Abstract
The development of breast cancer is a multistep process associated with complex changes in host gene expression patterns including inactivation of tumor suppressor genes and activation of oncogenes. Critically, hereditary predisposition plays a significant role in cancer susceptibility. However, mutation of the BRCA1 gene is found only in the minority of hereditary breast cancer, which indicates that there might be alternative, novel mechanisms contributing to inactivation of the BRCA1 gene. Studies have shown that aberrant methylation of genomic DNA plays an important role in carcinogenesis. The aim of this study was to investigate whether DNA methylation may be an alternative mechanism for the inactivation of BRCA1 as an epigenetic modification of the genome and whether hereditary breast cancer has a different BRCA1 methylation phenotype pattern than sporadic breast cancer. The pattern of CpG island methylation within the promoter region of BRCA1 was assessed by bisulfite sequencing DNA from peripheral blood cells of 72 patients with hereditary predisposition but without BRCA1 mutations and 30 sporadic breast cancer controls. The overall methylation level in patients with hereditary predisposition was significantly lower than that in the sporadic control group. However, patients with hereditary predisposition showed a significantly higher methylation susceptibility for the sites −518 when compared to controls. These results suggest that there might be different BRCA1 promoter methylation levels and patterns in sporadic and hereditary breast cancer in peripheral blood DNA. These findings may facilitate the early diagnosis of hereditary breast cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer is the single most common cause for cancer-related mortality in women worldwide. In 2008, 1.38 million new cases of breast cancer are diagnosed and 458,400 deaths occur [1]. Should these trends continue, it is estimated that 1.45 million new cases of breast cancer will be diagnosed globally in 2010 [2]. Studies have shown that inherited predisposition accounts for 25–30% of all breast cancers [3]. BRCA1 has been identified as the first major gene associated with familial breast cancer predisposition [4]. As a tumor suppressor genes, BRCA1 is involved in many critical biological processes, including DNA damage repair, cell cycle control, and transcriptional regulation [5]. Women who carry pathogenic BRCA1 mutations have been shown to carry a 50–80% risk of developing breast cancer by the age of 70 [6]. However, the majority of breast cancers that exhibit a BRCA1-like phenotype do not harbor detectable germline mutations in BRCA1 [7]. Some of this discordance may be due to epigenetic defects in BRCA1.
Cancer initiation and progression are driven by the accumulation of inherited or acquired DNA alterations, which may be genetic or epigenetic in nature. Since epigenetic silencing processes are mitotically heritable, they can play the same roles and undergo the same selective processes as genetic alterations in the development of cancer [8]. Epigenetics mechanisms include histone modifications and DNA methylation, which are all associated with cancer [9]. In this study, we focus exclusively on DNA methylation. Methylation occurs at cytosine residues within CpG dinucleotides. It has been estimated that 80% of CpGs are methylated in mammalian genomes. In contrast, CpGs in GC-rich regions such as 5′ regulatory sequences are usually unmethylated, which is an important feature in the promoter regions of genes and for the regulation of gene expression [10]. Apart from certain methylation boundaries, normal cells have virtually no methylation in the promoter regions [11], whereas cancer cells are characterized by an imbalance in methylation, in which global hypomethylation is accompanied by localized hypermethylation of promoter regions. Although further investigation is required, evidence suggests that methylation of promoter regions effectively inhibits transcription of many crucial downstream genes in the neoplastic process [12]. The BRCA1 promoter has previously been shown to be aberrantly methylated in sporadic breast cancer [13]. However, very few studies have examined the methylation status and patterns of the BRCA1 promoter in hereditary breast cancer, especially in Han Chinese woman.
