Abstract
Type I interferons (IFN-Is) are a very important group of cytokines that are produced by innate immune cells but also act on adaptive immune cells. IFN-Is possess antiviral, antitumor, and anti-proliferative effects, as well are associated with the initiation and maintenance of autoimmune disorders. Studies have shown that aberrantly expressed IFN-Is and/or type I IFN-inducible gene signatures in the serum or tissues of patients with autoimmune disorders are linked to their pathogenesis, clinical manifestations, and disease activity. Type I interferonopathies with mutations in genes impacting the type I IFN signaling pathway have shown symptoms and characteristics similar to those of systemic lupus erythematosus (SLE). Furthermore, both interventions in animal models and clinical trials of therapies targeting the type I IFN signaling pathway have shown efficacy in the treatment of autoimmune diseases. Our review aims to summarize the functions and targeted therapies (as well as clinical trials) of IFN-Is in both adult and pediatric autoimmune diseases, such as SLE, pediatric SLE (pSLE), rheumatoid arthritis (RA), juvenile idiopathic arthritis (JIA), juvenile dermatomyositis (JDM), Sjögren syndrome (SjS), and systemic sclerosis (SSc), discussing the potential abnormal regulation of transcription factors and epigenetic modifications and providing a potential mechanism for pathogenesis and therapeutic strategies for future clinical use.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Autoimmune diseases include a wide range of diseases, including systemic lupus erythematosus (SLE), rheumatoid arthritis (RA), Sjögren syndrome (SjS), and systemic sclerosis (SSc) [1]. They occur when the immune system loses the ability to maintain self-tolerance and produces overactive immunocytes and an abnormal balance of cytokines [3]. These lead to abnormally increased autoantibodies against self-tissues, resulting in inflammation and damage to multiple organs, such as the kidneys, joints, skin, and central nervous system. The conventional treatments of autoimmune disorders include anti-inflammatory and immunosuppressive agents, which often provide only partial amelioration of symptoms [4,5,6,7,8]. In addition, these agents possess numerous side effects, including infection, obesity, abnormal glucose metabolism, osteoporosis, and even avascular necrosis [4, 5]. Due to the complex and unclear mechanisms of the pathogenesis of autoimmune diseases, the effective therapies are lacking [6,7,8].
In 1979, IFNs were found to be increased in autoimmune disorders, and there is a positive correlation between the amount of IFNs in the sera and disease activity as well as the level of antibodies to DNA in patients with SLE [9]. In addition, after IFN-α therapy, autoimmune disorders such as autoimmune thyroid disease and SLE-like syndrome can occur in patients with hepatitis C virus or malignant carcinoid tumors [10, 11]. Similarly, type I IFN signatures are also increased in the patients with RA and dermatomyositis (DM), demonstrating a role of IFNs in promoting the development of autoimmune diseases [12, 13].
Type I IFNs are widely expressed cytokines that are involved in innate immunity and adaptive immune system and in particular play an important role in antiviral activity and cancer immunosurveillance [14,15,16]. IFN-Is are mainly produced by innate immune cells in mammals, such as macrophages and dendritic cells (DCs) [17]. IFN-β can be induced by most types of cells, whereas IFN-α is largely produced by plasmacytoid DC (pDCs) [17]. Once produced, type I IFNs interact with the interferon-α/β receptor (IFNAR, including subunits IFNAR1 and IFNAR2) expressed in most nucleated cells, and ultimately initiate the transcription of interferon-stimulated genes (ISGs) to produce type I IFN gene signature through JAK/signal transducers and activators of transcription (STAT) pathway [14, 17, 18]. Many molecules encoded by ISGs induce antiviral and antitumor activity but may also play a role in autoimmunity [12].
With more attention on the significance of type I IFN signaling pathway in autoimmune diseases, many monoclonal antibodies (mAbs) and small molecule inhibitors are designed to target the type I IFN signaling pathway for the treatment of autoimmune disorders, such as therapies targeting IFN-α or IFNARs, pDCs, Toll-like receptors (TLRs), and their downstream pathways. Clinical trials using these treatments have already produced positive results. The JAK inhibitors tofacitinib and baricitinib have been approved by the United States Food and Drug Administration (FDA) and the European Medicines Agency (EMA) for the treatment of RA [19]. A mAb against IFNAR, anifrolumab (formerly MEDI-546), has just completed phase III clinical trials in patients with SLE and has shown excellent efficacy and safety profiles [20]. The JAK1/JAK2 inhibitor baricitinib has also entered phase III trials for the treatment of SLE [21]. Furthermore, there are also many ongoing clinical trials and experiments about treatments blocking the type I IFN pathway in both humans and animals with SLE, pediatric SLE (pSLE), RA, JIA, SjS, and other autoimmune diseases. However, there are also some challenges that will be discussed in more detail below. We focus on type I IFNs (mainly IFN-α and IFN-β) and summarize their roles in the pathogenesis and treatment of autoimmune diseases. Our purpose is to find new potential pathogenesis and therapeutic strategies for type I IFNs for further studies.
The Family and Origin of Type I Interferons
The IFN family contains three groups: type I, type II, and type III IFNs. IFN-Is include IFN-α (13 distinct subtypes of IFN-α in humans and 14 in mice), IFN-β, IFN-δ, IFN-ε, IFN-κ, IFN-ω, and IFN-ζ [22]. Humans can express IFN-α, IFN-β, IFN-ε, IFN-κ, and IFN-ω [22]. The production of type I IFNs is induced when different stimuli are detected by pattern recognition receptors (PRRs) on the plasma membrane in endosomes and the cytoplasm of aforementioned type I IFN-producing cells [17]. Different stimuli consist of exogenous pathogens, apoptotic cells, apoptosis-derived membrane vesicles (AdMVs), NETotic neutrophils, endogenous self-nucleic acids, and related immune complexes (ICs) [23,24,25,26]. The PRRs, such as TLRs, retinoic acid-inducible gene-I (RIG-I)-like receptors (RLRs), and nod-like receptors (NLRs), can be divided into DNA or RNA sensors during the productions of type I IFNs [2].
TLR9 is an endosomal membrane sensor for DNA stimuli and is mainly expressed in pDCs, macrophages, B cells [24, 25, 27, 28]. Extracellular stimuli containing nucleic acids can be internalized by FcγRIIa on the surface of pDCs to form endosomes [23,24,25,26]. TLR9 in the endoplasmic reticulum (ER) can traffic into the endosome and deliver extracellular DNA signals. Upon recognizing unmethylated CpG DNA released by the above complexes, TLR9 interacts with myeloid differentiation primary response gene 88 (MyD88). MyD88 then recruits and activates interleukin-1 receptor-associated kinase 4 (IRAK4), triggering IRAK1, tumor necrosis factor receptor-associated factor 6 (TRAF6), and TRAF3 pathway to activate interferon regulatory family 7 (IRF7), which can translocate to the nucleus and initiate transcription of IFN-Is [23,24,25,26].
Cyclic GMP-AMP synthase (cGAS) is expressed in most cells and is the most important intracellular DNA sensor during the production of IFN-Is [25, 29, 30]. When activated by double-stranded (ds) DNA, cGAS initiates the synthesis of cyclic dinucleotide second messenger (cGAMP) and then activates stimulator of interferon genes (STING) in the ER [29]. STING then recruits TANK-binding kinase 1 (TBK1) to phosphorylate IRF3, which can translocate to the nucleus to promote IFN-β production [29]. Other intracellular receptors for DNA stimuli include interferon-induced protein 16 (IFI16), Z-DNA-binding protein 1 (ZBP1, also known as DAI), and DNA-dependent RNA polymerase III [25, 30, 31].
TLR3, TLR7, and TLR8 are endosomal membrane receptors expressed in many innate immune cells, and TLR7 is particularly highly expressed in pDCs [24, 27, 28, 32, 33]. TLR3 can detect dsRNA in endosomes internalized by FcγRIIa, and TLR7 and TLR8 can detect single-stranded (ss) RNA in endosomes [24, 33]. After activation, TLR3 can bind to Toll-IL-1 receptor domain-containing adaptor inducing IFN-β (TRIF) to activate TRAF3 and then recruit and activate TBK1 to phosphorylate IRF3 for IFN-β production [34]. Currently, cyclin-dependent kinases (CDKs) have been found to be critical in the translation of IFN-β, and CDK inhibition dramatically suppresses the transcription of ISGs [35]. Activated TLR7 and TLR8 induce the transcription of IFN-Is in the same manner as that of TLR9 [34].
RLRs include RIG-I, melanoma differentiation-associated gene 5 (MDA5), and laboratory of genetics and physiology 2 (LGP2) [33]. RIG-I and MDA5 are expressed in cytoplasm of fibroblasts, cDCs, macrophages [33, 35]. RIG-I detects short ds-5′ tri-phosphorylated RNA and ss-5′ tri-phosphorylated RNA, whereas MDA5 can recognize long dsRNA and ssRNA [33, 35, 36]. After activation, RIG-I and MDA5 bind to the adaptor mitochondrial antiviral signaling protein (MAVS) on the outer membrane of mitochondria and then activate IRF3 through TRAF3 and TBK1 [37]. TLR4 can sense extracellular lipopolysaccharide (LPS) on the cell membrane. Activated TLR4 recruits TRAM and then activates TRIF, TBK1, and IRF3 to initiate the transcription of IFN-Is [34].
Type I Interferon Signaling Pathway
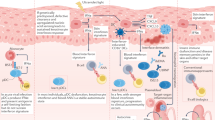
IFN-α and IFN-β exert their immune functions by ligating with their specific IFNARs that are expressed on the plasma membrane of most nucleated cells [18]. The interactions between IFNAR and IFN-α/β drive the dimerization of IFNARs, and its subunits IFNAR1 and IFNAR2 separately bind to and activate different types of Janus kinase (JAK) protein families: IFNAR1 activates tyrosine kinase 2 (TYK2) and IFNAR2 activates JAK1 [15]. Later, cytoplasmic transcription factors STAT1 and STAT2 can be phosphorylated by JAKs, and interferon regulatory family 9 (IRF9) can bind to STAT1/STAT2 heterodimers to form the heterotrimeric complex IFN-stimulated gene factor 3 (ISGF3), which can initiate transcription of ISGs by combining with IFN-stimulated response elements (ISREs, consensus sequence TTTCNNTTTC) [15, 17]. ISGs include a series of genes encoding antiviral and antitumor molecules, and transcription factors such as IRFs [12] (Fig. 1). Notably, IRF7 promotes the expression of ISGs that also include IRF7, forming a positive feedback loop within the type I IFN signaling pathway [38].
The production of type I interferons and type I interferon signaling pathway. Extracellular stimuli containing DNA can be internalized to form endosomes by FcγRIIa on the surface of pDCs, and then recognized by TLR9 to activate MyD88. MyD88 recruits IRAK4 and leads, through a pathway involving IRAK1, TRAF6, and TRAF3, to activation of IRF7, which can translocate into the nucleus and initiate transcription of IFN-Is. cGAS can be activated by intracellular dsDNA. cGAS phosphorylates IRF3 to translocate into the nucleus and induce IFN-β production through the cGAMP/STING/TBK1 pathway. CDKs are very critical in the translation of IFN-β. TLR3 can detect dsRNA and then activate IRF3 to induce IFN-β production by activating TRIF, TRAF3, and TBK1. TLR7 and 8 can recognize ssRNA in endosomes and then bind to MyD88 and interact with IRAK4, IRAK1, TRAF6, and TRAF3 to activate IRF7 to induce the transcription of IFN-Is. RIG-I and MDA5 mainly recognize dsRNA and then bind to the adaptor MAVS on the outer membrane of mitochondria, and MAVS then activates IRF3 through TRAF3 and TBK1. TLR4 can sense LPS on the cell membrane. Activated TLR4 recruits TRAM and then activates TRIF, TBK1, and IRF3 to initiate the transcription of IFN-Is. The interaction between IFN-α/β and IFNARs drives the dimerization of IFNARs, leading to the autophosphorylation and transphosphorylation of JAKs. TYK2 and JAK1 bind to IFNAR1 and IFNAR2, respectively. Type I IFN signals then pass to the inner side of cells via the JAK–STAT signaling pathway. Later, IFNAR1 can be phosphorylated by JAKs, leading to repositioning and binding of STAT2. Phosphorylated STAT2 can recruit and phosphorylate STAT1 to form STAT1/STAT2 heterodimers and separate from IFNAR. The heterotrimeric complex ISGF3 comprises IRF9 and STAT1/STAT2 heterodimers to initiate transcription of ISGs by combining with ISREs. AdMV: apoptosis-derived membrane vesicle; RIG-I: retinoic acid-inducible gene-I; MAVS: mitochondrial antiviral signaling protein; MDA5: melanoma differentiation-associated gene 5; TRAF3: tumor necrosis factor receptor-associated factor 3; TBK1: TANK-binding kinase 1; IRF3: interferon regulatory family 3; cGAS: cyclic GMP-AMP synthase; cGAMP: cyclic dinucleotide second messenger; STING: stimulator of IFN genes; LPS: lipopolysaccharide; TLR: Toll-like receptor; TRAM: TRIF-related adaptor molecule; TRIF: TIR-domain-containing adaptor protein-inducing IFN-β; MyD88: myeloid differentiation primary response gene 88; IRAK1: Interleukin-1 receptor-associated kinase 1; IFN: interferon; CDK: cyclin-dependent kinase; IFNAR1: interferon-α/β receptor 1; JAK: Janus kinase 1; TYK2: tyrosine kinase 2; STAT1: signal transducers and activators of transcription 1; ISGF3: IFN-stimulated gene factor 3; ISG: interferon-stimulated gene
The Regulation of the Type I Interferon Signaling Pathway
The Interferon Regulatory Family
Interferon regulatory family (IRF) family is the regulator of IFN signaling pathways and includes 9 members in human: IRF1, IRF2, IRF3, IRF4 (PIP/LSIRF/ICSAT), IRF5, IRF6, IRF7, IRF8 (ICSBP), and IRF9 (ISGF3γ) [14]. IRF proteins contain conserved amino-terminal DNA-binding domains (DBDs), carboxy-terminal IRF-association domain 1 (IAD1) and IAD2 [14]. The DBDs can detect and regulate ISRE, and the IADs can mediate homodimers and heterodimers with other IRFs and transcription factors [14].
The type I IFN signaling pathway begins with IRF3 being phosphorylated via signaling from PRRs, which leads to expression of IFN-β and IFN-α4 through dimerization, nuclear translocation, and formation of a multiprotein complex with coactivators CREB binding protein (CBP) and p300 [39]. IRF7 is poorly expressed in multiple cells but can be strongly induced by the type I IFN signaling pathway [38]. Type I IFNs induced by IRF3 then initiate the transcription and expression of ISGs that include IRF7, initiating a positive feedback loop of IRF7 [12]. At the later phase, both IRF3 and IRF7 can promote the expression of IFN-Is [39].
IRF1, IRF5, IRF8, and IRF9 also participate in the regulation of type I IFN signaling pathways [38]. IRF9 is a part of ISGF3 that is necessary for the transcription of ISGs [15, 17]. IRF8 has been proved to cooperatively promote the rapid production of IFN-β induced by IRF3 in human monocytes [40]. IRF1 and MyD88 are required for the expression of IFN-β in cDCs but not pDCs [41]. IRF5 promotes the transcriptions of specific ISREs that are mainly for encoding proinflammatory cytokines [38, 39]. Additionally, IRF4 and IRF8 are essential for DC subset development [39].
Post-Translational Modifications of the Type I Interferon Signaling Pathway
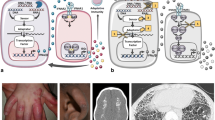
Post-translational modifications (PTMs), including phosphorylation, polyubiquitylation, methylation, acetylation, ISGylation, and SUMOylation, are emerging mechanisms for regulating the type I IFN signaling pathway to maintain the balance of the production of IFN-Is and transcription of ISGs [17, 42]. PTMs of type I IFN signaling pathway are summarized in Fig. 2 and Table 1.
