Abstract
CD4 T cells are an integral part of adaptive immunity. microRNAs have been identified as fundamental regulators of post-transcriptional programs and to play roles in T lymphocytes’ development, differentiation, and effector functions. To better understand the role of miRNAs in T cells and to identify potential therapeutic tools and targets, we have undertaken studies of miRNAs that modulate or are modulated by T-cell receptor signaling. We identified miR-181a as a key regulator of TCR signaling strength, and hence T-cell development, and the miR-17-92 cluster as an important player in CD4 T cells’ response against antigens. These discoveries, coupled with work by other researchers, reveal the power and importance of miRNA-mediated regulation in T-cell responses and offer new insights into the burgeoning field of immunoregulation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A properly functioning immune system must distinguish foreign and self-antigens and mount a response against the former with remarkable sensitivity and specificity [1]. T cells achieve this goal by specific antigen recognition through their antigen receptor (the TCR) and a highly regulated development and differentiation process both in the thymus and in secondary lymphoid tissues. The signal transduction machinery downstream of TCR:peptide major histocompatibility complex (pMHC) engagement has been heavily studied, and the detailed functions of several kinases, phosphatases, and adaptor/scaffold molecules in this pathway are well documented [2–5]. As we better understand these signaling networks involved in T-cell activation and differentiation, we are increasingly amazed by the elegant precision with which these networks are dynamically regulated to ensure the demanded functional outcome. However, many questions remain. For example, how is the signaling modulated to ensure proper development of thymocytes, such as maintaining the higher sensitivity of a DP T cell to facilitate positive and negative selection? Or, how is this network re-programmed when a T cell responds to its antigen, and how do pathogens or tumor cells utilize such regulatory mechanisms to favor their invasion? Or, how does this network integrate other cues from the microenvironment to turn on lineage-specific genes and silence the fate-unrelated genes when a T cell differentiates into diverse effector or regulatory subsets? While multiple approaches might be applied to clarify such questions and to perturb such networks for therapeutic benefits, our laboratory is focusing on investigating how these processes are regulated by microRNAs (miRNAs).
miRNAs are 22- to 24-nt small, non-coding RNAs that modulate gene expression post-transcriptionally through inducing mRNA degradation or translational repression [6]. As the overall importance of miRNAs for hematopoiesis [7] and immune regulation [8] has been now well recognized [9], illustrating the function and mechanisms of individual miRNAs in specific immune cells is gaining increasing interest. Using a high-throughput expression profiling platform developed in the laboratory, we identified a large group of miRNAs that are dynamically regulated during T-cell development, following antigen challenge, during differentiation, as well as during infectious disease progression. Specifically, we are currently working on elucidating the function and molecular mechanisms of several miRNAs in regulating these processes.
Regulating thymocyte development: mir-181a as an intrinsic antigen sensitivity rheostat
The elaborate T-cell responses to antigens are largely dictated by the affinity of TCR:pMHC complexes, particularly their dissociation rate. In general, peptides with slower dissociation rates elicit stronger TCR signals and lead to higher T-cell reactivity to antigenic peptides [10–13]. Variations in the antigenic peptide affinity to the TCR may lead to both quantitative and qualitative changes in its ability to elicit the TCR signaling cascade and T-cell responses [14]. With increasing dissociation rates, peptide ligands are ranked from agonists (ligands that evoke full T-cell responses) to weak agonists (ligands that are able to activate T cells in substantially higher concentrations), to antagonists (ligands that are unable to activate T cells by themselves and block TCR activation to otherwise stimulatory concentrations of agonist ligand) [15], to co-agonists (ligands that are unable to stimulate the TCR but can assist agonist signaling) [16], to null peptides (ligands that cannot participate in TCR signaling). Although a number of models have been proposed to explain the kinetic discrimination in T-cell activation, how T cells sense quantitative changes in antigenic peptide affinity through the TCR and yield both quantitatively and qualitatively different responses is still an intensive area of study.