A recent study revealed that two patients with multiple cancers typical of hereditary nonpolyposis colorectal cancer (HNPCC) had extensive promoter methylation of MLH1 in cell types derived from all three embryonic germ layers suggesting that the methylation was constitutional and possibly germline [14]. Suijkerbuijk and colleagues reported that BRCA1-associated breast cancers have less promoter methylation compared with sporadic breast carcinomas and increased promoter methylation than normal tissue [15]. All these studies indicated that methylation was likely to be hereditary and a different etiology in sporadic and hereditary breast cancer.
To sum up, maybe there is a possibility that the participation of BRCA1 in hereditary and sporadic breast carcinogenesis is mediated by epigenetic mechanisms distinct from the inheritance of germline mutations. To test this hypothesis, we analyze the methylation status and patterns of BRCA1 promoter methylation in two highly selected groups of breast cancer patients, familial versus sporadic breast cancer, which have no BRCA1 germline mutations. Our results show significant differences in the methylation levels and patterns between these two groups.
Results
Baseline characteristics of two cases
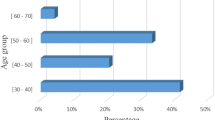
The mean ages of the hereditary study cases and sporadic controls at their peripheral blood sample collection were 40 years (ranging from 24 to 71 years) and 44 years (ranging from 38 to 59 years), respectively. There was not a significantly different mean age when comparing hereditary study cases and sporadic breast cancer controls (P = 0.058).
Tumors in most hereditary study cases were of the ductal type (69 of 72). The remaining tumor samples included medullary (2 of 72) and mammilliform (1 of 72) types. No lobular carcinomas were identified. Most of the sporadic cancers were also ductal carcinomas (28 of 30). The others were medullary cancer (1 of 30) and lobular carcinoma (1 of 30), there was no mammilliform type identified in the sporadic controls.
Our work has revealed that a significantly higher frequency of hereditary study cases are ER (P < 0.001), PR (P = 0.013), and HER2/neu negative (P = 0.046) compared with sporadic breast tumors and have a higher positive lymph node rate than sporadic cases (P = 0.037). All baseline detailed characteristics are shown in Table 1.
Comparing methylation levels in hereditary study cases with sporadic breast cancer controls
When the methylation data were analyzed as continuous variables, among the 102 women for whom methylation data were available, the mean cumulative methylation index (CMI, which means the global level of methylation) was significantly lower in the hereditary study case (1.10) than in the sporadic breast cancer controls (1.54, P = 0.002, as shown in Fig. 1). In a univariate logistic regression model, the CMI was still relatively low in the hereditary study case when age was adjusted as a confounding factor (OR 4.825, 95% CI: 1.889–12.329, P = 0.001). Furthermore, this result was still significant when all potential confounding factors were involved in a multivariate logistic regression model (OR 17.891, 95% CI: 3.207–99.800, P = 0.001). In addition, the CMI was not associated with ER, PR, and HER2/neu expression or positive lymph node rate.
Box plot illustration for in group comparison of the relative methylation of the BRCA1 promoter in the hereditary study case and sporadic control case. Note that 72 cases with hereditary breast cancer predisposition demonstrated relatively lower methylation levels than sporadic breast cancer controls. The mean CMI of hereditary study cases was 1.10 (range 0.3–3.1). The mean CMI of sporadic breast cancer controls was 1.54 (range 0.4–3.1). The mean CMI was significantly lower in the hereditary study case than in the sporadic breast cancer controls (P = 0.002)
Clustering analysis of the methylation pattern in the BRCA1 promoter
Hierarchical clustering of the 72 hereditary study cases and 30 sporadic control cases was performed to analyze BRCA1 promoter methylation status and pattern. As shown in Fig. 2, there was evidence of samples clustering into hereditary study and sporadic control groups on the basis of methylation patterns. When all samples were set into two clusters, the distribution of the hereditary study (5/43) and sporadic control cases (25/29) were significantly different (P < 0.001). Most of the hereditary study cases were distributed into the same cluster and exhibited a low methylation level overall, with a high frequency of methylation at sites −567, −565, −533, and −518. In contrast, the majority of sporadic breast cancer controls were found in the other cluster and showed a relatively higher level and more irregular pattern of methylation.
