Abstract
Among the particular immunomodulation properties of mesenchymal stem cells (MSCs), one relies on their capacity to regulatory T cell (Treg) induction from effector T cells. Stable expression of Foxp3 has a dominant role in suppressive phenotype and stability of induced regulatory T cells (iTregs). How MSCs induce stable Foxp3 expression in iTregs remains unknown. We previously showed MSCs could enhance demethylation of Treg-specific demethylated region (TSDR) in iTregs in cell-cell contact manner (unpublished data). Here, we evaluated the possible effect of MSCs on the mRNA expression of Runx complex genes (Runx1, Runx3, and CBFB) that perch on TSDR in iTregs and play the main role in suppressive properties of Tregs, a regulatory pathway that has not yet been explored by MSCs. Also, we investigated the mRNA expression of MBD2 that promotes TSDR demethylation in Tregs. We first showed that in vitro MSC-iTreg induction was associated with strong mRNA modifications of genes involved in Runx complex. We next injected high doses of MSCs in a murine model of C57BL/6 into Balb/C allogeneic skin transplantation to prolong allograft survival. When splenocytes of grafted mice were analyzed, we realized that the Foxp3 expression was increased at day 5 and 10 post-graft merely in MSC-treated mice. Furthermore, Foxp3 mRNA expression was associated with modified Runx complex mRNA expression comparable to what was shown in in vitro studies. Hence, our data identify a possible mechanism in which MSCs convert conventional T cells to iTreg through strong modifications of mRNA of genes that are involved in Runx complex of Foxp3.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bone marrow-derived mesenchymal stem cells (BM-MSCs) can modulate the immune system both in vitro and in vivo in different conditions like autoimmunity, graft rejection, and graft-versus-host disease [1,2,3,4]. This immunomodulation ability depends on the soluble inhibitory molecules and cytokines (i.e., IL-10, TGFβ, HLA-G, IDO, and PGE2) and cell-cell contact to induce regulatory immune cells such as regulatory T cells (Tregs) and/or tolerogenic dendritic cells (tDC) [2, 5,6,7,8] Particularly, maturation and differentiation of DCs are affected by MSCs through reduction of MHC class II, co-stimulatory molecules surface membrane expression, and reduced secretion of interleukin 12 (IL-12) and tumor necrosis factor-α (TNF-α) [6, 9, 10] . MSCs-induced tDCs can trigger the conversion of CD4 T cells toward anti-inflammatory phenotype attested by IL-10 secretion [9, 11].
Tregs play a crucial role in self-tolerance and homeostasis [12]. These cells are either generated in the thymus during embryonic states (nTregs) or in the periphery (iTregs) from effector CD4+ T cells [13]. Previous investigations proved that MSCs could favor iTregs conversion and enhance both iTregs and nTregs expansion and suppressive function both in vitro and in vivo. [3, 6, 14,15,16] Stable Foxp3 expression plays a major role in the phenotypic and functional stability of iTregs. Indeed, under specific inflammatory conditions, iTregs could lose their Foxp3 expression and convert into effector T cells [17].
It has been already shown that hypomethylation of the Treg-specific demethylated region (TSDR) in the Foxp3 promoter is required for stable Foxp3 expression [18]. TSDR is completely demethylated in nTregs, partially methylated in iTregs, and completely methylated in effector T cells [18]. Hence, removal of methyl groups from TSDR enhances the approachability for transcription factors to perch on TSDR and increase the Foxp3 expression [18]. The role of TSDR methylation in Foxp3 stability is supported by TSDR-null Treg cells, which lose Foxp3 expression. Methyl-CpG-binding domain protein 2 (MBD2) acts as a DNA demethylase by removing repressive methyl residues and thereby activating gene transcription [19]. Overexpression of MBD2 leads to demethylation of CpG regions and Foxp3 gene expression [20]. Studies have reported the correlations between low levels of MBD2 expression and DNA demethylation in several autoimmune diseases such as lupus erythematosus (SLE), dermatomyositis and systemic sclerosis, rheumatoid arthritis, psoriasis, and breast cancer [20,21,22,23]. Deletion of MBD2 in Tregs leads to methylated TSDR and decreases the suppressive function of Tregs both in vivo and in vitro [24].
Furthermore, the complex of Core-binding factor subunit beta (CBFB) and Runt-related transcription factor 1 and 3 (RUNX1 and 3) is necessary for Treg suppressive function [26]. CBFB, the non-DNA-binding factor, can attach to RUNX proteins and increases the DNA-binding affinity and stability of these proteins. RUNX1 and RUNX3 are up-regulated in iTregs and are required for their induction. Using siRNA for RUNX1 and RUNX3, a significant reduction in Foxp3 expression has been shown. This was not observed in GATA3 level in Th2 cells, T-bet level in Th1 cells, or RORγt mRNA expression in Th17 cells. The highest reduction of Foxp3 happens when RUNX1 and RUNX3 are silenced together [25]. It was shown that deficiency of CBFB could decrease Foxp3 expression in nTreg cells [25]. Moreover, using the inhibitor for RUNX proteins could interfere with Foxp3 induction even in the presence of exogenous IL-2 which is highly necessary for iTregs [25]. Hence, increasing RUNX1, RUNX3, and CBFB complex in hypo-methylated TSDR increases the stability and suppressive function of Treg.
Although the regulatory effect of MSCs on the immune system has been well studied, the molecular mechanism underlying this phenomenon is still a matter of question, and to our knowledge, the modulatory effect of MSCs on RUNX1, RUNX3, CBFB, and MBD2 gene expression in Tregs has not yet been explored.
We previously showed that MSCs could enhance demethylation of TSDR in iTregs in a cell-cell contact manner (unpublished data). Here, we first investigated the role of cell-cell contact and cytokine secretion of BM-MSCs on induction, stability, and suppressive function of Tregs in different experimental conditions. Second, we studied the effect of MSCs on RUNX1, RUNX3, CBFB, and MBD2 genes mRNA expression in CD4+ T cells in correlation with Foxp3 stability and suppressive function of iTregs. Third, we investigated the effect of ex vivo-expanded BM-MSCs on induction of transplant tolerance in a model of fully allogeneic skin transplantation and further analyzed the cytokine secretion pattern in grafted mice as well as interest gene expression on CD4+ T cells harvested from protected mice.
We demonstrate for the first time that MSCs could enhance suppressive phenotype and stability of Tregs through regulation of RUNX-CBFB complex but not MBD2 both in vitro and in vivo.
