Abstract
The multiple autoimmune syndromes (MAS) consist on the presence of three or more well-defined autoimmune diseases (ADs) in a single patient. The aim of this study was to analyze the clinical and genetic characteristics of a large series of patients with MAS. A cluster analysis and familial aggregation analysis of ADs was performed in 84 patients. A genome-wide microsatellite screen was performed in MAS families, and associated loci were investigated through the pedigree disequilibrium test. Systemic lupus erythematosus (SLE), autoimmune thyroid disease (AITD), and Sjögren’s syndrome together were the most frequent ADs encountered. Three main clusters were established. Aggregation for type 1 diabetes, AITD, SLE, and all ADs as a trait was found. Eight loci associated with MAS were observed harboring autoimmunity genes. The MAS represent the best example of polyautoimmunity as well as the effect of a single genotype on diverse phenotypes. Its study provides important clues to elucidate the common mechanisms of ADs (i.e., autoimmune tautology).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Autoimmune diseases (ADs) are chronic conditions initiated by the loss of immunological tolerance to self-antigens and represent a heterogeneous group of disorders that afflict specific target organs or multiple organ systems. The chronic nature of these diseases places a significant burden on the utilization of medical care, direct and indirect economic costs, and quality of life. A significant number of autoimmune disorders are more prevalent in women than men and are among the leading causes of disability for this group of patients.
The fact that ADs share several clinical signs and symptoms (i.e., subphenotypes), physiopathological mechanisms, and genetic factors has been called autoimmune tautology and indicates that they share common mechanisms [1]. There are two clinical conditions supporting this theory namely polyautoimmunity and familial autoimmunity. Polyautoimmunity is the presence of two or more ADs in a single patient while familial autoimmunity occurs when different relatives from a nuclear family present with diverse ADs [2–4]. These conditions indicate that similar genetic, epigenetic, and environmental factors influence ADs.
The multiple autoimmune syndromes (MAS) were described by Humbert and Dupond in 1988 as a syndrome(s) consisting of the presence of three or more well-characterized ADs in one patient [5]. While describing the syndrome, their observations led them to a rough classification of clusters based on the co-occurrence of ADs, which they named as type one through three. In MAS-1, the authors grouped together myasthenia gravis (MG), thymoma, dermatopolymyositis, and autoimmune myocarditis. For MAS-2, they grouped Sjögren’s syndrome (SS), rheumatoid arthritis (RA), primary biliary cirrhosis (PBC), systemic sclerosis (SSc), and autoimmune thyroid disease (AITD). Finally, MAS-3 consisted of AITD, MG and/or thymoma, SS, pernicious anemia (PA), idiopathic thrombocytopenic purpura (ITP), Addison’s disease, type 1 diabetes (T1D), vitiligo (VIT), autoimmune hemolytic anemia, and systemic lupus erythematosus (SLE) [5]. In spite of the great breakthrough achieved by Humbert and Dupond, providing a new taxonomy for these cocurrent phenotypes based on clinical evidence, the concept became subject to modifications. Further studies evaluating polyautoimmunity in patients having a primary AD found an association between other ADs not reported in the original report [4, 6].
Besides MAS, another well-known example of polyautoimmunity is the polyglandular autoimmune syndrome, type II (PASII). This syndrome is also universally known as Schmidt’s syndrome owing to a case reported by Schmidt in 1926 of two patients that presented Addison’s disease and chronic lymphocytic thyroiditis [7]. Afterward in 1964, Carpenter included in an extensive review of the literature the presence of T1D to the syndrome defining the classic triad for PASII [8]. The diagnosis of PASII is made by the presence of at least two of the previously described conditions in one patient. There are, nevertheless, three more types of PAS [9]. PAS type I is given by the presence of oral candidiasis, hypoparathyroidism, and Addison’s disease within an individual associated to mutations in the AIRE gene, a disorder that has an autosomic recessive inheritance. PAS type III is defined as the presence of AITD and other AD, including diabetes but does not typically include adrenal gland malfunction. Other diseases such as PA or celiac disease (CD) may also be observed. Finally, PAS type IV has been described as the association of two or more organ-specific ADs. There is, nevertheless, some controversy around this topic, and some authors argument that PAS type II, type III, and type IV are different manifestations of the same syndrome [9]. This disagreement has strong foundations, for there are several reports showing association between organ-specific ADs both endocrine and non-endocrine and systemic ADs like SLE, SS, or RA [4, 6]. In fact, the MAS as well as PASII, PASIII, and PASIV correspond to different forms of polyautoimmunity. The importance of polyautoimmunity lies in the relation of diverse phenotypes with a single genotype, which allow a better understanding of the autoimmune tautology. Herein, we report a clinical and genetic analysis of a large series of patients with MAS.
