Abstract
Dysregulation of microRNA-100 (miR-100) has been shown to be involved in cancer tumorigenesis and progression of several cancer types. However, its expression patterns in tumors are controversial. The aim of this study was to investigate the expression and clinical significance of miR-100 in colorectal cancer (CRC). Quantitative real-time PCR was used to analyze the expression of miR-100 in 138 pairs of human CRC and adjacent normal tissues. The prognostic values of miR-100 in CRC were also analyzed. The results showed that the miR-100 expression was significantly downregulated in CRC tissues when compared to adjacent normal tissues (P < 0.001). Also, low miR-100 expression was observed to be significantly correlated with larger tumor size (P = 0.023), higher incidence of lymph node metastasis (P = 0.009), and advanced TNM stage (P = 0.016). More importantly, Kaplan–Meier analysis showed that CRC patients with low miR-100 expression tended to have shorter overall survival. In multivariate analysis stratified for known prognostic variables, low miR-100 expression was identified as an independent prognostic factor for overall survival. In conclusion, our data indicated for the first time that the downregulation of miR-100 was associated with advanced clinical features and poor prognosis of CRC patients, suggesting that miR-100 downregulation may serve as an unfavorable prognostic biomarker in CRC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Colorectal cancer (CRC) is one of the leading causes of cancer-related death worldwide, with high possibilities of recurrence and metastasis [1]. The prognosis for CRC patients has shown little improvement despite advances in treatment approaches over the last decades. Consequently, there is an ongoing effort to identify novel and effective targets to improve therapeutic efficacy and clinical outcome for CRC.
MicroRNAs (miRNAs) are a class of small non-coding RNA molecules that regulate gene expression by binding to the 3′ untranslated regions (3′ UTR) of target miRNAs, repressing miRNA translation or cleaving target miRNA [2, 3]. As master regulators of gene expression, miRNAs are involved in modulating diverse biological processes, such as metabolism, survival, differentiation, and apoptosis [2]. The deregulation of expression of miRNAs has been shown to contribute to the multistep processes of carcinogenesis in human by modification of either oncogenic or tumor suppressor function [4]. Moreover, accumulating evidence indicated that miRNAs are potential biomarkers of diagnosis, treatment, and outcomes, which are highly tissue specific for cancer patients [5–9].
Dysregulated expression of miR-100 has been found in various types of cancers. MiR-100 can act as either oncogene or tumor suppressor in different cancers. For example, it has been demonstrated to be involved in nasopharyngeal cancer [10], bladder cancer [11], epithelial ovarian cancer [12], and hepatocellular carcinoma [13] by deregulation of its target genes as a tumor suppressor; it has also been found to promote tumorigenesis in renal cell carcinoma [14] and acute myeloid leukemia [15] as an oncogene. Recently, Peng et al. [16] reported that the enforced expression of miR-100 could markedly suppresses the proliferation of CRC cells by targeting RAP1B, indicating that downregulation of miR-100 might play an important role during CRC progression. However, the clinical significance of miR-100 in CRC has not yet been elucidated. In the present study, we investigated the expression level of miR-100 in human CRC tissues and analyzed its correlation of miR-100 expression with clinicopathological factors of patients. The prognostic value of miR-100 expression in CRC was also analyzed.
Materials and methods
Patients and tissue samples
Human CRC tissues (n = 138) and adjacent normal tissues (n = 138) were obtained through the Department of General Surgery at Zaozhuang Mining Group Central Hospital (Zaozhuang, China), between June 2005 and November 2008. None of the patients recruited in this study received preoperative treatment, such as radiotherapy or chemotherapy. The tissue specimens were immediately frozen in liquid nitrogen after surgical removal and stored at −80 °C until analysis. The diagnoses were confirmed by pathological findings. All tumors were classified according to the seventh edition of the UICC TNM staging system for CRC. Written informed consent was obtained from each patient, and research protocols were approved by the Ethical Committee of Zaozhuang Mining Group Central Hospital.
All CRC patients have been followed up at intervals of 3 months during the first 2 years and 6 months up to the fifth year, and the date of latest record retrieved was December 31, 2013. Overall survival was calculated from the date of the initial surgical operation to the date of death or last date of follow-up.