Hierarchical clustering of 72 breast cancer cases with inherited predisposition (indicated by a black rectangle on the vertical rightmost bar) and 30 sporadic breast cancer controls (gray rectangle on the vertical rightmost bar) based on the BRCA1 promoter methylation status in PBLs. Each row displays the methylation level of all 30 CpGs in one sample, with CpGs arranged horizontally from 5′ to 3′ in the promoter. The exact position of each CpG is given at the bottom of the figure
Methylation was analyzed as binary variables. The methylation frequency for every given CpG is shown in Fig. 3. The CpG sites in 30 sporadic breast cancer control samples were extensively methylated. However, intense methylation was not observed at specific sites apart from at −567 and −565, which displayed 0–90% methylation in hereditary breast cancer samples. Unlike the sporadic breast cancer controls, the CpG sites in the hereditary study cases were largely unmethylated except at 4 sites: −567, −565, −533, and −518, which displayed 0–90% methylation. In addition, methylation at the CpG site −518 was significantly higher in the hereditary study cases than the sporadic controls (64, 30%, P = 0.003).
Methylation was analyzed as binary variables and the methylation rate of every given CpG site is shown. The CpG site has a relatively high methylation frequency at the sites −567 and −565 in both cases. In addition, the sites −533 and −518 show a relatively high methylation frequency in cases with hereditary predisposition. In particular, at the site −518, 45 of 72 (64%) were methylated in cases with hereditary predisposition versus 9 of 30 (30%) in sporadic controls. The difference was significant (P = 0.03). The exact position of each CpG site is given at the bottom of the figure
Discussion
The large number of familial breast cancer without an identified causative gene mutation has led many groups to explore putative BRCAX gene(s) via different approaches, but ended up with no success [17]. Thus, an alternative mechanism was proposed suggesting that epigenetic defects in BRCA1 may contribute to breast cancer predisposition. This hypothesis was supported by a recent study showing that both familial and sporadic breast cancer samples harbored a hypermethylated promoter at BRCA1 [18]. Teresa et al. examined promoter methylation of the BRCA1 gene in 49 women with hereditary breast cancer. They found that the BRCA1 gene promoter was methylated in 51% of patients, among which 67% failed to express the corresponding protein [19]. This result indicated that DNA methylation represents an important alternative mechanism for the inactivation of BRCA1. Moreover, BRCA1 promoter methylation was reported to be associated with poor survival in patients with sporadic breast cancer [20]. Despite these scientific advances, there are many discrepancies in these previous observations including wide variability of methylation rates and patterns between sporadic and familial breast cancers. Furthermore, distinct pathologies may exist between BRCA1 methylated tumors and BRCA1 mutated tumors [21]. Many confounding factors, such as different criteria of patient selection, different tissues for testing, or different testing methods, may lead to different or even controversial conclusions. In our study, patients with hereditary predisposition had a low level of CpG island promoter methylation, whereas women without hereditary predisposition showed a high level of promotor methylation. This finding is consistent with a previous report from Suijkerbuijk group [15], whose study suggested that BRCA1-associated breast cancers show lower promoter methylation when compared with sporadic breast cancers.