Patients and Methods
Study Population
Cases with MAS were patients presenting with three or more well-characterized ADs fulfilling validated classification criteria as previously reported [10] and who were attending the Center for Autoimmune Diseases Research (CREA), at the University of Rosario in Bogotá and Medellin, Colombia; the Discipline of Rheumatology, Universidade do Estado do Rio de Janeiro, Brazil; the Department of Autoimmune Diseases, Hospital Clinic, Barcelona, Catalonia, Spain; and Internal Medicine Service, Hospital de São João, Porto, Portugal.
Family Collection
Only relatives of Colombian patients were included and interviewed using a standardized questionnaire that incorporated demographics and medical information including a check-point list of 18 ADs [10]. In order to avoid ascertainment bias, the diagnosis of AD was only considered reliable and consequently registered if made by a certified physician (i.e., internist, endocrinologist, or rheumatologist) and confirmed by chart review or verification during discussion with the relative. The Ethics Committee of the University of Rosario approved the present study.
Autoantibodies
Antinuclear antibodies were determined by indirect immunofluorescence on HEp-2 cells. The detection of the specific antibodies, such as rheumatoid factor, lupus anticoagulant, native anti-DNA, anti-RNP, anti-Sm, anti-Ro, anti-La, anticardiolipins, anti-thyroglobulin, and anti-thyroid peroxidase antibodies and other organ-specific autoantibodies, were determined by commercial kits and were performed at each participating center.
DNA Preparation and Genotyping
Genomic DNA from 34 patients and their relatives was extracted from 10 mL of an EDTA-anticoagulated blood sample using the classical salting out protocol. Three hundred ninety-three polymorphic Marshfield microsatellite markers with an average spacing of ~10 cM from Marshfield Screening Set 16 were genotyped by the Marshfield Center for Medical Genetics (http://research.marshfieldclinic.org/genetics/)
Statistical and Genetic Analysis
Data were managed and stored using the SPSS program (v17 for Windows, Chicago, IL, USA) and pedigree information including demographic and genotype data for each individual was stored in PROGENY v7 (http://www.progenygenetics.com/). Results are presented as means ± standard deviation and in percentages. Comparisons between means were performed by the Student´s t test, and those between percentages were done by the χ 2 test and two-sided Fisher’s exact test as appropriate. Crude odds ratios (OR) were calculated with 95 % confidence intervals (CI). A p value of less than 0.05 was considered as significant.
Clusters of ADs were examined by unweighted pair-group method using arithmetic averages (UPGMA) [11]. UPGMA employs a sequential clustering algorithm, in which local topological relationships are identified in order of similarity, and the taxonomic tree is built in a stepwise manner. Additionally bootstrapping was made taking into account 1,000 replicas in order to evaluate the empirical significance of the clustering. As rule of dumb, bootstrapping of 70 % was considered equivalent to a significance p value <0.05.
Familial aggregation (λ R) was calculated for first-degree relatedness (parent/offspring and sibling/sibling pairs) using the formula \( {\lambda_{{_{\text{R}}}}} = {{{{K_{\text{Relative}}}}} \left/ {K} \right.} \), where K Relative (K R) was the prevalence for a specific degree of relatedness in the sample, and K was the prevalence in the control pedigree samples or the mean prevalence in the population, as described elsewhere [10]. A historical- and ethnicity-matched control group, previously reported [10], served as control pedigree group.
The association analysis between genotypes and MAS taken as a trait was performed by the pedigree disequilibrium test (PDT), as described elsewhere [12, 13] by using UNPHASED v3.0.9 [14]. General settings were as follows: disease gene frequency (normal, 0.95; disease, 0.05); model, dominant; phenocopy rate, 0.02; and penetrance, 0.98. Significances were corrected by empirical permutation by randomizing the transmission status of the parental genotypes.