The post-translational modifications (PTMs) of type I interferon pathway. The gray boxes represent the negative regulators of type I IFN signaling pathway, and the orange arrows shows the path of type I IFN pathway and the blue one represents the negative webs. IFI35, RNF125, NLRC5, PCBP2, CYLD, TRIM13, and TRIM40 can negatively regulate the type I IFN pathway through sensing of RNA via RIG-I or MDA5 and signaling MAVS. A20, TRIP, TRIM8, TRIM11, TRIM26, TRIM27, TRIM56, and SHP-2 can suppress TBK1, and NLRC5, TRIM26, TRIM27, TRIM56, PLASy, ISG56, ASCC3, RAUL, SMAD2, and SMAD3 can inhibit IRF3. TRIM38 can induce SUMOylation of cGAS and STING to inhibit type I IFN response, and TRIM32 and TRIM56 can induce polyubiquitination of STING. The TRIF can be negatively regulated by TRIM32, TRIM38, SHP-2, and TRAF3 and TRAF6 through TLRs signaling can be inhibited by TRIM38, OTUB1, OTUB2, and DUBA. GBP4 can disrupt the interactions between TRAF6 and IRF7, and IRF7 can be directly suppressed by PLASy, ISG56, Nmi, OASL1, DDX24, ASCC3, and RAUL. The production of IFN-β can be directly inhibited via TRIP, ISG56, SMAD2, and SMAD3. PKD2 can regulate ubiquitination and degradation of IFNAR1, and USP18 can compete with JAK1 for IFNAR2. SHP-1, SHP-2, PTPN1, TC-PTP, SOCS1, SOCS3, PLAS1, PLASy, C1S, and DCST1 can suppress the JAK–STAT pathway by inhibiting JAKs or STATs. RIG-I: retinoic acid-inducible gene-I; MAVS: mitochondrial antiviral signaling protein; MDA5: melanoma differentiation-associated gene 5; TRAF3: tumor necrosis factor receptor–associated factor 3; TBK1: TANK-binding kinase 1; IRF3: interferon regulatory family 3; cGAS: cyclic GMP-AMP synthase; cGAMP: cyclic dinucleotide second messenger; STING: stimulator of IFN genes; TLR: Toll-like receptor; TRAM: TRIF-related adaptor molecule; TRIF: TIR-domain-containing adaptor protein-inducing IFN-β; MyD88: myeloid differentiation primary response gene 88; IRAK1: Interleukin-1 receptor-associated kinase 1; IFNAR1: interact with interferon-α/β receptor 1; JAK: Janus kinase 1; TYK2: tyrosine kinase 2; STAT1: signal transducers and activators of transcription 1; ISGF3: IFN-stimulated gene factor 3; ISG: interferon-stimulated gene; IFI35: interferon-induced protein 35; RNF125: ring-finger protein 125; NLRC5: NLR family CARD domain-containing 5; PCBP2: poly(rC)-binding protein 2; CYLD: cylindromatosis lysine 63 deubiquitinase; TRIP: TNF receptor-associated factor (TRAF)–interacting protein; TRIM: tripartite motif-containing; GBP4: guanylate binding protein 4; OTUB1: OTU domain-containing ubiquitin aldehyde binding protein 1; DUBA: deubiquitinating enzyme A; PLAS: protein inhibitor of activated STAT; ISG56: IFN-stimulated gene 56; IFIT1: interferon-induced protein with tetratricopeptide repeats 1; Nmi: N-Myc and STATs interactor; OASL1: oligoadenylate synthetase-like 1; DDX24: DEAD-box helicase 24; ASCC3: activating signal cointegrator complex 3; RAUL: RTA-associated ubiquitin ligase; SMAD2: SMAD family member 2; PKD2: protein kinase D2; USP18: ubiquitin-specific peptidase 18; SHP-1: SH2-containing protein tyrosine phosphatase 1; PTPN1: protein tyrosine phosphatase, non-receptor type 1; PTP1B: protein tyrosine phosphatase 1B; TC-PTP: T cell-protein tyrosine phosphatase; PTPN2: protein tyrosine phosphatase, non-receptor type 2; SOCS: suppressors of cytokine signaling; CIS: cytokine-induced SRC-homology 2 (SH2) protein; DCST1: DC-STAMP domain-containing 1
Epigenetic Modifications in the Regulation of the Type I Interferon Signaling Pathway
Epigenetic modifications generate hereditary alterations in gene expression without changing the DNA coding sequence [71]. Epigenetic modifications, including DNA methylation, histone modifications, microRNAs (miRNAs), and long non-coding RNAs (lncRNAs), are important in regulating the type I IFN pathway [71].
DNA methylation is involved in heritable DNA modification at the 5′-position of cytosine rings residing in the dinucleotide sequence CpG [72]. Decreased DNA methylation in IFN-induced genes have been found in patients with SLE, compared to healthy controls [73]. Several different methylation positions and genes have been identified in patients with RA, including MX dynamin-like GTPase 1 (MX1), IFN-induced protein 44-like (IFI44L), Deltex E3 Ubiquitin Ligase 3L (DTX3L), and polymerase family member 9 (PARP9), forming the IFN-inducible gene interaction network [74]. In addition, hypomethylation of IFN-regulated genes, including MX1, IFI44L, and PARP9, have also been identified in patients with primary SjS (pSjS) [75]. Many hypomethylations of type I IFN-associated genes have been identified in CD4+ and CD8+ methylated regions in patients with SSc [76]. Furthermore, inhibition of DNA methylation can induce type I IFN responses in cancer, demonstrating negative regulatory functions of DNA methylation in type I IFN pathway [77].
Histone modifications include methylation, acetylation, ubiquitination, and phosphorylation [78]. The dimethylation of histone H3 at lysine 9 (H3K9me2) has an inhibitory effect on IFN and ISG expression [79]. Methyltransferase SET domain-containing protein 2 (SETD2) can catalyze the trimethylation of H3K36 at certain ISG promoters, such as ISG15, to activate them, and SETD2 can also amplify the functions of IFN-Is by inducing STAT1 methylation [80]. Disruption of telomeric silencing-1 like (DOT1L) protein is a methyltransferase that can mediate the modification of H3K79me2/3 at Ifnb1 promoters, promoting the transcription and expression of IFN-β [81]. The transcription of IFN-β can also be regulated by the histone acetyltransferase proteins CBP and p300/CBP-associated factors (PCAF)/general control non-derepressible 5 (GCN5) [82]. The GCN5/PCAF complex is recruited to acetylate nucleosomes to accelerate the transcription of IFN-β through recruiting SWItch/sucrose non-fermentable (SWI/SNF) to bind to transcription initiation factor IID (TFIID) [83].
Histone deacetylase 1 (HDAC1) and HDAC8 can inhibit the production of IFN-β by inducing histone H3/H4 deacetylation of the Ifnb1 promoter [84]. However, HDAC has also been shown to promote the initiation of transcription or transcription elongation of ISGs by recruiting P-TEFb to ISG promoters through Brd4 [85]. Furthermore, small ubiquitin-like modifier (SUMO) can silence Ifnb1 at its promoter through sumoylation [86]. In addition, PARP9-DTX3L is also an E3 ubiquitin ligase and can bind to STAT1 to act on host histone H2BJ to promote the expression of ISGs [42, 87]. The human Bre1/RNF20 complex is necessary to promote global increased monoubiquitination of histone 2B at lysine 120 to activate the expression of ISGs [88].
miRNA and lncRNAs are also important in regulating type I IFN signaling pathways. miR-466l can directly suppress multiple IFN-α species via their 3′ untranslated region (3′UTR) [89]. miR-130b negatively regulates IFN-α in renal cells in a mouse model of lupus nephritis, whereas miR-618 can increase IFN-α production in patients with SSc [90, 91]. miR-146a can suppress IFN-β production and miRNA-302d can suppress ISG expression by targeting IRF9 [92, 93]. miR-279 can inhibit type I IFN signaling directly via inhibiting STAT [94]. miR-155 can inhibit suppressors of cytokine signaling 1 (SOCS1) and promote the phosphorylation of STAT1 and STAT3 in hepatocarcinoma cells [95]. Controversially, type I IFN signaling was found to be enhanced in miR-155-deficient CD8+ T cells [96]. In addition, miR-221 and miR-203 can activate JAK–STAT pathway by inhibiting SOCS1 and SOCS3 [97]. The miR-21 negatively regulates IFN-α signaling through targeting MyD88 and IRAK1 in patients with HCV infection [98]. Furthermore, lncRNA#32 has been shown to upregulate ISG expression and the antiviral functions of type I IFN [99]. lncLrrc55-AS can promote the production of IRF3 and IFN-I [100]. However, lncRNA-CMPK2 has been identified as a negative regulator of the type I IFN signaling pathway [101].
Susceptibility Genes in the Type I Interferon Signaling Pathway in Autoimmune Diseases
Several susceptibility genes have been found to be related to the production of type I IFN or type I IFN signaling pathway in autoimmune diseases. Susceptibility genes of SLE associated with type I IFNs include IRF5, IRF7, IRF8, IRAK1, tumor necrosis factor, alpha-induced protein 3 (TNFAIP3), TNFAIP3 interacting protein 1 (TNIP1), interferon induced with helicase C domain 1 (IFIH1), TYK2, STAT4, protein tyrosine phosphatase non-receptor type 22 (PTPN22), three prime repair exonuclease 1(TREX1), E26 transformation-specific 1 (ETS1), and ubiquitin-conjugating enzyme E2 L3 (UBE2L3) [102,103,104,105,106,107].
Large-scale association studies have shown a relationship between polymorphisms of IRF5, IRF7, IRF8, and SLE risk [102]. Several variants of IRF5, IRF7, and IRF8 can induce transcription and expression of proteins in regulating the type I IFN signaling pathway [102]. Notably, IRF5 and STAT4 are considered the most strongly SLE-associated loci except for the MHC region [103, 104]. A variant of STAT4 (rs7574865) is related to elevated sensitivity to IFN-α signaling [103]. STAT4 protein can be phosphorylated and activated by interacting with IFNAR2 cytoplasmic domain, and then translocate into nucleus to initiate the expression of IFN-γ and other proinflammatory molecules [108,109,110]. PTPN22 is the susceptibility gene for both SLE and RA in the European population [103, 105]. High IFN-α and low TNF-α in serum occur in patients with SLE with the rs2476601 risk allele of PTPN22 [103, 105]. PTPN22 protein can directly interact with and promote the ubiquitination of TRAF3 [111]. TNFAIP3 and TNIP1, which are proven to be susceptibility genes of SLE through GWAS, encode A20 and A20-binding protein, respectively, and are regulators in type I IFN pathway, NF-κB activation, and TNF-mediated apoptosis [102,103,104].
TREX1 encodes 3′-5′ DNA exonuclease TREX1 to clear aberrant DNA and is a susceptibility gene of SLE [103, 105]. IFIH1 encodes the dsRNA sensor MDA5 [102]. A missense allele of IFIH1 (rs1990760) is identified to be related to increased transcription and expression of IFIH1 and is involved in elevated expression of IFN-induced genes in anti-dsDNA-positive patients with SLE [102, 104]. TYK2 encodes TYK2, a member of the JAK–STAT pathway [105]. UBE2L3 has genome-wide significance in relation to SLE in Asian but not European populations [103]. UBE2L3 encodes an E2 ubiquitin-conjugating enzyme that can interact with Triad3A (also called RNF216) to regulate the degradation of TLR4 and TLR9 [112]. IRAK1 acts on the delivery of TLR signaling, and IRAK1 gene polymorphisms were identified to be related to the pathogenesis of SLE [106, 107]. ETS1 is a negative regulator in expression of IFN-induced genes by interacting with ISGs [102]. ETS1 is an SLE susceptibility locus in Asia, and the SNP rs6590330 is related to decreased expression of ETS1 [102, 105].
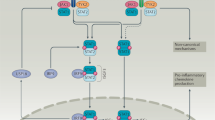
Many SLE risk susceptibility genes involved in the type I IFN pathway also participate in the pathogenesis of other autoimmune diseases, such as RA, JIA, SSc, and SjS. Susceptibility genes in RA related to the type I IFN pathway include PTPN22, STAT4, TNFAIP3, IRF5, UBE2L3, ETS1, IRF8, TYK2, IRAK1, and TRAF6 [105, 113]. IRF5, PTPN22, STAT4, TYK2, and UBE2L3 are associated with an increased risk of JIA [114, 115]. Susceptibility genes in SSc include IRF5, IRF7, IRF8, PTPN22, STAT4, TNFAIP3, TNIP1, and TYK2, while IRF5, STAT4 TNFAIP3, and TNIP1 are involved in SjS [116, 117]. The association between susceptibility genes and type I IFN signaling pathway is described in Fig. 3.
The susceptibility genes involved in type I interferon pathway in autoimmune diseases. The red words represent the susceptibility genes involved in type I IFN pathway and some of them also represent their encoded proteins. TREX1 encodes 3′-5’ DNA exonuclease TREX1, and TREX1 is related to the susceptibility of SLE by impacting the degradations and increasing the expression of IFN-Is. IFIH1 encodes dsRNA sensor MDA5 and is involved in increased transcription of IFIH1 and elevated expression of IFN-induced genes. UBE2L3 encodes an E2 ubiquitin-conjugating enzyme that involves in the degradation of TLR4 and TLR9. IRAK1 and TRAF6 are related to the production of IFN-Is and NF-κB, and the multiple SNPs with IRAK1 and TRAF6 are associated with onset and development of autoimmune diseases. PTPN22 can regulate type I IFN signatures by interacting with TRAF3, and the polymorphism in PTPN22 (rs2476601) is related to high serum IFN-α activity. IRF5, IRF7, and IRF8 play an important role in upregulating the type I IFN responses. Several polymorphisms of IRF5, IRF7, and IRF8 are related to their expression of proteins and increased IFN-Is. TYK2 encodes TYK2 in JAK–STAT pathway, and a variant (rs280519) of TYK2 is related to increased production of IFN-I. STAT4 encodes STAT4 protein that is associated with increased sensitivity to IFN-α signaling and elevated expression of IFN-γ. TNFAIP3 and TNIP1 encode A20 and A20-binding protein separately that are regulatory factors in type I IFN pathway, NF-κB activation. ETS1 can negatively regulate the expression of IFN-induced genes by interacting with ISGs. The SNP rs6590330 of ETS1 is related to the decreased expression of ETS1. MDA5: melanoma differentiation-associated gene 5; MAVS: mitochondrial antiviral signaling protein; TRAF3: tumor necrosis factor receptor–associated factor 3; TBK1: TANK-binding kinase 1; IRF3: interferon regulatory family 3; TLR: Toll-like receptor; TRAM: TRIF-related adaptor molecule; TRIF: TIR-domain-containing adaptor protein-inducing IFN-β; MyD88: myeloid differentiation primary response gene 88; IRAK1: Interleukin-1 receptor-associated kinase 1; IFNAR1: interact with interferon-α/β receptor 1; JAK: Janus kinase 1; TYK2: tyrosine kinase 2; STAT1: signal transducers and activators of transcription 1; ISGF3: IFN-stimulated gene factor 3; ISG: interferon-stimulated gene; TREX1: three prime repair exonuclease 1; IFIH1: interferon induced with helicase C domain 1; UBE2L3: ubiquitin-conjugating enzyme E2 L3; PTPN22: protein tyrosine phosphatase non-receptor type 22; TNFAIP3: tumor necrosis factor, alpha-induced protein 3; TNIP1: TNFAIP3 interacting protein 1
Type I Interferon Signaling Pathway and Autoimmune Diseases
Type I Interferonopathies
Type I interferonopathy is a type of Mendelian genetic disorder with diverse molecular defects and clinical features but with the same overproduction of type I IFNs [118, 119]. Type I interferonopathies include a wide range of genetic diseases, such as Aicardi-Goutières syndrome (AGS), Sting-associated vasculopathy with onset in infancy (SAVI), spondyloenchrondro-dysplasia (SPENCD), Singleton-Merten syndrome, and ISG15 deficiency [118, 119]. Type I interferonopathies usually occur in childhood and some of them possess lupus-like features, supporting the association between lupus and high level of IFN-Is [108]. Type I interferonopathies can be caused by mutations in multiple genes associated with type I IFN signaling pathway, such as TREX1, ribonuclease H2 subunit A (RNASEH2A), RNASEH2B, RNASEH2C, SAM, and HD domain-containing deoxynucleoside triphosphate triphosphohydrolase 1 (SAMHD1), adenosine deaminase RNA specific (ADAR1), and IFIH1 [119]. Type I interferonopathies are summarized in Table 2.
Due to the poor response of biologic disease–modifying antirheumatic drugs (DMARDs) in type I interferonopathies, there is increasing interest in targeted therapies related to the type I IFN signaling pathway [137]. JAK inhibition has shown promising results in type I interferonopathies [137,138,139,140]. Case reports of JAK inhibitors, such as baricitinib, ruxolitinib to treat ubiquitin-specific peptidase 18 (USP18) deficiency, SAVI, AGS, and TREX1 chilblain lupus, have been published [136,137,138,139,140,141,142]. Cases of chilblain lupus have also shown a good response to tofacitinib therapy [127]. Furthermore, in the NCT01724580 and NCT02974595 programs, 18 patients with CANDLE or SAVI treated with baricitinib also had good results [137]. The side effects of JAK inhibition seem to be mild and acceptable, with slight and reversible decreases in lymphocytes and platelets, elevated transaminases and serum creatinine, and herpes zoster infection [143, 144]. However, due to the extremely low incidence of type I interferonopathies, conducting clinical trials is difficult. In vitro experiments or the use of animal models may help identify additional JAK–STAT candidates.