Current studies have revealed that DNA methylation inhibits transcription by interfering with the initiation of transcription. This repression can arise through a variety of processes. One potential mechanism is a reduction in the binding affinity of sequence-specific transcription factors. In normal tissue, methylation of the functional BRCA1 promoter region is minimal, leaving transcription factor binding sites accessible to transcription binding proteins, thereby allowing transcription to proceed [22]. It is clear that abnormal methylation of the BRCA1 gene at the cytosine residue alters the spatial conformation of the BRCA1 promoter [23, 24]. As a result, the binding sites for the transcription binding proteins become inaccessible and transcription is blocked [25]. Methylated DNA is localized in the inactive chromatin. It has been suggested that by binding to specific methylated DNA, binding protein clusters of methylated CpGs may promote the formation of inactive chromatin and the exclusion of the transcription machinery [26]. For some transcription factors like E2F and CREB, methylation at specific CpGs has been shown to directly inhibit protein binding and thus inhibit transcription [27, 28]. A second possibility is the recruitment of transcriptional co-repressors by the methylated CpG sequences, which indirectly inhibit the binding of transcription factors [29]. Therefore, studies on specific methylation sites and their methylation frequency may unveil the mechanisms by which the transcription of some anti-oncogenes is inhibited in breast cancer. Our study shows that the level of methylation in breast cancers with inherited predisposition was significantly lower than that in sporadic breast cancers. Moreover, the methylation sites in the CpG island of the BRCA1 gene promoter were significantly different between these two groups. In breast cancer with hereditary predisposition, sites −567, −565, −533, and −518 are prone to be methylated. In sporadic controls, with the exception of methylation at sites −567 and −565, many methylated sites are distributed randomly in the region near the transcriptional start site. Since −567 and −565 are the boundary methylation sites in the BRCA1 promoter and are constitutively methylated in healthy subjects, our data suggest that site −533 and −518 is likely to be more susceptible to methylation in hereditary breast cancer.
A few mechanisms could explain the different BRCA1 methylation profiles between hereditary and sporadic breast cancer. First, the hereditary factor is methylation itself. Although complete erasure of epimutations during spermatogenesis has been observed [30], it is still possible that the methylation profile was maternally derived and maintained by DNA methyl-transferase 1 (DNMT1) [31]. Second, the hereditary factor may contain other factors, which regulate methylation, such as DNA methyl-transferase 3a, 3b (DNMT3a, 3b), and key enzymes of methyl-group metabolism that can adjust the methylation pattern by de novo methylation [32, 33]. Overexpression of DNMT1, 3a, and 3b has been demonstrated in a variety of tumors including bladder, colon, kidney, and pancreatic cancers [28]. In addition, it has been shown that expression of DNMT1 and DNMT3b is necessary for the survival of tumor cells in culture [34]. Perhaps, they can methylated certain sites according to their own blueprint, which is inherited from parents. However, our results disagree with the findings of Joseph et al., which revealed that the CpG sites in 43 hereditary breast cancer patients were prone to be methylated at −567, −565, −37, and −29, displaying 0–100% methylation [35]. This may be due to the fact that people of different races were recruited in these two studies or the limitation of the sample size.
A principal tenet of Darwin’s hypotheses for the evolution of species is that most germline mutations are deleterious, or of no functional significance; mutations giving rise to a specific advantage are selected in an evolving population. These same selective concepts can be applied to epigenetic events, which can occur at a much higher rate compared with mutations in somatic cells [8]. Moreover, methylation plays an important and early role in tumor development and is therefore a promising biomarker for the early detection of malignancy [36]. Although the precise mechanism of DNA methylation initiation and spread during tumorigenesis is unknown, our findings reveal that BRCA1 promoter methylation can be extensive in both hereditary and sporadic breast cancer although their methylation patterns can be different significantly. In conclusion, our sequence analysis suggests that certain CpG sites may be preferentially methylated in the BRCA1 promoter in hereditary breast tumors. These results may provide a potential means of detecting hereditary breast cancer at an early (possibly preinvasive) stage. In the near future, results obtained from ongoing clinical trials examining methylation in hereditary breast cancer may reveal the diagnostic and therapeutic potential of methylation as a major class of molecular markers.
