Abstract
NTRK1/3 rearrangements have been reported in 2.3–3.4% of papillary thyroid carcinoma (PTC) and are regarded as potential therapeutic targets. Recently, the application of immunohistochemistry (IHC) to detect NTRK rearrangements has been widely discussed. The current study aimed to characterize the clinicopathological features of PTC with NTRK1/3 fusions, to examine the utility of pan-TRK IHC, and to compare IHC with fluorescent in situ hybridization (FISH) and next-generation sequencing (NGS). In a cohort of 525 consecutive PTC cases, 60 BRAFV600E-negative cases underwent complete analyses of FISH, and 12 (2.3%) cases with NTRK1/3 break-apart were found. A novel ERC1-NTRK3 fusion was identified by NGS in one case. Pathological features of non-infiltrative tumor border, clear cell change, and reduced nuclear elongation and irregularity were significantly more common in NTRK1/3-rearranged PTC when compared with 48 BRAFV600E-negative non-NTRK1/3 PTC cases. In whole tissue sections, pan-TRK IHC was positive in 3/7 (42.9%) cases with an ETV6-NTRK3 rearrangement including 2 cases with low percentage of stained tumor cells, 2/3 (66.7%) with non-ETV6 NTRK3 rearrangements, and 2/2 (100%) with NTRK1 rearrangements. All FISH-negative cases were negative for pan-TRK in tissue microarray sections. As a result, pan-TRK IHC showed a sensitivity of 58.3% and specificity of 100% for NTRK1/3 rearrangements in BRAFV600E-negative PTC. In conclusion, NTRK1/3-rearranged PTC shared some unique morphologic features. Pan-TRK IHC showed high specificity and moderate sensitivity for NTRK1/3-rearranged PTC and should be interpreted with caution due to staining heterogeneity. Based on the above findings, we propose an algorithm integrating morphology, IHC, and molecular testing to detect NTRK1/3 rearrangements in PTC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Papillary thyroid carcinoma (PTC) is the most common malignancy of the thyroid gland and accounts for more than 80% of thyroid carcinoma [1]. Although the prognosis of PTC is generally excellent, a small portion of cases may still develop an aggressive clinical course and become refractory to conventional radioiodine therapy. In such cases, novel targeted therapeutic agents may be potentially beneficial. The tropomyosin receptor kinase (Trk) family, which is composed of TrkA, TrkB, and TrkC, is recently regarded as a potential target of cancer therapy. Chromosomal rearrangements involving the associated NTRK1/2/3 genes have been found in malignancies of various organs, including thyroid, brain, lung, colorectum, and soft tissue [2]. As for PTC, NTRK1/3 rearrangements were detected in 2.3–3.4% tumors in The Cancer Genome Atlas (TCGA) project and the Chinese population study [3, 4]. Various tyrosine kinase inhibitors against Trk family are currently under clinical trials, and two of these agents, larotrectinib and entrectinib, have demonstrated clinically significant antitumor activity [5, 6]. The U.S. Food and Drug Administration approval of these two agents for solid tumors harboring NTRK gene fusions in November 2018 and August 2019, respectively, further necessitates accurate and efficient screening strategy for these genetic alterations.
In routine pathology practice, the detection of NTRK gene fusions predominantly relies on fluorescence in situ hybridization (FISH), reverse transcription polymerase chain reaction (RT-PCR), or next-generation sequencing (NGS). However, these methodologies may be compromised by suboptimal RNA quality of the formalin-fixed paraffin-embedded (FFPE) tissue, high labor demand, and high cost. Recently, the application of immunohistochemistry (IHC) using anti-pan-TRK antibody to detect NTRK gene fusions in various tumor types has been widely discussed and mainly focused on secretory carcinoma of the salivary gland and soft tissue tumors [7,8,9,10,11,12,13,14,15,16,17,18]. PTC was included in only two of studies with a limited number of cases [19, 20]. On the other hand, although the histological features of PTC with NTRK gene fusions, especially with an ETV6-NTRK3 fusion, have been reported in previous studies [21,22,23,24,25,26,27], comprehensive morphologic comparison between NTRK-rearranged and non-NTRK PTC has not been systemically investigated. This study aimed to examine the frequency of NTRK1/3 rearrangements in a Taiwanese cohort of PTC, to further characterize the clinicopathological features of PTC with NTRK fusions, and to validate the sensitivity and specificity of pan-TRK immunohistochemistry as a screening tool.
Materials and Methods
Patient Selection
This study was approved by the Institutional Review Board (IRB) of Taipei Veterans General Hospital, which granted exemption of informed consent for tissue procurement through the Biobank of Taipei Veterans General Hospital after an unlinked anonymous process (IRB no.: 2019-07-001BC, Biobank no.: 10818). A retrospective search of consecutive PTC cases with tumor size larger than 0.5 cm between October 2015 and March 2019 from the pathology archives of Taipei Veterans General Hospital was performed. All cases had available information regarding the BRAFV600E status using a validated IHC assay (clone VE1, Spring Bioscience, Pleasanton, CA, USA) [28]. Cases of BRAFV600E-negative PTC were subjected to this study. All pathology slides were retrieved for microscopic review. Cases with pathological features diagnostic of cribriform-morular variant of PTC (CMV-PTC) were excluded due to its universal alterations in the WNT pathway-related genes [29].