Results
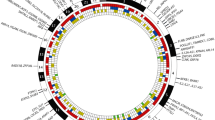
In this study, 84 patients with MAS were included (Table 1 and Fig. 1a). The mean age at onset of the first AD was 32.7 ± 14 years and the mean duration of the disease was 11.6 ± 8.7. The most frequent first ADs were SLE, AITD, and SS, presenting in 32.7, 21.43, and 10 %, respectively. The second more frequent ADs were SS, AITD, and SLE presenting in 11, 9, and 8 %, respectively. The third most frequent ADs were SS, AITD, and antiphospholipid syndrome, presenting in 16, 9, and 7 %, respectively (Fig. 1b). There were 20 (23.8 %) patients who presented a fourth AD, and one patient presenting five ADs. Cladistic analysis yielded 42 clusters with three ADs predominating: SLE, SS, and AITD (Fig. 1a).
Clusters of autoimmune diseases in patients with MAS and chromosomal loci associated. a The main clusters of autoimmune diseases. Bootstrapping was performed taking into account 1,000 replicas. b The frequencies of autoimmune diseases and mean age at diagnosis. The eight loci associated with autoimmunity are shown in c. p values were determined by 10,000 simulations. Their chromosomal locations are illustrated in d. Abbreviations: SLE systemic lupus erythematosus, RA rheumatoid arthritis, SS Sjögren’s syndrome, APS antiphospholipid syndrome, V vasculitis, SSc systemic sclerosis, AITD autoimmune thyroid disease, JRA juvenile rheumatoid arthritis, VIT vitiligo, DPM dermatopolymyositis, MG myasthenia gravis, IBD inflammatory bowel diseases including Crohn’s disease and ulcerative colitis, IBDS inflammatory biliary diseases including primary biliary cirrhosis and sclerosing cholangitis, PA pernicious anemia, AH autoimmune hepatitis, DAD demyelinating autoimmune diseases including transverse myelitis and multiple sclerosis, RP relapsing polychondritis, CD celiac disease
Of the 46 families of MAS cases, 12 (26 %) had at least one first-degree relative (FDR) with an AD as compared with 27 (21.8 %) of 124 families in the controls (Table 2). When families were stratified by the number of FDR, the risk of familial autoimmunity increased with the number of FDR affected with an AD (Table 2).
Specific ADs in FDR of MAS patients and controls are shown in Table 3. The percentage of FDRs with at least one AID in patients was 6.7 % as compared with 3.85 % in controls (OR 2.7, 95 % CI 1.50–4.88, p = 0.001).
The general statistics of the pedigrees for the MAS patients and controls are disclosed in Table 4. The mean pedigree size, standard deviation, as well as the total number of relative pairs were obtained in order to calculate the prevalence for each AD. The analyses were restricted to FDR.
The prevalence for each AD as well as for all ADs taken together for each pair of relatives (parent/offspring, sibling/sibling, and total FDR) is disclosed in Table 5. Previously reported prevalences were also taken into account [10]. These prevalences were used to calculate the familial aggregation for different degrees of relatives (Table 6). Values supporting familial aggregation (λ > 1.0) were observed using both the data on pedigrees and putative published prevalences. Familial aggregation of AITD, SLE, T1D, and all AID taken together (Table 3) was observed in families of MAS patients.
Results of PDT yielded eight loci associated with MAS at chromosomes 1p35.1, 6p22.3, 7p15.3, 13q31.1, 14q32.12, 16p12.1, 18q21.32, and 21q22.2 (Fig. 1c, d). Only marker wise analyses were performed because the moderate size of the families precludes confident haplotype definition.
Discussion
In this study, we analyzed the characteristics of a large series of patients with MAS. Our results disclosed (1) three main clusters of polyautoimmunity, (2) aggregation of ADs in families of patients with MAS (i.e., familial autoimmunity), and (3) eight loci associated with autoimmunity.
One of the first descriptions of MAS corresponded to a young woman presenting with RA, ITP, PA, Hashimoto's thyroiditis, SSc, and CD [15]. Her family study revealed RA, PA, and T1D among her FDRs. She carried the HLA-A1, HLA-A2, HLA-B5, HLA-B8, HLA-DR3, and HLA-DR4 groups [15]. This case well illustrates the relationship between familial autoimmunity and polyautoimmunity in the proband and the presence of a single genotype responsible for diverse ADs. Somers et al. [16] by using the UK General Practice Research Database demonstrated coexistence of RA, AITD, and T1D at higher than expected rates but reduced comorbidity between RA and multiple sclerosis (MS). Current data support a tendency for ADs to co-occur at greater than expected rates within proband patients and their families [17].
