Abstract
With the advent of computational genomics, an intensive search is underway for unique biomarkers for Homo sapiens that could be used to differentiate taxa within the Hominoidea, in particular to distinguish Homo from the apes (Pan, Gorilla, Pongo, and Hylobates) and species or subspecies within the genus Homo (H. sapiens, H. heidelbergensis, H. neanderthalensis, H. erectus, and the Denisovans). Here, we suggest that the |-FAM72–SRGAP2-| (family with sequence similarity 72/SLIT-ROBO Rho GTPase activating protein 2) gene pair is a unique molecular biomarker for the genus Homo that could also help to place Australopithecus at its most appropriate place within the phylogenetic tree and may explain the distinctive higher brain cognitive functions of humans.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The evolutionary relationships between modern Homo sapiens and other hominin species are controversial. Many scientists believe that H. sapiens evolved from populations of H. heidelbergensis in Africa (~150 and 200 ka) and later (~40–50 ka) migrated out of Africa, replacing all populations of H. neanderthalensis in Europe and H. heidelbergensis elsewhere. Others, however, believe that archaic populations evolved into H. sapiens in each region of the world, and therefore consider H. neanderthalensis to be a subspecies of H. sapiens. This multiregional hypothesis permits the evolution of regional characteristics that distinguish populations in different regions, while still allowing universally favorable traits to spread across regions by gene flow. Supporters of this idea believe that interbreeding between H. neanderthalensis and H. sapiens was more widespread than is traditionally accepted by the opposing “Out of Africa” perspective and that H. neanderthalensis made important genetic contributions to living H. sapiens. The recent complete sequencing of the H. neanderthalensis genome indicates that the levels of interbreeding between H. neanderthalensis and H. sapiens were in fact greater than previously thought, and therefore the question of whether or not H. sapiens and H. neanderthalensis were different species becomes philosophical (Becoming human: Homo sapiens 2016; Encyclopedia Britannica: Homo sapiens 2016; Encyclopedia of Life: Homo sapiens - Modern Human 2016; Institute of human origins: Human Origins 2016; Smithsonian National Museum of Natural History: What does it mean to be human 2016).
For this review, all the chromosome (chr) datasets were downloaded from the public database of the National Center for Biotechnology Information (NCBI). We relied on the taxonomy offered by NCBI and reconstructed the simplified taxonomic tree from publically available databases. Additionally, we retrieved the genomes of species within the taxonomy from assemblies offered by NCBI and all data retrieval occurred automatically by evaluating database-information available on NCBI-servers as described previously (Kutzner et al. 2015).
A Simplified Phylogenetic Tree of the Hominoidea
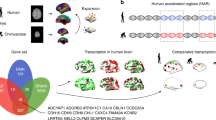
Evolutionary and molecular anthropologists are trying to explain the gap between humans (genus Homo) and the remainder of the Hominoidea by focusing their studies on whole genomes to understand the evolution of uniquely human traits in a phylogenetic context (Fig. 1). Rapid progress in biotechnology has led to the development of “omics” research (e.g., genomics, proteomics, metabolomics) in healthcare science to help understand the causes and mechanisms of complex human diseases (Sun and Hu 2016; Suravajhala et al. 2016; Yugi et al. 2016).
A simplified phylogenetic tree explaining the development of modern humans based on current evolutionary theories. Thus far, Australopithecus is considered to be one of the common ancestors of the genus Homo, although clear-cut sequencing (e.g., |-FAM72–SRGAP2-| evidence for the correct categorization of Australopithecus as a subspecies of Homo, as a separate genus within the Hominini, or even as a subspecies of Pan is lacking (Wikipedia: Australopithecus 2016)
Human Genomics
Whole genome studies allow for the exploration of genomic commonalities and differences between the human and ape subfamilies. Meanwhile, despite high-throughput next generation sequencing (NGS) efforts associated with the 1000 Genomes Project C et al. (2012)–Genomes Project C et al. 2015, and The International Genome Sample Resource [IGSR]) (1000 Genomes Project 2016), comprehensive data are lacking, and the high costs of genome sequencing forced researchers to cease further investigations. Sequencing costs limit the accuracy of genome reconstruction by limiting genome depth (x-fold of genome coverage) (Sims et al. 2014). Although the final 1000 Genomes data set comprises 2504 present-day human individuals from 26 populations around the world, only low coverage (2-6x) whole-genome sequencing (WGS) data, and mainly exome sequence data, are available for all individuals, while data for only 24 individuals were subjected to high coverage sequencing for validation purposes (Genomes Project C et al. 2012, 2015; Sankararaman et al. 2014; Sudmant et al. 2015).