Clinicopathological Evaluation and Tissue Microarray Preparation
Clinical characters including age, sex, tumor size, and TNM stage were retrieved from the medical charts. Histological features including the presence of extrathyroidal extension, lymphatic invasion, tumor border, and proportions of growth patterns (follicular, papillary, and solid/trabecular/insular) were recorded. Tumor border was classified into three types: well-circumscribed, multinodular permeative (invasive growth in a multinodular fashion), and infiltrative (irregular and spiculated tumor invasion sometimes with desmoplastic change). Cytological features including clear cell change, reverse nuclear polarity (nuclei that are not basally oriented), and nuclear shape were also evaluated. The presence of reduced nuclear elongation and irregularity was considered significant if such nuclear features were observed in a majority of tumor cells in contrast to typical nuclear changes of classical PTC. Cases with sufficient FFPE tumor materials were subjected to tissue microarrays (TMA) construction using the MTA Booster OI manual tissue arrayer (Alphalys, Plaisir, France). One 2-mm tissue core obtained from the tumorous area in each case was applied.
Fluorescence In Situ Hybridization
To evaluate NTRK1/3 rearrangements, dual-color break-apart FISH probes for NTRK1 and NTRK3 (ZytoVision, Bremerhaven, Germany) were performed on TMA sections. Given that ETV6 gene is the most frequent fusion partner of NTRK3 gene in PTC [3, 4], dual-color break-apart FISH probes for ETV6 (ZytoVision) were also assessed in this study. Case showing more than 20% of nuclei positive for break-apart signals, namely two separate green and orange signals with distance more than one signal diameter, was defined as positive.
RNA Extraction, Targeted Next-Generation Sequencing, and Reverse-Transcriptase PCR
Cases with NTRK1 rearrangements or isolated non-ETV6 NTRK3 rearrangements were subjected to targeted next-generation sequencing to identify the fusion partner genes. Total RNA was extracted from the representative FFPE tumor tissue using the RNeasy FFPE Kit (Qiagen, Germantown, MD, USA). The library was prepared using ArcherDX FusionPlex Comprehensive Thyroid and Lung Panel (ArcherDX, Boulder, CO, USA). The sample was sequenced on Illumina NextSeq 500 sequencer (Illumina, San Diego, CA, USA). Potential fusions were analyzed using Archer Analysis bioinformatics platform. For the novel ERC1-NTRK3 fusion detected in one of our cases, the rearrangement was further validated with reverse-transcriptase PCR and Sanger sequencing using primers (forward: 5′-GCACAAGGAACAGGTGGAA-3′, reverse: 5′-CGAAGAGAACCACCAACAGG-3′) flanking the fusion site.
Immunohistochemistry
Immunohistochemical stain for anti-pan-TRK antibody (clone EPR17341, dilution 1:50, Abcam, Cambridge, MA, USA) was performed on 4-μm-thick sections of the TMA blocks. An automated protocol including sample de-paraffinization, antigen retrieval (in ER2 solution [EDTA-based, pH 9.0] at 100 °C for 20 min), primary antibody incubation, and signal visualization was conducted on a Leica Bond-Max autostainer (Leica Biosystems, Buffalo Grove, IL, USA). Tissue from secretory carcinoma of the salivary gland with a confirmed ETV6-NTRK3 fusion and positive pan-TRK staining was applied as positive control. The IHC protocol had been optimized to balance the intensity of positive staining of control tissue and non-specific background staining. Positive staining was defined as any unequivocal immunoreactivity on cytoplasm and/or nuclei with clear contrast with surrounding non-tumorous tissue. For cases showing positive pan-TRK IHC and/or positive FISH on TMA materials, pan-TRK IHC was repeated on the whole tissue tumor sections to evaluate staining heterogeneity. Percentage of positively stained cells, staining intensity, and staining pattern in both TMA and whole tissue sections were recorded. Additional IHC for TTF-1 (clone 8G7G3/1, Dako, Carpinteria, CA, USA) was performed to exclude the possibility of secretary carcinoma of salivary gland type.
Statistical Analysis
A comparison of clinicopathological characteristics between NTRK1/3-rearranged PTC and BRAFV600E-negative non-NTRK1/3 PTC excluding CMV-PTC was performed. Continuous variables and categorical variables were analyzed using two-tailed t test and Fisher’s exact test, respectively. Statistical analyses were performed using the R software version 3.5.1 (R Foundation for Statistical Computing, Vienna, Austria).