Systemic Lupus Erythematosus and Pediatric Systemic Lupus Erythematosus
In 1999, genes induced by IFN-Is were found to be upregulated in peripheral blood mononuclear cells (PBMCs) of approximately two-thirds of adult patients with SLE through microarray gene expression analysis [145]. The level of type I IFNs is also positively related to the disease activities and severity of symptoms (such as skin rash, fever, and leucopenia) in patients with SLE [146]. In addition, ANA and anti-dsDNA antibodies appear in patients after treatment with IFN-Is in patients with malignant carcinoid tumors [11]. Moreover, TLRs have been identified to be crucial in the maintaining SLE [147].
IFN-α is a powerful inducer for the differentiation or activation of DCs [148]. The activation of T cells and the suppression of Treg cells can be induced by IFN-Is in lupus-prone mice after UVB exposure [149]. IFN-Is participate in early transitional B cells and promote the expansion of autoantibody-secreting cells [150]. In patients with SLE, IFN-Is can lead to a strong inflammatory response by suppressing the maturation of miR-146a [151]. IFN-β is increased in circulating B cells in SLE patients, especially those with nephritis and autoantibodies [152]. Moreover, increased IFN-κ promotes the development of CLE after ultraviolet light exposure [146, 153, 154].
pSLE or childhood-onset SLE (cSLE) occurs during childhood with more severe damage to internal organs (especially to kidney) than adult SLE, revealing its stronger association with genetics [155]. The lupus-like phenotypes in children, also called “monogenic lupus,” is one of type I interferonopathies and can be caused by mutation of genes involved in type I IFN signaling pathway, such as DNase1L3, TREX1, SAMHD1, RNASEH2ABC, ADAR1, IFIH1, ISG15 [130, 155]. In a study assessing type I IFN signaling in pediatric autoimmune diseases, 82% (64 of 78) of samples from patients with juvenile SLE (JSLE) and 75% (76 of 101) of samples from children with juvenile DM (JDM) had positive IFN scores [156]. Upregulated expression of ISGs was also observed in cSLE, and it can be suppressed with TBK1 inhibitors BX795 [157]. The expression of TLRs in JSLE is also increased [158]. TLRs can be activated by apoptotic neutrophils and the IFN-α expression can be suppressed after MyD88 and TRIF inhibitors in PBMC of JSLE patients [158]. However, more clinical research on inhibitors involved in type I IFN pathway is urgently needed [157, 158].
With better understanding of the role of IFN-I, many promising treatments targeting type I IFN signaling pathway are undergoing clinical studies in SLE [159]. Rontalizumab, a humanized IgG1 anti-IFN-α mAb, has an acceptable safety profile and induces decreased expression of IRGs in phase I studies of patients with SLE [160, 161]. However, due to the unmet endpoints in phase II trials, the development of rontalizumab was terminated [160]. Sifalimumab (formerly MEDI-545) is a fully humanized IgG1 anti-IFN-α mAb [162]. In a phase IIb trial in 2016, SLE patients treated with sifalimumab met the primary endpoint and their other observed indexes and symptoms were also significantly improved [162]. AGS-009 is a humanized IgG4 anti-IFN-α mAb that was shown to be generally tolerated by patients with SLE [163]. However, there were no subsequent trials. Interferon-α-kinoid (IFN-K) is a therapeutic vaccine targeting IFN-α [164, 165]. In a phase IIb trial, IFN-K significantly decreased IFN gene signatures and achieved the biological coprimary endpoint but not the clinical coprimary endpoint, revealing a need for further evaluation [165].
An anti-IFNAR1 mAb, anifrolumab, showed significant effects in decreasing the disease activity of SLE in phase II trials, and another phase IIb study in patients with moderate to severe SLE was considered very successful [166, 167]. Two phase III trials (known as TULIP–1 and TULIP–2) in patients with SLE were completed in 2019. One of the endpoints, British Isles Lupus Disease Activity Group (BILAG)–based Composite Lupus Assessment (BICLA), but not SLE response index (SRI), was met in the first phase III trial, TULIP-1, and both were achieved in TULIP-2 [20, 168]. Regardless of the seemingly conflicting results, anifrolumab is the most successful and promising IFN pathway-targeted therapy for SLE and awaits use in clinical practice [20, 168].
BIIB059 is a mAb that binds to blood DC antigen 2 (BDCA2), an inhibitory receptor on pDCs, and induces its rapid internalization to inhibit the production of IFN-Is [169]. BIIB059 was seemingly beneficial to patients with SLE with cutaneous manifestations [169]. Venetoclax (also called ABT-199) is a Bcl-2 antagonist that can kill pDCs in lupus-prone mice and has shown a good clinical response in patients with SLE in a phase I trial [170,171,172].
Hydroxychloroquine, a TLR9, TLR7, and TLR8 antagonist, is a conventional antimalarial drug for SLE [6]. The preclinical data of CPG-52364, an inhibitor of TLR7, TLR8, and TLR9, showed that it can enhance the effectiveness of hydroxychloroquine [173]. Dynavax posted a news article about a phase 1b/2a trial of DV1179, a TLR7 and TLR9 inhibitor, in patients with SLE, but it did not meet the pharmacodynamic endpoints. A small molecular TLR9 inhibitor ST2825 has demonstrated great potential, with promising safety and efficacy profiles in patients with SLE [173].
The IRAK4 inhibitor BMS-986126, and Bruton’s tyrosine kinase (BTK) inhibitor evobrutinib (also called M2591, MSC2364447C) have been shown to suppress the type 1 IFN signaling pathway in preclinical experiments of SLE, but the results of clinical trials are either lacking or unpublished [174, 175]. Baricitinib (also called LY3009104) is a selective inhibitor of JAK1/JAK2 and has shown significant efficacy in patients with SLE in phase trials [21]. Tofacitinib, an inhibitor of JAK1/JAK3, is approved by the FDA for the treatment of RA, psoriatic arthritis, and ulcerative colitis [176]. However, clinical trials of tofacitinib for SLE and CLE are still in progress [176]. Filgotinib, a highly selective JAK1 inhibitor, has just finished phase II clinical trials. R333, a topical JAK/SYK inhibitor for DLE treatment, failed to significantly improve lesion activity in Phase II trials [177]. For nucleic acid-targeted therapies, rhDNase seemed to be unsuccessful, whereas RSLV-132 showed good results in a phase I trial and has entered a phase II trial for SLE (Table 3) [176].
Rheumatoid Arthritis
Case reports show that IFN-β1, a therapy used in multiple sclerosis (MS), can lead to the development of RA [179]. Type I IFN signatures can be detected in the peripheral blood of patients with RA, even in the preclinical phase [12]. Data from clinical trials show that high IFN-I signatures can predict RA as a biomarker in at-risk individuals [180]. In addition, synovial DCs have been shown to promote the inflammatory response and differentiation of Th1 and Th17 cells [7]. Furthermore, some susceptibility genes in RA are related to the type I IFN signaling pathway, such as IRF5, STAT4, and PTPN22 [105, 113].
Due to the unsatisfactory response to DMARDs, inhibitors targeting the type I IFN pathway have entered clinical trials of patients with RA. The selective JAK1/JAK3 inhibitor tofacitinib and the selective JAK1/JAK2 inhibitor baricitinib are effective and are approved by the FDA and EMA for the treatment of RA, especially for those intolerant or unresponsive to DMARDs [19]. Upadacitinib and filgotinib are selective JAK1 inhibitors, and both have shown good results in patients with RA in phase II and III studies [19, 181, 182]. The JAK1/JAK3 inhibitor, peficitinib, has also been shown to be effective in patients with RA, but the data from phase III trials are still pending [183, 184]. On the other hand, decernotinib, a JAK3 inhibitor, did not show clear dose–response curves and had a higher risk of toxic effects at higher doses [185]. The JAK1/JAK2 inhibitor ruxolitinib is also a promising candidate and is entering clinical trials [178]. JAK inhibitors are generally tolerated in patients with RA, with a risk of herpes zoster but not serious infections or malignancies [186]. Currently, Syk and BTK inhibitors have also entered clinical trials for RA [187, 188].
Although INF-Is have been implicated in the pathogenesis of RA, intra-articular injections with IFN-α and intraperitoneal injection with IFN-β can prevent the occurrence or development of RA in wild-type mice or RA mouse models [189]. IFN-β injected daily can also relieve the symptoms of RA in rhesus monkeys [189]. However, in clinical trials, treatment with subcutaneous recombinant IFN-β showed no improvement in active patients in a phase II clinical trial (Table 4) [190].
Juvenile Idiopathic Arthritis
Juvenile idiopathic arthritis (JIA) is the most common pediatric rheumatic disease with persistent arthritis in children or teenagers under 16 years of age [191]. Both IFN-γ and IFN-Is contribute to the pathogenesis of JIA [192, 193]. Type I IFN-producing cells were identified to be enriched in synovial fluid and the IFN-α producing cells were found in the lymph-follicular-like structures, but the expression of IFN-α was not upregulated in the serum and synovial fluid [193]. In enthesitis-associated JIA, the expression of TLR2 and TLR4 was elevated in the peripheral blood and synovial fluid monocytes [194]. IRF5 (rs2004640) and PTPN22 1858C > T have been reported to be involved in an increased risk of JIA [113, 195].
More and more clinical studies involving inhibitors suppressing the type I IFN pathway have been conducted in patients with JIA. Selective JAK inhibitors, such as tofacitinib, have shown promising results in the treatment of JIA. In a phase I, open-label, and multicenter study (NCT01513902) in 2017, tofacitinib was well tolerated in patients with JIA [196]. Tofacitinib has been shown to be effective in the treatment of refractory JIA in case reports [197]. Pfizer posted that the phase III clinical trial (NCT02592434) for tofacitinib was completed in 2019 with good results, supporting the use of tofacitinib in patients with JIA. Clinical studies involving baricitinib (NCT4088396; NCT03773965; NCT03773978) and upadacitinib (NCT03725007) in patients with JIA are still in the recruiting stage, and there are no results yet.
Juvenile Dermatomyositis
Juvenile dermatomyositis (JDM) is a rare pediatric autoimmune disease with an average age at disease onset of 7.5 years. JDM is characterized by proximal muscle weakness and pathognomonic skin rashes [198]. Overexpressed type I IFN-inducible transcripts and activated type I IFN signatures are observed in patients with DM [13]. Similarly, serum IFN-α activity is increased for a short time in newly diagnosed patients with JDM and is associated with high serum muscle–derived enzymes [199]. The expression of IFN-α/β-inducible genes in muscle biopsy and proteins induced by type I IFNs such as myxovirus resistance protein A (MxA) in PBMC, affected muscle and skin, are elevated in patients with JDM [200,201,202]. TLR3 was also found to be elevated in vascular endothelial cells, myeloid DCs (mDCs), and regenerating myofibers in patients with JDM [203]. Type I IFN signatures are increased in B cells of JDM, and the activation of TLR7 and IFN-α may lead to the expanded immature transitional B cell population [204]. In addition, in vivo and in vitro studies have shown that myogenic precursor cells are also a source of IFN-Is [205].
The JAK inhibitors, ruxolitinib and tofacitinib, have been reported to be effective in the treatment of adult DM [206, 207]. In addition, a case report showed that refractory patients with JDM responded to the JAK1/JAK2 inhibitor, baricitinib [208]. A phase 1b clinical study (NCT00533091) observed that the anti-IFN-α mAb, sifalimumab inhibited type I IFN signatures in patients with DM, but there is as yet no pediatric data [209]. A clinical trial (NCT02612857) of the TLR 7, 8, 9 inhibitor, IMO-8400, has been completed in patients with DM, but again, no pediatric data is available. Clinical trials on the treatment of JDM by inhibiting type I IFN signatures are sorely needed.
Sjögren Syndrome
Primary Sjögren syndrome (pSjS) is an inflammatory autoimmune disorder with the loss of secretory function in exocrine glands, causing dryness of the mucosal surfaces, especially in the mouth (salivary) and eyes (lacrimal glands) [210]. IFN-α expression levels at the transcript or protein levels have been found to be increased in the labial salivary glands and blood of patients with pSjS [210, 211]. In addition, a positive correlation has been identified between genes induced by IFN-Is and titers of anti-Ro/SSA, anti-La/SSB autoantibodies, EULAR SjS disease activity index (ESSDAI), biological markers of activity, and BAFF mRNA in monocytes of patients with SjS, demonstrating a key role of IFN-Is in pSjS [210, 212]. pDCs, the main source of IFN-Is, have been found in the salivary gland of patients with pSjS, whereas they are decreased in the peripheral blood [210, 212,213,214]. This phenomenon also exists in SLE, and both may be caused by the re-translocation of pDCs from peripheral blood to diseased tissue [210, 212].
A study (NCT03100942) involving the JAK1 inhibitor filgotinib, Syk inhibitor lanraplenib, and BTK inhibitor tirabrutinib has been completed, but results are pending [8]. The JAK inhibitor filgotinib-treated mouse models with SjS showed significantly increased salivary flow rates and remarkable reductions in lymphocytic infiltration of the salivary gland and IFN-I signatures [215]. The JAK1/2 inhibitor AG490 and ruxolitinib and the TBK1 inhibitor BX795 also exhibited good responses in animal models with SjS [216, 217]. On the other hand, hydroxychloroquine, an inhibitor of TLRs, decreased IFN-I signatures but did not improve the symptoms of SjS in a clinical trial (NCT00632866) [218]. Moreover, IFN-α administered via the oromucosal route in patients with pSjS was demonstrated to be effective and safe in the improvement of salivary output in a phase II clinical trial [219]. However, another trial of 497 patients with pSjS with the treatment of either IFN-α or placebo failed to meet the primary endpoints [219]. Therefore, more evidence is needed to further understand the role of IFN-Is in the pathogenesis and treatment of SjS.
Systemic Sclerosis
Systemic sclerosis (SSc), also called scleroderma, is characterized by fibrosis and dysfunction of the skin, internal organs, and a vasculopathy [8]. Case reports have shown that SSc occurs in patients after using IFN-α to treat chronic myelogenous leukemia or IFN-β to treat hepatitis C [220]. IFN signatures can also be found in peripheral blood and affected skin of patients with SSc, even during the early phase of disease [221]. In addition, higher IFN signatures are positively correlated with anti-topoisomerase antibodies or anti-U1-RNP antibodies but negatively correlated with anti-RNA polymerase III antibodies in patients with SSc [220, 222]. The level of type I IFNs is also related to severe symptoms in the skin, lung, and skeletal muscle of patients with SSc [223]. Moreover, disease activity can be suppressed after treatment with the anti-IFNAR mAb anifrolumab in patients with SSc, demonstrating an important role of type I IFNs in the pathogenesis of SSc [224].
A phase 1, multicenter, open-label study (NCT00930683) in which patients with SSc were treated with the anti-IFN-α mAb anifrolumab has shown good results and supported its further clinical development [225]. pDCs can directly promote the development of skin fibrosis, and depleting pDCs can prevent the progression of the fibrotic process in mouse models of SSc [226]. Clinical trials (NCT03817424) of VIB7734, an inhibitor of pDCs, in patients with SSc are still recruiting, and another clinical study (NCT04206644) of JAK inhibitors is also ongoing.
Perspectives
Autoimmune diseases consist of a wide range of systemic diseases that affect up to 10% of the world’s population [227]. Not only are the symptoms potentially disabling, but the treatments are often not much better, with many associated with significant side effects [4,5,6,7]. The type I IFN signaling pathway is of great significance in antiviral and antitumor immunity [15, 17]. In addition, as mentioned above, the type I IFN signaling pathway is essential for the initiation and development of autoimmune diseases [15, 17]. Several inhibitors blocking specific molecules in the type I IFN signaling pathway, such as tofacitinib and baricitinib for the treatment of RA, are licensed by the FDA/EMA, and more are currently undergoing clinical trials [19]. There are encouraging signs that many targeted inhibitors have completed or entered phase III clinical trials and seem to be very promising, including the anti-IFNAR1 mAb anifrolumab for SLE [20, 168]; JAK1/JAK2 inhibitor baricitinib for SLE [21]; and the JAK1 inhibitors upadacitinib and filgotinib for RA [181, 182]. Tofacitinib has completed a phase III clinical trial in patients with JIA and shown positive results [198]. Other studies are still not completed so results are unavailable. More data and evidence are needed to support targeted inhibitors that have not entered phase III clinical trials but which have shown good responses in case reports or animal models.