Materials and methods
Patient selection
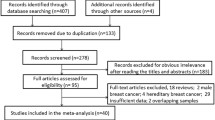
This study was approved by the IRB Committee of Harbin Medical University. This research was completed in compliance with the Helsinki Declaration. All patients in this study were diagnosed with breast cancer and underwent surgery from 2007 to 2010 at the Harbin Medical University Affiliated Tumor Hospital. Seventy-eight women included in the familial breast cancer group to be collected and finally 72 patients agreed to participate. These patients did not receive chemotherapy or radiotherapy before surgical operation. All patients in this study were tested negative for BRCA1 mutations by full-length sequencing. The 72 women included in the familial breast cancer group fit at least one of the following criteria: (1) patient with at least one first or second degree relative to breast and ovarian cancer; or (2) patient having a previous personal history of ovarian cancer. Thirty-two sporadic breast cancer patients who had no familial history were recruited but at last 30 patients agreed to take part in this study as the control group.
Genomic DNA isolation
Written consent forms were obtained from each patient. All patients signed an agreement form for the genetic analysis of their DNA. Peripheral blood samples were collected into EDTA vials. Genomic DNA was extracted from peripheral blood cells using a standard phenol–chloroform extraction method.
Sodium bisulfite genomic sequencing of the BRCA1 promoter
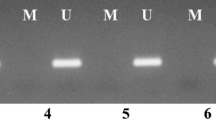
Bisulfate modification of DNA was performed using the EZ DNA Methylation-Gold Kit according to the manufacturer’s instructions (Zymo Research, USA). The BRCA1 promoter CpG island was amplified from the bisulfite-modified DNA by two rounds of PCR using nested primers specific to the bisulfite-modified sequence of the BRCA1 promoter [16]. The following primers were applied: primer 1 (nt 895 to nt 915), 5′-GGGGTTGGATGGGAATTGTAG-3′; primer 2 (nt 1,687 to nt 1,710), 5′-CTCTACTACCTTTACCCAAAAACA-3′; primer 3 (nt 989 to nt 1,013), 5′-GTTTATAATTGTTGATAAGTATAAG-3′; primer 4 (nt 1,627 to nt 1,646), 5′-AAAACCCCACAACCTATCCC-3′. Primers 1 and 2 were used in the first round of amplification under the following conditions: 95°C for 10 min followed by 35 cycles of 95°C for 1 min, 56°C for 3 min, 72°C for 1 min, with a final extension of 72°C for 5 min and rapid cooling to 4°C. One to ten percent of the first round PCR product was used for the second round of PCR using primers 3 and 4 under the following conditions as follows: 95°C for 5 min followed by 35 cycles of 95°C for 1 min, 56°C for 1.5 min, 72°C for 1 min; ending with a final extension of 72°C for 5 min and rapid cooling to 4°C.
The bisulfite-polymerase chain reaction (PCR) product was subcloned into a TA vector (Promega, USA), and 10 individual clones were sequenced per sample using the ABI Prism 3730 DNA Analyzer. The methylation status of individual CpG sites was determined by comparison of the sequence obtained with the known BRCA1 promoter sequence. The number of methylated CpGs at a specific site was divided by the number of clones analyzed (n = 10 in all cases) to yield a percent methylation for each site.
Statistical analysis
The data were analyzed in two ways: (1) as binary variables: CpG was considered methylated when more than 10% of the sampled clones were methylated; and (2) as continuous variables: CMI was calculated as the sum of the proportion of methylated clones for every given CpG in each sample.
The Chi-square test was used for comparing proportions. The two-sample t-test was used for continuous variables. The potential confounding effect of other factors known to influence methylation among breast cancer patients was evaluated by adjustment in the univariate and multivariate logistic regression model. These factors include age at diagnosis, the expression of estrogen receptor (ER), progestogen receptor (PR) and HER2/neu, as well as, positive lymph node rate. Hierarchical clustering analysis was used to group the samples into subclasses (Euclidian distance, Ward method).
All statistical analyses were performed using SPSS software (version 13.0). The significant α level of 0.05 was used.