RNA isolation
Total RNA was extracted from tissue samples using TRIzol (Invitrogen, Carlsbad, CA) according to the manufacturer’s protocol. The concentration and purity of RNA were measured spectrophotometrically at 260 and 280 nm, whereas RNA integrity was evaluated by agarose gel electrophoresis. Only the samples with the OD A260/A280 ratio close to value of 2.0, which indicates that the RNA is pure, were subsequently analyzed.
MiRNA detection by real-time quantitative RT-PCR (qRT-PCR)
cDNA were generated from 1 μl total RNA using One Step PrimeScript miRNA cDNA Synthesis Kit (Takara, Japan) according to the manufacture’s instructions. Real-time PCR was performed using SYBR Premix Ex TaqTM II (Takara, Japan) on an ABI 7500 Real-Time PCR System (Applied Biosystems, USA) with the amplification steps as follows: 95 °C for 10 s, and 40 cycles at 95 °C for 5 s and 60 °C for 34 s. At the end of the PCR cycles, melting curve analyses were performed in order to validate the specific generation of the expected PCR product. Each sample was run in duplicates for analysis. The expression levels of miR-100 were normalized to U6 small nuclear RNA (U6) and were calculated utilizing the 2−ΔΔC method [17].
Statistical analysis
The Kruskal–Wallis test or the Mann–Whitney U test was performed to compare miRNAs levels between groups. The chi-square or Fisher’s exact probability test was used to examine possible correlations between miR-100 expression and clinical features. Survival curves were constructed with the Kaplan–Meier method and compared by the log-rank test. The influence of each variable on survival was examined by the Cox multivariate regression analysis. All statistical analyses were carried out by using SPSS 13.0 software (SPSS Inc., IL, USA), and a P < 0.05 was considered statistically significant.
Results
MiR-100 expression in CRC patients
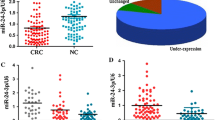
MiR-100 expression was detected in 138 pairs of CRC tissues and adjacent normal tissues by qRT-PCR. Expression of miR-100 was normalized with U6, and the values obtained were compared. The results showed that miR-100 expression in human CRC tissues was significantly lower than in adjacent normal tissues (Fig. 1a, P < 0.001). In addition, the miR-100 expression was found to be decreased at least twofold compared with adjacent normal tissues in 57.2 % (79/138) of CRC cases (Fig. 1b). The median expression level of miR-100 (1.26) was used as a cutoff point to divide all 138 patients into two groups: CRC patients expressing miR-100 at levels less than the cutoff value were assigned to the low expression group (median expression value 0.61, n = 69), and those samples with expression equal or above the cutoff value were assigned to the high expression group (median expression value 2.13, n = 69).
MiR-100 expression is downregulated in CRC tissues than adjacent normal tissues (P < 0.001). a MiR-100 expression is examined using qRT-PCR in 138 pairs of human CRC tissues (Cancer) and adjacent normal tissues (Normal), and its expression is normalized to the level of U6 in each sample. Data are shown separately in human samples. b Fold changes of miR-100 of each individual paired sample. The data are represented as log2-fold change (Cancer/Normal), which was defined as >1 (overexpression) and, <1 (underexpression); the remaining fold changes are defined as unchanged
Association of miR-100 expression with clinicopathological characteristics of CRC patients
Table 1 summarized the association between miR-100 expression and clinicopathological characteristics of CRC patients. Low miR-100 expression was observed to be closely correlated with larger tumor size (P = 0.023), higher incidence of lymph node metastasis (P = 0.009), and advanced TNM stage (P = 0.016). In contrast, there was no association between miR-100 expression and other clinical factors, such as gender, age, tumor location, local invasion, distant metastasis, and histology grade (all P > 0.05).
Association of miR-100 expression with prognosis of CRC patients
The prognostic value of miR-100 expression was investigated using the Kaplan–Meier method and log-rank test. As shown in Fig. 2, there was a significant correlation between miR-100 expression and 5-year overall survival of CRC patients (P = 0.006, log-rank test). The cumulative 5-year overall survival rate of CRC patients with low miR-100 expression was significantly lower than that of those patients with high miR-100 expression. Univariate Cox proportional hazards regressions model analysis demonstrated that local invasion, lymph node metastasis, distant metastasis, TNM stage, histology grade, and low miR-100 expression were statistically significant risk factors affecting the overall survival of patients with CRC (Table 2). No significant associations were found for gender, age at diagnosis, location, tumor size, and patient outcome. Multivariate analysis using the Cox proportional hazard model for all variables that were significant in the univariate analysis confirmed that the status of lymph node metastasis (P = 0.031), distant metastasis (P = 0.028), and the level of miR-100 expression (P = 0.017) were independent prognostic factors for patients with CRC (Table 2). These findings imply that miR-100 affects the prognosis of CRC patients and that low miR-100 expression might be a poor prognostic factor.