Polyautoimmunity has been reported in most of the ADs including SLE (41 %) [18], SSc (38 %) [19], PBC (32 %) [20], VIT (27 %) [21], MG (13 %) [22], Graves' disease (10 %), Hashimoto's thyroiditis (14 %) [23], and MS (6 %) [24] among others. The main difference between polyautoimmunity and the overlap syndromes lies in the fact that the former is the presence of two or more well-defined autoimmune conditions fulfilling validated classification criteria while the later is the partial presence of signs and symptoms of diverse ADs. Most of the cases of overlap syndromes have been described in cross-sectional studies. As our results show, there is a lag time interval between the first and the second AD (Table 1). Just as in the mixed connective tissue disease (MCTD), the classical overlap syndrome, in which some patients will develop during the course of the disease SLE, SSc, or RA, and some, will be presenting with a longstanding MCTD [25].
We have reproduced a cluster analysis for MAS by estimating uncorrected general distances from a categorical character matrix, which depicted the MAS disease profile (Fig. 1a). Three major clusters were obtained, from which three main conditions are represented and might be considered as the “chaperones” of the autoimmunity namely SLE, AITD, and SS. The fourth cluster deserves further investigation since it corresponded to a series of young patients. However, this fourth cluster was characterized by the presence of CD. It is probably that other clusters exist and that ours reflect those related mainly to autoimmune rheumatic diseases. However, it is noteworthy that the above-mentioned clusters which were obtained in our own cases are similar to those previously reported from the literature [6, 17].
Aggregation of autoimmunity was observed in families of MAS patients (Table 3), mainly AITD and SLE. Two different estimates of excess risk are possible when dealing with family data; first, to treat the presence of the disease in relatives as a risk factor for the development of the disease in probands (type I RR) or to evaluate whether the relatives of an affected index person have an excess risk of disease compared to the general population (type II RR) [26]. Familial autoimmunity has been consistently associated with polyautoimmunity [17–19]. Unlike familial AD (i.e., the same AD in a nuclear family), familial autoimmunity uses the term “autoimmune” as a trait that encircles all accepted pathologies for which evidence suggests an autoimmune origin. There is increased evidence for aggregation of diverse ADs in families of patients with an AD [4].
ADs are entities frequently observed in genetically susceptible individuals in whom their clinical expression is modified by permissive and protective environments occurring over time. These are complex diseases, meaning that their inheritance do not follow a single-gene dominant or single-gene recessive Mendelian law, and thus that they are polygenic. The impact of genetic predisposition on susceptibility to ADs was first identified by the analysis of disease concordance rates in monozygotic twins (concordance rates ranged from about 15 to 57 %) [27]. The decrease in the concordance rates of siblings compared with the rate among monozygotic twins supports the presence of multiple genes contributing to the autoimmune phenotype onset. The common variants/multiple disease hypothesis has been invoked to underlie the pathogenesis of several complex diseases [28, 29]. This hypothesis states that complex phenotypes are not unique entities but are mosaics of common disease-specific alleles and non-disease-specific modifying alleles in the population influenced by a vast array of environmental factors [28]. Becker et al. [30] on the basis of previous AD linkage studies reported 18 common non-major histocompatibility complex loci clusters and hypothesized a shared genetic basis for the autoimmune trait. Since then, several reports have confirmed that autoimmune phenotypes might represent pleiotropic outcomes of non-specific disease genes [1, 31–34]. To date, in contrast to the numerous genome-wide association studies for common variants, there have been only a few studies reporting rare variants of large effect predisposing individuals to ADs [35].