References
Ahmedin J, Freddie B, Melissa M, Jacques F, Elizabeth W, David F. Global cancer statistics. CA Cancer J Clin. 2011;61(2):69–90.
James CR, Quinn JE, Mullan PB, Johnston PG, Harkin DP. BRCA1, a potential predictive biomarker in the treatment of breast cancer. Oncologist. 2007;12(2):142–50.
Pasche B. Recent advances in breast cancer genetics. Cancer Treat Res. 2008;141:1–10.
Miki Y, Swensen J, Shattuck-Eidens D, Futreal PA, Harshman K, Tavtigian S. A strong candidate for the breast and ovarian cancer susceptibility gene BRCA1. Science. 1994;266(5182):66–71.
Andrieu N, Goldgar DE, Easton DF, Rookus M, Brohet R, Antoniou AC. Pregnancies, breast-feeding, and breast cancer risk in the international BRCA1/2 carrier cohort study (IBCCS). J Natl Cancer Inst. 2006;98(8):535–44.
Gorski JJ, Kennedy RD, Hosey AM, Harkin DP. The complex relationship between BRCA1 and ERα in hereditary breast cancer. Clin Cancer Res. 2009;15(5):1514–8.
Xu X, Gammon MD, Zhang Y, et al. BRCA1 promoter methylation is associated with increased mortality among women with breast cancer. Breast Cancer Res Treat. 2009;115(2):397–404.
Jones PA, Baylin SB. The epigenomics of cancer. Cell. 2007;128(4):683–92.
Brooks J, Cairns P, Zeleniuch-Jacquotte A. Promoter methylation and the detection of breast cancer. Cancer Causes Control. 2009;20(9):1539–50.
Bird A. DNA methylation patterns and epigenetic memory. Genes Dev. 2002;16(1):6–21.
Rice JC, Ozcelik H, Maxeiner P, Andrulis I, Futscher BW. Methylation of the BRCA1 promoter is associated with decreased BRCA1 mRNA levels in clinical breast cancer specimens. Carcinogenesis. 2000;21(9):1761–5.
Attwood JT, Yung RL, Richardson BC. DNA methylation and the regulation of gene transcription. Cell Mol Life Sci. 2002;59(2):241–57.
Birgisdottir V, Stefansson OA, Bodvarsdottir SK, Hilmarsdottir H, Jonasson JG, Eyfjord JE. Epigenetic silencing and deletion of the BRCA1 gene in sporadic breast cancer. Breast Cancer Res. 2006;8(4):R38.
Suter CM, Martin DI, Ward RL. Germline epimutation of MLH1 in individuals with multiple cancers. Nat Genet. 2004;36(5):497–501.
Suijkerbuijk KP, Fackler MJ, Sukumar S, et al. Methylation is less abundant in BRCA1-associated compared with sporadic breast cancer. Ann Oncol. 2008;19(11):1870–4.
Rice JC, Massey-Brown KS, Futscher BW. Aberrant methylation of the BRCA1 CpG island promoter is associated with decreased BRCA1 mRNA in sporadic breast cancer cells. Oncogene. 1998;17(14):1807–12.
Honrado E, Osorio A, Milne RL, et al. Immunohistochemical classification of non-BRCA1/2 tumors identifies different groups that demonstrate the heterogeneity of BRCAX families. Mod Pathol. 2007;20(12):1298–306.
Ali AB, Iau PT, Sng JH. Cancer-specific methylation in the BRCA1 promoter in sporadic breast tumours. Med Oncol. 2010. [Epub ahead of print].
Tapia T, Smalley SV, Kohen P, Muñoz A, Solis LM, Corvalan A. Promoter hypermethylation of BRCA1 correlates with absence of expression in hereditary breast cancer tumors. Epigenetics. 2008;3(3):157–63.
Chen Y, Zhou J, Xu Y, Li Z, Wen X, Yao L, Xie Y, Deng D. BRCA1 promoter methylation associated with poor survival in Chinese patients with sporadic breast cancer. Cancer Sci. 2009;100:1663–7.