The role of IFN-Is in the pathogenesis of autoimmune diseases is still unclear. Under most circumstances, IFN-Is are proinflammatory, but type I IFNs also seem to have a protective function in RA and MS based on some studies [189, 228]. Some have proposed that conflicting results may be a result of the varying immunomodulatory and proinflammatory functions of interacting cytokines in autoimmune disorders that are predominately driven by Th1 cells or Th17 cells [229]. Furthermore, the results of anifrolumab in the phase III TULIP-1 and TULIP-2 trials are partially inconsistent [20, 168]. The end point BICLA, but not SRI, was met in TULIP-1, but both endpoints were met in TULIP-2 [20, 168]. Heterogeneity of the disease state and genetics may be one of the reasons that different types of patients with SLE may have varying responses to specific therapies. The establishment of subtypes and personalized approaches based on immune genotypes is needed for solving disease heterogeneity and efficiently achieving better targeted therapies [6]. In addition, the anti-IFN-α mAbs rontalizumab and AGS-009 did not achieve the expected results as anti-IFNAR mAbs, which may be caused by the 13 different subtypes of IFN-α in humans, making it difficult to block all types of IFN-α [160]. Besides, IFN-β may continue to play a role in disease progression even in the presence of anti-IFN-α therapies.
Overall, type I IFN signaling pathway is of great significance in the pathogenesis and treatment of both adult and pediatric autoimmune diseases. Many inhibitors targeting the type I IFN signaling pathway have shown positive results in clinical trials of autoimmune diseases. Thus, it is very important to further explore the role of the type I IFN signaling pathway in autoimmune diseases to develop more effective and safe therapies.
Abbreviations
- AGS:
-
Aicardi-Goutières syndrome
- BTK:
-
Bruton’s tyrosine kinase
- cGAS:
-
Cyclic GMP-AMP synthase
- IFN-I:
-
Type I interferon
- IRAK:
-
Interleukin-1 receptor-associated kinase
- IRF:
-
Interferon regulatory family
- ISG:
-
Interferon-stimulated gene
- JAK:
-
Janus kinase
- JDM:
-
Juvenile dermatomyositis
- JIA:
-
Juvenile idiopathic arthritis
- MAVS:
-
Mitochondrial antiviral signaling protein
- MDA:
-
Melanoma differentiation-associated gene
- MyD:
-
Myeloid differentiation primary response gene
- pDC:
-
Plasmacytoid dendritic cell
- PRR:
-
Pattern recognition receptor
- pSjS:
-
primary Sjögren syndrome
- pSLE:
-
Pediatric systemic lupus erythematosus
- RA:
-
Rheumatoid arthritis
- RIG-I:
-
Retinoic acid-inducible gene-I
- SSc:
-
Systemic sclerosis
- STAT:
-
Signal transducers and activators of transcription
- STING:
-
Stimulator of interferon
- Syk:
-
Spleen tyrosine kinase
- TBK:
-
TANK-binding kinase
- TLR:
-
Toll-like receptor
- TRAF:
-
Tumor necrosis factor receptor-associated factor
- TREX:
-
Three prime repair exonuclease
- TRIF:
-
Toll-IL-1 receptor domain-containing adaptor inducing IFN-β
- TYK:
-
Tyrosine kinase
References
Wahren-Herlenius M, Dorner T (2013) Immunopathogenic mechanisms of systemic autoimmune disease. Lancet (London, England) 382(9894):819–831. https://doi.org/10.1016/S0140-6736(13)60954-X
Yan N, Chen ZJ (2012) Intrinsic antiviral immunity. Nat Immunol 13(3):214–222. https://doi.org/10.1038/ni.2229
Abramson J, Husebye ES (2016) Autoimmune regulator and self-tolerance - molecular and clinical aspects. Immunol Rev 271(1):127–140. https://doi.org/10.1111/imr.12419
Seibel MJ, Cooper MS, Zhou H (2013) Glucocorticoid-induced osteoporosis: mechanisms, management, and future perspectives. Lancet Diabetes Endocrinol 1(1):59–70. https://doi.org/10.1016/S2213-8587(13)70045-7
Bakshi J, Segura BT, Wincup C, Rahman A (2018) Unmet needs in the pathogenesis and treatment of systemic lupus erythematosus. Clin Rev Allergy Immunol 55(3):352–367. https://doi.org/10.1007/s12016-017-8640-5
Davis LS, Reimold AM (2017) Research and therapeutics-traditional and emerging therapies in systemic lupus erythematosus. Rheumatology (Oxford, England) 56(suppl_1):i100–i113. https://doi.org/10.1093/rheumatology/kew417
Cheung TT, McInnes IB (2017) Future therapeutic targets in rheumatoid arthritis? Semin Immunopathol 39(4):487–500. https://doi.org/10.1007/s00281-017-0623-3
Mavragani CP, Moutsopoulos HM (2019) Sjogren’s syndrome: old and new therapeutic targets J autoimmun:102364. https://doi.org/10.1016/j.jaut.2019.102364
Hooks JJ, Moutsopoulos HM, Geis SA, Stahl NI, Decker JL, Notkins AL (1979) Immune interferon in the circulation of patients with autoimmune disease. N Engl J Med 301(1):5–8. https://doi.org/10.1056/NEJM197907053010102
Prummel MF, Laurberg P (2003) Interferon-alpha and autoimmune thyroid disease. Thyroid 13(6):547–551. https://doi.org/10.1089/105072503322238809
Ronnblom LE, Alm GV, Oberg KE (1990) Possible induction of systemic lupus erythematosus by interferon-alpha treatment in a patient with a malignant carcinoid tumour. J Intern Med 227(3):207–210. https://doi.org/10.1111/j.1365-2796.1990.tb00144.x
Barrat FJ, Crow MK, Ivashkiv LB (2019) Interferon target-gene expression and epigenomic signatures in health and disease. Nat Immunol 20(12):1574–1583. https://doi.org/10.1038/s41590-019-0466-2
Higgs BW, Liu Z, White B, Zhu W, White WI, Morehouse C, Brohawn P, Kiener PA, Richman L, Fiorentino D, Greenberg SA, Jallal B, Yao Y (2011) Patients with systemic lupus erythematosus, myositis, rheumatoid arthritis and scleroderma share activation of a common type I interferon pathway. Ann Rheum Dis 70(11):2029–2036. https://doi.org/10.1136/ard.2011.150326
Negishi H, Taniguchi T, Yanai H (2018) The interferon (IFN) class of cytokines and the IFN regulatory factor (IRF) transcription factor family. Cold Spring Harb Perspect Biol 10(11). https://doi.org/10.1101/cshperspect.a028423
Lazear HM, Schoggins JW, Diamond MS (2019) Shared and distinct functions of type I and type III interferons. Immunity 50(4):907–923. https://doi.org/10.1016/j.immuni.2019.03.025
Hillion S, Arleevskaya MI, Blanco P, Bordron A, Brooks WH, Cesbron JY, Kaveri S, Vivier E, Renaudineau Y (2020) The innate part of the adaptive immune system. Clin Rev Allergy Immunol 58(2):151–154. https://doi.org/10.1007/s12016-019-08740-1
Ivashkiv LB, Donlin LT (2014) Regulation of type I interferon responses. Nat Rev Immunol 14(1):36–49. https://doi.org/10.1038/nri3581
Schreiber G (2017) The molecular basis for differential type I interferon signaling. J Biol Chem 292(18):7285–7294. https://doi.org/10.1074/jbc.R116.774562
Taylor PC (2019) Clinical efficacy of launched JAK inhibitors in rheumatoid arthritis. Rheumatology (Oxford, England) 58(Suppl 1):i17–i26. https://doi.org/10.1093/rheumatology/key225
Morand EF, Furie R, Tanaka Y, Bruce IN, Askanase AD, Richez C, Bae SC, Brohawn PZ, Pineda L, Berglind A, Tummala R, Investigators T-T (2020) Trial of anifrolumab in active systemic lupus erythematosus. N Engl J Med 382(3):211–221. https://doi.org/10.1056/NEJMoa1912196
Kubo S, Nakayamada S, Tanaka Y (2019) Baricitinib for the treatment of rheumatoid arthritis and systemic lupus erythematosus: a 2019 update. Expert Rev Clin Immunol 15(7):693–700. https://doi.org/10.1080/1744666X.2019.1608821
Pestka S, Krause CD, Walter MR (2004) Interferons, interferon-like cytokines, and their receptors. Immunol Rev 202:8–32. https://doi.org/10.1111/j.0105-2896.2004.00204.x
Kato Y, Park J, Takamatsu H, Konaka H, Aoki W, Aburaya S, Ueda M, Nishide M, Koyama S, Hayama Y, Kinehara Y, Hirano T, Shima Y, Narazaki M, Kumanogoh A (2018) Apoptosis-derived membrane vesicles drive the cGAS-STING pathway and enhance type I IFN production in systemic lupus erythematosus. Ann Rheum Dis 77(10):1507–1515. https://doi.org/10.1136/annrheumdis-2018-212988
Baccala R, Hoebe K, Kono DH, Beutler B, Theofilopoulos AN (2007) TLR-dependent and TLR-independent pathways of type I interferon induction in systemic autoimmunity. Nat Med 13(5):543–551. https://doi.org/10.1038/nm1590
Lou H, Pickering MC (2018) Extracellular DNA and autoimmune diseases. Cell Mol Immunol 15(8):746–755. https://doi.org/10.1038/cmi.2017.136
Raftery N, Stevenson NJ (2017) Advances in anti-viral immune defence: revealing the importance of the IFN JAK/STAT pathway. Cell Mol Life Sci 74(14):2525–2535. https://doi.org/10.1007/s00018-017-2520-2
Lester SN, Li K (2014) Toll-like receptors in antiviral innate immunity. J Mol Biol 426(6):1246–1264. https://doi.org/10.1016/j.jmb.2013.11.024
Lee CC, Avalos AM, Ploegh HL (2012) Accessory molecules for toll-like receptors and their function. Nat Rev Immunol 12(3):168–179. https://doi.org/10.1038/nri3151
Zevini A, Olagnier D, Hiscott J (2017) Crosstalk between cytoplasmic RIG-I and STING sensing pathways. Trends Immunol 38(3):194–205. https://doi.org/10.1016/j.it.2016.12.004
Briard B, Place DE, Kanneganti TD (2020) DNA sensing in the innate immune response. Physiology (Bethesda) 35(2):112–124. https://doi.org/10.1152/physiol.00022.2019
Jonsson KL, Laustsen A, Krapp C, Skipper KA, Thavachelvam K, Hotter D, Egedal JH, Kjolby M, Mohammadi P, Prabakaran T, Sorensen LK, Sun C, Jensen SB, Holm CK, Lebbink RJ, Johannsen M, Nyegaard M, Mikkelsen JG, Kirchhoff F, Paludan SR, Jakobsen MR (2017) IFI16 is required for DNA sensing in human macrophages by promoting production and function of cGAMP. Nat Commun 8:14391. https://doi.org/10.1038/ncomms14391
Arleevskaya MI, Larionova RV, Brooks WH, Bettacchioli E, Renaudineau Y (2020) Toll-like receptors, infections, and rheumatoid arthritis. Clin Rev Allergy Immunol 58(2):172–181. https://doi.org/10.1007/s12016-019-08742-z
Brisse M, Ly H (2019) Comparative structure and function analysis of the RIG-I-like receptors: RIG-I and MDA5. Front Immunol 10:1586. https://doi.org/10.3389/fimmu.2019.01586
O'Neill LA, Bowie AG (2007) The family of five: TIR-domain-containing adaptors in toll-like receptor signalling. Nat Rev Immunol 7(5):353–364. https://doi.org/10.1038/nri2079
Cingoz O, Goff SP (2018) Cyclin-dependent kinase activity is required for type I interferon production. Proc Natl Acad Sci U S A 115(13):E2950–E2959. https://doi.org/10.1073/pnas.1720431115
Dias Junior AG, Sampaio NG, Rehwinkel J (2019) A balancing act: MDA5 in antiviral immunity and autoinflammation. Trends Microbiol 27(1):75–85. https://doi.org/10.1016/j.tim.2018.08.007
Yoneyama M, Fujita T (2009) RNA recognition and signal transduction by RIG-I-like receptors. Immunol Rev 227(1):54–65. https://doi.org/10.1111/j.1600-065X.2008.00727.x
Taniguchi T, Ogasawara K, Takaoka A, Tanaka N (2001) IRF family of transcription factors as regulators of host defense. Annu Rev Immunol 19:623–655. https://doi.org/10.1146/annurev.immunol.19.1.623
Tailor P, Tamura T, Ozato K (2006) IRF family proteins and type I interferon induction in dendritic cells. Cell Res 16(2):134–140. https://doi.org/10.1038/sj.cr.7310018
Li P, Wong JJ, Sum C, Sin WX, Ng KQ, Koh MB, Chin KC (2011) IRF8 and IRF3 cooperatively regulate rapid interferon-beta induction in human blood monocytes. Blood 117(10):2847–2854. https://doi.org/10.1182/blood-2010-07-294272
Mancuso G, Gambuzza M, Midiri A, Biondo C, Papasergi S, Akira S, Teti G, Beninati C (2009) Bacterial recognition by TLR7 in the lysosomes of conventional dendritic cells. Nat Immunol 10(6):587–594. https://doi.org/10.1038/ni.1733
Chen K, Liu J, Cao X (2017) Regulation of type I interferon signaling in immunity and inflammation: a comprehensive review. J Autoimmun 83:1–11. https://doi.org/10.1016/j.jaut.2017.03.008
Das A, Dinh PX, Panda D, Pattnaik AK (2014) Interferon-inducible protein IFI35 negatively regulates RIG-I antiviral signaling and supports vesicular stomatitis virus replication. J Virol 88(6):3103–3113. https://doi.org/10.1128/JVI.03202-13
Arimoto K, Takahashi H, Hishiki T, Konishi H, Fujita T, Shimotohno K (2007) Negative regulation of the RIG-I signaling by the ubiquitin ligase RNF125. Proc Natl Acad Sci U S A 104(18):7500–7505. https://doi.org/10.1073/pnas.0611551104
Cui J, Zhu L, Xia X, Wang HY, Legras X, Hong J, Ji J, Shen P, Zheng S, Chen ZJ, Wang RF (2010) NLRC5 negatively regulates the NF-kappaB and type I interferon signaling pathways. Cell 141(3):483–496. https://doi.org/10.1016/j.cell.2010.03.040
Catrysse L, Vereecke L, Beyaert R, van Loo G (2014) A20 in inflammation and autoimmunity. Trends Immunol 35(1):22–31. https://doi.org/10.1016/j.it.2013.10.005
Li X, Fu Z, Liang H, Wang Y, Qi X, Ding M, Sun X, Zhou Z, Huang Y, Gu H, Li L, Chen X, Li D, Zhao Q, Liu F, Wang H, Wang J, Zen K, Zhang CY (2018) H5N1 influenza virus-specific miRNA-like small RNA increases cytokine production and mouse mortality via targeting poly(rC)-binding protein 2. Cell Res 28(2):157–171. https://doi.org/10.1038/cr.2018.3
Maelfait J, Beyaert R (2012) Emerging role of ubiquitination in antiviral RIG-I signaling. Microbiol Mol Biol Rev 76(1):33–45. https://doi.org/10.1128/MMBR.05012-11
Zhang M, Wang L, Zhao X, Zhao K, Meng H, Zhao W, Gao C (2012) TRAF-interacting protein (TRIP) negatively regulates IFN-beta production and antiviral response by promoting proteasomal degradation of TANK-binding kinase 1. J Exp Med 209(10):1703–1711. https://doi.org/10.1084/jem.20120024
van Gent M, Sparrer KMJ, Gack MU (2018) TRIM proteins and their roles in antiviral host defenses. Annu Rev Virol 5(1):385–405. https://doi.org/10.1146/annurev-virology-092917-043323
Ozato K, Shin DM, Chang TH, Morse HC 3rd (2008) TRIM family proteins and their emerging roles in innate immunity. Nat Rev Immunol 8(11):849–860. https://doi.org/10.1038/nri2413