Discussion
CRC remains one of the leading causes of cancer-related death worldwide4, so finding new molecular targets for its diagnosis, prognosis, and treatment has the potential to improve the clinical strategy and outcome of this disease [18, 19]. In this retrospective study of 138 patients with newly diagnosed CRC, there are three significant findings according to our results: Firstly, miR-100 expression in CRC tissues was significantly lower than that in adjacent normal tissues; the reduced expression of miR-100 was significantly associated with the advanced clinical features of CRC patients; and both univariate and multivariate analyses revealed that the expression level of miR-100 was a significant risk factor affecting the overall survival of patients with CRC. These results suggest that miR-100 expression could be a valuable biomarker of the progression and the prognosis in CRC. To the best of our knowledge, this is the first study to analyze the expression patterns and clinical significance of miR-100 in a large number of CRC patients.
MiRNAs are small endogenous non-coding RNAs that bind to partially complementary recognition sequences of target mRNAs, causing either degradation or preventing their translation [2, 3]. Since their discovery in 1993, altered expressions of miRNAs have been associated with a variety of human diseases, including cancer [5, 20]. To our interests, recent studies have witnessed dysregulation of multiple miRNAs in CRC. For example, Asangani et al. [21] reported that miR-21 expression was significantly increased in CRC tissues and cell lines and that miR-21 contributed to the malignant potential such as cell proliferation, migration, and invasion in CRC. CRC patients with high miR-21 expression tended to have a poorer prognosis than those with low miR-21 expression [22]. Chen et al. [23] found that miR-103 targets the known metastasis suppressors death-associated protein kinase and Kruppel-like factor four in CRC cells, resulting in increased cell motility and cell–matrix adhesion. Overexpression of miR-106a could increase CRC cell migration and invasion by inhibiting transforming growth factor-β receptor [24]. Zhang et al. [25] also reported that miR-143 was downregulated in CRC tissues and cell lines and that enforced expression of miR-143 inhibited cell invasion and migration of CRC cells. These findings suggested that deregulation of miRNAs may play a vital role in the tumorigenesis and progression of CRC. More extensive investigations are required to elucidate the roles of miRNAs in the development of CRC so as to identify those miRNAs that may serve as novel prognosis predictor or as therapeutic targets for CRC.
MiR-100, as a potential tumor related miRNA, has been reported to be involved in tumorigenesis and tumor development [26]. Notably, the roles of miR-100 in different cancers are quite contradictory as it can behave either as an oncogene or a tumor suppressor gene, depending on the tumor type examined. For example, Shi et al. [10] reported that underexpressed miR-100 leads to Plk1 overexpression, which in turn contributes to nasopharyngeal cancer progression. Oliveira et al. [11] also confirmed that miRNA-100 acts as a tumor suppressor in human bladder carcinoma 5,637 cells. Peng et al. [12] demonstrated that low miR-100 expression may be an independent poor prognostic factor for human epithelial ovarian cancer. Huang et al. [27] found that downregulation of miR-100 was significantly associated with advanced FIGO stage, the presence of lymph node metastasis, and reduced survival of small-cell carcinoma of the cervix patients. In addition to these, the downregulation of miR-100 was also shown in hepatocellular carcinoma [13], oral cancer [28], and adrenocortical cancer [29]. In contrast, an upregulation of miR-100 has been observed in renal cell carcinoma, acute myeloid leukemia, and prostate cancer [14, 26, 30]. In particular, in CRC, the previous study of Peng et al. [16] found the decreased expression of miR-100 in CRC cells. In line with this finding, our data also validated the downregulation of miR-100 in a large number of clinical CRC cases. In addition, we found that the decreased expression of miR-100 was significantly associated with larger tumor size, higher incidence of lymph node metastasis, and advanced TNM stage, which raises the possibility that miR-100 might have an important role in the development or pathogenesis of CRC. More importantly, we proved that miR-100 expression was significantly associated with overall survival of patients with CRC. In support of this, Kaplan–Meier analysis of overall survival showed that patients whose tumors had low miR-100 expression tend to have a significantly worse overall survival than those with high miR-100 expression, indicating that low miR-100 level is a marker of poor prognosis for patients with CRC. Cox proportional hazards model adjusted for known prognostic variables such as lymph node metastasis and distant metastasis proved that miR-100 was an independent prognostic marker for CRC. Thus, miR-100 could be used as a molecular prognostic marker additive to the known prognostic indicator, identifying patients who are more likely to have higher risk of death, thus, should receive more aggressive treatment. Of course, as the size of tissue samples in this study is still small, multicenter, randomized studies should be necessary to testify the prognostic values of miR-100 expression.