Results
Patient Characteristics and Clinical Features
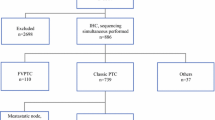
A total of 525 consecutive PTC cases between October 2015 and March 2019 were identified. There were 397 (75.6%) female patients and 128 (24.4%) male patients with the mean age of 49.7 years (range: 11–86 years) at diagnosis. When grouped by BRAFV600E mutation status, the cases were classified into 456 (86.9%) BRAFV600E-positive PTC and 69 (13.1%) BRAFV600E-negative PTC. Four cases of CMV-PTC and 5 cases with insufficient or suboptimal tissues for additional testing were excluded. A total of 60 BRAFV600E-negative PTC underwent complete evaluation of IHC and FISH in TMA sections. None of the 60 patients had a prior history of radiation. Twelve (2.3%) cases with NTRK1 or NTRK3 rearrangements were detected by FISH, and 6 of them were further confirmed by targeted NGS. The 12 NTRK1/3-rearranged PTC cases were composed of 10 (83.3%) female patients and 2 (16.7%) male patients (Table 1). The mean age at diagnosis was 43.6 years (range: 20–68 years). The average tumor size was 2.3 cm (range: 0.9–4.5 cm), and none of the 12 cases showed extrathyroidal extension according to the 8th edition of the American Joint Committee on Cancer (AJCC) staging system. Metastasis to regional lymph nodes was noted in five (41.7%) patients and lung metastasis was present in another one (8.3%) patient at diagnosis. The AJCC stage I disease was seen in nine (75%) cases, stage II in two (16.7%) cases, and stage IVB in one (8.3%) case. When compared with non-NTRK1/3 PTC, the NTRK1/3-rearranged PTC cases more commonly showed stage II-IVB disease (25% versus 4%, p = 0.0499), but there was no significant difference in gender, age, tumor size, extrathyroidal extension, and lymph node metastasis. Eight (66.7%) cases received total thyroidectomy followed by radioactive iodine treatment, while the other four (33.3%) cases underwent lobectomy only. None of the 12 cases received TRK inhibitor therapy. The case with lung metastasis remained stable disease status. No recurrence was observed in all the other cases during a median follow-up period of 26 months (12 to 52 months).
Pathological Findings
The 12 NTRK1/3-rearranged PTC were classified into 8 (66.7%) conventional PTC, 2 (16.7%) follicular variant of PTC, and 2 (16.7%) solid variant of PTC (Table 2). As for the border between the tumor and surrounding parenchyma, 4 (33.3%) cases showed infiltrative growth sometimes with scar-like stromal reaction, while the other 8 (66.7%) cases had a well-circumscribed or multinodular permeative margin (Fig. 1a). The two (16.7%) cases showing a well-circumscribed tumor border were precluded from the diagnosis of noninvasive follicular thyroid neoplasm with papillary-like nuclear features (NIFTP) based on the presence of distinct papillary areas. Seven (58.3%) cases showed a predominance of follicular growth pattern (Fig. 1b), 3 (25%) showed a predominance of papillary growth pattern, while 2 (16.7%) showed a predominance of solid/trabecular/insular growth patterns (Fig. 1c). Reverse nuclear polarity (Fig. 1d) was observed exclusively in the three papillary pattern-predominant cases. Clear cell change was discerned in nine (75%) cases either as small foci commonly located at the periphery of the tumors (Fig. 1e) or in a more diffuse form (up to 70% of tumor cells in case 12). Although all the cases showed a nuclear score of ≥ 2 [30], reduced nuclear elongation and irregularity of tumor cells were observed in 6 (50%) cases (Fig. 1f). Of note, all three papillary pattern-predominant cases harbored an ETV6-NTRK3 fusion.
Pathological features of NTRK1/3-rearranged papillary thyroid carcinoma (PTC) (H&E stain). a, b Follicular variant of PTC with multinodular permeative tumor border (case 8, 20× and 200×). c Solid variant of PTC (case 10, 200×). d Papillary pattern-predominant PTC with reverse nuclear polarity (case 6, 200×). e Clear cell change of tumor cells (case 11, 400×). f Reduced nuclear elongation and irregularity of tumor cells in an area of papillary growth pattern (case 7, 400×)
In comparison between 12 NTRK1/3-rearranged PTC and 48 BRAFV600E-negative non-NTRK1/3 PTC cases, the NTRK1/3-rearranged PTC showed significantly more common non-infiltrative (well-circumscribed/multinodular permeative) tumor border (66.7% versus, 33.3% p = 0.0498), clear cell change (75% versus 37.5%, p = 0.0258), and reduced nuclear elongation and irregularity (50% versus 16.7%, p = 0.0238). At least one of these three morphologic features was seen in all of 12 NTRK1/3-rearranged PTC but only in 26 (54.2%) of 48 non-NTRK1/3 PTC. There was also a tendency for NTRK1/3-rearranged PTC to harbor solid/trabecular/insular growth patterns in more than 70% of tumor areas (16.7% versus 2.1%, p = 0.0990).
Molecular Testing
FISH studies revealed 6 PTC with an ETV6-NTRK3 rearrangement (Fig. 2a, b), 4 with isolated non-ETV6 NTRK3 rearrangements, and 2 with NTRK1 rearrangements (Fig. 2c). The 6 cases with NTRK1 rearrangements or isolated non-ETV6 NTRK3 rearrangements were sent for targeted NGS. An additional case with an ETV6exon4-NTRK3exon14 rearrangement was detected. For the remaining cases, TPM3exon7-NTRK1exon10, TPRexon21-NTRK1exon10, ERC1exon12-NTRK3exon13, SQSTM1exon4-NTRK3exon14, and VIMexon7-NTRK3exon14 fusions were identified in one case each. The kinase domain of NTRK1/3 genes was preserved in all cases. Among these rearrangements, ERC1-NTRK3 (Fig. 3a) is a novel fusion that has not been reported in the literature. This novel fusion was further confirmed using reverse-transcriptase PCR and Sanger sequencing (Fig. 3b).