Materials and methods
Mesenchymal stem cell culture
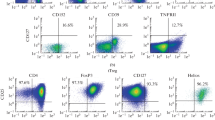
MSCs were collected and cultured by flushing from femoral and tibia bone marrow of six to 8-week-old Balb/C female mice. The mice were purchased from the central animal laboratory (Shiraz University of Medical Sciences, Iran). This research was approved by the Committee of Ethics in Animal Experiments (CEEA) of Shiraz Medical Sciences University. All methods and procedures were performed in accordance with the relevant guidelines and regulations. The cells were cultured in 25-cm2 flasks with Dulbecco’s modified Eagle medium (DMEM), with a low glucose concentration, Glutamax I, 10% heat-inactivated FBS, 1% penicillin, and streptomycin (all from Gibco, Germany). The cells were incubated at 37 °C in a 5% CO2 atmosphere. Non-adherent cells were removed every 8 h; pure MSCs were obtained after 4 to 5 weeks. Mice MSCs derived from bone marrow express surface markers such as CD44 and Sca-1 and do not express specific hematopoietic markers such as CD34 and CD45 (PE-conjugated anti-Sca-1, anti-CD44, anti-CD45 antibodies, and FITC-conjugated anti-CD34 all from eBioscience, USA). Unstained and proper isotype was used as a control for analyzing data by FlowJo software. The second passage of MSCs was used for experiments.
BM-MSCS can differentiate into osteocyte or adipocyte when cultured in an appropriate differentiation condition (DMEM supplemented with 0.5-μM ascorbic phosphate, 1-μM dexamethasone, and 200-μM indomethacin) for 10 days. Then, cells were stained with 0.5% oil red for 10 min. To differentiate isolated cells into osteocytes, they were cultured in a specific differentiation medium (DMEM supplemented with 0.5-μM ascorbic phosphate, 10-mM β-glycerophosphate, and 1-μM dexamethasone) for 21 days and stained with 1% alizarin red for 10 min.
Isolation of dendritic cells and CD4+ T cell
The spleen of Balb/C mice was removed for DC isolation. The most practical way to enrich the DC fraction is to use a density gradient. Nycodenz has successfully been used for enrichment of DCs obtained from different sources 78, 79. First, DCs were isolated by nycodenz and then CD11c positive selection kit (Miltenyi Biotechnology, Germany) was used for DC isolation according to the manufacturer’s guide. CD4+ T cell negative selection kit (Miltenyi Biotechnology, Germany) was used for isolation of total CD4+ T cells from C57BL/6 spleen. CD4+CD25+ cells were depleted from CD4+ T cell pool that showed more than 95% purity of CD4+CD25− T cells; it was performed by the CD4+CD25+ regulatory T cell isolation kit (Miltenyi Biotechnology, Germany). These CD4+CD25− T cells were cultured with MSCs. The CD4+CD25+ regulatory T cell isolation kit was used for isolation of Tregs from conditions after culture with MSCs. Magnetic-activated cell sorting (MACS) method was used for all cells isolation.
In vitro study design
Indeed, we reproduced the entire already studied MSC culture models leading to Tregs induction [8, 26,27,28,29]. In the first line, we sought to know whether MSCs could induce Tregs in all these conditions via the same mechanisms. MSCs could influence T cells directly or indirectly. In the direct mechanism, MSCs effect on T cells could be through cell-cell contact or soluble factors, while the effect of MSCs on T cells might occur indirectly via modulating antigen-presenting cell (APCs) such as DCs, resulting in altered cytokine expression and impaired antigen presentation [30,31,32].
-
1-
In MSC + TC condition, we added 2 × 106 CD4+CD25− T cells directly to2 × 105 allogeneic MSCs.
-
2-
In MSC + MLR condition, we added both 2 × 106 CD4+CD25− T cells (C57BL/6) and 2 × 105 DCs (Balb/C) to 2 × 105 MSCs (Balb/C).
-
3-
In MSC + MLR + LPS condition, we added both 2 × 106 CD4+CD25− T cells (C57BL/6) and 2 × 105 DCs (Balb/C) to 2 × 105 MSCs (Balb/C) in addition to LPS. LPS (200 ng/mL) (Sigma, USA) was directly added to each well.
T cells were isolated by CD4+ isolation kit (MACS) from DCs in conditions 2 and 3.
-
4-
In MSC-DC condition, first, freshly isolated 2 × 106 DCs (Balb/C) were cultured with 2 × 105 autologous MSCs for 24 h. Then, after 24 h, 2 × 105 DCs were isolated and irradiated (30 Gy) and added to 2 × 106 freshly isolated CD4+CD25− allogeneic T cells.
Co-culture and transwell approach to T cells and mesenchymal stem cells
The second passage of Balb/C MSCs was seeded into 6-well plates. MSCs were incubated for 3 h in a complete DMEM. Then, freshly isolated C57BL/6 mice T cells (106 cells per well) were added to the MSCs at a 1:10 ratio. All co-culture experiments were performed in a DMEM containing 10% FBS, 1% penicillin, and 1% streptomycin. T cells were harvested at different times: 6 h, 12 h, 24 h, 48 h, 72 h, and 5 days after co-culture. Also, transwell experiments were performed at the same ratio, conditions, and times; 106 T cells were cultured in the lower chamber and MSCs were seeded in the upper chamber of transwell plates (0.8-μm pore-size membrane, SPL, USA).
For experiments using MSC-DC, freshly isolated Balb/C mice DCs (106 cells per well) were added to the MSCs (1:10) in co-culture and transwell conditions. DCs were harvested after 24 h and added to freshly isolated C57BL/6 T cells to perform MLR. We used Mouse Regulatory T Cell Staining Kit #3 (ebioscience, USA) to investigate FOXP3 proteins in all conditions after 72 h and 5 days.
Allogeneic mixed lymphocyte reactions (MLR) were performed with 106 T cells (C57BL/6) and 105 DCs (Balb/C) in co-culture or transwell conditions with MSCs (with or without LPS) in RPMI-1640 supplemented with 10% FBS and 1% penicillin/streptomycin.
Mix lymphocyte reaction and negative control
DCs were irradiated (30 Gy). Then, cells were washed with PBS for three times and resuspended in RPMI supplemented with 10% heat-inactivated FBS, 1% penicillin, and streptomycin. MLR was set up in 96-well round-bottom cell culture plates (Nunc, Denmark). 105 untreated DCs (BALB/c), as stimulator cells, were co-cultured with 106 CD4+CD25− T cells (C57BL/ 6), as responder cells in a total volume of 200 μL. CD4−CD25+ effector T cells of this condition were isolated by CD4+CD25− T cell isolation kit after 5 days and used as a negative control.