Matros E, Wang ZC, Lodeiro G, Miron A, Iglehart JD, Richardson AL. BRCA1 promoter methylation in sporadic breast tumors: relationship to gene expression profiles. Breast Cancer Res Treat. 2005;91:179–86.
Zhao Z, Han L. CpG islands: algorithms and applications in methylation studies. Biochem Biophys Res Commun. 2009;382(4):643–5.
Ehrlich M. DNA methylation in cancer: too much, but also too little. Oncogene. 2002;21(35):5400–13.
Baylin SB, Herman JG. DNA hypermethylation in tumorigenesis: epigenetics joins genetics. Trends Genet. 2000;16(4):168–74.
Zhu WG, Srinivasan K, Dai Z, Duan W, et al. Methylation of adjacent CpG sites affects Sp1/Sp3 binding and activity in the p21 (Cip1) promoter. Mol Cell Biol. 2003;23(12):4056–65.
Clark SJ, Harrison J, Molloy PL. Sp1 binding is inhibited by mCpmCpG methylation. Gene. 1997;195(1):67–71.
Watt F, Molloy PL. Cytosine methylation prevents binding to DNA of a HeLa cell transcription factor required for optimal expression of the adenovirus major late promoter. Genes Dev. 1988;2(9):1136–43.
Mancini DN, Rodenhiser DI, Ainsworth PJ, O’Malley FP, Singh SM, Xing W. CpG methylation within the 5′ regulatory region of the BRCA1 gene is tumor specific and includes a putative CREB binding site. Oncogene. 1998;16(9):1161–9.
Catteau A, Joanna R. Morris. BRCA1 methylation: a significant role in tumour development. Semin Cancer Biol. 2002;12(5):359–71.
Hitchins MP, Ward RL. Erasure of MLH1 methylation in spermatozoa- implications for epigenetic inheritance. Nat Genet. 2007;39:1289.
Dupont C, Armant DR, Brenner CA. Epigenetics: definition, mechanisms and clinical perspective. Semin Reprod Med. 2009;27(5):351–7.
Okano M, Xie S, Li E. Cloning and characterization of a family of novel mammalian DNA (cytosine-5) methyltransferases. Nat Genet. 1998;19(3):219–20.
Macis D, Maisonneuve P, Johansson H, Bonanni B, Botteri E, Iodice S. Methylenetetrahydrofolate reductase (MTHFR) and breast cancer risk: a nested-case-control study and a pooled meta-analysis. Breast Cancer Res Treat. 2007;106(2):263–71.
Abreu PA, Dellamora-Ortiz G, Leão-Ferreira LR, et al. DNA mrthylation: a promising target for the twenty-first century. Expert Opin Ther Targets. 2008;12(8):1035–47.
Chen Y, Toland AE, McLennan J, et al. Lack of germ-line promoter methylation in BRCA1-negative families with familial breast cancer. Genet Test. 2006;10(4):281–4.
Yazici H, Terry MB, Cho YH, et al. Aberrant methylation of RASSF1A in plasma DNA before breast cancer diagnosis in the breast cancer family registry. Cancer Epidemiol Biomarkers Prev. 2009;18(10):2723–5.
Acknowledgments
The authors are grateful to the insightful work provided by Dr. Yashuang Zhao and all of the fellows and students who gave me the possibility to complete this thesis. We thank Dr. Shihui Yu and Dr. QingBin Song for critical reading of the manuscript, graphics assistance and word processing assistance.
Conflict of interest
The authors declare that they have no competing financial interests.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Pang, D., Zhao, Y., Xue, W. et al. Methylation profiles of the BRCA1 promoter in hereditary and sporadic breast cancer among Han Chinese. Med Oncol 29, 1561–1568 (2012). https://doi.org/10.1007/s12032-011-0100-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12032-011-0100-0