More interestingly, T-cell responses and TCR signaling to a specific antigen also vary in T cells of different developmental stages, suggesting that T-cell sensitivity to antigen is regulated during development [17–20]. Qualitatively, in immature CD4 CD8 double positive thymocytes, low-affinity antigenic peptides that are unable to activate mature effector T cells are sufficient to induce strong activation and clonal deletion [17]; antagonists, normally inhibitory to effector T cells, can induce positive selection [21, 22] or even negative selection [8]. Quantitatively, while mature CD4 [23, 24] and CD8 [25] T cells demand 10 agonist pMHCs in order to form a mature immunological synapse and execute a sustained effector response, two are sufficient to trigger apoptosis in DP thymocytes [26]. These observations demonstrate that T-cell sensitivity is intrinsically regulated to ensure the proper development of specificity and sensitivity to foreign antigens while avoiding self recognition at the mature stage. A few studies have described factors that may help explain this fascinating phenomenon. For example, CD5 could influence the fate of DP cells by acting as a negative regulator of TCR-mediated signal transduction [27]; alternative sialylation of cell surface molecules could partially alter TCR sensitivity during development [28]; and very recently, the reciprocal action of CD45 and Csk tightly regulates the function of Lck, the key kinase initiating the whole signaling cascade, and therefore the responsiveness of a T cell [29]. However, other molecular programs are clearly at play.
Our initial interest was the elucidation of the cell-intrinsic factors responsible for modulating the sensitivity of the TCR throughout development, which led us into the miRNA field. miR-181a had previously been shown to impact both B- and T-cell development when expressed in hematopoietic progenitor cells [7] and also had high expression in the thymus [7]. Upon profiling of the expression levels in T cells throughout thymic development, we observed a dynamic regulation of miR-181a levels with a gradual decrease in copy number as maturation progressed [8]. When one considers that the cells are also changing in size, the relative cytosolic concentration matches what one would predict for a modulator of TCR signaling strength, given the sensitivity pattern outlined previously. Using a transgenic TCR system, we showed that mature T cells expressing ectopic miR-181a required only two peptides to elicit a half-maximal calcium response, an ~two-fold increase over the 5 peptides normally required. More strikingly, this increase also allows miR-181a to effectively convert an antagonist to an agonist, eliciting TCR signaling. The mechanism involves the downregulation of a network of phosphatases responsible for negatively regulating distinct steps of TCR signaling: PTPN22, which removes activating phosphorylation on Lck and ZAP70 [30, 31]; SHP2, an effector for inhibitory transmembrane adapter proteins (TRAPs) such as SIT and TRIM, which have also been shown to influence thymocyte sensitivity and tolerance [32]; DUSP5, which dephosphorylates activated ERK in the nucleus; and DUSP6, which targets cytosolic ERK1/2 [33]. Notably, miR-181a only moderately represses each target (a decrease in protein level as small as 40%), and the synergistic repression of multiple targets in the network is required. The summed reduction in the negative regulatory network results in a dramatic increase in both steady-state and TCR-signaling-induced levels of activated Lck and ERK1/2.
Based on these molecular aspects, one would expect miR-181a to play a key role during the selection process in the thymus. Using fetal thymic organ cultures (FTOC), we demonstrated that elimination of miR-181a via a miR-181a-specific antagomir increased the number of T cells coming out of negative selection and decreased the number in a positive selection setting [8]. The direct prediction from the former effect is the generation of mature self-reactive T cells and a skewing of the TCR repertoire toward higher affinities. In support of this notion, SP cells developed ex vivo in the presence of antagomir-181a inhibition contain a population that responds to syngeneic APCs with constant elevation of CD69, a marker for early T-cell activation, and production of proinflammatory cytokines such as IFNγ, TNF, and IL-17. Furthermore, to test whether there is an evidential repertoire shift generating autoresponsive T cells specific to particular self-peptides of notably high affinity, we made use of the two self-peptides that we had previously found to have the highest capacity to stimulate the 5C.C7 TCR: GP, and ATPase 11c. By fixing the 5C.C7β chain but allowing the TCRα chain to vary as normal, we hoped to generate a pre-selection repertoire that would be biased toward recognizing these two peptides. We then allowed the 5C.C7β-transgenic thymocytes to mature in thymi in which miR-181a was inhibited. Using GP-I-Ek and ATPase-I-Ek tetramers, we could then see that miR-181a-inhibited thymi had allowed the maturation of 30-fold more GP- and ATPase-reactive T cells, when compared to otherwise unmanipulated 5C.C7β-transgenic thymi [34]. Taken together, these results strongly suggest that, in addition to the spectrum of presented endogenous peptides, the abundance of certain self antigen, and the structural characteristics (e.g. affinities) of recombined TCRs on each DP thymocytes, the net “readiness” (with respect to both quantity and quality) of key signaling intermediates represents the fourth layer of complexity in determining the outcome of selection.