Hu Y, Wang J, Yang B, Zheng N, Qin M, Ji Y, Lin G, Tian L, Wu X, Wu L, Sun B (2011) Guanylate binding protein 4 negatively regulates virus-induced type I IFN and antiviral response by targeting IFN regulatory factor 7. J Immunol 187(12):6456–6462. https://doi.org/10.4049/jimmunol.1003691
Liu B, Mink S, Wong KA, Stein N, Getman C, Dempsey PW, Wu H, Shuai K (2004) PIAS1 selectively inhibits interferon-inducible genes and is important in innate immunity. Nat Immunol 5(9):891–898. https://doi.org/10.1038/ni1104
Kubota T, Matsuoka M, Xu S, Otsuki N, Takeda M, Kato A, Ozato K (2011) PIASy inhibits virus-induced and interferon-stimulated transcription through distinct mechanisms. J Biol Chem 286(10):8165–8175. https://doi.org/10.1074/jbc.M110.195255
Li Y, Li C, Xue P, Zhong B, Mao AP, Ran Y, Chen H, Wang YY, Yang F, Shu HB (2009) ISG56 is a negative-feedback regulator of virus-triggered signaling and cellular antiviral response. Proc Natl Acad Sci U S A 106(19):7945–7950. https://doi.org/10.1073/pnas.0900818106
John SP, Sun J, Carlson RJ, Cao B, Bradfield CJ, Song J, Smelkinson M, Fraser IDC (2018) IFIT1 exerts opposing regulatory effects on the inflammatory and interferon gene programs in LPS-activated human macrophages. Cell Rep 25(1):95–106 e106. https://doi.org/10.1016/j.celrep.2018.09.002
Wang J, Yang B, Hu Y, Zheng Y, Zhou H, Wang Y, Ma Y, Mao K, Yang L, Lin G, Ji Y, Wu X, Sun B (2013) Negative regulation of Nmi on virus-triggered type I IFN production by targeting IRF7. J Immunol 191(6):3393–3399. https://doi.org/10.4049/jimmunol.1300740
Lee MS, Kim B, Oh GT, Kim YJ (2013) OASL1 inhibits translation of the type I interferon-regulating transcription factor IRF7. Nat Immunol 14(4):346–355. https://doi.org/10.1038/ni.2535
Ma Z, Moore R, Xu X, Barber GN (2013) DDX24 negatively regulates cytosolic RNA-mediated innate immune signaling. PLoS Pathog 9(10):e1003721. https://doi.org/10.1371/journal.ppat.1003721
Li J, Ding SC, Cho H, Chung BC, Gale M Jr, Chanda SK, Diamond MS (2013) A short hairpin RNA screen of interferon-stimulated genes identifies a novel negative regulator of the cellular antiviral response. mBio 4(3):e00385-00313. https://doi.org/10.1128/mBio.00385-13
Yu Y, Hayward GS (2010) The ubiquitin E3 ligase RAUL negatively regulates type i interferon through ubiquitination of the transcription factors IRF7 and IRF3. Immunity 33(6):863–877. https://doi.org/10.1016/j.immuni.2010.11.027
Sugiyama Y, Kakoi K, Kimura A, Takada I, Kashiwagi I, Wakabayashi Y, Morita R, Nomura M, Yoshimura A (2012) Smad2 and Smad3 are redundantly essential for the suppression of iNOS synthesis in macrophages by regulating IRF3 and STAT1 pathways. Int Immunol 24(4):253–265. https://doi.org/10.1093/intimm/dxr126
Zheng H, Qian J, Varghese B, Baker DP, Fuchs S (2011) Ligand-stimulated downregulation of the alpha interferon receptor: role of protein kinase D2. Mol Cell Biol 31(4):710–720. https://doi.org/10.1128/MCB.01154-10
Arimoto KI, Lochte S, Stoner SA, Burkart C, Zhang Y, Miyauchi S, Wilmes S, Fan JB, Heinisch JJ, Li Z, Yan M, Pellegrini S, Colland F, Piehler J, Zhang DE (2017) STAT2 is an essential adaptor in USP18-mediated suppression of type I interferon signaling. Nat Struct Mol Biol 24(3):279–289. https://doi.org/10.1038/nsmb.3378
Yasukawa H, Sasaki A, Yoshimura A (2000) Negative regulation of cytokine signaling pathways. Annu Rev Immunol 18:143–164. https://doi.org/10.1146/annurev.immunol.18.1.143
An H, Hou J, Zhou J, Zhao W, Xu H, Zheng Y, Yu Y, Liu S, Cao X (2008) Phosphatase SHP-1 promotes TLR- and RIG-I-activated production of type I interferon by inhibiting the kinase IRAK1. Nat Immunol 9(5):542–550. https://doi.org/10.1038/ni.1604
An H, Zhao W, Hou J, Zhang Y, Xie Y, Zheng Y, Xu H, Qian C, Zhou J, Yu Y, Liu S, Feng G, Cao X (2006) SHP-2 phosphatase negatively regulates the TRIF adaptor protein-dependent type I interferon and proinflammatory cytokine production. Immunity 25(6):919–928. https://doi.org/10.1016/j.immuni.2006.10.014
Bourdeau A, Dube N, Tremblay ML (2005) Cytoplasmic protein tyrosine phosphatases, regulation and function: the roles of PTP1B and TC-PTP. Curr Opin Cell Biol 17(2):203–209. https://doi.org/10.1016/j.ceb.2005.02.001
Alexander WS (2002) Suppressors of cytokine signalling (SOCS) in the immune system. Nat Rev Immunol 2(6):410–416. https://doi.org/10.1038/nri818
Nair S, Bist P, Dikshit N, Krishnan MN (2016) Global functional profiling of human ubiquitome identifies E3 ubiquitin ligase DCST1 as a novel negative regulator of type-I interferon signaling. Sci Rep 6:36179. https://doi.org/10.1038/srep36179
Jeronimo C, Bastian PJ, Bjartell A, Carbone GM, Catto JW, Clark SJ, Henrique R, Nelson WG, Shariat SF (2011) Epigenetics in prostate cancer: biologic and clinical relevance. Eur Urol 60(4):753–766. https://doi.org/10.1016/j.eururo.2011.06.035
Szyf M (2003) DNA methylation and cancer therapy. Drug Resist Updat 6(6):341–353. https://doi.org/10.1016/j.drup.2003.10.002
Imgenberg-Kreuz J, Carlsson Almlof J, Leonard D, Alexsson A, Nordmark G, Eloranta ML, Rantapaa-Dahlqvist S, Bengtsson AA, Jonsen A, Padyukov L, Gunnarsson I, Svenungsson E, Sjowall C, Ronnblom L, Syvanen AC, Sandling JK (2018) DNA methylation mapping identifies gene regulatory effects in patients with systemic lupus erythematosus. Ann Rheum Dis 77(5):736–743. https://doi.org/10.1136/annrheumdis-2017-212379
Zhu H, Wu LF, Mo XB, Lu X, Tang H, Zhu XW, Xia W, Guo YF, Wang MJ, Zeng KQ, Wu J, Qiu YH, Lin X, Zhang YH, Liu YZ, Yi NJ, Deng FY, Lei SF (2019) Rheumatoid arthritis-associated DNA methylation sites in peripheral blood mononuclear cells. Ann Rheum Dis 78(1):36–42. https://doi.org/10.1136/annrheumdis-2018-213970
Imgenberg-Kreuz J, Sandling JK, Almlof JC, Nordlund J, Signer L, Norheim KB, Omdal R, Ronnblom L, Eloranta ML, Syvanen AC, Nordmark G (2016) Genome-wide DNA methylation analysis in multiple tissues in primary Sjogren’s syndrome reveals regulatory effects at interferon-induced genes. Ann Rheum Dis 75(11):2029–2036. https://doi.org/10.1136/annrheumdis-2015-208659
Ding W, Pu W, Wang L, Jiang S, Zhou X, Tu W, Yu L, Zhang J, Guo S, Liu Q, Ma Y, Chen S, Wu W, Reveille J, Zou H, Jin L, Wang J (2018) Genome-wide DNA methylation analysis in systemic sclerosis reveals hypomethylation of IFN-associated genes in CD4(+) and CD8(+) T cells. J Investig Dermatol 138(5):1069–1077. https://doi.org/10.1016/j.jid.2017.12.003
Chiappinelli KB, Strissel PL, Desrichard A, Li H, Henke C, Akman B, Hein A, Rote NS, Cope LM, Snyder A, Makarov V, Budhu S, Slamon DJ, Wolchok JD, Pardoll DM, Beckmann MW, Zahnow CA, Merghoub T, Chan TA, Baylin SB, Strick R (2015) Inhibiting DNA methylation causes an interferon response in cancer via dsRNA including endogenous retroviruses. Cell 162(5):974–986. https://doi.org/10.1016/j.cell.2015.07.011
Shilatifard A (2008) Molecular implementation and physiological roles for histone H3 lysine 4 (H3K4) methylation. Curr Opin Cell Biol 20(3):341–348. https://doi.org/10.1016/j.ceb.2008.03.019
Fang TC, Schaefer U, Mecklenbrauker I, Stienen A, Dewell S, Chen MS, Rioja I, Parravicini V, Prinjha RK, Chandwani R, MacDonald MR, Lee K, Rice CM, Tarakhovsky A (2012) Histone H3 lysine 9 di-methylation as an epigenetic signature of the interferon response. J Exp Med 209(4):661–669. https://doi.org/10.1084/jem.20112343
Chen K, Liu J, Liu S, Xia M, Zhang X, Han D, Jiang Y, Wang C, Cao X (2017) Methyltransferase SETD2-mediated methylation of STAT1 is critical for interferon antiviral activity. Cell 170(3):492–506 e414. https://doi.org/10.1016/j.cell.2017.06.042
Chen X, Liu X, Zhang Y, Huai W, Zhou Q, Xu S, Chen X, Li N, Cao X (2020) Methyltransferase Dot1l preferentially promotes innate IL-6 and IFN-beta production by mediating H3K79me2/3 methylation in macrophages. Cell Mol Immunol 17(1):76–84. https://doi.org/10.1038/s41423-018-0170-4
Munshi N, Agalioti T, Lomvardas S, Merika M, Chen G, Thanos D (2001) Coordination of a transcriptional switch by HMGI(Y) acetylation. Science 293(5532):1133–1136. https://doi.org/10.1126/science.293.5532.1133
Agalioti T, Lomvardas S, Parekh B, Yie J, Maniatis T, Thanos D (2000) Ordered recruitment of chromatin modifying and general transcription factors to the IFN-beta promoter. Cell 103(4):667–678. https://doi.org/10.1016/s0092-8674(00)00169-0
Meng J, Liu X, Zhang P, Li D, Xu S, Zhou Q, Guo M, Huai W, Chen X, Wang Q, Li N, Cao X (2016) Rb selectively inhibits innate IFN-beta production by enhancing deacetylation of IFN-beta promoter through HDAC1 and HDAC8. J Autoimmun 73:42–53. https://doi.org/10.1016/j.jaut.2016.05.012
Marie IJ, Chang HM, Levy DE (2018) HDAC stimulates gene expression through BRD4 availability in response to IFN and in interferonopathies. J Exp Med 215(12):3194–3212. https://doi.org/10.1084/jem.20180520
Decque A, Joffre O, Magalhaes JG, Cossec JC, Blecher-Gonen R, Lapaquette P, Silvin A, Manel N, Joubert PE, Seeler JS, Albert ML, Amit I, Amigorena S, Dejean A (2016) Sumoylation coordinates the repression of inflammatory and anti-viral gene-expression programs during innate sensing. Nat Immunol 17(2):140–149. https://doi.org/10.1038/ni.3342
Zhang Y, Mao D, Roswit WT, Jin X, Patel AC, Patel DA, Agapov E, Wang Z, Tidwell RM, Atkinson JJ, Huang G, McCarthy R, Yu J, Yun NE, Paessler S, Lawson TG, Omattage NS, Brett TJ, Holtzman MJ (2015) PARP9-DTX3L ubiquitin ligase targets host histone H2BJ and viral 3C protease to enhance interferon signaling and control viral infection. Nat Immunol 16(12):1215–1227. https://doi.org/10.1038/ni.3279
Fonseca GJ, Thillainadesan G, Yousef AF, Ablack JN, Mossman KL, Torchia J, Mymryk JS (2012) Adenovirus evasion of interferon-mediated innate immunity by direct antagonism of a cellular histone posttranslational modification. Cell Host Microbe 11(6):597–606. https://doi.org/10.1016/j.chom.2012.05.005
Li Y, Fan X, He X, Sun H, Zou Z, Yuan H, Xu H, Wang C, Shi X (2012) MicroRNA-466l inhibits antiviral innate immune response by targeting interferon-alpha. Cell Mol Immunol 9(6):497–502. https://doi.org/10.1038/cmi.2012.35
Han X, Wang Y, Zhang X, Qin Y, Qu B, Wu L, Ma J, Zhou Z, Qian J, Dai M, Tang Y, Chan EK, Harley JB, Zhou S, Shen N (2016) MicroRNA-130b ameliorates murine lupus nephritis through targeting the type I interferon pathway on renal mesangial cells. Arthritis Rheumatol (Hoboken, NJ) 68(9):2232–2243. https://doi.org/10.1002/art.39725
Rossato M, Affandi AJ, Thordardottir S, Wichers CGK, Cossu M, Broen JCA, Moret FM, Bossini-Castillo L, Chouri E, van Bon L, Wolters F, Marut W, van der Kroef M, Silva-Cardoso S, Bekker CPJ, Dolstra H, van Laar JM, Martin J, van Roon JAG, Reedquist KA, Beretta L, Radstake T (2017) Association of MicroRNA-618 expression with altered frequency and activation of plasmacytoid dendritic cells in patients with systemic sclerosis. Arthritis Rheumatol (Hoboken, NJ) 69(9):1891–1902. https://doi.org/10.1002/art.40163
Ho BC, Yu IS, Lu LF, Rudensky A, Chen HY, Tsai CW, Chang YL, Wu CT, Chang LY, Shih SR, Lin SW, Lee CN, Yang PC, Yu SL (2014) Inhibition of miR-146a prevents enterovirus-induced death by restoring the production of type I interferon. Nat Commun 5:3344. https://doi.org/10.1038/ncomms4344
Smith S, Fernando T, Wu PW, Seo J, Ni Gabhann J, Piskareva O, McCarthy E, Howard D, O'Connell P, Conway R, Gallagher P, Molloy E, Stallings RL, Kearns G, Forbess L, Ishimori M, Venuturupalli S, Wallace D, Weisman M, Jefferies CA (2017) MicroRNA-302d targets IRF9 to regulate the IFN-induced gene expression in SLE. J Autoimmun 79:105–111. https://doi.org/10.1016/j.jaut.2017.03.003
Yoon WH, Meinhardt H, Montell DJ (2011) miRNA-mediated feedback inhibition of JAK/STAT morphogen signalling establishes a cell fate threshold. Nat Cell Biol 13(9):1062–1069. https://doi.org/10.1038/ncb2316
Kohanbash G, Okada H (2012) MicroRNAs and STAT interplay. Semin Cancer Biol 22(1):70–75. https://doi.org/10.1016/j.semcancer.2011.12.010
Gracias DT, Stelekati E, Hope JL, Boesteanu AC, Doering TA, Norton J, Mueller YM, Fraietta JA, Wherry EJ, Turner M, Katsikis PD (2013) The microRNA miR-155 controls CD8(+) T cell responses by regulating interferon signaling. Nat Immunol 14(6):593–602. https://doi.org/10.1038/ni.2576
Navarro A, Pairet S, Alvarez-Larran A, Pons A, Ferrer G, Longaron R, Fernandez-Rodriguez C, Camacho L, Monzo M, Besses C, Bellosillo B (2016) miR-203 and miR-221 regulate SOCS1 and SOCS3 in essential thrombocythemia. Blood. Cancer J 6:e406. https://doi.org/10.1038/bcj.2016.10
Chen Y, Chen J, Wang H, Shi J, Wu K, Liu S, Liu Y, Wu J (2013) HCV-induced miR-21 contributes to evasion of host immune system by targeting MyD88 and IRAK1. PLoS Pathog 9(4):e1003248. https://doi.org/10.1371/journal.ppat.1003248
Nishitsuji H, Ujino S, Yoshio S, Sugiyama M, Mizokami M, Kanto T, Shimotohno K (2016) Long noncoding RNA #32 contributes to antiviral responses by controlling interferon-stimulated gene expression. Proc Natl Acad Sci U S A 113(37):10388–10393. https://doi.org/10.1073/pnas.1525022113
Zhou Y, Li M, Xue Y, Li Z, Wen W, Liu X, Ma Y, Zhang L, Shen Z, Cao X (2019) Interferon-inducible cytoplasmic lncLrrc55-AS promotes antiviral innate responses by strengthening IRF3 phosphorylation. Cell Res 29(8):641–654. https://doi.org/10.1038/s41422-019-0193-0
Kambara H, Niazi F, Kostadinova L, Moonka DK, Siegel CT, Post AB, Carnero E, Barriocanal M, Fortes P, Anthony DD, Valadkhan S (2014) Negative regulation of the interferon response by an interferon-induced long non-coding RNA. Nucleic Acids Res 42(16):10668–10680. https://doi.org/10.1093/nar/gku713
Cui Y, Sheng Y, Zhang X (2013) Genetic susceptibility to SLE: recent progress from GWAS. J Autoimmun 41:25–33. https://doi.org/10.1016/j.jaut.2013.01.008