Ancillary testing of NTRK1/3-rearranged papillary thyroid carcinoma (PTC). a, b Positive break-apart fluorescence in situ hybridization (FISH) signals for ETV6 and NTRK3 rearrangements, respectively (case 6). c Positive break-apart FISH signals for NTRK1 rearrangement (case 11). d Pan-TRK immunohistochemistry (IHC) with weak cytoplasmic and nuclear staining in PTC with an ETV6-NTRK3 fusion (case 6, 200×). e Pan-TRK IHC with strong cytoplasmic and membranous staining in PTC with a TPM3-NTRK1 fusion (case 11, 200×). f Pan-TRK IHC with strong cytoplasmic staining in PTC with a VIM-NTRK3 fusion (case 10, 200×)
Immunohistochemistry
In TMA sections, Pan-TRK IHC was positive in one case with an ETV6-NTRK3 rearrangement (1/7, 14.3%), 2 with other NTRK3 rearrangements (2/3, 66.7%), 2 with NTRK1 rearrangements (2/2, 100%), and was negative in all FISH-negative cases. The case with an ETV6-NTRK3 rearrangement showed both cytoplasmic and nuclear staining (Fig. 2d) while another case with a TPM3-NTRK1 rearrangement showed both cytoplasmic and membranous staining patterns (Fig. 2e). The other three positive cases were stained in the cytoplasm only (Fig. 2f). Overall, pan-TRK IHC showed a sensitivity of 41.7% (5/12), a specificity of 100%, a positive predictive value of 100%, and a negative predictive value of 87.3% for detecting NTRK1/3 rearrangements in BRAFV600E-negative PTC in TMA materials. Further pan-TRK staining for whole tissue sections in all FISH-positive cases revealed the presence of staining heterogeneity in a subset of IHC-positive cases in TMA (Table 2). Moreover, weak cytoplasmic and nuclear staining on a small population of tumor cells was detected in two additional ETV6-NTRK3 cases (5% and 10%, respectively). Therefore, the sensitivity of pan-TRK IHC was raised to 50% (5/10) for NTRK3-rearranged PTC and 58.3% (7/12) for all NTRK1/3-rearranged PTC in whole tissue sections. All the 12 cases with NTRK1/3 rearrangements showed positive TTF-1 staining, which excluded the diagnosis of secretory carcinoma of salivary gland type.
Discussion
NTRK1/3 gene fusions have been reported as the third most common gene rearrangements in TCGA PTC project and the second most common gene rearrangements in the Chinese PTC study [3, 4]. The frequency of these fusions in our cohort was 2.3%, similar to that in previous reports [3, 4]. Although higher frequency (18.3–25.9%) of NTRK1/3 rearrangements had been observed in the pediatric group [23, 27], there was no significant difference in age at diagnosis between NTRK1/3-rearranged and BRAFV600E-negative non-NTRK1/3 PTC in our study cohort. This could be partly attributed to that the patient population at our hospital is mainly adult. The prognostic significance of NTRK1/3 rearrangements remains unclear due to a limited number of cases. NTRK1/3 rearrangements were associated with higher AJCC stages in BRAFV600E-negative PTC in our study. The frequency of metastasis to lymph nodes and distant organs of NTRK1/3-rearranged PTC in our study was comparable with that in a previous series of PTC with an ETV6-NTRK3 fusion [24]. More frequent metastasis to regional lymph nodes and distant organs was observed in one recent NTRK1/3-rearranged PTC series [26], while a lower frequency (1.0%) of NTRK1/3 rearrangements was reported in a large-scale genetic analysis of advanced PTC [31]. The conflict of the survival data in the literature might be due to case selection in different studies.
Histological features of NTRK1/3-rearranged PTC have been reported in several prior studies [21,22,23,24,25,26,27]. In the current series, we identified significantly higher frequencies of clear cell change and non-infiltrative (well-circumscribed/multinodular permeative) tumor border in NTRK1/3-rearranged PTC, which aligned with the previous description in a series of ETV6-NTRK3-positive PTC [24]. We also identified reduced nuclear elongation and irregularity of tumor cells as a novel pathological feature for NTRK1/3-rearranged PTC in contrast to the oval-shaped nuclei in the classical PTC. Although the aforementioned features may be affected by tissue processing, staining, and interobserver variability, these are probably still helpful when applied in combinations. We did not find a significant difference in follicular or solid/trabecular/insular growth patterns between NTRK1/3-rearranged and non-NTRK1/3 PTC regardless of different cut-offs applied. Although these histological patterns were previously found in some of NTRK1/3-rearranged PTC [21,22,23,24,25,26,27], the difference in our study might be blunted by the presence of other translocation-associated PTC in the non-NTRK1/3 group (e.g., RET and ALK translocation) that might also harbor a significant proportion of follicular or solid/trabecular/insular patterns [32, 33].