A question, however, remains: how is miR-181a expression modulated throughout T-cell development? One provocative explanation is that TCR signaling itself provides direct feedback to downregulate miR-181a expression levels. By forming this loop of mutual regulation, the sensitivity of the TCR is strongly enforced in pre-selection T cells and conveniently restrained in post-selection ones. We have seen that miR-181a is rapidly downregulated within 1 h following TCR engagement by either negatively or positively selecting ligands [34]. Although the mechanism mediating this process is still under vigorous investigation, and we cannot rule out other entirely distinct regulatory mechanisms, the rapid change of miR-181a levels upon TCR engagement would eloquently explain the attenuated response of T cells throughout their thymic development. Given that DN3/4 T cells have higher expression of miR-181a than DP cells and that miR-181a enhances DN3/4 to DP progression in OP9-DL1 co-cultures [35], one might also speculate that the higher sensitivity (perhaps even hypersensitivity) in DNs might allow for the sensing of signals from the oligomerization of pre-TCR [36], which could conceivably then downregulate miR-181a to the level seen in DP cells. Furthermore, it could provide another layer of modulation to affirm peripheral tolerance. It was reported that T cells selected by high-affinity, positively selecting ligands appear to be attenuated in their responsiveness in the periphery [37]. If relatively stronger TCR signals in the thymus induce greater miR-181a downregulation, and if this low miR-181a expression is maintained during the remainder of the T-cell’s maturation, then this low miR-181a might render the resulting mature T-cell population hyporesponsive, even though they carry higher-affinity TCRs.
The elucidation of the function of miR-181a and mechanisms regulating its expression will not only offer further insight into T-cell development, especially with respect to the role of TCR signaling, but should also help fill in some gaps related to the development of autoimmune diseases. Autoimmunity is an undesirable immune response that occurs when the immune system goes awry and attacks its host. In many cases, it seems to be the result of a breakdown in T-cell tolerance. Central tolerance evolved to enable each mature T cell to discriminate self from non-self in the context of MHC and ideally there would be no self-reactive TCRs in the peripheral mature T-cell repertoire [38]. However, there are self-reactive T cells that can escape negative selection and enter the peripheral T-cell pool despite expression of their cognate antigen in the thymus [39, 40]. Under normal circumstances, this potential risk is well buffered by the mechanisms of peripheral tolerance [41], but, as in any buffer system, buffering capacity is finite. One of the challenges to this delicate balance is the avidity of T-cell responses. The sensitivity of a mature T cell directly determines its avidity. Some evidence suggests that T cells with high avidity toward self antigen are a pathogenically relevant population in peripheral tissues [42]. In contrast, other research indicates that lower avidity T cells are abundant at earlier stages of disease [43]. The proposed mechanism is that high-avidity T cells require less antigen and co-stimulation and therefore will be the first and fastest to expand [44]. Such expansion will initiate a wave of inflammation and consequently create a microenvironment more favorable for the activation of low-avidity T cells at a later stage[45]. However, when we consider that T-cell sensitivity cannot be “overgeneralized” (i.e. considered only in terms of TCR affinity), an alternative model that reconciles such seemingly disparate evidence emerges: T cells’ readiness for responses is heterogenic. As discussed previously, the “individualism” of the T cell, enforced by its internal regulatory machinery (such as miR-181a, CD45, or Csk), shapes the characteristics of T-cells’ responses to antigen. Such a model allows T cells with high-affinity antigen receptor but a low level of response readiness to escape death during thymic selection; it could also enable T cells with low-affinity receptor but a high level of response readiness to become a violent initiator of self-destruction. Taking the model one step further, the dynamic regulation on the cytosolic level of miR-181a can be considered a mechanism of active tolerance in the periphery. Nevertheless, much remains to be determined regarding the extent to which TCR signaling strength, not solely the TCR affinity, impacts the onset and progression of autoimmunity, and miRNAs certainly represent one of many avenues to explore.