To close the genomic gap and identify differences between taxa such as Pan and Homo, continuing solid and well-grounded research-based substantiated genome data analyses are urgently needed. Current state-of-the-art NGS technology, combined with steady improvements in big data computing whole genome analysis, could provide the background for (i) coordinated present-day human genome studies across the globe under the auspices of IGSR to ensure that existing population diversity gaps are filled, and (ii) genome comparisons across the entire phylogenetic tree to gain further insight into the relationships among the Hominidae (Fig. 1).
Only with full access to entire representative genomes of modern human population diversity across the globe will we be able to detect variations within present-day humans, which in turn would then allow us to make more reasonable statements with regards to archaic human taxa such as Neanderthals or Denisovans (Meyer et al. 2016; Sudmant et al. 2015).
Considering the many genomic variations detected in the 1000 Genomes Project data, including larger deletions, a recent study compared the mitochondrial genome of one individual from the Sima de los Huesos (SH) to only one Neanderthal (Altai), one Denisovan, and one present-day modern human (MButi) from Africa. This study is limited by problems in terms of genome quantity (“extremely small amounts of data available”) and quality (degradation and contamination with modern human and/or microbial DNA) (Meyer et al. 2016). Thus, conclusions drawn from such studies require due diligence.
Biomarkers for the Genus Homo
The use of omics “big data” analysis may aid in the search for possible biomarkers distinguishing the genus Homo and may help to further unravel the significant commonalities and differences between modern humans and the apes.
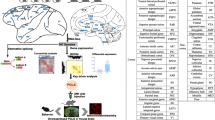
The human brain, with its higher cognitive functions, distinguishes humans from all other hominoid species. Therefore, the human brain may serve as a special source of possible genomic biomarkers to clearly differentiate humans among hominoids. A recently described unique gene pair, FAM72 (controls neuronal stem-cell proliferation/development (Benayoun et al. 2014))–SRGAP2 (neuronal development, differentiation, synaptic, and cerebral plasticity (Charrier et al. 2012; Rincic et al. 2016)), could explain the distinctive nature of human higher brain development, brain plasticity, and cognitive functions (Dennis et al. 2012; Kutzner et al. 2015). Therefore, this gene pair is a potential H. sapiens biomarker (Fig. 2).
The FAM72 (A–D) and SRGAP (A–D) gene pairs in apes and human. While four |-FAM72–SRGAP2-| gene pairs are found in Homo (including e.g., Neanderthals and Denisovans), only one gene pair is present in the apes. The mechanism underlying the human |-FAM72–SRGAP2-| gene pair duplication and amplification (arrows indicate the different options in the schematic) remains to be elucidated. When comparing the master gene |-FAM72–SRGAP2-| on chr 1 in apes and chr 1 in human, non-preserved areas are observed in the middle of preserved areas. It is still not known how |-FAM72A–SRGAP2A-| was transferred and amplified to the other gene pairs |-FAM72D–SRGAP2B-|, |-FAM72C–SRGAP2D-| and |-FAM72B–SRGAP2C-| in Homo during evolution, while no species have been observed with two or three |-FAM72–SRGAP2-| gene pairs (Kutzner et al. 2015). For clarity, official gene symbols are used instead of gene names in Italics style and protein names in Roman style
Evolutionary Perspective
From an evolutionary perspective, the single |-FAM72–SRGAP2-| gene pair defines the notochord-containing vertebrates, whereas the four paralogous |-FAM72–SRGAP2-| gene pair couples seem to be a distinctive feature of H. sapiens only among the Hominidae, a finding that is perhaps associated with higher (e.g., explicit) cognitive capabilities, language activity, and consciousness (Geschwind and Konopka 2012; Geschwind and Rakic 2013; Kutzner et al. 2015). It might be possible to consider the |-FAM72–SRGAP2-| gene pair as one master gene. While the cell activates FAM72 in neuronal progenitor cells (and keeps SRGAP2 switched off), it turns SRGAP2 on (and FAM72 off) during differentiation and neuron maturation. However, it remains an enigma why no species contains two or three |-FAM72–SRGAP2-| gene pairs; the other great apes (chimpanzees and gorillas) also contain only one such gene pair.