Following the development of targeted therapeutic agents against NTRK gene fusions, pan-TRK immunohistochemistry has been regarded as a promising screen tool and examined in several types of tumors [7,8,9,10, 12, 13, 16, 18,19,20, 34, 35]. Variable performance of pan-TRK IHC was observed possibly due to different tumor types, staining protocols, and interpretation criteria ranging from any staining to moderate staining in more than 50% of tumor cells. Despite its high specificity (pooled analysis: 5241/5549, 94.4%), lower sensitivity and percentage of positively stained tumor cells were seen in tumors with NTRK3 rearrangements (Table 3). The lower sensitivity of pan-TRK IHC was also demonstrated in some types of tumors including thyroid carcinoma (sensitivity of 50–81.8%), particularly in PTC with an ETV6-NTRK3 fusion (sensitivity of 60%) [19, 20]. Predominance of such fusions in our study cohort may partly explain the lower sensitivity in comparison with prior studies [19, 20]. This finding is important because ETV6-NTRK3 is the most common type of NTRK1/3 gene rearrangements in PTC. Given that NTRK1/3 rearrangements are rare in PTC, pan-TRK IHC may show limited value as a screening tool when used singly.
In our study, all pan-TRK IHC-positive PTC with an ETV6-NTRK3 fusion showed a cytoplasmic and nuclear staining pattern and the only PTC with a TPM3-NTRK1 fusion showed a cytoplasmic and membranous staining pattern. Cases with other fusion types were only stained in the cytoplasm. Similar staining localization has been also reported in other tumor types [10, 12, 13, 20, 36, 37]. It is also worth noting that staining heterogeneity of pan-TRK IHC was observed in a subset of our cases. Two cases with an ETV6-NTRK3 fusion showed a low percentage of stained tumor cells in whole tissue sections (5% and 10%) and were interpreted as negative in TMA sections. On the other hand, all pan-TRK positive cases with ≥ 60% of staining cells in whole tissue slides showed diffuse staining in TMA slides. The former phenomenon has been mentioned in a series of secretory carcinoma of the salivary gland, in which more than half of cases had less than 20% of positively stained cells [8]. This indicated that one should be alerted to possible false negativity when evaluating pan-TRK IHC in a small biopsy specimen.
Based on our findings, we propose a testing algorithm to detect NTRK rearrangements in PTC (Fig. 4). PTC cases can be firstly triaged based on BRAFV600E status, followed by pan-TRK IHC in BRAFV600E-negative cases. Whole tissue slides are more favored than small biopsy specimens for IHC. BRAFV600E-positive PTC is unlikely to harbor NTRK gene fusions, whereas positive pan-TRK IHC is very specific for NTRK gene fusions in BRAFV600E-negative PTC. Histological review is useful to select cases negative for both BRAFV600E mutation and pan-TRK IHC for additional work-up. Further molecular testing may be beneficial for cases showing any suggestive morphologic features, including non-infiltrative tumor border, clear cell change, and reduced nuclear elongation and irregularity. Following this algorithm, molecular testing for NTRK1/3 rearrangements can be avoided in nearly half of BRAFV600E-negative non-NTRK PTC.
There are some limitations in our study. First, this is a retrospective study and none of our patients had used the TRK inhibitor to obtain the information regarding treatment response. Second, we did not test our cohort for NTRK2 gene rearrangements. Although NTRK2 gene rearrangements have not been reported in PTC so far, the presence of NTRK2 fusions cannot be excluded unless a thorough genetic analysis is performed for all cases. Third, we obviated BRAFV600E-positive PTC in our study population from further molecular testing. On extremely rare occasions, BRAFV600E mutation may coexist with NTRK gene rearrangements [21]. Finally, the rarity of pediatric patients in our study population prevented us from providing a more comprehensive clinicopathological investigation among both children and adults.
In conclusion, we detected NTRK1/3 gene rearrangements in 12 (2.3%) PTC cases in our study cohort and found a novel ERC1-NTRK3 gene fusion in one case. We identified three probably useful morphologic features, including non-infiltrative tumor border, clear cell change, and reduced nuclear elongation and irregularity, for the diagnosis of NTRK1/3-rearranged PTC. Pan-TRK IHC showed high specificity for both NTRK1 and NTRK3 fusions, but lower sensitivity for NTRK3-rearranged PTC. Moreover, limited staining was seen in two cases with an ETV6-NTRK3 fusion. Accordingly, pan-TRK IHC should be interpreted with caution for PTC due to staining heterogeneity. Based on the above findings, we propose a testing algorithm integrating morphology, immunohistochemistry, and molecular testing to detect NTRK1/3 rearrangements in PTC.