Importance of miRNAs for T-cell effector function: mir-17-92 as a comprehensive promoter of the CD4 T-cell antigen response
A typical T-cell response against foreign or altered self-antigens involves robust clonal expansion followed by a contraction phase to maintain immune homeostasis [46, 47]. Activation also drives CD4 T cells to differentiate into particular effector lineages [48]: TH1 cells are responsible for clearance of intracellular infection and are implicated as the effectors in various autoimmune malignancies, TH2 cells control extracellular microbe infection as well as mediate chronic inflammation and allergic responses, and a recently identified third subset, TH17 cells, has been linked to a growing list of autoimmune disorders. Naïve CD4 T cells can also differentiate into a subset of cells having inhibitory functions: regulatory T cells (TRegs). Proliferation, differentiation, and programmed cell death constitute the basic aspects of a typical T-cell response and proper regulation ensures the clearance of specific pathogens or tumors. Identifying miRNAs that are actively regulating these effector functions will facilitate the development of immune therapeutic methods.
Work by other researchers has already identified several miRNAs important for T-cell effector function and differentiation. For example, miR-146a is upregulated following TCR stimulation and functions to suppress IL-2 production and protect T cells from activation-induced cell death (AICD) [49]. It is also required for TReg-mediated suppression of TH1 responses by targeting Stat1 [50] and was identified, along with miR-150 and miR-155, as a miRNA upregulated during the differentiation of TH17 cells [51]. miR-150 was identified as miRNA expressed in both B and T cells and blocks development in the former when expressed prematurely [52]. More recently, it was demonstrated to inhibit the development of central memory CD8 T cells [53]. miR-155 was demonstrated to be important for balancing TH1/TH2 cell differentiation [54] and for the germinal center reaction, specifically, T-cell-dependent antibody responses [55]. Recently, it was found that the level of miR-155 expression is under the control of Foxp3 protein [56, 57], and, reciprocally, miR-155 is required for maintaining TReg identity [58, 59] by targeting suppressor of cytokine signaling 1 (SOCS1) protein [58].
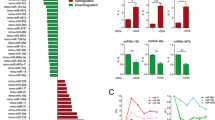
Following our investigations into how miR-181a controls the sensitivity of TCR signaling, we wondered if antigen challenge might regulate the levels of miRNAs globally and reasoned that upregulated miRNAs would likely help modulate the effector response. We performed miRNA expression profiling and observed, strikingly, that around 70% of miRNAs rapidly decline in CD4 T cells following antigen stimulation. A few, however, showed significant upregulation, and the miR-17-92 cluster, with its six miRNAs, represented a significant proportion of upregulated miRNAs and formed a distinct group, although each miRNA within the cluster exhibited individual expression dynamics (unpublished data).
The miR-17-92 cluster (miR-17, miR-18a, miR-19a, miR-20a, miR-19b, and miR-92a) [60, 61] was originally described as an “oncomiR” because it was shown to collaborate with c-Myc to induce B-cell lymphomas [62]. Further work in B cells highlighted a role for the cluster in driving B-cell development from the pro-B to pre-B stage via downregulation of the proapoptotic protein Bim [63]. Transgenic mice overexpressing miR-17-92 in lymphocytes develop autoimmunity, an unsurprising phenotype given that the lymphocytes are hyperproliferative and resistant to activation-induced cell death (AICD) [64]. The data begged for an exploration of the function of the whole cluster as well as the individual miRNA functions in the CD4 T-cell antigen response.
Utilizing an in vitro retroviral overexpression system and the miR-17-92 T-cell-specific knockout mice, we found that miR-17-92 is a comprehensive and indispensable positive regulator of CD4 T-cell response to tumor challenge (unpublished data). We noted that mice with a conditional deletion of miR-17-92 in T cells were quite vulnerable to B16 melanoma challenge. Detailed analysis of miR-17-92-deficient CD4 T cells revealed a significant proliferative, survival, and TH1 differentiation disadvantage, explaining the tumor susceptibility phenotype. We further noted that miR-17-92 is critical in inhibiting the induction of iTRegs, which are often induced within the tumor microenvironment and then function in tumor maintenance [65]. Intriguingly, the individual miRNAs within the miR-17-92 cluster have distinct functions. Specifically, miR-19b is the key element promoting CD4 T-cell proliferation, survival, and IFNγ production through targeting PTEN, while miR-17 blocks iTReg differentiation and inhibits AICD by downregulating TGFβRII and CREB1 expression. Quite surprisingly, miR-18a and miR-92a seem to play antagonizing functions against the whole cluster, as they promote apoptosis of effector CD4 T cells upon antigen rechallenge. Elucidation of the detailed mechanism and direct targets of these two miRNAs is currently in progress.