In conclusion, this is the first report demonstrating that the downregulation of miR-100 was associated with advanced clinical features and poor prognosis of CRC patients, suggesting that miR-100 downregulation may be used as an unfavorable prognostic biomarker in CRC. Further studies are needed to investigate the precise molecular mechanism of miR-100 in the tumorigenesis and progression of CRC and illustrate whether miR-100 may be used as a potential therapeutic target for the treatment of CRC patients.
References
Siegel R, Naishadham D, Jemal A. Cancer statistics, 2013. CA Cancer J Clin. 2013;63(1):11–30. doi:10.3322/caac.21166.
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116(2):281–97.
He L, Hannon GJ. MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet. 2004;5(7):522–31. doi:10.1038/nrg1379.
Zhang B, Pan X, Cobb GP, Anderson TA. microRNAs as oncogenes and tumor suppressors. Dev Biol. 2007;302(1):1–12. doi:10.1016/j.ydbio.2006.08.028.
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, et al. MicroRNA expression profiles classify human cancers. Nature. 2005;435(7043):834–8. doi:10.1038/nature03702.
Garzon R, Marcucci G, Croce CM. Targeting microRNAs in cancer: rationale, strategies and challenges. Nat Rev Drug Discov. 2010;9(10):775–89. doi:10.1038/nrd3179.
Zhou XJ, Dong ZG, Yang YM, Du LT, Zhang X, Wang CX. Limited diagnostic value of microRNAs for detecting colorectal cancer: a meta-analysis. Asian Pac J Cancer Prev. 2013;14(8):4699–704.
Nair VS, Maeda LS, Ioannidis JP. Clinical outcome prediction by microRNAs in human cancer: a systematic review. J Natl Cancer Inst. 2012;104(7):528–40. doi:10.1093/jnci/djs027.
Ma XL, Liu L, Liu XX, Li Y, Deng L, Xiao ZL, et al. Prognostic role of microRNA-21 in non-small cell lung cancer: a meta-analysis. Asian Pac J Cancer Prev. 2012;13(5):2329–34.
Shi W, Alajez NM, Bastianutto C, Hui AB, Mocanu JD, Ito E, et al. Significance of Plk1 regulation by miR-100 in human nasopharyngeal cancer. Int J Cancer. 2010;126(9):2036–48. doi:10.1002/ijc.24880.
Oliveira JC, Brassesco MS, Morales AG, Pezuk JA, Fedatto PF, da Silva GN, et al. MicroRNA-100 acts as a tumor suppressor in human bladder carcinoma 5637 cells. Asian Pac J Cancer Prev. 2011;12(11):3001–4.
Peng DX, Luo M, Qiu LW, He YL, Wang XF. Prognostic implications of microRNA-100 and its functional roles in human epithelial ovarian cancer. Oncol Rep. 2012;27(4):1238–44. doi:10.3892/or.2012.1625.
Cairo S, Wang Y, de Reynies A, Duroure K, Dahan J, Redon MJ, et al. Stem cell-like micro-RNA signature driven by Myc in aggressive liver cancer. Proc Natl Acad Sci USA. 2010;107(47):20471–6. doi:10.1073/pnas.1009009107.
Wang G, Chen L, Meng J, Chen M, Zhuang L, Zhang L. Overexpression of microRNA-100 predicts an unfavorable prognosis in renal cell carcinoma. Int Urol Nephrol. 2013;45(2):373–9. doi:10.1007/s11255-012-0374-y.
Zheng YS, Zhang H, Zhang XJ, Feng DD, Luo XQ, Zeng CW, et al. MiR-100 regulates cell differentiation and survival by targeting RBSP3, a phosphatase-like tumor suppressor in acute myeloid leukemia. Oncogene. 2012;31(1):80–92. doi:10.1038/onc.2011.208.