TGFβ-induced Tregs
To generate iTregs, we used Fantini’s protocol [33]. Briefly, CD4+CD25− T cells were isolated. Then, we coated them in 24-well plates with 10 μg/ml of anti-CD3 antibody in PBS at 37 °C for 2 h. Following this, we washed the plate once with PBS before cell plating. We plated 1 ml of 2 × 106 ml−1 CD4+CD25− cells re-suspended in an X-Vivo15 serum-free medium in the anti-CD3-precoated wells in the absence of antibiotics. Anti-CD28 antibody (2 μ/g ml) and TGFβ (5 ng ml) were immediately added and the cells were incubated for 5 days. iTregs were isolated from this condition by CD4+CD25+ regulatory T cell isolation kit and use as a positive control.
In vivo study design and skin transplantation
1.5 cm2 full thickness of back-skin of C57BL/6 mice was transplanted to the back of Balb/C mice fixed with 4–6 stitches, covered with a proper bandage. Bandages were removed 7 days after transplantation and the skin graft was monitored every day, and rejected graft before day 7 was considered as a technical error and excluded from the study. Grafts were considered as a rejected graft when they showed at least 90% necrosis.
The study was designed for 10 groups with 3 kinds of treatment: some mice received 106 MSCs intravenously on day 1 and 0 after completion of the skin graft procedure and on day 2 and day 4 post-transplantation. Also, some mice received CsA (Gavage, 50 mg/kg/day) from 7 days before transplantation to the time of sacrifice. The control group received PBS. Every group contains 5 mice. The mice were sacrificed at days 5, 10, and 15 and CD4+ T cells were isolated from splenocytes of grafted mice for biological analysis.
Suppression assay
For in vitro suppressive assay, we used the basic protocol described by Collison LW [34]. Briefly, MSCs and T cells were cultured for 72 h in four in vitro conditions. 104 MSC-cultured T cells (C57BL/6) of each condition were added to 105 CD4+CD25− T cells (C57BL/6) (ratio 1:10) in the plate coated with 104 allogeneic DCs (Ballb/C) and incubated for 48 h at 37 °C. Then, BrdU labeling was added to cultures and incubated for another 24 h. Then, labeling medium was removed by centrifuge and flicking off and dried at 60 °C for 1 h. The cells were fixed by fixDenat and stored at 25 °C for 1 h and then fixDenat was removed by flacking off and tapping. Anti-BrdU-POD was added to cells and incubated at 25 °C for 90 °C. Conjugated antibody was removed by flicking and rinsing by PBS three times. After substrate solutions and color developing, the absorbance of samples was measured by ELISA reader at 370 nm (reference wavelength 429 nm). For in vivo suppressive assay, CD4+ T cells were isolated from skin-transplanted mice and added to effector T cells and DCs as in vitro.
Gene expression analysis by real-time PCR
RNA was isolated from CD+DC25+ Tregs in vitro and CD+ T cells in the spleen of skin-grafted mice after the mentioned times using Trizol (Invitrogen, USA). Then, cDNA synthesis was performed using a reverse transcription kit (Takara, Japan). Quantitative RT-PCR experiments were performed using ABI step one plus (Applied Biosystems, USA) and GAPDH were used as a housekeeping gene. The Specific primers were designed by AlleleID software (Version 7.5) (Table 1).
Statistical analysis
In vitro and in vivo data were analyzed by Prism 5th version (GraphPad software). Student t test or one-way ANOVA with post hoc and also two-way ANOVA analyses were performed depending on the number of comparatives. The data are represented as mean ± SEM, n = 4 independent experiments. ns, non-significant; *P < .05, **P < .01, ***P < .001.
*P < .05, **P < .01, ***P < .001. Correlation coefficient is significant at 0.8 < CC < 1, P***, 0.8 < CC < 0.6 P**, 0.6 < CC < 0.4 P* and non-significant at CC > 0.4. The minus before the significant correlation coefficient is the sign of negative correlation.
Results
Mesenchymal stem cells can trigger the conversion of effector T cells toward induced Foxp3+-expressing Tregs
In the present study, using four in vitro experimental conditions, we investigated the capacity of BM-MSCs to convert CD4+ T cells to iTregs. MSCs were obtained from bone marrow cells of Balb/C mice as described in “Materials and methods.” MSC phenotype was attested by membrane expression of Sca-1 and CD44 and absence of CD34 and CD45 markers (Supplementary Fig. S1a). Moreover, their capacity to differentiate into chondrocytes and adipocytes under appropriate differentiation conditions were tested (Supplementary Fig. S1b). CD4+CD25− T cells and DCs were isolated from C57BL/6 mice and cultured alone or with allogenic MSCs in cell-cell contact and in transwell conditions for 5 days. Thereafter, the expression of Foxp3+ cells among total CD4+CD25+ T cells was evaluated at 6 h, 12 h, 24 h, 48 h, 72 h, and 5 days after culture by RT-PCR. These cells were compared with iTregs that are classically obtained by in vitro T cell activation in the presence of TGFβ and IL-2 after 5 days of culture (Fig. 1a, b). We observed that MSCs could induce Foxp3 mRNA expression in co-culture and transwell which is in accordance with previous reports [2, 3, 6, 14, 15, 35,36,37].
MSCs can induce effector T cells to Foxp3-expressing Tregs with strong immunosuppressive capacity. CD4+ effector T cells and DCs were co-cultured with allogeneic MSCs in four conditions; as described in “Materials and methods.” FOXP3 mRNA of MSC-cultured T cells was measured after 6 h, 12 h, 24 h, 48 h, 72 h, and 5 days by RT-PCR. Allogeneic MLR was performed and CD4+CD25− effector T cells isolated after 5 days after MLR and used as a negative control and TGFβ- induced Treg cells were used as a positive control. The samples were normalized by GAPDH and compared with the negative control in a co-culture and b transwell conditions. c For the suppressive assay, MSC-cultured T cells were isolated after 48 h and added to T cells that were stimulated with anti-CD3 and anti-CD28. After 48 h, BrdU was used to measure the proliferation of CD4+ T cells and compared with MLR. Data are represented as mean ± SEM; n = 4 independent experiments and significant results as *P < .05, **P < .01, ***P < .001
In order to investigate whether these BM-MSC-induced Tregs are as potent as TGFβ-iTregs to suppress effector T cells, all CD4+ T cells were collected from previous co-culture/transwell conditions and were added in a new suppression assay in a 1/10 ratio. We observed MSC-cultured CD4+ T cells were able to dramatically suppress responder CD4+CD25− T cells compared to controls with the highest suppression rate for the MSC + DC condition after 48 h (Fig. 1).