miR-17-92 is the first identified miRNA cluster that promotes cancer progression, and it does so at multiple stages and in multiple cell types [66]. Our study indicates that individual miRNAs within this cluster possess individual or even antagonizing functions, even though several of them belong to the same family, and our current knowledge fails to predict divergence of their targeting. To treat these six miRNAs as an indivisible unit became obviously misleading for our comprehension and dangerous for therapeutic design. On the cancer biology front, this complexity calls for detailed functional and mechanistic dissection to optimize antitumor therapy. On the immunology front, the functions of these miRNAs are carried out through multiple protein targets, and the suppression of each individual target is at a relatively moderate level. This provides a unique means to effectively modulate the overall quality of T-cell responses, whose decision making usually involves multiple pathways and multiple genes in each pathway. Since it is unlikely that the physiological function of other types of cells depends upon the same combination of targets, treatments targeting miRNAs may have minimal side effects.
Although the role of the miR-17-92 cluster in CD4 T cells has been extensively characterized by us and others, little is known about its function in CD8 T cells. Given the potent antitumor phenotype observed in CD4 T cells, members of the miR-17-92 cluster may strongly augment or suppress CD8 T-cell effector responses in antitumor immunity. To answer this question and better understand the therapeutic potential of miR-17-92-mediated immune therapies, we are currently working to dissect the role of the miR-17-92 cluster in CD8 T cells.
Besides regulations following TCR engagement, relevant signaling events also regulate miRNA expression in a variety of other cell types. Using miR-17-92 as an example, estrogen receptor α signaling [67], stress-induced senescence [68], and hypoxia [69] have all been shown to regulate miR-17-92 levels. Taken together, such data indicate that the regulation of miRNAs plays an important role in the phenotype switching following the sensing of extrinsic signals by the cell. Indeed, it may even be that messages interpreted by one cell type result in phenotypic changes in other cell types via the organized secretion of miRNAs by the sensing cell. A recent study has demonstrated that monocytes secrete a variety of miRNAs (e.g. miR-150) in targeted microvesicles (MVs) and that the fusion of such MVs with target cells results in protein level decreases within the target cell [70]. The implication here is rather dramatic: not only do cells store miRNAs intracellularly for directed secretion, but the regulatory potential of a given miRNA stretches beyond the cell in which it is produced. Such targeted regulation outside the cell of origin could affect rapid phenotypic changes in cells otherwise oblivious to a particular signal, perhaps even priming them to better receive or more strongly react to upcoming signaling events.
Closing remarks: a new frontier in immunoregulation
Our work on miR-181a demonstrated the first molecular mechanism of miRNA immunoregulation [8, 34]. Mounting evidence indicates that miRNAs regulate at least 20–30% of protein coding genes in vertebrates at a significant level [71–73], representing a fundamental layer of the post-transcriptional program modulating immune cell development, immune responses to infection, and the incidence of autoimmune disease and allergy [9]. Hence, miRNAs can be used as effective tools to manipulate a specific immune response and for discovery. Identification of miRNAs’ functionally relevant targets may uncover novel proteins (or novel functions of known proteins) as immediate targets for immune therapy. Furthermore, unlike the lengthy and costly process of protein-based drug development, designing and delivering an oligonucleotide-based miRNA mimic or inhibitor is simple and effective [74–76]. miRNAs are opening many new areas for exploration and add a further layer of complexity to the regulation of cellular responses. Given the power of miRNA manipulation, proper dissection of miRNA function in the context of immune regulation should provide much simpler and more controllable clinical tools for the treatment of both cancer and autoimmunity.