Peng H, Luo J, Hao H, Hu J, Xie SK, Ren D, et al. MicroRNA-100 regulates SW620 colorectal cancer cell proliferation and invasion by targeting RAP1B. Oncol Rep. 2014;31(5):2055–62. doi:10.3892/or.2014.3075.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) method. Methods. 2001;25(4):402–8. doi:10.1006/meth.2001.1262.
Woolf SH. The best screening test for colorectal cancer—a personal choice. N Engl J Med. 2000;343(22):1641–3. doi:10.1056/nejm200011303432211.
Weitz J, Koch M, Debus J, Hohler T, Galle PR, Buchler MW. Colorectal cancer. Lancet. 2005;365(9454):153–65. doi:10.1016/s0140-6736(05)17706-x.
Perera RJ, Ray A. MicroRNAs in the search for understanding human diseases. BioDrugs. 2007;21(2):97–104.
Asangani IA, Rasheed SA, Nikolova DA, Leupold JH, Colburn NH, Post S, et al. MicroRNA-21 (miR-21) post-transcriptionally downregulates tumor suppressor Pdcd4 and stimulates invasion, intravasation and metastasis in colorectal cancer. Oncogene. 2008;27(15):2128–36. doi:10.1038/sj.onc.1210856.
Schetter AJ, Leung SY, Sohn JJ, Zanetti KA, Bowman ED, Yanaihara N, et al. MicroRNA expression profiles associated with prognosis and therapeutic outcome in colon adenocarcinoma. JAMA. 2008;299(4):425–36. doi:10.1001/jama.299.4.425.
Chen HY, Lin YM, Chung HC, Lang YD, Lin CJ, Huang J, et al. miR-103/107 promote metastasis of colorectal cancer by targeting the metastasis suppressors DAPK and KLF4. Cancer Res. 2012;72(14):3631–41. doi:10.1158/0008-5472.can-12-0667.
Feng B, Dong TT, Wang LL, Zhou HM, Zhao HC, Dong F, et al. Colorectal cancer migration and invasion initiated by microRNA-106a. PloS One. 2012;7(8):e43452. doi:10.1371/journal.pone.0043452.
Zhang Y, Wang Z, Chen M, Peng L, Wang X, Ma Q, et al. MicroRNA-143 targets MACC1 to inhibit cell invasion and migration in colorectal cancer. Mol Cancer. 2012;11:23. doi:10.1186/1476-4598-11-23.
Zeng Y, Qu X, Li H, Huang S, Wang S, Xu Q, et al. MicroRNA-100 regulates osteogenic differentiation of human adipose-derived mesenchymal stem cells by targeting BMPR2. FEBS Lett. 2012;586(16):2375–81. doi:10.1016/j.febslet.2012.05.049.
Huang L, Lin JX, Yu YH, Zhang MY, Wang HY, Zheng M. Downregulation of six microRNAs is associated with advanced stage, lymph node metastasis and poor prognosis in small cell carcinoma of the cervix. PloS One. 2012;7(3):e33762. doi:10.1371/journal.pone.0033762.
Henson BJ, Bhattacharjee S, O’Dee DM, Feingold E, Gollin SM. Decreased expression of miR-125b and miR-100 in oral cancer cells contributes to malignancy. Genes Chromosomes Cancer. 2009;48(7):569–82. doi:10.1002/gcc.20666.
Doghman M, El Wakil A, Cardinaud B, Thomas E, Wang J, Zhao W, et al. Regulation of insulin-like growth factor-mammalian target of rapamycin signaling by microRNA in childhood adrenocortical tumors. Cancer Res. 2010;70(11):4666–75. doi:10.1158/0008-5472.can-09-3970.
Leite KR, Tomiyama A, Reis ST, Sousa-Canavez JM, Sanudo A, Camara-Lopes LH, et al. MicroRNA expression profiles in the progression of prostate cancer–from high-grade prostate intraepithelial neoplasia to metastasis. Urol Oncol. 2013;31(6):796–801. doi:10.1016/j.urolonc.2011.07.002.
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chen, P., Xi, Q., Wang, Q. et al. Downregulation of microRNA-100 correlates with tumor progression and poor prognosis in colorectal cancer. Med Oncol 31, 235 (2014). https://doi.org/10.1007/s12032-014-0235-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-014-0235-x