Modification of RUNX complex (RUNX1, RUNX3, and CBFB) gene expression in mesenchymal stem cell-induced Tregs
The high expression of RUNX complex and its binding to TSDR lead to high and stable Foxp3 expression, therefore, maintaining tolerance. Three RUNX binding sites were reported in the Foxp3 promoter that was conserved between human, mouse, and rat [38]. RUNX complex binds to TSDR to trigger Foxp3 expression. The highest suppressive and stable phenotype was observed when this complex binds to demethylated TSDR [25, 39]. Since we observed an enhanced demethylation of TSDR in MSC-induced Tregs (unpublished data) and due to the important role of RUNX complex in the suppressive phenotype of Tregs, we investigated whether BM-MSCs can enhance RUNX complex mRNA expression or not. We thus investigated the expression level of Runx1, Runx3, and CBFB genes, implicated in stable Foxp3 expression in the four experimental conditions previously described in “Materials and methods.”
After 6 h, 12 h, 24 h, 48 h, 72 h, and 5 days of culture, mRNA of MSC-induced Tregs were extracted and compared to effector CD4+CD25− T cells (control) and TGFβ-induced Tregs. The samples were normalized by expression of an endogenous housekeeping gene (GAPDH). TGFβ-iTregs expressed higher amounts of Runx1, Runx3, and CBFB, as compared to the control group (Fig. 2). These observations were even more marked with MSC-induced Tregs compared to TGFβ-induced Tregs and increased over the time of co-culture regardless of the Tregs induction method (Fig. 2).
Modification of Runx1, Runx3, and CBFB gene expression in MSC-induced Tregs. CD4+ effector T cell and DCs were isolated and co-cultured with allogeneic MSCs in four conditions; as described in “Materials and methods.” Total mRNA of MSC-cultured T cells was extracted after 6 h, 12 h, 24 h, 48 h, 72 h, and 5 d and expression of Runx1, Runx3, and CBFB was assessed by quantitative RT-PCR. Allogeneic MLR was performed and CD4+CD25− effector T cells isolated after 5 days after MLR and used as a negative control and TGFβ- induced Treg cells were used as a positive control. The samples were normalized by expression of an endogenous housekeeping gene (GAPDH) and compared with the negative control. Data are represented as mean ± SEM; n = 4 independent experiments and significant result as *P < .05, **P < .01, ***P < .001. Correlation of each gene with FOXP3 was shown under its expression graph. Spearman correlation coefficient r and significance levels were shown on the top of each graph. The significance level for correlations is represented as 0.8 < CC < 1, P***, 0.8 < CC < 0.6 P**, and 0.6 < CC < 0.4 P* and CC > 0.4 is considered non-significant
We observed a modest induction of RUNX1, RUNX3, and CBFB in MSC + MLR conditions up to 72 h of co-culture compared to MSC + TC and MSC + DC conditions (Fig. 2). However, the expression of mentioned genes was not stable in MSC + TC group as they decreased at day 5 post-co-culture. In contrast, the expression of these genes was stable or continued to increase between 72 h and 5 days in other conditions. When the isolated DCs were co-cultured with allogeneic MSCs for 24 h and then added to total CD4+ T cells, we detected the highest amount of RUNX1, RUNX3, and CBFB expression from 72 h and after 5 days of co-culture (Fig. 2). Interestingly, we always observed a strong correlation for each studied gene with Foxp3 expression (Fig. 2). All these observations were confirmed by performing the same experiments using transwell conditions (Supplementary Fig. S2).
Bone marrow-derived mesenchymal stem cell could not induce MBD2 in mesenchymal stem cell-induced Treg
Due to the role of MBD2 in TSDR demethylation, we investigated whether demethylation of TSDR is related to upregulation of MBD2 by MSCs or not.
As described before, after 6 h, 12 h, 24 h, 48 h, 72 h, and 5 days of co-culture, MBD2 mRNA of MSC-induced Tregs in all in vitro conditions were compared to effector CD4+CD25− T cells (control) and TGFβ-induced Tregs. We did not observe any significant increase in MBD2 expression regardless of co-culture conditions (Fig. 3) or transwell conditions (Supplementary Fig. S3). Moreover, the correlation analysis did not show any significant correlation between Foxp3 and MBD2 expression.
Modification of Mbd2 gene expression in MSC-induced Tregs in a co-culture system. CD4+ effector T cells and DCs were isolated and cultured with allogeneic MSCs in co-culture system in four conditions, as described in “Materials and methods.” Total mRNA of MSC-cultured T cells was extracted after 6 h, 24 h, 24 h, 48 h, 72 h, and 5 days and expression of Mbd2 was assessed by quantitative RT-PCR. Allogeneic MLR was performed and CD4+CD25− effector T cells isolated after 5 days after MLR and used as a negative control and TGFβ- induced Treg cells were used as a positive control. The samples were normalized by expression of an endogenous housekeeping gene (GAPDH) and compared with the negative control. Data are represented as mean ± SEM; n = 4 independent experiments and significant result as *P < .05, **P < .01, and ***P < .001. Correlation of each gene with FOXP3 was shown under its expression graph. Spearman correlation coefficient r and significance levels were shown on the top of each graph. The significance level for correlations is represented as 0.8 < CC < 1, P***, 0.8 < CC < 0.6 P**, and 0.6 < CC < 0.4 P* and CC > 0.4 is considered non-significant
Bone marrow-derived mesenchymal stem cells infusion delays skin graft rejection and enhances RUNX1, RUNX3, and CBFB expression
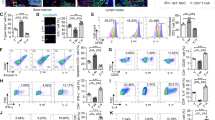
We next evaluated the capacity of BM-MSCs to suppress effector T cells in vivo and their ability to prolong graft survival in a mouse model of fully allogeneic skin transplantation (C57BL/6 into Balb/C) as described in the method. In parallel, we compared their effects with those of CsA treatment or mice treated with PBS. The fourth group of mice underwent autologous skin transplantation. We did not observe any graft rejection in autologous control mice as expected, whereas PBS-treated mice rapidly rejected the skin graft (MST = 10 days); MSC- and CsA-treated mice significantly enhanced graft survival (respectively MST = 17 and MST = 20.50) compared to PBS-treated mice (Fig. 4a).