References
Lim H, Devesa SS, Sosa JA, Check D, Kitahara CM (2017) Trends in thyroid cancer incidence and mortality in the United States, 1974-2013. JAMA 317:1338-1348. https://doi.org/10.1001/jama.2017.2719
Amatu A, Sartore-Bianchi A, Siena S (2016) NTRK gene fusions as novel targets of cancer therapy across multiple tumour types. ESMO Open 1:e000023. https://doi.org/10.1136/esmoopen-2015-000023
Cancer Genome Atlas Research Network (2014) Integrated genomic characterization of papillary thyroid carcinoma. Cell 159:676-690. https://doi.org/10.1016/j.cell.2014.09.050
Liang J, Cai W, Feng D, Teng H, Mao F, Jiang Y, Hu S, Li X, Zhang Y, Liu B, Sun ZS (2018) Genetic landscape of papillary thyroid carcinoma in the Chinese population. J Pathol 244:215-226. https://doi.org/10.1002/path.5005
Drilon A, Laetsch TW, Kummar S, DuBois SG, Lassen UN, Demetri GD, Nathenson M, Doebele RC, Farago AF, Pappo AS, Turpin B, Dowlati A, Brose MS, Mascarenhas L, Federman N, Berlin J, El-Deiry WS, Baik C, Deeken J, Boni V, Nagasubramanian R, Taylor M, Rudzinski ER, Meric-Bernstam F, Sohal DPS, Ma PC, Raez LE, Hechtman JF, Benayed R, Ladanyi M, Tuch BB, Ebata K, Cruickshank S, Ku NC, Cox MC, Hawkins DS, Hong DS, Hyman DM (2018) Efficacy of larotrectinib in TRK fusion-positive cancers in adults and children. N Engl J Med 378:731-739. https://doi.org/10.1056/NEJMoa1714448
Doebele RC, Drilon A, Paz-Ares L, Siena S, Shaw AT, Farago AF, Blakely CM, Seto T, Cho BC, Tosi D, Besse B, Chawla SP, Bazhenova L, Krauss JC, Chae YK, Barve M, Garrido-Laguna I, Liu SV, Conkling P, John T, Fakih M, Sigal D, Loong HH, Buchschacher GL, Jr., Garrido P, Nieva J, Steuer C, Overbeck TR, Bowles DW, Fox E, Riehl T, Chow-Maneval E, Simmons B, Cui N, Johnson A, Eng S, Wilson TR, Demetri GD, trail investigators (2020) Entrectinib in patients with advanced or metastatic NTRK fusion-positive solid tumours: integrated analysis of three phase 1-2 trials. Lancet Oncol 21:271-282. https://doi.org/10.1016/S1470-2045(19)30691-6
Hung YP, Jo VY, Hornick JL (2019) Immunohistochemistry with a pan-TRK antibody distinguishes secretory carcinoma of the salivary gland from acinic cell carcinoma. Histopathology 75:54-62. https://doi.org/10.1111/his.13845
Bell D, Ferrarotto R, Liang L, Goepfert RP, Li J, Ning J, Broaddus R, Weber RS, El-Naggar AK (2020) Pan-Trk immunohistochemistry reliably identifies ETV6-NTRK3 fusion in secretory carcinoma of the salivary gland. Virchows Arch 476:295-305. https://doi.org/10.1007/s00428-019-02640-7
Guibourg B, Cloarec E, Conan-Charlet V, Quintin-Roue I, Grippari JL, Le Flahec G, Marcorelles P, Uguen A (2020) EPR17341 and A7H6R pan-TRK immunohistochemistry result in highly different staining patterns in a series of salivary gland tumors. Appl Immunohistochem Mol Morphol https://doi.org/10.1097/PAI.0000000000000825
Xu B, Haroon Al Rasheed MR, Antonescu CR, Alex D, Frosina D, Ghossein R, Jungbluth AA, Katabi N (2020) Pan-Trk immunohistochemistry is a sensitive and specific ancillary tool for diagnosing secretory carcinoma of the salivary gland and detecting ETV6-NTRK3 fusion. Histopathology 76:375-382. https://doi.org/10.1111/his.13981
Chiang S, Cotzia P, Hyman DM, Drilon A, Tap WD, Zhang L, Hechtman JF, Frosina D, Jungbluth AA, Murali R, Park KJ, Soslow RA, Oliva E, Iafrate AJ, Benayed R, Ladanyi M, Antonescu CR (2018) NTRK fusions define a novel uterine sarcoma subtype with features of fibrosarcoma. Am J Surg Pathol 42:791-798. https://doi.org/10.1097/PAS.0000000000001055
Hung YP, Fletcher CDM, Hornick JL (2018) Evaluation of pan-TRK immunohistochemistry in infantile fibrosarcoma, lipofibromatosis-like neural tumour and histological mimics. Histopathology 73:634-644. https://doi.org/10.1111/his.13666
Rudzinski ER, Lockwood CM, Stohr BA, Vargas SO, Sheridan R, Black JO, Rajaram V, Laetsch TW, Davis JL (2018) Pan-Trk immunohistochemistry identifies NTRK rearrangements in pediatric mesenchymal tumors. Am J Surg Pathol 42:927-935. https://doi.org/10.1097/PAS.0000000000001062