Current theories cannot adequately explain this gap between humans and the other great apes (Dennis et al. 2012; Dewey 2011; Geschwind and Konopka 2012; Geschwind and Rakic 2013; Wolfe 2000). It also remains a point of special interest whether any species can be identified within the genus Homo that carries or carried two or three |-FAM72–SRGAP2-| gene pairs. With Australopithecus considered as one of the common ancestors of the genus Homo, A. africanus probably evolved into A. sediba, which some scientists think may have evolved into H. habilis, H. erectus, and eventually modern humans, H. sapiens (Fig. 1) (Bruxelles et al. 2014; Granger et al. 2015; Kivell 2015; Zihlman et al. 1978). Other species in question include H. naledi (Berger et al. 2015; Dirks et al. 2015; Harcourt-Smith et al. 2015; Stringer 2015) and H. floresiensis (Brown and Maeda 2009; Daegling et al. 2014; Jungers et al. 2009; Orr et al. 2013). Thus far, clear-cut proven genomic sequencing evidence does not exist.
Conclusion
Qualitative assessments and impartial communications regarding the strength of genomics data across related species are essential for the long-term management of predictive uncertainty and for the successful application of human genomics in systematics. The master gene |-FAM72–SRGAP2-| constitutes a potential human biomarker defining H. sapiens with higher brain cognitive functions, and its malfunction could lead to pathological conditions (Guo et al. 2008; Heese 2013; Kutzner et al. 2015; Nehar et al. 2009; Pramanik et al. 2015).
References
Becoming human: Homo sapiens (Accessed 20 May 2016). http://becominghuman.org/
Benayoun BA et al (2014) H3K4me3 breadth is linked to cell identity and transcriptional consistency. Cell 158:673–688. doi:10.1016/j.cell.2014.06.027
Berger LR et al (2015) Homo Naledi, a new species of the genus homo from the Dinaledi chamber. South Africa. Elife 4. doi:10.7554/eLife.09560
Brown P, Maeda T (2009) Liang Bua Homo floresiensis mandibles and mandibular teeth: a contribution to the comparative morphology of a new hominin species. J Hum Evol 57:571–596. doi:10.1016/j.jhevol.2009.06.002
Bruxelles L, Clarke RJ, Maire R, Ortega R, Stratford D (2014) Stratigraphic analysis of the Sterkfontein StW 573 Australopithecus skeleton and implications for its age. J Hum Evol 70:36–48. doi:10.1016/j.jhevol.2014.02.014
Charrier C et al (2012) Inhibition of SRGAP2 function by its human-specific paralogs induces neoteny during spine maturation. Cell 149:923–935. doi:10.1016/j.cell.2012.03.034
Daegling DJ, Patel BA, Jungers WL (2014) Geometric properties and comparative biomechanics of Homo floresiensis mandibles. J Hum Evol 68:36–46. doi:10.1016/j.jhevol.2014.01.001
Dennis MY et al (2012) Evolution of human-specific neural SRGAP2 genes by incomplete segmental duplication. Cell 149:912–922. doi:10.1016/j.cell.2012.03.033
Dewey CN (2011) Positional orthology: putting genomic evolutionary relationships into context. Brief Bioinform 12:401–412. doi:10.1093/bib/bbr040
Dirks PH et al (2015) Geological and taphonomic context for the new hominin species Homo naledi from the Dinaledi Chamber, South Africa. Elife 4. doi:10.7554/eLife.09561
Encyclopedia Britannica: Homo sapiens (Accessed 20 May 2016). http://globalbritannicacom/topic/Homo-sapiens
Encyclopedia of Life: Homo sapiens - Modern Human (Accessed 20 May 2016). http://eolorg/pages/327955/overview
Genomes Project (Accessed 20 May 2016). http://www.1000genomes.org/
Genomes Project C et al (2012) An integrated map of genetic variation from 1,092 human genomes. Nature 491:56–65. doi:10.1038/nature11632
Genomes Project C et al (2015) A global reference for human genetic variation. Nature 526:68–74. doi:10.1038/nature15393
Geschwind DH, Konopka G (2012) Neuroscience: genes and human brain evolution. Nature 486:481–482. doi:10.1038/nature11380
Geschwind DH, Rakic P (2013) Cortical evolution: judge the brain by its cover. Neuron 80:633–647. doi:10.1016/j.neuron.2013.10.045
Granger DE, Gibbon RJ, Kuman K, Clarke RJ, Bruxelles L, Caffee MW (2015) New cosmogenic burial ages for Sterkfontein member 2 Australopithecus and member 5 Oldowan. Nature 522:85–88. doi:10.1038/nature14268