We identified eight loci associated with MAS (Fig. 1c, d). All these loci harbor plausible genes for autoimmunity [34, 36]. For instance, 1p35.1 has been associated with Crohn's disease and SLE [37, 38]. 6p22.3 is related to the major histocompatibility complex and HLA genes, which is so far the strongest locus associated with ADs [36]. A vulnerability locus to MS maps to 7p15, a region that has been linked to multiple common ADs [28, 39]. 13q31.1 locus has been associated with MS and primary sclerosing cholangitis [40, 41], as well as with RA [42]. However, one should be cautious about common associations between loci and ADs because for some of them the association could be not certainly the same and even opposite as it is the case of some shared polymorphisms between RA and MS [43]. 14q32 is a common locus for AITD and T1D, two common associated diseases [44, 45]. 16p12.1 locus has been associated with SLE [46] and linked to RA [47]. 18q21.32 harbor CD226 gene which encodes the lymphocyte cell surface CD226 (DNAM1) receptor. A non-synonymous polymorphism in exon 7 of this gene, Gly307Ser (rs763361), has been identified as conferring risk to many ADs [48] including a MAS (PASII) [49]. Finally, we observed that 21q22 was also associated with MAS (Fig. 1c, d). This locus has been previously associated with SLE and T1D [50, 51].
It is noteworthy that not all ADs share the same genetic susceptibility. Thus, the genetic risk factors for ADs consist of two forms: those common to many ADs and those specific to a given disorder. Combinations of common and disease-specific alleles at HLA and non-HLA genes, in interaction with epigenetic and environmental factors over time (i.e., gluten, tobacco, Epstein–Barr virus, cytomegalovirus, and so on), will determine the final clinical autoimmune phenotype [1]. Yet only around 20 % of the inherited risk for ADs can be explained at present [1, 52]. Since MAS represent the best example of polyautoimmunity [17] as well as the effect of a single genotype on diverse autoimmune phenotypes, its study will provide important clues to elucidate the common mechanisms of ADs [53].
References
Anaya JM (2010) The autoimmune tautology. Arthritis Res Ther 12:147
Lorber M, Gershwin ME, Shoenfeld Y (1994) The coexistence of systemic lupus erythematosus with other autoimmune diseases: the kaleidoscope of autoimmunity. Semin Arthritis Rheum 24:105–113
Shoenfeld Y (1993) The kaleidoscope of autoimmunity. Autoimmunity 15:245–252
Anaya JM, Corena R, Castiblanco J, Rojas-Villarraga A, Shoenfeld Y (2007) The kaleidoscope of autoimmunity. Multiple autoimmune syndromes and familial autoimmunity. Expert Rev Clin Immunol 3:623–635
Humbert P, Dupond JL (1988) Les syndromes auto-immuns multiples. Ann Med Interne (Paris) 139:159–168
Anaya JM, Castiblanco J, Rojas-Villarraga A. (2008) The multiple autoimmune syndromes. In: Diagnostic criteria in autoimmune diseases. Shoenfeld Y et al. (eds.) Humana, Totowa, 65-69
Schmidt MB (1926) Eine biglandulare Erkrankung (Nebennieren und Schilddrüse) bei Morbus Addisonii. Verh Dtsch Ges Pathol: 21
Carpenter CC, Solomon N, Silverberg SG, Bledsoe T, Northcutt RC, Klinenberg JR et al (1964) Schmidt's syndrome (thyroid and adrenal insufficiency). A Review of the Literature and a report of fifteen new cases including ten instances of coexistent diabetes mellitus. Med (Baltimore) 43:153–180
Eisenbarth GS, Gottlieb PA (2004) Autoimmune polyendocrine syndromes. N Engl J Med 350:2068–2079
Anaya JM, Tobon GJ, Vega P, Castiblanco J (2006) Autoimmune disease aggregation in families with primary Sjögren’s syndrome. J Rheumatol 33:2227–2234
Madison WP, Madison DR (2011) Mesquite: a modular system for evolutionary analysis. Version 2.75. Available at http://mesquiteproject.org