BM-MSC infusion allows for delayed skin graft rejection via increased FOXP3 expression and Runx1, Runx3, and CBFB but not Mbd2 gene expression in iTregs. Balb/C mice were grafted with full-thickness allogeneic back skin from C57BL/6 mice and treated with MSCs, cyclosporine, or PBS as mentioned before. a Graft survival time. (b) The mice were scarifcied in days 5, 10, and 15 and total mRNA of splenic CD4+ T cells was extracted. The expression of Runx1, Runx3, CBFB, and Mbd2 was assessed by quantitative RT-PCR. The samples were normalized by expression of an endogenous housekeeping gene (GAPDH) and compared with the PBS-treated group. The data are represented as mean ± SEM; n = 5 in each group. Significant results were also showed as *P < .05, **P < .01, and ***P < .001
In vitro, we were able to demonstrate that induction of Tregs by MSCs was associated with increased Foxp3 expression as well as modified RUNX1, RUNX3, and CBFB gene expression in iTregs. Here, we investigated whether the same mechanisms were involved when MSCs were administered directly in vivo. We thus reproduced allogeneic skin graft experiments, while we sacrificed the mice and harvested T cells from their splenocytes at day 5, 10, and 15 post-transplantation. Furthermore, we extracted total mRNA and measured the expression of Foxp3, RUNX1, RUNX3, CBFB, and MBD2. The expression of Foxp3 gene was elevated merely in MSC-treated mice and not in the two other groups (Fig. 4b). Moreover, at all the three time-points, Foxp3 expression was associated with modified gene expression of RUNX1, RUNX3, and CBFB but not MBD2 comparable to what was observed in vitro experiments (Fig. 4b). Therefore, here for the first time, we reveal the association of Tregs induction by MSCs and strong RUNX complex expression modification.
Discussion
The multi-lineage differentiation and immunomodulation capacities of MSCs have made this somatic progenitor cells an interesting target for cell therapy. Immunomodulating property of MSCs was studied in several animal models of autoimmune diseases such as experimental autoimmune encephalomyelitis (EAE) [40] and collagen-induced arthritis (CIA) [41]. In addition, they were used in different tissue and organ transplantations clinical trials including GVHD [42, 43], islet transplantation [44], liver transplantation [45], and renal transplantation [46] with already promising results. Around 500 MSC-related clinical trials were registered on NIH Clinical Trial Database as of 2016 (https://clinicaltrials.gov/); nearly half of which are based on immunomodulatory effects of MSCs.
The MSCs ability to induce effector T cells to iTregs was previously reported [2, 3, 6, 14, 15, 35,36,37, 47]. Moreover, it was previously observed that stability and suppressive features of iTregs strongly depend on stable Foxp3 expression. In parallel, RUNX complex and epigenetic mechanisms were demonstrated to be implicated in the stability of Foxp3 [37, 48, 49]. Here, we show for the first time that induction of Tregs by MSC is directly associated with strong modifications of RUNX complex genes involved in the stable Foxp3 expression.
In this work, we used four different in vitro conditions that allow MSCs to generate iTregs. Depending on the experimental model used, we observed that variable amounts of iTregs were produced. When we looked at the 3 genes (RUNX1, RUNX3, CBFB) implied in expression and suppressive feature of the Foxp3, we observed that the transcripts of all these genes were globally modified in the same way, all leading to expression of the Foxp3 and consequently, its stability. Indeed, the addition of MSCs to T cell culture can remarkably increase the expression of RUNX1, RUNX3, and CBFB all implied in the Foxp3 expression process, whereas the expression of MBD2 implied in TSDR demethylation expression remains unchanged. These convergent data obtained from four different experimental conditions highlight the robustness of our observations. Furthermore, compared to direct contact of MSCs to T cells, the addition of DCs previously co-cultured with MSCs induced more stable Foxp3 expression at day 5, which is associated with strong immunosuppressive capacities. This proposes the significant capacity of MSCs to modify antigen-presenting cell (APCs) that are then capable of inducing Tregs from CD4+ T cells. Here again, it is in this MSC-DC condition that we observed the more important modifications for RUNX1, RUNX3, and CFBF expression, suggesting that the more these genes are expressed, the more suppressive iTregs are. In contrast, we also observed an expression reduction in target genes in MSC + TC condition on day 5. This might be due to the absence of TCR-induced T cell signaling in the absence of DC stimulation.
In line with this, enhanced demethylation of TSDR and enhanced expression of RUNX complex allow more and stable Foxp3 expression that consequently leads to more suppressive MSC-induced Tregs.
In 2013, Wang et al. showed that MBD2 can demethylate TSDR in regulatory T cells [24]. Here, we observed that MBD2 mRNA expression does not enhance in MSC-induced Tregs. It seems that MSCs promote TSDR demethylation via other methyl transferases but not MBD2.
In order to evaluate the importance of our results, it was essential to assess whether the mechanisms we identified in vitro were also confirmed in vivo. To this end, we used a more immunogenic experimental model that consists of transplanting allogeneic skin into immune-competent mice. Although less remarkable than CsA-treated mice, MSC administration statistically significantly delayed graft rejection compared to PBS-treated mice. This effect is accompanied by greater in vitro immunosuppressive properties of the T cells collected in the spleens of mice treated with MSCs compared to those in the CsA-treated mice (data not shown). This was convergent with an increased mRNA Foxp3 expression as well as RUNX complex gene modifications improved in mice treated with MSCs compared to mice of the two other controls. In accordance with our result, the team of Choi et al. showed that cyclosporine can block RUNX1 expression. It seems that MSCs prolong skin graft via enhancing expression of RUNX complex and stabilizing Foxp3 [50].
The immunosuppressive capacities of MSCs make these cells a therapeutic tool of great potential to control immune pathologies. In the context of alloreactivity, we have shown that MSCs have the capacity to induce Tregs from conventional T cells. The mechanisms behind this effect are through keeping the stability of the Foxp3 gene. We identified for the first time that RUNX1, RUNX3, and CBFB as master genes in maintaining Treg stability play an indispensable role in this action. Since both in vitro and in vivo data are quite similar, we assume that they reflect a possible mechanism that remains to be demonstrated in autoimmunity. Although we observed significant increases in the mentioned gene in the presence of MSCs, more investigations such as blocking these gene expression and protein investigation are needed to completely confirm this mechanism.