References
Davis MM, et al. T cells as a self-referential, sensory organ. Annu Rev Immunol. 2007;25:681–95.
Kane LP, Lin J, Weiss A. Signal transduction by the TCR for antigen. Curr Opin Immunol. 2000;12:242–9.
Vang T, et al. Protein tyrosine phosphatases in autoimmunity. Annu Rev Immunol. 2008;26:29–55.
Samelson, L.E.: Signal transduction mediated by the T cell antigen receptor: the role of adapter proteins. Annu Rev Immunol 2002; 20: 371-394.
Shaw AS, Filbert EL. Scaffold proteins and immune-cell signalling. Nat Rev Immunol. 2009;9:47–56.
Fabian MR, Sonenberg N, Filipowicz W. Regulation of mRNA translation and stability by microRNAs. Annu Rev Biochem. 2010;79:351–79.
Chen CZ, Li L, Lodish HF, Bartel DP. microRNAs modulate hematopoietic lineage differentiation. Science. 2004;303:83–6.
Li QJ, et al. miR-181a Is an Intrinsic Modulator of T Cell Sensitivity and Selection. Cell. 2007;129:147–61.
Baltimore D, Boldin MP, O’Connell RM, Rao DS, Taganov KD. microRNAs: new regulators of immune cell development and function. Nat Immunol. 2008;9:839–45.
Matsui K, Boniface JJ, Steffner P, Reay PA, Davis MM. Kinetics of T-cell receptor binding to peptide/I-Ek complexes: correlation of the dissociation rate with T-cell responsiveness. Proc Natl Acad Sci USA. 1994;91:12862–6.
Davis MM, et al. Ligand recognition by alpha beta T cell receptors. Annu Rev Immunol. 1998;16:523–44.
Holler PD, Kranz DM. Quantitative analysis of the contribution of TCR/pepMHC affinity and CD8 to T cell activation. Immunity. 2003;18:255–64.
Holler PD, Kranz DM. T cell receptors: affinities, cross-reactivities, and a conformer model. Mol Immunol. 2004;40:1027–31.
Evavold BD, Allen PM. Separation of IL-4 production from Th cell proliferation by an altered T cell receptor ligand. Science. 1991;252:1308–10.
Evavold BD, Sloan-Lancaster J, Allen PM. Antagonism of superantigen-stimulated helper T-cell clones and hybridomas by altered peptide ligand. Proc Natl Acad Sci USA. 1994;91:2300–4.
Krogsgaard M, et al. Agonist/endogenous peptide-MHC heterodimers drive T cell activation and sensitivity. Nature. 2005;434:238–43.
Pircher H, Rohrer UH, Moskophidis D, Zinkernagel RM, Hengartner H. Lower receptor avidity required for thymic clonal deletion than for effector T-cell function. Nature. 1991;351:482–5.
Davey GM, et al. Preselection thymocytes are more sensitive to T cell receptor stimulation than mature T cells. J Exp Med. 1998;188:1867–74.
Lucas B, Stefanova I, Yasutomo K, Dautigny N, Germain RN. Divergent changes in the sensitivity of maturing T cells to structurally related ligands underlies formation of a useful T cell repertoire. Immunity. 1999;10:367–76.
Curtsinger JM, Lins DC, Mescher MF. CD8+ memory T cells (CD44high, Ly-6C+) are more sensitive than naive cells to (CD44low, Ly-6C-) to TCR/CD8 signaling in response to antigen. J Immunol. 1998;160:3236–43.
Hogquist KA, et al. T cell receptor antagonist peptides induce positive selection. Cell. 1994;76:17–27.
Kao H, Allen PM. An antagonist peptide mediates positive selection and CD4 lineage commitment of MHC class II-restricted T cells in the absence of CD4. J Exp Med. 2005;201:149–58.
Irvine DJ, Purbhoo MA, Krogsgaard M, Davis MM. Direct observation of ligand recognition by T cells. Nature. 2002;419:845–9.
Li QJ, et al. CD4 enhances T cell sensitivity to antigen by coordinating Lck accumulation at the immunological synapse. Nat Immunol. 2004;5:791–9.
Purbhoo MA, Irvine DJ, Huppa JB, Davis MM. T cell killing does not require the formation of a stable mature immunological synapse. Nat Immunol. 2004;5:524–30.