Davis JL, Lockwood CM, Stohr B, Boecking C, Al-Ibraheemi A, DuBois SG, Vargas SO, Black JO, Cox MC, Luquette M, Turpin B, Szabo S, Laetsch TW, Albert CM, Parham DM, Hawkins DS, Rudzinski ER (2019) Expanding the spectrum of pediatric NTRK-rearranged mesenchymal tumors. Am J Surg Pathol 43:435-445. https://doi.org/10.1097/PAS.0000000000001203
Suurmeijer AJ, Dickson BC, Swanson D, Zhang L, Sung YS, Huang HY, Fletcher CD, Antonescu CR (2019) The histologic spectrum of soft tissue spindle cell tumors with NTRK3 gene rearrangements. Genes Chromosom Cancer 58:739-746. https://doi.org/10.1002/gcc.22767
Rabban JT, Devine P, Sangoi AR, Poder L, Alvarez E, Davis JL, Rudzinski E, Garg K, Bean GR (2020) NTRK fusion cervical sarcoma: a report of 3 cases, emphasizing morphological and immunohistochemical distinction from other uterine sarcomas, including adenosarcoma. Histopathology. https://doi.org/10.1111/his.14069
Wong DD, Vargas AC, Bonar F, Maclean F, Kattampallil J, Stewart C, Sulaiman B, Santos L, Gill AJ (2020) NTRK-rearranged mesenchymal tumours: diagnostic challenges, morphological patterns and proposed testing algorithm. Pathology 52:401-409. https://doi.org/10.1016/j.pathol.2020.02.004
Yamamoto H, Nozaki Y, Kohashi K, Kinoshita I, Oda Y (2020) Diagnostic utility of pan-Trk immunohistochemistry for inflammatory myofibroblastic tumours. Histopathology 76:774-778. https://doi.org/10.1111/his.14010
Gatalica Z, Xiu J, Swensen J, Vranic S (2019) Molecular characterization of cancers with NTRK gene fusions. Mod Pathol 32:147-153. https://doi.org/10.1038/s41379-018-0118-3
Solomon JP, Linkov I, Rosado A, Mullaney K, Rosen EY, Frosina D, Jungbluth AA, Zehir A, Benayed R, Drilon A, Hyman DM, Ladanyi M, Sireci AN, Hechtman JF (2020) NTRK fusion detection across multiple assays and 33,997 cases: diagnostic implications and pitfalls. Mod Pathol 33:38-46. https://doi.org/10.1038/s41379-019-0324-7
Leeman-Neill RJ, Kelly LM, Liu P, Brenner AV, Little MP, Bogdanova TI, Evdokimova VN, Hatch M, Zurnadzy LY, Nikiforova MN, Yue NJ, Zhang M, Mabuchi K, Tronko MD, Nikiforov YE (2014) ETV6-NTRK3 is a common chromosomal rearrangement in radiation-associated thyroid cancer. Cancer 120:799-807. https://doi.org/10.1002/cncr.28484
Picarsic JL, Buryk MA, Ozolek J, Ranganathan S, Monaco SE, Simons JP, Witchel SF, Gurtunca N, Joyce J, Zhong S, Nikiforova MN, Nikiforov YE (2016) Molecular characterization of sporadic pediatric thyroid carcinoma with the DNA/RNA ThyroSeq v2 next-generation sequencing assay. Pediatr Dev Pathol 19:115-122. https://doi.org/10.2350/15-07-1667-OA.1
Prasad ML, Vyas M, Horne MJ, Virk RK, Morotti R, Liu Z, Tallini G, Nikiforova MN, Christison-Lagay ER, Udelsman R, Dinauer CA, Nikiforov YE (2016) NTRK fusion oncogenes in pediatric papillary thyroid carcinoma in northeast United States. Cancer 122:1097-1107. https://doi.org/10.1002/cncr.29887
Seethala RR, Chiosea SI, Liu CZ, Nikiforova M, Nikiforov YE (2017) Clinical and morphologic features of ETV6-NTRK3 translocated papillary thyroid carcinoma in an adult population without radiation exposure. Am J Surg Pathol 41:446-457. https://doi.org/10.1097/PAS.0000000000000814
Bastos AU, de Jesus AC, Cerutti JM (2018) ETV6-NTRK3 and STRN-ALK kinase fusions are recurrent events in papillary thyroid cancer of adult population. Eur J Endocrinol 178:83-91. https://doi.org/10.1530/EJE-17-0499
Chu YH, Dias-Santagata D, Farahani AA, Boyraz B, Faquin WC, Nose V, Sadow PM (2020) Clinicopathologic and molecular characterization of NTRK-rearranged thyroid carcinoma (NRTC). Mod Pathol https://doi.org/10.1038/s41379-020-0574-4
Pekova B, Sykorova V, Dvorakova S, Vaclavikova E, Moravcova J, Katra R, Astl J, Vlcek P, Kodetova D, Vcelak J, Bendlova B (2020) RET, NTRK, ALK, BRAF and MET fusions in a large cohort of pediatric papillary thyroid carcinomas, Thyroid. https://doi.org/10.1089/thy.2019.0802
Hang JF, Li AF, Chang SC, Liang WY (2016) Immunohistochemical detection of the BRAF V600E mutant protein in colorectal cancers in Taiwan is highly concordant with the molecular test. Histopathology 69:54-62. https://doi.org/10.1111/his.12903