Abbreviations
- MSCs:
-
Mesenchymal stem cells
- iTregs:
-
Induced regulatory T cells
- DCs:
-
Dendritic cells
- NKs:
-
Natural killer cells
- IL-12:
-
Interleukin-12
- TNFα:
-
Tumor necrosis factor-α
- nTregs:
-
Natural Tregs
References
Wang Y, Zhang A, Ye Z, Xie H, Zheng S. Bone marrow-derived mesenchymal stem cells inhibit acute rejection of rat liver allografts in association with regulatory T-cell expansion. Transplant. Proc. [Internet]. 2009;41:4352–6. [cited 2017 Feb 7] Available from: http://linkinghub.elsevier.com/retrieve/pii/S0041134509013669
Ge W, Jiang J, Arp J, Liu W, Garcia B, Wang H. Regulatory T-cell generation and kidney allograft tolerance induced by mesenchymal stem cells associated with indoleamine 2,3-dioxygenase expression. Transplantation [Internet]. 2010;90:1312–20. [cited 2017 Feb 7] Available from: http://content.wkhealth.com/linkback/openurl?sid=WKPTLP:landingpage&an=00007890-201012270-00014
Casiraghi F, Azzollini N, Cassis P, Imberti B, Morigi M, Cugini D, et al. Pretransplant infusion of mesenchymal stem cells prolongs the survival of a semiallogeneic heart transplant through the generation of regulatory T cells. J Immunol [Internet]. 2008;181:3933–46. [cited 2017 Feb 7] Available from: http://www.ncbi.nlm.nih.gov/pubmed/18768848
Cohen JL, Sudres M. A role for mesenchymal stem cells in the control of graft-versus-host disease. Transplantation [Internet]. 2009;87:S53–4. [cited 2017 Feb 13] Available from: http://www.ncbi.nlm.nih.gov/pubmed/19424007
Parekkadan B, Tilles AW, Yarmush ML. Bone marrow-derived mesenchymal stem cells ameliorate autoimmune enteropathy independently of regulatory T cells. Stem Cells [Internet]. 2008;26:1913–9. [cited 2017 Feb 7] Available from: http://doi.wiley.com/10.1634/stemcells.2007-0790
Duffy MM, Ritter T, Ceredig R, Griffin MD. Mesenchymal stem cell effects on T-cell effector pathways. Stem Cell Res Ther [Internet]. 2011;2:34. [cited 2017 Feb 7] Available from: http://www.ncbi.nlm.nih.gov/pubmed/21861858
Reinders ME, Hoogduijn MJ. NK cells and MSCs: possible implications for MSC therapy in renal transplantation. J Stem Cell Res Ther [Internet]. 2014;4:1000166. Europe PMC Funders; [cited 2017 Feb 7] Available from: http://www.ncbi.nlm.nih.gov/pubmed/24900946
Cahill EF, Tobin LM, Carty F, Mahon BP, English K. Jagged-1 is required for the expansion of CD4+ CD25+ FoxP3+ regulatory T cells and tolerogenic dendritic cells by murine mesenchymal stromal cells. Stem Cell Res Ther [Internet]. 2015;6:19. BioMed Central; [cited 2017 Feb 7] Available from: http://www.ncbi.nlm.nih.gov/pubmed/25890330
Djouad F, Charbonnier L-M, Bouffi C, Louis-Plence P, Bony C, Apparailly F, et al. Mesenchymal stem cells inhibit the differentiation of dendritic cells through an interleukin-6-dependent mechanism. Stem Cells [Internet]. 2007;25:2025–32. [cited 2017 Feb 7] Available from: http://doi.wiley.com/10.1634/stemcells.2006-0548
Ramasamy R, Fazekasova H, Lam EW-F, Soeiro I, Lombardi G, Dazzi F. Mesenchymal stem cells inhibit dendritic cell differentiation and function by preventing entry into the cell cycle. Transplantation [Internet]. 2007;83:71–6. [cited 2017 Feb 7] Available from: http://content.wkhealth.com/linkback/openurl?sid=WKPTLP:landingpage&an=00007890-200701150-00014
Li Y-P, Paczesny S, Lauret E, Poirault S, Bordigoni P, Mekhloufi F, et al. Human mesenchymal stem cells license adult CD34+ hemopoietic progenitor cells to differentiate into regulatory dendritic cells through activation of the Notch pathway. J Immunol [Internet]. 2008;180:1598–608. [cited 2017 Feb 14] Available from: http://www.ncbi.nlm.nih.gov/pubmed/18209056
Sakaguchi S, Yamaguchi T, Nomura T, Ono M, Regulatory T. Cells and immune tolerance. Cell. 2008;133(5):775–87. https://doi.org/10.1016/j.cell.2008.05.009.