Martin ER, Monks SA, Warren LL, Kaplan NL (2000) A test for linkage and association in general pedigrees: the pedigree disequilibrium test. Am J Hum Genet 67:146–154
Arcos-Burgos M, Castellanos FX, Konecki D, Lopera F, Pineda D, Palacio JD et al (2004) Pedigree disequilibrium test (PDT) replicates association and linkage between DRD4 and ADHD in multigenerational and extended pedigrees from a genetic isolate. Mol Psychiatry 9:252–259
Dudbridge F (2006) UNPHASED user guide. Technical Report 2006/5, MRC Biostatistics Unit, Cambridge, UK
Sheehan NJ, Stanton-King K (1993) Polyautoimmunity in a young woman. Br J Rheumatol 32:254–256
Somers EC, Thomas SL, Smeeth L, Hall AJ (2009) Are individuals with an autoimmune disease at higher risk of a second autoimmune disorder? Am J Epidemiol 169:749–755
Rojas-Villarraga A, Amaya-Amaya J, Rodriguez-Rodriguez A, Mantilla RD, Anaya JM (2012) Introducing polyautoimmunity: secondary autoimmune diseases no longer exist. Autoimmune Dis 254319
Rojas-Villarraga A, Toro CE, Espinosa G, Rodríguez-Velosa Y, Duarte-Rey C, Mantilla RD et al (2010) Factors influencing polyautoimmunity in systemic lupus erythematosus. Autoimmun Rev 9:229–232
Hudson M, Rojas-Villarraga A, Coral-Alvarado P, López-Guzmán S, Mantilla RD, Chalem P et al (2008) Polyautoimmunity and familial autoimmunity in systemic sclerosis. J Autoimmun 31:156–159
Gershwin ME, Selmi C, Worman HJ, Gold EB, Watnik M, Utts J et al (2005) Risk factors and comorbidities in primary biliary cirrhosis: a controlled interview-based study of 1032 patients. Hepatology 42:1194–1202
Amerio P, Di Rollo D, Carbone A, Auriemma M, Marra ME, De Remigis P et al (2010) Polyglandular autoimmune diseases in a dermatological clinical setting: vitiligo-associated autoimmune diseases. Eur J Dermatol 20:354–358
Mao ZF, Yang LX, Mo XA, Qin C, Lai YR, He NY et al (2011) Frequency of autoimmune diseases in myasthenia gravis: a systematic review. Int J Neurosci 121:121–129
Boelaert K, Newby PR, Simmonds MJ, Holder RL, Carr-Smith JD, Heward JM et al (2010) Prevalence and relative risk of other autoimmune diseases in subjects with autoimmune thyroid disease. Am J Med 123:183, e1-9
Alemany-Rodríguez MJ, Aladro Y, Amela-Peris R, Pérez-Viéitez MC, Reyes-Yáñez MP, Déniz-Naranjo MC et al (2005) Enfermedades autoinmunes y esclerosis múltiple. Rev Neurol 40:594–597
Venables PJ (2006) Mixed connective tissue disease. Lupus 15:132–137
Susser E, Susser M (1989) Familial aggregation studies. A note on their epidemiologic properties. Am J Epidemiol 129:23–30
Wandstrat A, Wakeland E (2001) The genetics of complex autoimmune diseases: non-MHC susceptibility genes. Nat Immunol 2:802–809
Becker KG (2001) The common genetic hypothesis of autoimmune/inflammatory disease. Curr Opin Allergy Clin Immunol 1:399–405
Yang Q, Khoury MJ, Friedman J, Little J, Flanders WD (2005) How many genes underlie the occurrence of common complex diseases in the population? Int J Epidemiol 34:1129–1137
Becker KG, Simon RM, Bailey-Wilson JE, Freidlin B, Biddison WE, McFarland HF et al (1998) Clustering of non-major histocompatibility complex susceptibility candidate loci in human autoimmune diseases. Proc Natl Acad Sci U S A 95:9979–9984
Anaya JM, Gómez L, Castiblanco J (2006) Is there a common genetic basis for autoimmune diseases? Clin Dev Immunol 13:185–195
Forabosco P, Bouzigon E, Ng MY, Hermanowski J, Fisher SA, Criswell LA et al (2009) Meta-analysis of genome-wide linkage studies across autoimmune diseases. Eur J Hum Genet 17:236–243
Invernizzi P, Gershwin ME (2009) The genetics of human autoimmune disease. J Autoimmun 33:290–299
Cotsapas C, Voight BF, Rossin E, Lage K, Neale BM, Wallace C et al (2011) Pervasive sharing of genetic effects in autoimmune disease. PLoS Genet 7:e1002254