Guo C et al (2008) Ugene, a newly identified protein that is commonly overexpressed in cancer and binds uracil DNA glycosylase. Cancer Res 68:6118–6126. doi:10.1158/0008-5472.CAN-08-1259
Harcourt-Smith WE et al (2015) The foot of Homo naledi. Nat Commun 6:8432. doi:10.1038/ncomms9432
Heese K (2013) The protein p17 signaling pathways in cancer. Tumour Biol 34:4081–4087. doi:10.1007/s13277-013-0999-1
Institute of human origins: Human Origins (Accessed 20 May 2016). https://ihoasuedu/
Jungers WL et al (2009) The foot of Homo floresiensis. Nature 459:81–84. doi:10.1038/nature07989
Kivell TL (2015) Evidence in hand: recent discoveries and the early evolution of human manual manipulation. Philos Trans R Soc Lond Ser B Biol Sci 370. doi:10.1098/rstb.2015.0105
Kutzner A, Pramanik S, Kim PS, Heese K (2015) All-or-(N)One—an epistemological characterization of the human tumorigenic neuronal paralogous FAM72 gene loci. Genomics 106:278–285. doi:10.1016/j.ygeno.2015.07.003
Meyer M et al (2016) Nuclear DNA sequences from the Middle Pleistocene Sima de los Huesos hominins. Nature 531:504–507. doi:10.1038/nature17405
Nehar S, Mishra M, Heese K (2009) Identification and characterisation of the novel amyloid-beta peptide-induced protein p17. FEBS Lett 583:3247–3253. doi:10.1016/j.febslet.2009.09.018
Orr CM et al (2013) New wrist bones of Homo Floresiensis from Liang Bua (Flores, Indonesia). J Hum Evol 64:109–129. doi:10.1016/j.jhevol.2012.10.003
Pramanik S, Kutzner A, Heese K (2015) Lead discovery and in silico 3D structure modeling of tumorigenic FAM72A (p17). Tumour Biol 36:239–249. doi:10.1007/s13277-014-2620-7
Rincic M, Rados M, Krsnik Z, Gotovac K, Borovecki F, Liehr T, Brecevic L (2016) Complex intrachromosomal rearrangement in 1q leading to 1q32.2 microdeletion: a potential role of SRGAP2 in the gyrification of cerebral cortex. Mol Cytogenet 9:19. doi:10.1186/s13039-016-0221-4
Sankararaman S et al (2014) The genomic landscape of Neanderthal ancestry in present-day humans. Nature 507:354–357. doi:10.1038/nature12961
Sims D, Sudbery I, Ilott NE, Heger A, Ponting CP (2014) Sequencing depth and coverage: key considerations in genomic analyses. Nat Rev Genet 15:121–132. doi:10.1038/nrg3642
Smithsonian National Museum of Natural History: What does it mean to be human (Accessed 20 May 2016). http://humanoriginssiedu/
Stringer C (2015) The many mysteries of Homo naledi. Elife 4. doi:10.7554/eLife.10627
Sudmant PH et al (2015) An integrated map of structural variation in 2,504 human genomes. Nature 526:75–81. doi:10.1038/nature15394
Sun YV, Hu YJ (2016) Integrative analysis of multi-omics data for discovery and functional studies of complex human diseases. Adv Genet 93:147–190. doi:10.1016/bs.adgen.2015.11.004
Suravajhala P, Kogelman LJ, Kadarmideen HN (2016) Multi-omic data integration and analysis using systems genomics approaches: methods and applications in animal production, health and welfare. Genet Sel Evol 48:38. doi:10.1186/s12711-016-0217-x
Wikipedia: Australopithecus (Accessed 20 May 2016). https://enwikipediaorg/wiki/Australopithecus - cite_note-Reardon2012–2
Wolfe K (2000) Robustness—it's not where you think it is. Nat Genet 25:3–4. doi:10.1038/75560
Yugi K, Kubota H, Hatano A, Kuroda S (2016) Trans-omics: how to reconstruct biochemical networks across multiple ‘Omic’ layers. Trends Biotechnol 34:276–290. doi:10.1016/j.tibtech.2015.12.013
Zihlman AL, Cronin JE, Cramer DL, Sarich VM (1978) Pygmy chimpanzee as a possible prototype for the common ancestor of humans, chimpanzees and gorillas. Nature 275:744–746
Acknowledgements
This study was supported by Hanyang University by providing a scholarship to Ms. Nguyen Thi Thanh Ho.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of Interest
The authors declare that they have no conflicts of interest.
Rights and permissions
About this article
Cite this article
Ho, N.T.T., Kim, PS., Kutzner, A. et al. Cognitive Functions: Human vs. Animal – 4:1 Advantage |-FAM72–SRGAP2-|. J Mol Neurosci 61, 603–606 (2017). https://doi.org/10.1007/s12031-017-0901-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-017-0901-5