Deng Y, Tsao BP (2010) Genetic susceptibility to systemic lupus erythematosus in the genomic era. Nat Rev Rheumatol 6(12):683–692. https://doi.org/10.1038/nrrheum.2010.176
Ghodke-Puranik Y, Niewold TB (2015) Immunogenetics of systemic lupus erythematosus: a comprehensive review. J Autoimmun 64:125–136. https://doi.org/10.1016/j.jaut.2015.08.004
Delgado-Vega A, Sanchez E, Lofgren S, Castillejo-Lopez C, Alarcon-Riquelme ME (2010) Recent findings on genetics of systemic autoimmune diseases. Curr Opin Immunol 22(6):698–705. https://doi.org/10.1016/j.coi.2010.09.002
Zhai Y, Xu K, Leng RX, Cen H, Wang W, Zhu Y, Zhou M, Feng CC, Ye DQ (2013) Association of interleukin-1 receptor-associated kinase (IRAK1) gene polymorphisms (rs3027898, rs1059702) with systemic lupus erythematosus in a Chinese Han population. Inflamm Res 62(6):555–560. https://doi.org/10.1007/s00011-013-0607-2
Jacob CO, Zhu J, Armstrong DL, Yan M, Han J, Zhou XJ, Thomas JA, Reiff A, Myones BL, Ojwang JO, Kaufman KM, Klein-Gitelman M, McCurdy D, Wagner-Weiner L, Silverman E, Ziegler J, Kelly JA, Merrill JT, Harley JB, Ramsey-Goldman R, Vila LM, Bae SC, Vyse TJ, Gilkeson GS, Gaffney PM, Moser KL, Langefeld CD, Zidovetzki R, Mohan C (2009) Identification of IRAK1 as a risk gene with critical role in the pathogenesis of systemic lupus erythematosus. Proc Natl Acad Sci U S A 106(15):6256–6261. https://doi.org/10.1073/pnas.0901181106
Tyler DR, Persky ME, Matthews LA, Chan S, Farrar JD (2007) Pre-assembly of STAT4 with the human IFN-alpha/beta receptor-2 subunit is mediated by the STAT4 N-domain. Mol Immunol 44(8):1864–1872. https://doi.org/10.1016/j.molimm.2006.10.006
Hagberg N, Joelsson M, Leonard D, Reid S, Eloranta ML, Mo J, Nilsson MK, Syvanen AC, Bryceson YT, Ronnblom L (2018) The STAT4 SLE risk allele rs7574865[T] is associated with increased IL-12-induced IFN-gamma production in T cells from patients with SLE. Ann Rheum Dis 77(7):1070–1077. https://doi.org/10.1136/annrheumdis-2017-212794
Hagberg N, Ronnblom L (2019) Interferon-alpha enhances the IL-12-induced STAT4 activation selectively in carriers of the STAT4 SLE risk allele rs7574865[T]. Ann Rheum Dis 78(3):429–431. https://doi.org/10.1136/annrheumdis-2018-213836
Wang Y, Shaked I, Stanford SM, Zhou W, Curtsinger JM, Mikulski Z, Shaheen ZR, Cheng G, Sawatzke K, Campbell AM, Auger JL, Bilgic H, Shoyama FM, Schmeling DO, Balfour HH Jr, Hasegawa K, Chan AC, Corbett JA, Binstadt BA, Mescher MF, Ley K, Bottini N, Peterson EJ (2013) The autoimmunity-associated gene PTPN22 potentiates toll-like receptor-driven, type 1 interferon-dependent immunity. Immunity 39(1):111–122. https://doi.org/10.1016/j.immuni.2013.06.013
Chuang TH, Ulevitch RJ (2004) Triad3A, an E3 ubiquitin-protein ligase regulating toll-like receptors. Nat Immunol 5(5):495–502. https://doi.org/10.1038/ni1066
Yarwood A, Huizinga TW, Worthington J (2016) The genetics of rheumatoid arthritis: risk and protection in different stages of the evolution of RA. Rheumatology (Oxford, England) 55(2):199–209. https://doi.org/10.1093/rheumatology/keu323
Hinks A, Cobb J, Marion MC, Prahalad S, Sudman M, Bowes J, Martin P, Comeau ME, Sajuthi S, Andrews R, Brown M, Chen WM, Concannon P, Deloukas P, Edkins S, Eyre S, Gaffney PM, Guthery SL, Guthridge JM, Hunt SE, James JA, Keddache M, Moser KL, Nigrovic PA, Onengut-Gumuscu S, Onslow ML, Rose CD, Rich SS, Steel KJ, Wakeland EK, Wallace CA, Wedderburn LR, Woo P, Boston Children's JIAR, British Society of P, Adolescent Rheumatology Study G, Childhood Arthritis Prospective S, Childhood Arthritis Response to Medication S, German Society for Pediatric R, Study JIAGE, Registry NJG, Study T, United Kingdom Juvenile Idiopathic Arthritis Genetics C, Bohnsack JF, Haas JP, Glass DN, Langefeld CD, Thomson W, Thompson SD (2013) Dense genotyping of immune-related disease regions identifies 14 new susceptibility loci for juvenile idiopathic arthritis. Nat Genet 45(6):664–669. https://doi.org/10.1038/ng.2614
Nordang GB, Viken MK, Amundsen SS, Sanchez ES, Flato B, Forre OT, Martin J, Kvien TK, Lie BA (2012) Interferon regulatory factor 5 gene polymorphism confers risk to several rheumatic diseases and correlates with expression of alternative thymic transcripts. Rheumatology (Oxford, England) 51(4):619–626. https://doi.org/10.1093/rheumatology/ker364
Harris VM, Scofield RH, Sivils KL (2019) Genetics in Sjogren’s syndrome: where we are and where we go. Clin Exp Rheumatol 37 Suppl 118(3):234–239
Burbelo PD, Ambatipudi K, Alevizos I (2014) Genome-wide association studies in Sjogren’s syndrome: what do the genes tell us about disease pathogenesis? Autoimmun Rev 13(7):756–761. https://doi.org/10.1016/j.autrev.2014.02.002
Crow YJ (2015) Type I interferonopathies: mendelian type I interferon up-regulation. Curr Opin Immunol 32:7–12. https://doi.org/10.1016/j.coi.2014.10.005
Lee-Kirsch MA, Wolf C, Kretschmer S, Roers A (2015) Type I interferonopathies--an expanding disease spectrum of immunodysregulation. Semin Immunopathol 37(4):349–357. https://doi.org/10.1007/s00281-015-0500-x
Crow YJ, Hayward BE, Parmar R, Robins P, Leitch A, Ali M, Black DN, van Bokhoven H, Brunner HG, Hamel BC, Corry PC, Cowan FM, Frints SG, Klepper J, Livingston JH, Lynch SA, Massey RF, Meritet JF, Michaud JL, Ponsot G, Voit T, Lebon P, Bonthron DT, Jackson AP, Barnes DE, Lindahl T (2006) Mutations in the gene encoding the 3′-5' DNA exonuclease TREX1 cause Aicardi-Goutieres syndrome at the AGS1 locus. Nat Genet 38(8):917–920. https://doi.org/10.1038/ng1845
Crow YJ, Leitch A, Hayward BE, Garner A, Parmar R, Griffith E, Ali M, Semple C, Aicardi J, Babul-Hirji R, Baumann C, Baxter P, Bertini E, Chandler KE, Chitayat D, Cau D, Dery C, Fazzi E, Goizet C, King MD, Klepper J, Lacombe D, Lanzi G, Lyall H, Martinez-Frias ML, Mathieu M, McKeown C, Monier A, Oade Y, Quarrell OW, Rittey CD, Rogers RC, Sanchis A, Stephenson JB, Tacke U, Till M, Tolmie JL, Tomlin P, Voit T, Weschke B, Woods CG, Lebon P, Bonthron DT, Ponting CP, Jackson AP (2006) Mutations in genes encoding ribonuclease H2 subunits cause Aicardi-Goutieres syndrome and mimic congenital viral brain infection. Nat Genet 38(8):910–916. https://doi.org/10.1038/ng1842
Rice GI, Bond J, Asipu A, Brunette RL, Manfield IW, Carr IM, Fuller JC, Jackson RM, Lamb T, Briggs TA, Ali M, Gornall H, Couthard LR, Aeby A, Attard-Montalto SP, Bertini E, Bodemer C, Brockmann K, Brueton LA, Corry PC, Desguerre I, Fazzi E, Cazorla AG, Gener B, Hamel BC, Heiberg A, Hunter M, van der Knaap MS, Kumar R, Lagae L, Landrieu PG, Lourenco CM, Marom D, McDermott MF, van der Merwe W, Orcesi S, Prendiville JS, Rasmussen M, Shalev SA, Soler DM, Shinawi M, Spiegel R, Tan TY, Vanderver A, Wakeling EL, Wassmer E, Whittaker E, Lebon P, Stetson DB, Bonthron DT, Crow YJ (2009) Mutations involved in Aicardi-Goutieres syndrome implicate SAMHD1 as regulator of the innate immune response. Nat Genet 41(7):829–832. https://doi.org/10.1038/ng.373
Coquel F, Neumayer C, Lin YL, Pasero P (2019) SAMHD1 and the innate immune response to cytosolic DNA during DNA replication. Curr Opin Immunol 56:24–30. https://doi.org/10.1016/j.coi.2018.09.017
Rice GI, Kasher PR, Forte GM, Mannion NM, Greenwood SM, Szynkiewicz M, Dickerson JE, Bhaskar SS, Zampini M, Briggs TA, Jenkinson EM, Bacino CA, Battini R, Bertini E, Brogan PA, Brueton LA, Carpanelli M, De Laet C, de Lonlay P, del Toro M, Desguerre I, Fazzi E, Garcia-Cazorla A, Heiberg A, Kawaguchi M, Kumar R, Lin JP, Lourenco CM, Male AM, Marques W Jr, Mignot C, Olivieri I, Orcesi S, Prabhakar P, Rasmussen M, Robinson RA, Rozenberg F, Schmidt JL, Steindl K, Tan TY, van der Merwe WG, Vanderver A, Vassallo G, Wakeling EL, Wassmer E, Whittaker E, Livingston JH, Lebon P, Suzuki T, McLaughlin PJ, Keegan LP, O'Connell MA, Lovell SC, Crow YJ (2012) Mutations in ADAR1 cause Aicardi-Goutieres syndrome associated with a type I interferon signature. Nat Genet 44(11):1243–1248. https://doi.org/10.1038/ng.2414
Oda H, Nakagawa K, Abe J, Awaya T, Funabiki M, Hijikata A, Nishikomori R, Funatsuka M, Ohshima Y, Sugawara Y, Yasumi T, Kato H, Shirai T, Ohara O, Fujita T, Heike T (2014) Aicardi-Goutieres syndrome is caused by IFIH1 mutations. Am J Hum Genet 95(1):121–125. https://doi.org/10.1016/j.ajhg.2014.06.007
Pelzer N, Hoogeveen ES, Haan J, Bunnik R, Poot CC, van Zwet EW, Inderson A, Fogteloo AJ, Reinders MEJ, Middelkoop HAM, Kruit MC, van den Maagdenberg A, Ferrari MD, Terwindt GM (2019) Systemic features of retinal vasculopathy with cerebral leukoencephalopathy and systemic manifestations: a monogenic small vessel disease. J Intern Med 285(3):317–332. https://doi.org/10.1111/joim.12848
Konig N, Fiehn C, Wolf C, Schuster M, Cura Costa E, Tungler V, Alvarez HA, Chara O, Engel K, Goldbach-Mansky R, Gunther C, Lee-Kirsch MA (2017) Familial chilblain lupus due to a gain-of-function mutation in STING. Ann Rheum Dis 76(2):468–472. https://doi.org/10.1136/annrheumdis-2016-209841
Liu Y, Jesus AA, Marrero B, Yang D, Ramsey SE, Sanchez GAM, Tenbrock K, Wittkowski H, Jones OY, Kuehn HS, Lee CR, DiMattia MA, Cowen EW, Gonzalez B, Palmer I, DiGiovanna JJ, Biancotto A, Kim H, Tsai WL, Trier AM, Huang Y, Stone DL, Hill S, Kim HJ, St Hilaire C, Gurprasad S, Plass N, Chapelle D, Horkayne-Szakaly I, Foell D, Barysenka A, Candotti F, Holland SM, Hughes JD, Mehmet H, Issekutz AC, Raffeld M, McElwee J, Fontana JR, Minniti CP, Moir S, Kastner DL, Gadina M, Steven AC, Wingfield PT, Brooks SR, Rosenzweig SD, Fleisher TA, Deng Z, Boehm M, Paller AS, Goldbach-Mansky R (2014) Activated STING in a vascular and pulmonary syndrome. N Engl J Med 371(6):507–518. https://doi.org/10.1056/NEJMoa1312625
Navarro V, Scott C, Briggs TA, Barete S, Frances C, Lebon P, Maisonobe T, Rice GI, Wouters CH, Crow YJ (2008) Two further cases of spondyloenchondrodysplasia (SPENCD) with immune dysregulation. Am J Med Genet A 146A(21):2810–2815. https://doi.org/10.1002/ajmg.a.32518
Costa-Reis P, Sullivan KE (2017) Monogenic lupus: it’s all new! Curr Opin Immunol 49:87–95. https://doi.org/10.1016/j.coi.2017.10.008
Rutsch F, MacDougall M, Lu C, Buers I, Mamaeva O, Nitschke Y, Rice GI, Erlandsen H, Kehl HG, Thiele H, Nurnberg P, Hohne W, Crow YJ, Feigenbaum A, Hennekam RC (2015) A specific IFIH1 gain-of-function mutation causes Singleton-Merten syndrome. Am J Hum Genet 96(2):275–282. https://doi.org/10.1016/j.ajhg.2014.12.014
Jang MA, Kim EK, Now H, Nguyen NT, Kim WJ, Yoo JY, Lee J, Jeong YM, Kim CH, Kim OH, Sohn S, Nam SH, Hong Y, Lee YS, Chang SA, Jang SY, Kim JW, Lee MS, Lim SY, Sung KS, Park KT, Kim BJ, Lee JH, Kim DK, Kee C, Ki CS (2015) Mutations in DDX58, which encodes RIG-I, cause atypical Singleton-Merten syndrome. Am J Hum Genet 96(2):266–274. https://doi.org/10.1016/j.ajhg.2014.11.019
Shipman L (2014) Cytokines: ISG15--human deficiency reveals new proficiency. Nat Rev Immunol 14(12):780–781. https://doi.org/10.1038/nri3767
Cavalcante MP, Brunelli JB, Miranda CC, Novak GV, Malle L, Aikawa NE, Jesus AA, Silva CA (2016) CANDLE syndrome: chronic atypical neutrophilic dermatosis with lipodystrophy and elevated temperature-a rare case with a novel mutation. Eur J Pediatr 175(5):735–740. https://doi.org/10.1007/s00431-015-2668-4
Brehm A, Liu Y, Sheikh A, Marrero B, Omoyinmi E, Zhou Q, Montealegre G, Biancotto A, Reinhardt A, Almeida de Jesus A, Pelletier M, Tsai WL, Remmers EF, Kardava L, Hill S, Kim H, Lachmann HJ, Megarbane A, Chae JJ, Brady J, Castillo RD, Brown D, Casano AV, Gao L, Chapelle D, Huang Y, Stone D, Chen Y, Sotzny F, Lee CC, Kastner DL, Torrelo A, Zlotogorski A, Moir S, Gadina M, McCoy P, Wesley R, Rother KI, Hildebrand PW, Brogan P, Kruger E, Aksentijevich I, Goldbach-Mansky R (2015) Additive loss-of-function proteasome subunit mutations in CANDLE/PRAAS patients promote type I IFN production. J Clin Invest 125(11):4196–4211. https://doi.org/10.1172/JCI81260
Alsohime F, Martin-Fernandez M, Temsah MH, Alabdulhafid M, Le Voyer T, Alghamdi M, Qiu X, Alotaibi N, Alkahtani A, Buta S, Jouanguy E, Al-Eyadhy A, Gruber C, Hasan GM, Bashiri FA, Halwani R, Hassan HH, Al-Muhsen S, Alkhamis N, Alsum Z, Casanova JL, Bustamante J, Bogunovic D, Alangari AA (2020) JAK inhibitor therapy in a child with inherited USP18 deficiency. N Engl J Med 382(3):256–265. https://doi.org/10.1056/NEJMoa1905633
Sanchez GAM, Reinhardt A, Ramsey S, Wittkowski H, Hashkes PJ, Berkun Y, Schalm S, Murias S, Dare JA, Brown D, Stone DL, Gao L, Klausmeier T, Foell D, de Jesus AA, Chapelle DC, Kim H, Dill S, Colbert RA, Failla L, Kost B, O'Brien M, Reynolds JC, Folio LR, Calvo KR, Paul SM, Weir N, Brofferio A, Soldatos A, Biancotto A, Cowen EW, Digiovanna JJ, Gadina M, Lipton AJ, Hadigan C, Holland SM, Fontana J, Alawad AS, Brown RJ, Rother KI, Heller T, Brooks KM, Kumar P, Brooks SR, Waldman M, Singh HK, Nickeleit V, Silk M, Prakash A, Janes JM, Ozen S, Wakim PG, Brogan PA, Macias WL, Goldbach-Mansky R (2018) JAK1/2 inhibition with baricitinib in the treatment of autoinflammatory interferonopathies. J Clin Invest 128(7):3041–3052. https://doi.org/10.1172/JCI98814
Fremond ML, Rodero MP, Jeremiah N, Belot A, Jeziorski E, Duffy D, Bessis D, Cros G, Rice GI, Charbit B, Hulin A, Khoudour N, Caballero CM, Bodemer C, Fabre M, Berteloot L, Le Bourgeois M, Reix P, Walzer T, Moshous D, Blanche S, Fischer A, Bader-Meunier B, Rieux-Laucat F, Crow YJ, Neven B (2016) Efficacy of the Janus kinase 1/2 inhibitor ruxolitinib in the treatment of vasculopathy associated with TMEM173-activating mutations in 3 children. J Allergy Clin Immunol 138(6):1752–1755. https://doi.org/10.1016/j.jaci.2016.07.015