Lin X, Chen M, Liu Y, Guo Z, He X, Brand D, et al. Advances in distinguishing natural from induced Foxp3(+) regulatory T cells. Int J Clin Exp Pathol [Internet]. 2013;6:116–23. e-Century Publishing Corporation; [cited 2017 Feb 14] Available from: http://www.ncbi.nlm.nih.gov/pubmed/23329997
Kavanagh H, Mahon BP. Allogeneic mesenchymal stem cells prevent allergic airway inflammation by inducing murine regulatory T cells. Allergy [Internet]. 2011;66:523–31. [cited 2017 Feb 11] Available from: http://www.ncbi.nlm.nih.gov/pubmed/21091718
Tatara R, Ozaki K, Kikuchi Y, Hatanaka K, Oh I, Meguro A, et al. Mesenchymal stromal cells inhibit Th17 but not regulatory T-cell differentiation. Cytotherapy [Internet]. 2011;13:686–94. [cited 2017 Feb 11] Available from: http://www.ncbi.nlm.nih.gov/pubmed/21171824
Ge W, Jiang J, Arp J, Liu W, Garcia B, Wang H. Regulatory T-cell generation and kidney allograft tolerance induced by mesenchymal stem cells associated with indoleamine 2,3-dioxygenase expression. Transplantation [Internet]. 2010;90:1312–20. [cited 2017 Feb 11] Available from: http://www.ncbi.nlm.nih.gov/pubmed/21042238
Rudensky AY. Regulatory T cells and Foxp3. Immunol Rev [Internet]. 2011;241:260–8. [cited 2017 Feb 7] Available from: http://doi.wiley.com/10.1111/j.1600-065X.2011.01018.x
Kim H-P, Leonard WJ. CREB/ATF-dependent T cell receptor–induced FoxP3 gene expression: a role for DNA methylation. J Exp Med [Internet]. 2007;204:1543–51. [cited 2017 Apr 14] Available from: http://www.ncbi.nlm.nih.gov/pubmed/17591856
Szyf M, Bhattacharya SK, Ramchandani S, Cervoni N. A mammalian protein with specific demethylase activity for mCpG DNA. Nature [Internet]. 1999;397:579–83. [cited 2017 Apr 30] Available from: http://www.ncbi.nlm.nih.gov/pubmed/10050851
Balada E, Ordi-Ros J, Serrano-Acedo S, Martinez-Lostao L, Vilardell-Tarres M. Transcript overexpression of the MBD2 and MBD4 genes in CD4+ T cells from systemic lupus erythematosus patients. J Leukoc Biol [Internet]. 2007;81:1609–16. [cited 2017 Apr 30] Available from: http://www.ncbi.nlm.nih.gov/pubmed/17360956
Lei W, Luo Y, Lei W, Luo Y, Yan K, Zhao S, et al. Abnormal DNA methylation in CD4+ T cells from patients with systemic lupus erythematosus, systemic sclerosis, and dermatomyositis. Scand J Rheumatol [Internet]. 2009;38:369–74. [cited 2017 Apr 30] Available from: http://www.ncbi.nlm.nih.gov/pubmed/19444718
Liu C-C, Fang T-J, Ou T-T, Wu C-C, Li R-N, Lin Y-C, et al. Global DNA methylation, DNMT1, and MBD2 in patients with rheumatoid arthritis. Immunol Lett [Internet]. 2011;135:96–9. [cited 2017 Apr 30] Available from: http://www.ncbi.nlm.nih.gov/pubmed/20937307
Zhang P, Su Y, Chen H, Zhao M, Lu Q. Abnormal DNA methylation in skin lesions and PBMCs of patients with psoriasis vulgaris. J Dermatol Sci [Internet]. 2010;60:40–2. [cited 2017 Apr 30] Available from: http://www.ncbi.nlm.nih.gov/pubmed/20800455
Wang L, Liu Y, Han R, Beier UH, Thomas RM, Wells AD, et al. Mbd2 promotes Foxp3 demethylation and T-regulatory-cell function. Mol Cell Biol [Internet]. 2013;33:4106–15. [cited 2017 Apr 30] Available from: http://www.ncbi.nlm.nih.gov/pubmed/23979593
Rudra D, Egawa T, Chong MMW, Treuting P, Littman DR, Rudensky AY. Runx-CBFβ complexes control expression of the transcription factor Foxp3 in regulatory T cells. Nat Immunol [Internet]. 2009;10:1170–7. [cited 2017 Apr 30] Available from: http://www.ncbi.nlm.nih.gov/pubmed/19767756
Moravej A, Karimi M-H, Geramizadeh B, Hossein Aghdaie M, Kohi-Hoseinabadi O, Ebrahimnezhad S. Effect of mesenchymal stem cells on ilt3 expression in the splenocytes of skin graft recipient mice. Iran J Immunol [Internet]. 2016;13:274–88. [cited 2017 Jun 23] Available from: http://www.ncbi.nlm.nih.gov/pubmed/27999239
English K, Ryan JM, Tobin L, Murphy MJ, Barry FP, Mahon BP. Cell contact, prostaglandin E2 and transforming growth factor beta 1 play non-redundant roles in human mesenchymal stem cell induction of CD4+CD25High forkhead box P3+ regulatory T cells. Clin Exp Immunol [Internet]. 2009;156:149–60. [cited 2017 Apr 14] Available from: http://www.ncbi.nlm.nih.gov/pubmed/19210524
Gonzalez-Rey E, Gonzalez MA, Varela N, O’Valle F, Hernandez-Cortes P, Rico L, et al. Human adipose-derived mesenchymal stem cells reduce inflammatory and T cell responses and induce regulatory T cells in vitro in rheumatoid arthritis. Ann Rheum Dis [Internet]. 2010;69:241–8. [cited 2017 Apr 14] Available from: http://www.ncbi.nlm.nih.gov/pubmed/19124525
Luz-Crawford P, Kurte M, Bravo-Alegría J, Contreras R, Nova-Lamperti E, Tejedor G, et al. Mesenchymal stem cells generate a CD4+CD25+Foxp3+ regulatory T cell population during the differentiation process of Th1 and Th17 cells. Stem Cell Res Ther [Internet]. 2013;4:65. [cited 2017 Apr 14] Available from: http://www.ncbi.nlm.nih.gov/pubmed/23734780
Cutler AJ, Limbani V, Girdlestone J, Navarrete CV. Umbilical cord-derived mesenchymal stromal cells modulate monocyte function to suppress T cell proliferation. J Immunol [Internet]. 2010;185:6617–23. [cited 2017 Apr 14] Available from: http://www.ncbi.nlm.nih.gov/pubmed/20980628
Wang Q, Sun B, Wang D, Ji Y, Kong Q, Wang G, et al. Murine bone marrow mesenchymal stem cells cause mature dendritic cells to promote T-cell tolerance. Scand J Immunol [Internet]. 2008;68:607–15. [cited 2017 Apr 14] Available from: http://doi.wiley.com/10.1111/j.1365-3083.2008.02180.x
English K, Ryan JM, Tobin L, Murphy MJ, Barry FP, Mahon BP. Cell contact, prostaglandin E2 and transforming growth factor beta 1 play non-redundant roles in human mesenchymal stem cell induction of CD4+CD25High forkhead box P3+ regulatory T cells. Clin Exp Immunol [Internet]. 2009;156:149–60. [cited 2017 Feb 11] Available from: http://www.ncbi.nlm.nih.gov/pubmed/19210524
Fantini MC, Dominitzki S, Rizzo A, Neurath MF, Becker C. In vitro generation of CD4+CD25+ regulatory cells from murine naive T cells. Nat Protoc [Internet]. 2007;2:1789–94. Nature Publishing Group; [cited 2017 Feb 8] Available from: http://www.nature.com/doifinder/10.1038/nprot.2007.258