Ebert PJ, Ehrlich LI, Davis MM. Low ligand requirement for deletion and lack of synapses in positive selection enforce the gauntlet of thymic T cell maturation. Immunity. 2008;29:734–45.
Tarakhovsky A, et al. A role for CD5 in TCR-mediated signal transduction and thymocyte selection. Science. 1995;269:535–7.
Starr TK, Daniels MA, Lucido MM, Jameson SC, Hogquist KA. Thymocyte sensitivity and supramolecular activation cluster formation are developmentally regulated: a partial role for sialylation. J Immunol. 2003;171:4512–20.
Zikherman J, et al. CD45-Csk phosphatase-kinase titration uncouples basal and inducible T cell receptor signaling during thymic development. Immunity. 2010;32:342–54.
Wu J, et al. Identification of substrates of human protein-tyrosine phosphatase PTPN22. J Biol Chem. 2006;281:11002–10.
Zikherman J, et al. PTPN22 deficiency cooperates with the CD45 E613R allele to break tolerance on a non-autoimmune background. J Immunol. 2009;182:4093–106.
Koelsch U, Schraven B, Simeoni L. SIT and TRIM determine T cell fate in the thymus. J Immunol. 2008;181:5930–9.
Theodosiou, A. & Ashworth, A.: MAP kinase phosphatases. Genome Biol 2002; 3: REVIEWS3009.
Ebert, P.J., Jiang, S., Xie, J., Li, Q.J. & Davis, M.M.: An endogenous positively selecting peptide enhances mature T cell responses and becomes an autoantigen in the absence of microRNA miR-181a. Nat Immunol 2009.
Liu G, Min H, Yue S, Chen CZ. Pre-miRNA loop nucleotides control the distinct activities of mir-181a–1, mir-181c in early T cell development. PLoS One. 2008;3:e3592.
Yamasaki S, et al. Mechanistic basis of pre-T cell receptor-mediated autonomous signaling critical for thymocyte development. Nat Immunol. 2006;7:67–75.
Stefanski HE, Mayerova D, Jameson SC, Hogquist KA. A low affinity TCR ligand restores positive selection of CD8+ T cells in vivo. J Immunol. 2001;166:6602–7.
Hogquist KA, Baldwin TA, Jameson SC. Central tolerance: learning self-control in the thymus. Nat Rev Immunol. 2005;5:772–82.
Reddy J, et al. Detection of autoreactive myelin proteolipid protein 139–151-specific T cells by using MHC II (IAs) tetramers. J Immunol. 2003;170:870–7.
Danke NA, Koelle DM, Yee C, Beheray S, Kwok WW. Autoreactive T cells in healthy individuals. J Immunol. 2004;172:5967–72.
Goodnow, C.C.: Multistep pathogenesis of autoimmune disease. Cell 2007; 130: 25-35.
Bielekova B, et al. Expansion and functional relevance of high-avidity myelin-specific CD4+ T cells in multiple sclerosis. J Immunol. 2004;172:3893–904.
Amrani A, et al. Progression of autoimmune diabetes driven by avidity maturation of a T-cell population. Nature. 2000;406:739–42.
Savage PA, Boniface JJ, Davis MM. A kinetic basis for T cell receptor repertoire selection during an immune response. Immunity. 1999;10:485–92.
Tian J, Gregori S, Adorini L, Kaufman DL. The frequency of high avidity T cells determines the hierarchy of determinant spreading. J Immunol. 2001;166:7144–50.
Williams MA, Bevan MJ. Effector and memory CTL differentiation. Annu Rev Immunol. 2007;25:171–92.
van Leeuwen EM, Sprent J, Surh CD. Generation and maintenance of memory CD4(+) T Cells. Curr Opin Immunol. 2009;21:167–72.
Zhu J, Yamane H, Paul WE. Differentiation of effector CD4 T cell populations (*). Annu Rev Immunol. 2010;28:445–89.
Curtale G, et al. An emerging player in the adaptive immune response: microRNA-146a is a modulator of IL-2 expression and activation-induced cell death in T lymphocytes. Blood. 2010;115:265–73.
Lu LF, et al. Function of miR-146a in controlling Treg cell-mediated regulation of Th1 responses. Cell. 2010;142:914–29.
