-96 detected by RT-qPCR.
In conclusion, the results of this study revealed that miRNA-96 expression level in urine sample is a potentially useful urinary biomarker for early diagnosis of bladder cancer including bilharzial bladder cancer and it improves the sensitivity of urine cytology.
References
Avritscher EB, Cooksley CD, Grossman HB, Sabichi AL, Hamblin L, Dinney CP, Elting LS. Clinical model of lifetime cost of treating bladder cancer and associated complications. Urology. 2006;68:549–53.
Cortez MA, Bueso-Ramos CB, Ferdin J, Lopez-Berestein G, Sood AK, Calin GA. MicroRNAs in body fluids the mix of hormones and biomarkers. Nat Rev Clin Oncol. 2011;8:467–77.
Dyrskjøt L, Ostenfeld MS, Bramsen JB, Silahtaroglu AN, Lamy P, Ramanathan R, Fristrup N, Jensen JL, Andersen CL, Zieger K, Kauppinen S, Ulhøi BP, Kjems J, Borre M, Orntoft TF. Genomic profiling of microRNAs in bladder cancer: miR-129 is associated with poor outcome and promotes cell death in vitro. Cancer Res. 2010;69:4851–60.
Edge SB, Byrd DR, Compton CC. AJCC cancer staging manual. 7th ed. New York: Springer; 2010. p. 497–505.
Gui M, Idris MA, Shi YE, Muhling A, Ruppel A. Reactivity of Schistosoma japonicum and S. mansoni antigen preparations in indirect haemagglutination (IHA) with sera of patients with homologous and heterogonous schistosomiasis. Ann Trop Med Parasitol. 1991;85:599–604.
Guttilla IK, White BA. Coordinate regulation of FOXO1 by miR-27a, miR-96, and miR-182 in breast cancer cells. J Biol Chem. 2009;284:23204–16.
Han Y, Chen J, Zhao X, Liang C, Wang Y, Sun L, Jiang Z, Zhang Z, Yang R, Chen J. MicroRNA expression signatures of bladder cancer revealed by deep sequencing. PLoS One. 2011;6(3):e18286.
Konety BR. Molecular markers in bladder cancer: a critical appraisal. Urol Oncol. 2006;24(4):326–37.
Kosaka N, Iguchi H, Ochiya T. Circulating microRNA in body fluid: a new potential biomarker for cancer diagnosis and prognosis. Cancer Sci. 2010;101:2087–92.
Lee EJ, Gusev Y, Jiang J, Nuovo GJ, Lerner MR, Frankel WL, Morgan DL, Postier RG, Brackett DJ, Schmittgen TD. Expression profiling identifies microRNA signature in pancreatic cancer. Int J Cancer. 2007;120:1046–54.
Liu C, Yu J, Yu S, Lavker RM, Cai L, Liu W, Yang K, He X, Chen S. MicroRNA-21 acts as an oncomir through multiple targets in human hepatocellular carcinoma. J Hepatol. 2010;53:98–107.
Lotan Y, Svatek RS, Sagalowsky AI. Should we screen for bladder cancer in a high-risk population: a cost per life-year saved analysis. Cancer. 2006;107:982–90.
Murakami Y, Yasuda T, Saigo K, Urashima T, Toyoda H, Okanoue T, Shimotohno K. Comprehensive analysis of microRNA expression patterns in hepatocellular carcinoma and non-tumorous tissues. Oncogene. 2006;25:2537–45.
Myatt SS, Wang J, Monteiro LJ. Definition of microRNAs that repress expression of the tumor suppressor gene FOXO1 in endometrial cancer. Cancer Res. 2010;70:367–77.
Papanicolaou GN, Marshall VF. Urine sediment smears as a diagnostic procedure in cancers of the urinary tract. Science. 1945;101:519.
Pedriali M, Fabbri M, Campiglio M, Ménard S, Palazzo JP, Rosenberg A, Musiani P, Volinia S, Nenci I, Calin GA, Querzoli P, Negrini M, Croce CM. MicroRNA gene expression deregulation in human breast cancer. Cancer Res. 2005;65:7065–70.
Proctor I, Stoeber K, Williams GH. Biomarkers in bladder cancer. Histopathology. 2010;57:1–13.
Sobin LH, Wittkind CH. UICC TNM classification of malignant tumors. 5th ed. New York: Wiley Liss; 1997. p. 187.
Taylor DD, Gercel-Taylor C. MicroRNA signatures of tumor-derived exosomes as diagnostic biomarkers of ovarian cancer. Gynecol Oncol. 2008;110:13–21.
Wang Y, Luo H, Li Y, Chen T, Wu S, Yang L. hsa-miR-96 up-regulates (MAP4K1 and IRS1 and may function as a promising diagnostic marker in human bladder urothelial carcinomas. Mol Med Rep. 2012;5(1):260–5.
Yamada Y, Enokida H, Kojima S, Kawakami K, Chiyomaru T, Tatarano S, Yoshino H, Kawahara K, Nishiyama K, Seki N, Nakagawa M. MiR-96 and miR-183 detection in urine serve as potential tumor markers of urothelial carcinoma: correlation with stage and grade, and comparison with urinary cytology. Cancer Sci. 2011;102:522–9.
Yan B, Fu Q, Lai L, Tao X, Fei Y, Shen J, Chen Z, Wang Q. Down regulation of microRNA 99a in oral squamous cell carcinomas contributes to the growth and survival of oral cancer cells. Mol Med Rep. 2012;6:675–81.
Yanaihara N, Caplen N, Bowman E, Seike M, Kumamoto K, Yi M, Stephens RM, Okamoto A, Yokota J, Tanaka T, Calin GA, Liu CG, Croce CM, Harris CC. Unique microRNA molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell. 2006;9:189–98.
Yu S, Lu Z, Liu C, Meng Y, Ma Y, Zhao W, Liu J, Yu J, Chen J. miRNA-96 suppresses KRAS and functions as a tumor suppressor gene in pancreatic cancer. Cancer Res. 2010;70:6015–25.
Acknowledgments
This work was supported by Ain Shams University Research Projects 2013-14. The authors acknowledge Prof. Nahla Awad, at the early cancer detection unit, Faculty of Medicine, Ain Shams University, for her help in the cytological examinations of urine samples and Dr. Yosif Kotb at Urology department.
Conflict of interest
All authors declare nothing to disclose.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Eissa, S., Habib, H., Ali, E. et al. Evaluation of urinary miRNA-96 as a potential biomarker for bladder cancer diagnosis. Med Oncol 32, 413 (2015). https://doi.org/10.1007/s12032-014-0413-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-014-0413-x