Tungler V, Konig N, Gunther C, Engel K, Fiehn C, Smitka M, von der Hagen M, Berner R, Lee-Kirsch MA (2016) Response to: 'JAK inhibition in STING-associated interferonopathy' by Crow et al. cJ. Clin Invest 128(7):2760–2762. https://doi.org/10.1172/JCI121526
Meesilpavikkai K, Dik WA, Schrijver B, van Helden-Meeuwsen CG, Versnel MA, van Hagen PM, Bijlsma EK, Ruivenkamp CAL, Oele MJ, Dalm V (2019) Efficacy of baricitinib in the treatment of chilblains associated with Aicardi-Goutieres syndrome, a type I interferonopathy. Arthritis Rheumatol (Hoboken, NJ) 71(5):829–831. https://doi.org/10.1002/art.40805
Briand C, Fremond ML, Bessis D, Carbasse A, Rice GI, Bondet V, Duffy D, Chatenoud L, Blanche S, Crow YJ, Neven B (2019) Efficacy of JAK1/2 inhibition in the treatment of chilblain lupus due to TREX1 deficiency. Ann Rheum Dis 78(3):431–433. https://doi.org/10.1136/annrheumdis-2018-214037
Zimmermann N, Wolf C, Schwenke R, Luth A, Schmidt F, Engel K, Lee-Kirsch MA, Gunther C (2019) Assessment of clinical response to Janus kinase inhibition in patients with familial chilblain lupus and TREX1 mutation. JAMA Dermatol 155(3):342–346. https://doi.org/10.1001/jamadermatol.2018.5077
Fragoulis GE, McInnes IB, Siebert S (2019) JAK-inhibitors. New players in the field of immune-mediated diseases, beyond rheumatoid arthritis. Rheumatology (Oxford, England) 58(Suppl 1):i43–i54. https://doi.org/10.1093/rheumatology/key276
Winthrop KL (2017) The emerging safety profile of JAK inhibitors in rheumatic disease. Nat Rev Rheumatol 13(4):234–243. https://doi.org/10.1038/nrrheum.2017.23
Bronson PG, Chaivorapol C, Ortmann W, Behrens TW, Graham RR (2012) The genetics of type I interferon in systemic lupus erythematosus. Curr Opin Immunol 24(5):530–537. https://doi.org/10.1016/j.coi.2012.07.008
Psarras A, Emery P, Vital EM (2017) Type I interferon-mediated autoimmune diseases: pathogenesis, diagnosis and targeted therapy. Rheumatology (Oxford, England) 56(10):1662–1675. https://doi.org/10.1093/rheumatology/kew431
Guiducci C, Gong M, Xu Z, Gill M, Chaussabel D, Meeker T, Chan JH, Wright T, Punaro M, Bolland S, Soumelis V, Banchereau J, Coffman RL, Pascual V, Barrat FJ (2010) TLR recognition of self nucleic acids hampers glucocorticoid activity in lupus. Nature 465(7300):937–941. https://doi.org/10.1038/nature09102
Rizza P, Moretti F, Capone I, Belardelli F (2015) Role of type I interferon in inducing a protective immune response: perspectives for clinical applications. Cytokine Growth Factor Rev 26(2):195–201. https://doi.org/10.1016/j.cytogfr.2014.10.002
Wolf SJ, Estadt SN, Theros J, Moore T, Ellis J, Liu J, Reed TJ, Jacob CO, Gudjonsson JE, Kahlenberg JM (2019) Ultraviolet light induces increased T cell activation in lupus-prone mice via type I IFN-dependent inhibition of T regulatory cells. J Autoimmun 103:102291. https://doi.org/10.1016/j.jaut.2019.06.002
Liu M, Guo Q, Wu C, Sterlin D, Goswami S, Zhang Y, Li T, Bao C, Shen N, Fu Q, Zhang X (2019) Type I interferons promote the survival and proinflammatory properties of transitional B cells in systemic lupus erythematosus patients. Cell Mol Immunol 16(4):367–379. https://doi.org/10.1038/s41423-018-0010-6
Dieudonne Y, Gies V, Guffroy A, Keime C, Bird AK, Liesveld J, Barnas JL, Poindron V, Douiri N, Soulas-Sprauel P, Martin T, Meffre E, Anolik JH, Korganow AS (2019) Transitional B cells in quiescent SLE: An early checkpoint imprinted by IFN. J Autoimmun 102:150–158. https://doi.org/10.1016/j.jaut.2019.05.002
Hamilton JA, Hsu HC, Mountz JD (2019) Autoreactive B cells in SLE, villains or innocent bystanders? Immunol Rev 292(1):120–138. https://doi.org/10.1111/imr.12815
Chyuan IT, Tzeng HT, Chen JY (2019) Signaling pathways of type I and type III interferons and targeted therapies in systemic lupus erythematosus. Cells 8(9). https://doi.org/10.3390/cells8090963
Sarkar MK, Hile GA, Tsoi LC, Xing X, Liu J, Liang Y, Berthier CC, Swindell WR, Patrick MT, Shao S, Tsou PS, Uppala R, Beamer MA, Srivastava A, Bielas SL, Harms PW, Getsios S, Elder JT, Voorhees JJ, Gudjonsson JE, Kahlenberg JM (2018) Photosensitivity and type I IFN responses in cutaneous lupus are driven by epidermal-derived interferon kappa. Ann Rheum Dis 77(11):1653–1664. https://doi.org/10.1136/annrheumdis-2018-213197
Lo MS (2018) Insights gained from the study of pediatric systemic lupus erythematosus. Front Immunol 9:1278. https://doi.org/10.3389/fimmu.2018.01278
Rice GI, Melki I, Fremond ML, Briggs TA, Rodero MP, Kitabayashi N, Oojageer A, Bader-Meunier B, Belot A, Bodemer C, Quartier P, Crow YJ (2017) Assessment of type I interferon signaling in pediatric inflammatory disease. J Clin Immunol 37(2):123–132. https://doi.org/10.1007/s10875-016-0359-1
Wahadat MJ, Bodewes ILA, Maria NI, van Helden-Meeuwsen CG, van Dijk-Hummelman A, Steenwijk EC, Kamphuis S, Versnel MA (2018) Type I IFN signature in childhood-onset systemic lupus erythematosus: a conspiracy of DNA- and RNA-sensing receptors? Arthritis Res Therapy 20(1):4. https://doi.org/10.1186/s13075-017-1501-z
Midgley A, Thorbinson C, Beresford MW (2012) Expression of toll-like receptors and their detection of nuclear self-antigen leading to immune activation in JSLE. Rheumatology (Oxford, England) 51(5):824–832. https://doi.org/10.1093/rheumatology/ker400
Fava A, Petri M (2019) Systemic lupus erythematosus: diagnosis and clinical management. J Autoimmun 96:1–13. https://doi.org/10.1016/j.jaut.2018.11.001
Kalunian KC, Merrill JT, Maciuca R, McBride JM, Townsend MJ, Wei X, Davis JC Jr, Kennedy WP (2016) A phase II study of the efficacy and safety of rontalizumab (rhuMAb interferon-alpha) in patients with systemic lupus erythematosus (ROSE). Ann Rheum Dis 75(1):196–202. https://doi.org/10.1136/annrheumdis-2014-206090
McBride JM, Jiang J, Abbas AR, Morimoto A, Li J, Maciuca R, Townsend M, Wallace DJ, Kennedy WP, Drappa J (2012) Safety and pharmacodynamics of rontalizumab in patients with systemic lupus erythematosus: results of a phase I, placebo-controlled, double-blind, dose-escalation study. Arthritis Rheum 64(11):3666–3676. https://doi.org/10.1002/art.34632
Khamashta M, Merrill JT, Werth VP, Furie R, Kalunian K, Illei GG, Drappa J, Wang L, Greth W, investigators CDs (2016) Sifalimumab, an anti-interferon-alpha monoclonal antibody, in moderate to severe systemic lupus erythematosus: a randomised, double-blind, placebo-controlled study. Ann Rheum Dis 75 (11):1909–1916. https://doi.org/10.1136/annrheumdis-2015-208562
Chasset F, Arnaud L (2018) Targeting interferons and their pathways in systemic lupus erythematosus. Autoimmun Rev 17(1):44–52. https://doi.org/10.1016/j.autrev.2017.11.009
Ducreux J, Houssiau FA, Vandepapeliere P, Jorgensen C, Lazaro E, Spertini F, Colaone F, Roucairol C, Laborie M, Croughs T, Grouard-Vogel G, Lauwerys BR (2016) Interferon alpha kinoid induces neutralizing anti-interferon alpha antibodies that decrease the expression of interferon-induced and B cell activation associated transcripts: analysis of extended follow-up data from the interferon alpha kinoid phase I/II study. Rheumatology (Oxford, England) 55(10):1901–1905. https://doi.org/10.1093/rheumatology/kew262
Houssiau FA, Thanou A, Mazur M, Ramiterre E, Gomez Mora DA, Misterska-Skora M, Perich-Campos RA, Smakotina SA, Cerpa Cruz S, Louzir B, Croughs T, Tee ML (2020) IFN-alpha kinoid in systemic lupus erythematosus: results from a phase IIb, randomised, placebo-controlled study. Ann Rheum Dis 79(3):347–355. https://doi.org/10.1136/annrheumdis-2019-216379
Furie R, Khamashta M, Merrill JT, Werth VP, Kalunian K, Brohawn P, Illei GG, Drappa J, Wang L, Yoo S, Investigators CDS (2017) Anifrolumab, an anti-interferon-alpha receptor monoclonal antibody, in moderate-to-severe systemic lupus erythematosus. Arthritis & rheumatology (Hoboken, NJ) 69(2):376–386. https://doi.org/10.1002/art.39962
Morand EF, Trasieva T, Berglind A, Illei GG, Tummala R (2018) Lupus low disease activity state (LLDAS) attainment discriminates responders in a systemic lupus erythematosus trial: post-hoc analysis of the phase IIb MUSE trial of anifrolumab. Ann Rheum Dis 77(5):706–713. https://doi.org/10.1136/annrheumdis-2017-212504
Salmon JE, Niewold TB (2020) A successful trial for lupus - how good is good enough? N Engl J Med 382(3):287–288. https://doi.org/10.1056/NEJMe1915490
Furie R, Werth VP, Merola JF, Stevenson L, Reynolds TL, Naik H, Wang W, Christmann R, Gardet A, Pellerin A, Hamann S, Auluck P, Barbey C, Gulati P, Rabah D, Franchimont N (2019) Monoclonal antibody targeting BDCA2 ameliorates skin lesions in systemic lupus erythematosus. J Clin Invest 129(3):1359–1371. https://doi.org/10.1172/JCI124466
Zhan Y, Carrington EM, Ko HJ, Vikstrom IB, Oon S, Zhang JG, Vremec D, Brady JL, Bouillet P, Wu L, Huang DC, Wicks IP, Morand EF, Strasser A, Lew AM (2015) Bcl-2 antagonists kill plasmacytoid dendritic cells from lupus-prone mice and dampen interferon-alpha production. Arthritis & rheumatology (Hoboken, NJ) 67(3):797–808. https://doi.org/10.1002/art.38966
Lu P, Fleischmann R, Curtis C, Ignatenko S, Clarke SH, Desai M, Wong SL, Grebe KM, Black K, Zeng J, Stolzenbach J, Medema JK (2018) Safety and pharmacodynamics of venetoclax (ABT-199) in a randomized single and multiple ascending dose study in women with systemic lupus erythematosus. Lupus 27(2):290–302. https://doi.org/10.1177/0961203317719334
Nader A, Minocha M, Othman AA (2020) Exposure-response analyses of the effects of venetoclax, a selective BCL-2 inhibitor, on B-lymphocyte and total lymphocyte counts in women with systemic lupus erythematosus. Clin Pharmacokinet 59(3):335–347. https://doi.org/10.1007/s40262-019-00818-5
Li J, Wang X, Zhang F, Yin H (2013) Toll-like receptors as therapeutic targets for autoimmune connective tissue diseases. Pharmacol Ther 138(3):441–451. https://doi.org/10.1016/j.pharmthera.2013.03.003
Dudhgaonkar S, Ranade S, Nagar J, Subramani S, Prasad DS, Karunanithi P, Srivastava R, Venkatesh K, Selvam S, Krishnamurthy P, Mariappan TT, Saxena A, Fan L, Stetsko DK, Holloway DA, Li X, Zhu J, Yang WP, Ruepp S, Nair S, Santella J, Duncia J, Hynes J, McIntyre KW, Carman JA (2017) Selective IRAK4 inhibition attenuates disease in murine lupus models and demonstrates steroid sparing activity. J Immunol 198(3):1308–1319. https://doi.org/10.4049/jimmunol.1600583
Haselmayer P, Camps M, Liu-Bujalski L, Nguyen N, Morandi F, Head J, O'Mahony A, Zimmerli SC, Bruns L, Bender AT, Schroeder P, Grenningloh R (2019) Efficacy and pharmacodynamic modeling of the BTK inhibitor evobrutinib in autoimmune disease models. J Immunol 202(10):2888–2906. https://doi.org/10.4049/jimmunol.1800583
Klavdianou K, Lazarini A, Fanouriakis A (2020) Targeted biologic therapy for systemic lupus erythematosus: emerging pathways and drug pipeline. BioDrugs. https://doi.org/10.1007/s40259-020-00405-2
You H, Xu D, Zhao J, Li J, Wang Q, Tian X, Li M, Zeng X (2020) JAK inhibitors: prospects in connective tissue diseases. Clin Rev Allergy Immunol. https://doi.org/10.1007/s12016-020-08786-6
Malemud CJ (2018) The role of the JAK/STAT signal pathway in rheumatoid arthritis. Ther Adv Musculoskelet Dis 10(5–6):117–127. https://doi.org/10.1177/1759720X18776224
Hojjati SM, Heidari B, Babaei M (2016) Development of rheumatoid arthritis during treatment of multiple sclerosis with interferon beta 1-a. coincidence of two conditions or a complication of treatment: a case report. J Adv Res 7(5):611–613. https://doi.org/10.1016/j.jare.2016.06.004
Lubbers J, Brink M, van de Stadt LA, Vosslamber S, Wesseling JG, van Schaardenburg D, Rantapaa-Dahlqvist S, Verweij CL (2013) The type I IFN signature as a biomarker of preclinical rheumatoid arthritis. Ann Rheum Dis 72(5):776–780. https://doi.org/10.1136/annrheumdis-2012-202753
Nader A, Stodtmann S, Friedel A, Mohamed MF, Othman AA (2020) Pharmacokinetics of Upadacitinib in healthy subjects and subjects with rheumatoid arthritis, Crohn’s disease, ulcerative colitis, or atopic dermatitis: population analyses of phase 1 and 2 clinical trials. J Clin Pharmacol 60(4):528–539. https://doi.org/10.1002/jcph.1550
Genovese MC, Kalunian K, Gottenberg JE, Mozaffarian N, Bartok B, Matzkies F, Gao J, Guo Y, Tasset C, Sundy JS, de Vlam K, Walker D, Takeuchi T (2019) Effect of filgotinib vs placebo on clinical response in patients with moderate to severe rheumatoid arthritis refractory to disease-modifying antirheumatic drug therapy: the FINCH 2 randomized clinical trial. JAMA 322(4):315–325. https://doi.org/10.1001/jama.2019.9055
Takeuchi T, Tanaka Y, Tanaka S, Kawakami A, Iwasaki M, Katayama K, Rokuda M, Izutsu H, Ushijima S, Kaneko Y, Shiomi T, Yamada E, van der Heijde D (2019) Efficacy and safety of peficitinib (ASP015K) in patients with rheumatoid arthritis and an inadequate response to methotrexate: results of a phase III randomised, double-blind, placebo-controlled trial (RAJ4) in Japan. Ann Rheum Dis 78(10):1305–1319. https://doi.org/10.1136/annrheumdis-2019-215164
Genovese MC, Greenwald M, Codding C, Zubrzycka-Sienkiewicz A, Kivitz AJ, Wang A, Shay K, Wang X, Garg JP, Cardiel MH (2017) Peficitinib, a JAK inhibitor, in combination with limited conventional synthetic disease-modifying antirheumatic drugs in the treatment of moderate-to-severe rheumatoid arthritis. Arthritis & rheumatology (Hoboken, NJ) 69(5):932–942. https://doi.org/10.1002/art.40054
Westhovens R (2019) Clinical efficacy of new JAK inhibitors under development. Just more of the same? Rheumatology (Oxford, England) 58(Suppl 1):i27–i33. https://doi.org/10.1093/rheumatology/key256
Harigai M (2019) Growing evidence of the safety of JAK inhibitors in patients with rheumatoid arthritis. Rheumatology (Oxford, England) 58(Suppl 1):i34–i42. https://doi.org/10.1093/rheumatology/key287