Niimoto T, et al. microRNA-146a expresses in interleukin-17 producing T cells in rheumatoid arthritis patients. BMC Musculoskelet Disord. 2010;11:209.
Zhou B, Wang S, Mayr C, Bartel DP, Lodish HF. miR-150, a microRNA expressed in mature B and T cells, blocks early B cell development when expressed prematurely. Proc Natl Acad Sci USA. 2007;104:7080–5.
Almanza G, et al. Selected microRNAs define cell fate determination of murine central memory CD8 T cells. PLoS One. 2010;5:e11243.
Rodriguez A, et al. Requirement of bic/microRNA-155 for normal immune function. Science. 2007;316:608–11.
Thai TH, et al. Regulation of the germinal center response by microRNA-155. Science. 2007;316:604–8.
Marson A, et al. Foxp3 occupancy and regulation of key target genes during T-cell stimulation. Nature. 2007;445:931–5.
Zheng Y, et al. Genome-wide analysis of Foxp3 target genes in developing and mature regulatory T cells. Nature. 2007;445:936–40.
Lu LF, et al. Foxp3-dependent microRNA155 confers competitive fitness to regulatory T cells by targeting SOCS1 protein. Immunity. 2009;30:80–91.
Kohlhaas S, et al. Cutting edge: the Foxp3 target miR-155 contributes to the development of regulatory T cells. J Immunol. 2009;182:2578–82.
Tanzer A, Stadler PF. Molecular evolution of a microRNA cluster. J Mol Biol. 2004;339:327–35.
Ota A, et al. EIdentification and characterization of a novel gene, C13orf25, as a target for 13q31–q32 amplification in malignant lymphoma. Cancer Res. 2004;64:3087–95.
He L, et al. A microRNA polycistron as a potential human oncogene. Nature. 2005;435:828–33.
Ventura A, et al. Targeted deletion reveals essential and overlapping functions of the miR-17 through 92 family of miRNA clusters. Cell. 2008;132:875–86.
Xiao C, et al. Lymphoproliferative disease and autoimmunity in mice with increased miR-17–92 expression in lymphocytes. Nat Immunol. 2008;9:405–14.
Nishikawa H, Sakaguchi S. Regulatory T cells in tumor immunity. Int J Cancer. 2010;127:759–67.
Olive V, Jiang I, He L. Mir-17–92, a cluster of miRNAs in the midst of the cancer network. Int J Biochem Cell Biol. 2010;42:1348–54.
Castellano L, et al. The estrogen receptor-alpha-induced microRNA signature regulates itself and its transcriptional response. Proc Natl Acad Sci USA. 2009;106:15732–7.
Li G, Luna C, Qiu J, Epstein DL, Gonzalez P. Alterations in microRNA expression in stress-induced cellular senescence. Mech Ageing Dev. 2009;130:731–41.
Yan HL, et al. Repression of the miR-17-92 cluster by p53 has an important function in hypoxia-induced apoptosis. EMBO J. 2009;28:2719–32.
Zhang Y, et al. Secreted monocytic miR-150 enhances targeted endothelial cell migration. Mol Cell. 2010;39:133–44.
Baek, D. et al.: The impact of microRNAs on protein output. Nature 2008.
Selbach, M. et al.: Widespread changes in protein synthesis induced by microRNAs. Nature 2008.
Chi SW, Zang JB, Mele A, Darnell RB. Argonaute HITS-CLIP decodes microRNA-mRNA interaction maps. Nature. 2009;460:479–86.
Krutzfeldt J, et al. Silencing of microRNAs in vivo with ‘antagomirs’. Nature. 2005;438:685–9.
Thum, T. et al.: MicroRNA-21 contributes to myocardial disease by stimulating MAP kinase signalling in fibroblasts. Nature 2008.
Kota J, et al. Therapeutic microRNA delivery suppresses tumorigenesis in a murine liver cancer model. Cell. 2009;137:1005–17.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lykken, E.A., Li, QJ. microRNAs at the regulatory frontier: an investigation into how microRNAs impact the development and effector functions of CD4 T cells. Immunol Res 49, 87–96 (2011). https://doi.org/10.1007/s12026-010-8196-4
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12026-010-8196-4