Cameselle-Teijeiro JM, Peteiro-Gonzalez D, Caneiro-Gomez J, Sanchez-Ares M, Abdulkader I, Eloy C, Melo M, Amendoeira I, Soares P, Sobrinho-Simoes M (2018) Cribriform-morular variant of thyroid carcinoma: a neoplasm with distinctive phenotype associated with the activation of the WNT/beta-catenin pathway. Mod Pathol 31:1168-1179. https://doi.org/10.1038/s41379-018-0070-2
Seethala RR, Baloch ZW, Barletta JA, Khanafshar E, Mete O, Sadow PM, LiVolsi VA, Nikiforov YE, Tallini G, Thompson LD (2018) Noninvasive follicular thyroid neoplasm with papillary-like nuclear features: a review for pathologists. Mod Pathol 31:39-55. https://doi.org/10.1038/modpathol.2017.130
Pozdeyev N, Gay LM, Sokol ES, Hartmaier R, Deaver KE, Davis S, French JD, Borre PV, LaBarbera DV, Tan AC, Schweppe RE, Fishbein L, Ross JS, Haugen BR, Bowles DW (2018) Genetic analysis of 779 advanced differentiated and anaplastic thyroid cancers. Clin Cancer Res 24:3059-3068. https://doi.org/10.1158/1078-0432.CCR-18-0373
Nikiforov YE (2002) RET/PTC rearrangement in thyroid tumors. Endocr Pathol 13:3-16. https://doi.org/10.1385/ep:13:1:03
Chou A, Fraser S, Toon CW, Clarkson A, Sioson L, Farzin M, Cussigh C, Aniss A, O'Neill C, Watson N, Clifton-Bligh RJ, Learoyd DL, Robinson BG, Selinger CI, Delbridge LW, Sidhu SB, O'Toole SA, Sywak M, Gill AJ (2015) A detailed clinicopathologic study of ALK-translocated papillary thyroid carcinoma. Am J Surg Pathol 39:652-659. https://doi.org/10.1097/PAS.0000000000000368
Harrison BT, Fowler E, Krings G, Chen YY, Bean GR, Vincent-Salomon A, Fuhrmann L, Barnick SE, Chen B, Hosfield EM, Hornick JL, Schnitt SJ (2019) Pan-TRK immunohistochemistry: a useful diagnostic adjunct for secretory carcinoma of the breast. Am J Surg Pathol 43:1693-1700. https://doi.org/10.1097/PAS.0000000000001366
Remoue A, Conan-Charlet V, Bourhis A, Flahec GL, Lambros L, Marcorelles P, Uguen A (2019) Non-secretory breast carcinomas lack NTRK rearrangements and TRK protein expression. Pathol Int 69:94-96. https://doi.org/10.1111/pin.12766
Hechtman JF, Benayed R, Hyman DM, Drilon A, Zehir A, Frosina D, Arcila ME, Dogan S, Klimstra DS, Ladanyi M, Jungbluth AA (2017) Pan-Trk immunohistochemistry is an efficient and reliable screen for the detection of NTRK fusions. Am J Surg Pathol 41:1547-1551. https://doi.org/10.1097/PAS.0000000000000911
Lasota J, Chlopek M, Lamoureux J, Christiansen J, Kowalik A, Wasag B, Felisiak-Golabek A, Agaimy A, Biernat W, Canzonieri V, Centonze G, Chmielik E, Daum O, Dubova M, Dziuba I, Goertz S, Gozdz S, Guttmejer-Nasierowska A, Haglund C, Halon A, Hartmann A, Inaguma S, Izycka-Swieszewska E, Kaczorowski M, Kita P, Kolos M, Kopczynski J, Michal M, Milione M, Okon K, Peksa R, Pyzlak M, Ristimaki A, Rys J, Szostak B, Szpor J, Szumilo J, Teresinski L, Waloszczyk P, Wejman J, Wesolowski W, Miettinen M (2020) Colonic adenocarcinomas harboring NTRK fusion genes: a clinicopathologic and molecular genetic study of 16 cases and review of the literature. Am J Surg Pathol 44:162-173. https://doi.org/10.1097/PAS.0000000000001377
Acknowledgments
The authors would like to thank Ms. Shu-Ying Wang and Ms. Yun-Hsin Liang for technical assistance and the Biobank, Taipei Veterans General Hospital for the assistance with sample preparation in this study.
Funding
The study was supported by the research grants from Taipei Veterans General Hospital (Grant No.: V109B-029), Taipei Veterans General Hospital-National Yang-Ming University Excellent Physician Scientists Cultivation Program (Grant No.: 109-V-B-002 and 109-V-B-003), and Taipei Institute of Pathology (Grant No.: TIP-108-004).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This study was approved by the IRB of Taipei Veterans General Hospital, which granted exemption of informed consent for tissue procurement through the Biobank of Taipei Veterans General Hospital after an unlinked anonymous process (IRB no.: 2019-07-001BC, Biobank no.: 10818).
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Lee, YC., Chen, JY., Huang, CJ. et al. Detection of NTRK1/3 Rearrangements in Papillary Thyroid Carcinoma Using Immunohistochemistry, Fluorescent In Situ Hybridization, and Next-Generation Sequencing. Endocr Pathol 31, 348–358 (2020). https://doi.org/10.1007/s12022-020-09648-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12022-020-09648-9