Kjelgaard-Petersen CF, Platt A, Braddock M, Jenkins MA, Musa K, Graham E, Gantzel T, Slynn G, Weinblatt ME, Karsdal MA, Thudium CS, Bay-Jensen AC (2018) Translational biomarkers and ex vivo models of joint tissues as a tool for drug development in rheumatoid arthritis. Arthritis & rheumatology (Hoboken, NJ) 70(9):1419–1428. https://doi.org/10.1002/art.40527
Schafer PH, Kivitz AJ, Ma J, Korish S, Sutherland D, Li L, Azaryan A, Kosek J, Adams M, Capone L, Hur EM, Hough DR, Ringheim GE (2020) Spebrutinib (CC-292) affects markers of B cell activation, chemotaxis, and osteoclasts in patients with rheumatoid arthritis: results from a mechanistic study. Rheumatology and therapy 7(1):101–119. https://doi.org/10.1007/s40744-019-00182-7
Nehmar R, Mariotte A, de Cauwer A, Sibilia J, Bahram S, Georgel P (2018) Therapeutic perspectives for interferons and plasmacytoid dendritic cells in rheumatoid arthritis. Trends Mol Med 24(4):338–347. https://doi.org/10.1016/j.molmed.2018.02.001
van Holten J, Pavelka K, Vencovsky J, Stahl H, Rozman B, Genovese M, Kivitz AJ, Alvaro J, Nuki G, Furst DE, Herrero-Beaumont G, McInnes IB, Musikic P, Tak PP (2005) A multicentre, randomised, double blind, placebo controlled phase II study of subcutaneous interferon beta-1a in the treatment of patients with active rheumatoid arthritis. Ann Rheum Dis 64(1):64–69. https://doi.org/10.1136/ard.2003.020347
Mauro A, Rigante D, Cimaz R (2017) Investigational drugs for treatment of juvenile idiopathic arthritis. Expert Opin Investig Drugs 26(4):381–387. https://doi.org/10.1080/13543784.2017.1301929
Bracaglia C, de Graaf K, Pires Marafon D, Guilhot F, Ferlin W, Prencipe G, Caiello I, Davi S, Schulert G, Ravelli A, Grom AA, de Min C, De Benedetti F (2017) Elevated circulating levels of interferon-gamma and interferon-gamma-induced chemokines characterise patients with macrophage activation syndrome complicating systemic juvenile idiopathic arthritis. Ann Rheum Dis 76(1):166–172. https://doi.org/10.1136/annrheumdis-2015-209020
Gattorno M, Chicha L, Gregorio A, Ferlito F, Rossi F, Jarrossay D, Lanzavecchia A, Martini A, Manz MG (2007) Distinct expression pattern of IFN-alpha and TNF-alpha in juvenile idiopathic arthritis synovial tissue. Rheumatology (Oxford, England) 46(4):657–665. https://doi.org/10.1093/rheumatology/kel346
Myles A, Aggarwal A (2011) Expression of toll-like receptors 2 and 4 is increased in peripheral blood and synovial fluid monocytes of patients with enthesitis-related arthritis subtype of juvenile idiopathic arthritis. Rheumatology (Oxford, England) 50(3):481–488. https://doi.org/10.1093/rheumatology/keq362
Stanford SM, Bottini N (2014) PTPN22: the archetypal non-HLA autoimmunity gene. Nat Rev Rheumatol 10(10):602–611. https://doi.org/10.1038/nrrheum.2014.109
Ruperto N, Brunner HI, Zuber Z, Tzaribachev N, Kingsbury DJ, Foeldvari I, Horneff G, Smolewska E, Vehe RK, Hazra A, Wang R, Mebus CA, Alvey C, Lamba M, Krishnaswami S, Stock TC, Wang M, Suehiro R, Martini A, Lovell DJ, Pediatric Rheumatology International Trials O, Pediatric Rheumatology Collaborative Study G (2017) Pharmacokinetic and safety profile of tofacitinib in children with polyarticular course juvenile idiopathic arthritis: results of a phase 1, open-label, multicenter study. Pediatr Rheumatol Online J 15(1):86. https://doi.org/10.1186/s12969-017-0212-y
Huang Z, Lee PY, Yao X, Zheng S, Li T (2019) Tofacitinib treatment of refractory systemic juvenile idiopathic arthritis. Pediatrics 143(5). https://doi.org/10.1542/peds.2018-2845
Feldman BM, Rider LG, Reed AM, Pachman LM (2008) Juvenile dermatomyositis and other idiopathic inflammatory myopathies of childhood. Lancet (London, England) 371(9631):2201–2212. https://doi.org/10.1016/S0140-6736(08)60955-1
Niewold TB, Kariuki SN, Morgan GA, Shrestha S, Pachman LM (2009) Elevated serum interferon-alpha activity in juvenile dermatomyositis: associations with disease activity at diagnosis and after thirty-six months of therapy. Arthritis Rheum 60(6):1815–1824. https://doi.org/10.1002/art.24555
Moneta GM, Pires Marafon D, Marasco E, Rosina S, Verardo M, Fiorillo C, Minetti C, Bracci-Laudiero L, Ravelli A, De Benedetti F, Nicolai R (2019) Muscle expression of type I and type II interferons is increased in juvenile dermatomyositis and related to clinical and histologic features. Arthritis & rheumatology (Hoboken, NJ) 71(6):1011–1021. https://doi.org/10.1002/art.40800
O'Connor KA, Abbott KA, Sabin B, Kuroda M, Pachman LM (2006) MxA gene expression in juvenile dermatomyositis peripheral blood mononuclear cells: association with muscle involvement. Clin Immunol 120(3):319–325. https://doi.org/10.1016/j.clim.2006.05.011
Baechler EC, Bilgic H, Reed AM (2011) Type I interferon pathway in adult and juvenile dermatomyositis. Arthritis research & therapy 13(6):249. https://doi.org/10.1186/ar3531
Cappelletti C, Baggi F, Zolezzi F, Biancolini D, Beretta O, Severa M, Coccia EM, Confalonieri P, Morandi L, Mora M, Mantegazza R, Bernasconi P (2011) Type I interferon and toll-like receptor expression characterizes inflammatory myopathies. Neurology 76(24):2079–2088. https://doi.org/10.1212/WNL.0b013e31821f440a
Piper CJM, Wilkinson MGL, Deakin CT, Otto GW, Dowle S, Duurland CL, Adams S, Marasco E, Rosser EC, Radziszewska A, Carsetti R, Ioannou Y, Beales PL, Kelberman D, Isenberg DA, Mauri C, Nistala K, Wedderburn LR (2018) CD19(+)CD24(hi)CD38(hi) B cells are expanded in juvenile dermatomyositis and exhibit a pro-inflammatory phenotype after activation through toll-like receptor 7 and interferon-alpha. Front Immunol 9:1372. https://doi.org/10.3389/fimmu.2018.01372
Gitiaux C, Latroche C, Weiss-Gayet M, Rodero MP, Duffy D, Bader-Meunier B, Glorion C, Nusbaum P, Bodemer C, Mouchiroud G, Chelly J, Germain S, Desguerre I, Chazaud B (2018) Myogenic progenitor cells exhibit type I interferon-driven proangiogenic properties and molecular signature during juvenile dermatomyositis. Arthritis & rheumatology (Hoboken, NJ) 70(1):134–145. https://doi.org/10.1002/art.40328
Ladislau L, Suarez-Calvet X, Toquet S, Landon-Cardinal O, Amelin D, Depp M, Rodero MP, Hathazi D, Duffy D, Bondet V, Preusse C, Bienvenu B, Rozenberg F, Roos A, Benjamim CF, Gallardo E, Illa I, Mouly V, Stenzel W, Butler-Browne G, Benveniste O, Allenbach Y (2018) JAK inhibitor improves type I interferon induced damage: proof of concept in dermatomyositis. Brain 141(6):1609–1621. https://doi.org/10.1093/brain/awy105
Moghadam-Kia S, Charlton D, Aggarwal R, Oddis CV (2019) Management of refractory cutaneous dermatomyositis: potential role of Janus kinase inhibition with tofacitinib. Rheumatology (Oxford, England) 58(6):1011–1015. https://doi.org/10.1093/rheumatology/key366
Papadopoulou C, Hong Y, Omoyinmi E, Brogan PA, Eleftheriou D (2019) Janus kinase 1/2 inhibition with baricitinib in the treatment of juvenile dermatomyositis. Brain 142(3):e8. https://doi.org/10.1093/brain/awz005
Higgs BW, Zhu W, Morehouse C, White WI, Brohawn P, Guo X, Rebelatto M, Le C, Amato A, Fiorentino D, Greenberg SA, Drappa J, Richman L, Greth W, Jallal B, Yao Y (2014) A phase 1b clinical trial evaluating sifalimumab, an anti-IFN-alpha monoclonal antibody, shows target neutralisation of a type I IFN signature in blood of dermatomyositis and polymyositis patients. Ann Rheum Dis 73(1):256–262. https://doi.org/10.1136/annrheumdis-2012-202794
Yao Y, Liu Z, Jallal B, Shen N, Ronnblom L (2013) Type I interferons in Sjogren’s syndrome. Autoimmun Rev 12(5):558–566. https://doi.org/10.1016/j.autrev.2012.10.006
Zheng L, Zhang Z, Yu C, Tu L, Zhong L, Yang C (2009) Association between IFN-alpha and primary Sjogren’s syndrome. Oral Surg Oral Med Oral Pathol Oral Radiol Endod 107(1):e12–e18. https://doi.org/10.1016/j.tripleo.2008.09.015
Brkic Z, Maria NI, van Helden-Meeuwsen CG, van de Merwe JP, van Daele PL, Dalm VA, Wildenberg ME, Beumer W, Drexhage HA, Versnel MA (2013) Prevalence of interferon type I signature in CD14 monocytes of patients with Sjogren’s syndrome and association with disease activity and BAFF gene expression. Ann Rheum Dis 72(5):728–735. https://doi.org/10.1136/annrheumdis-2012-201381
Gottenberg JE, Cagnard N, Lucchesi C, Letourneur F, Mistou S, Lazure T, Jacques S, Ba N, Ittah M, Lepajolec C, Labetoulle M, Ardizzone M, Sibilia J, Fournier C, Chiocchia G, Mariette X (2006) Activation of IFN pathways and plasmacytoid dendritic cell recruitment in target organs of primary Sjogren’s syndrome. Proc Natl Acad Sci U S A 103(8):2770–2775. https://doi.org/10.1073/pnas.0510837103
Bave U, Nordmark G, Lovgren T, Ronnelid J, Cajander S, Eloranta ML, Alm GV, Ronnblom L (2005) Activation of the type I interferon system in primary Sjogren’s syndrome: a possible etiopathogenic mechanism. Arthritis Rheum 52(4):1185–1195. https://doi.org/10.1002/art.20998
Lee J, Lee J, Kwok SK, Baek S, Jang SG, Hong SM, Min JW, Choi SS, Lee J, Cho ML, Park SH (2018) JAK-1 inhibition suppresses interferon-induced BAFF production in human salivary gland: potential therapeutic strategy for primary Sjogren’s syndrome. Arthritis & rheumatology (Hoboken, NJ) 70(12):2057–2066. https://doi.org/10.1002/art.40589
Charras A, Arvaniti P, Le Dantec C, Arleevskaya MI, Zachou K, Dalekos GN, Bordon A, Renaudineau Y (2020) JAK inhibitors suppress innate epigenetic reprogramming: a promise for patients with Sjogren’s syndrome. Clin Rev Allergy Immunol 58(2):182–193. https://doi.org/10.1007/s12016-019-08743-y
Bodewes ILA, Huijser E, van Helden-Meeuwsen CG, Tas L, Huizinga R, Dalm V, van Hagen PM, Groot N, Kamphuis S, van Daele PLA, Versnel MA (2018) TBK1: a key regulator and potential treatment target for interferon positive Sjogren’s syndrome, systemic lupus erythematosus and systemic sclerosis. J Autoimmun 91:97–102. https://doi.org/10.1016/j.jaut.2018.02.001
Bodewes ILA, Gottenberg JE, van Helden-Meeuwsen CG, Mariette X, Versnel MA (2020) Hydroxychloroquine treatment downregulates systemic interferon activation in primary Sjogren's syndrome in the JOQUER randomized trial. Rheumatology (Oxford, England) 59(1):107–111. https://doi.org/10.1093/rheumatology/kez242
Meijer JM, Pijpe J, Bootsma H, Vissink A, Kallenberg CG (2007) The future of biologic agents in the treatment of Sjogren’s syndrome. Clin Rev Allergy Immunol 32(3):292–297. https://doi.org/10.1007/s12016-007-8005-6
Skaug B, Assassi S (2019) Type I interferon dysregulation in systemic sclerosis Cytokine:154635. https://doi.org/10.1016/j.cyto.2018.12.018
Wu M, Assassi S (2013) The role of type 1 interferon in systemic sclerosis. Front Immunol 4:266. https://doi.org/10.3389/fimmu.2013.00266
Assassi S, Mayes MD, Arnett FC, Gourh P, Agarwal SK, McNearney TA, Chaussabel D, Oommen N, Fischbach M, Shah KR, Charles J, Pascual V, Reveille JD, Tan FK (2010) Systemic sclerosis and lupus: points in an interferon-mediated continuum. Arthritis Rheum 62(2):589–598. https://doi.org/10.1002/art.27224
Liu X, Mayes MD, Tan FK, Wu M, Reveille JD, Harper BE, Draeger HT, Gonzalez EB, Assassi S (2013) Correlation of interferon-inducible chemokine plasma levels with disease severity in systemic sclerosis. Arthritis Rheum 65(1):226–235. https://doi.org/10.1002/art.37742
Guo X, Higgs BW, Bay-Jensen AC, Karsdal MA, Yao Y, Roskos LK, White WI (2015) Suppression of T cell activation and collagen accumulation by an anti-IFNAR1 mAb, anifrolumab, in adult patients with systemic sclerosis. The Journal of investigative dermatology 135(10):2402–2409. https://doi.org/10.1038/jid.2015.188
Goldberg A, Geppert T, Schiopu E, Frech T, Hsu V, Simms RW, Peng SL, Yao Y, Elgeioushi N, Chang L, Wang B, Yoo S (2014) Dose-escalation of human anti-interferon-alpha receptor monoclonal antibody MEDI-546 in subjects with systemic sclerosis: a phase 1, multicenter, open label study. Arthritis research & therapy 16(1):R57. https://doi.org/10.1186/ar4492
Ah Kioon MD, Tripodo C, Fernandez D, Kirou KA, Spiera RF, Crow MK, Gordon JK, Barrat FJ (2018) Plasmacytoid dendritic cells promote systemic sclerosis with a key role for TLR8. Sci Transl Med 10(423). https://doi.org/10.1126/scitranslmed.aam8458
Cattalini M, Soliani M, Caparello MC, Cimaz R (2019) Sex differences in pediatric rheumatology. Clin Rev Allergy Immunol 56(3):293–307. https://doi.org/10.1007/s12016-017-8642-3
Axtell RC, Raman C (2012) Janus-like effects of type I interferon in autoimmune diseases. Immunol Rev 248(1):23–35. https://doi.org/10.1111/j.1600-065X.2012.01131.x
Axtell RC, Raman C, Steinman L (2013) Type I interferons: beneficial in Th1 and detrimental in Th17 autoimmunity. Clin Rev Allergy Immunol 44(2):114–120. https://doi.org/10.1007/s12016-011-8296-5
Acknowledgments
We thank Xing Wang, Meixi Liu, Zhenwei Tang, and Bowen Li for their guidance in revising the manuscript.
Author information
Authors and Affiliations
Contributions
Concept and design: Qianjin Lu, Haijing Wu, Ming Zhao, Jiao Jiang; Drafting of the manuscript: Jiao Jiang; Revising of the manuscript: Qianjin Lu, Haijing Wu, Christopher Chang, Ming Zhao, Jiao Jiang.
Funding information
This work was supported by the National Natural Science Foundation of China (No. 81972943 and No. 81830097) and the Hunan Talent Young Investigator (No. 2019RS2012).
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Jiang, J., Zhao, M., Chang, C. et al. Type I Interferons in the Pathogenesis and Treatment of Autoimmune Diseases. Clinic Rev Allerg Immunol 59, 248–272 (2020). https://doi.org/10.1007/s12016-020-08798-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12016-020-08798-2