Collison LW, Vignali DAA. In vitro Treg suppression assays. Methods Mol Biol [Internet]. 2011;21–37. [cited 2017 Feb 8] Available from: http://www.ncbi.nlm.nih.gov/pubmed/21287326
Madec AM, Mallone R, Afonso G, Abou Mrad E, Mesnier A, Eljaafari A, et al. Mesenchymal stem cells protect NOD mice from diabetes by inducing regulatory T cells. Diabetologia [Internet]. 2009;52:1391–9. [cited 2017 Feb 11] Available from: http://www.ncbi.nlm.nih.gov/pubmed/19421731
Patel SA, Meyer JR, Greco SJ, Corcoran KE, Bryan M, Rameshwar P. Mesenchymal stem cells protect breast cancer cells through regulatory T cells: role of mesenchymal stem cell-derived TGF-β. J Immunol [Internet]. 2010;184:5885–94. [cited 2017 Feb 11] Available from: http://www.ncbi.nlm.nih.gov/pubmed/20382885
van Loosdregt J, Fleskens V, Fu J, Brenkman AB, Bekker CPJ, Pals CEGM, et al. Stabilization of the transcription factor Foxp3 by the deubiquitinase USP7 increases treg-cell-suppressive capacity. Immunity [Internet]. 2013;39:259–71. [cited 2017 Feb 8] Available from: http://www.ncbi.nlm.nih.gov/pubmed/23973222
Durst KL, Hiebert SW. Role of RUNX family members in transcriptional repression and gene silencing. Oncogene [Internet]. 2004;23:4220–4. [cited 2017 Apr 30] Available from: http://www.ncbi.nlm.nih.gov/pubmed/15156176
Ono M, Yaguchi H, Ohkura N, Kitabayashi I, Nagamura Y, Nomura T, et al. Foxp3 controls regulatory T-cell function by interacting with AML1/Runx1. Nature [Internet]. 2007;446:685–9. [cited 2017 Apr 30] Available from: http://www.ncbi.nlm.nih.gov/pubmed/17377532
Bowles AC, Scruggs BA, Bunnell BA. Mesenchymal stem cell-based therapy in a mouse model of experimental autoimmune encephalomyelitis (EAE). Methods Mol Biol [Internet]. 2014;303–19. [cited 2017 Feb 11] Available from: http://www.ncbi.nlm.nih.gov/pubmed/25173393
Park K-H, Mun CH, Kang M-I, Lee S-W, Lee S-K, Park Y-B. Treatment of collagen-induced arthritis using immune modulatory properties of human mesenchymal stem cells. Cell Transplant [Internet]. 2015;25:1057–72. [cited 2017 Feb 11] Available from: http://www.ncbi.nlm.nih.gov/pubmed/25853338
Le Blanc K, Frassoni F, Ball L, Locatelli F, Roelofs H, Lewis I, et al. Mesenchymal stem cells for treatment of steroid-resistant, severe, acute graft-versus-host disease: a phase II study. Lancet. 2008;371(9624):1579–86. https://doi.org/10.1016/S0140-6736(08)60690-X.
Le Blanc K, Rasmusson I, Sundberg B, Götherström C, Hassan M, Uzunel M, et al. Treatment of severe acute graft-versus-host disease with third party haploidentical mesenchymal stem cells. Lancet. 2004;363(9419):1439–41. https://doi.org/10.1016/S0140-6736(04)16104-7.
English K. Mesenchymal stem cells to promote islet transplant survival. Curr Opin Organ Transplant [Internet]. 2016;21:568–73. [cited 2017 Feb 11] Available from: http://content.wkhealth.com/linkback/openurl?sid=WKPTLP:landingpage&an=00075200-201612000-00005
Tian Y, Wang J, Wang W, Ding Y, Sun Z, Zhang Q, et al. Mesenchymal stem cells improve mouse non-heart-beating liver graft survival by inhibiting Kupffer cell apoptosis via TLR4-ERK1/2-Fas/FasL-caspase3 pathway regulation. Stem Cell Res Ther [Internet]. 2016;7:157. BioMed Central; [cited 2017 Feb 11] Available from: http://stemcellres.biomedcentral.com/articles/10.1186/s13287-016-0416-y
Casiraghi F, Perico N, Cortinovis M, Remuzzi G. Mesenchymal stromal cells in renal transplantation: opportunities and challenges. Nat Rev Nephrol [Internet]. 2016;12:241–53. Nature Research [cited 2017 Feb 11] Available from: http://www.nature.com/doifinder/10.1038/nrneph.2016.7
Khosravi M, Karimi MH, Hossein Aghdaie M, Kalani M, Naserian S, Bidmeshkipour A. Mesenchymal stem cells can induce regulatory T cells via modulating miR-126a but not miR-10a. Gene [Internet]. 2017;627:327–36. [cited 2017 Jul 14] Available from: http://www.ncbi.nlm.nih.gov/pubmed/28600182
Barbi J, Pardoll DM, Pan F. Ubiquitin-dependent regulation of Foxp3 and Treg function. Immunol Rev [Internet]. 2015;266:27–45. [cited 2017 Feb 11] Available from: http://www.ncbi.nlm.nih.gov/pubmed/26085205
Chen Z, Barbi J, Bu S, Yang H-Y, Li Z, Gao Y, et al. The ubiquitin ligase Stub1 negatively modulates regulatory T cell suppressive activity by promoting degradation of the transcription factor Foxp3. Immunity [Internet]. 2013;39:272–85. [cited 2017 Feb 8] Available from: http://linkinghub.elsevier.com/retrieve/pii/S1074761313003348
Choi S-C, Lee H, Choi J-H, Kim J-H, Park C-Y, Joo H-J, et al. Cyclosporin A induces cardiac differentiation but inhibits hemato-endothelial differentiation of P19 cells. Aalto-Setala K, editor. PLoS One [Internet]. 2015;10:e0117410. [cited 2017 Apr 30] Available from: http://www.ncbi.nlm.nih.gov/pubmed/25629977.
Acknowledgements
MK would like to express her deep gratitude to Professor José L. Cohen, Shiraz Organ Transplant Research Center, IFRES-INT, and INSERM U1197 team 1, for constant supporting.
Author information
Authors and Affiliations
Contributions
MK built and performed the experiments, analyzed the data, and wrote the manuscript. MHK and AB contributed to building the research and revised the manuscript. AM performed the experiments and revised the manuscript. SHA analyzed the data and revised the manuscript. SN assisted with experimental revisions and wrote the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
SN and MHK are co-last authors
Electronic supplementary material
ESM 1
(DOCX 1935 kb)
Rights and permissions
About this article
Cite this article
Khosravi, M., Bidmeshkipour, A., Moravej, A. et al. Induction of CD4+CD25+Foxp3+ regulatory T cells by mesenchymal stem cells is associated with RUNX complex factors. Immunol Res 66, 207–218 (2018). https://doi.org/10.1007/s12026-017-8973-4
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12026-017-8973-4