Nejentsev S, Walker N, Riches D, Egholm M, Todd JA (2009) Rare variants of IFIH1, a gene implicated in antiviral responses, protect against type 1 diabetes. Science 324:387–389
Cho JH, Gregersen PK (2011) Genomics and the multifactorial nature of human autoimmune disease. N Engl J Med 365:1612–1623
Shugart YY, Silverberg MS, Duerr RH, Taylor KD, Wang MH, Zarfas K et al (2008) An SNP linkage scan identifies significant Crohn's disease loci on chromosomes 13q13.3 and, in Jewish families, on 1p35.2 and 3q29. Genes Immun 9:161–167
International MHC and Autoimmunity Genetics Network, Rioux JD, Goyette P, Vyse TJ, Hammarström L, Fernando MM et al (2009) Mapping of multiple susceptibility variants within the MHC region for 7 immune-mediated diseases. Proc Natl Acad Sci U S A 106:18680–18685
Coppin H, Ribouchon MT, Fontaine B, Edan G, Clanet M, Roth MP, French Multiple Sclerosis Genetics Group (2004) A vulnerability locus to multiple sclerosis maps to 7p15 in a region syntenic to an EAE locus in the rat. Genes Immun 5:72–75
Lorentzen AR, Melum E, Ellinghaus E, Smestad C, Mero IL, Aarseth JH et al (2010) Association to the Glypican-5 gene in multiple sclerosis. J Neuroimmunol 226:194–197
Karlsen TH, Franke A, Melum E, Kaser A, Hov JR, Balschun T et al (2010) Genome-wide association analysis in primary sclerosing cholangitis. Gastroenterology 138:1102–1111
Allen AS, Satten GA (2009) Genome-wide association analysis of rheumatoid arthritis data via haplotype sharing. BMC Proc 15(Suppl 7):S30
Sirota M, Schaub MA, Batzoglou S, Robinson WH, Butte AJ (2009) Autoimmune disease classification by inverse association with SNP alleles. PLoS Genet 5:e1000792
Tomer Y, Ban Y, Concepcion E, Barbesino G, Villanueva R, Greenberg DA et al (2003) Common and unique susceptibility loci in Graves and Hashimoto diseases: results of whole-genome screening in a data set of 102 multiplex families. Am J Hum Genet 73:736–747
Wallace C, Smyth DJ, Maisuria-Armer M, Walker NM, Todd JA, Clayton DG (2010) The imprinted DLK1-MEG3 gene region on chromosome 14q32.2 alters susceptibility to type 1 diabetes. Nat Genet 42:68–71
Trevisan GL, Rassi DM, Baião AM, Sandrin-Garcia P, Mello SS, Tamia-Ferreira MC et al (2006) Using cDNA microarrays to identify human CD19(+) B cell gene products (ESTs) originated from systemic lupus erythematosus susceptibility loci. Autoimmun Rev 5:319–323
Criswell LA, Chen WV, Jawaheer D, Lum RF, Wener MH, Gu X et al (2007) Dissecting the heterogeneity of rheumatoid arthritis through linkage analysis of quantitative traits. Arthritis Rheum 56:58–68
Maiti AK, Kim-Howard X, Viswanathan P, Guillén L, Qian X, Rojas-Villarraga A et al (2010) Non-synonymous variant (Gly307Ser) in CD226 is associated with susceptibility to multiple autoimmune diseases. Rheumatol (Oxford) 49:1239–1244
Gan EH, Mitchell AL, Macarthur K, Pearce SH (2011) The role of a nonsynonymous CD226 (DNAX-accessory molecule-1) variant (Gly 307Ser) in isolated Addison's disease and autoimmune polyendocrinopathy type 2 pathogenesis. Clin Endocrinol (Oxf) 75:165–168
Clancy RM, Marion MC, Kaufman KM, Ramos PS, Adler A, International Consortium on Systemic Lupus Erythematosus Genetics et al (2010) Identification of candidate loci at 6p21 and 21q22 in a genome-wide association study of cardiac manifestations of neonatal lupus. Arthritis Rheum 62:3415–3424
Cooper JD, Walker NM, Smyth DJ, Downes K, Healy BC, Todd JA et al (2009) Follow-up of 1715 SNPs from the Wellcome Trust Case Control Consortium genome-wide association study in type I diabetes families. Genes Immun 10(Suppl 1):S85–94
Manolio TA, Collins FS, Cox NJ, Goldstein DB, Hindorff LA, Hunter DJ et al (2009) Finding the missing heritability of complex diseases. Nature 461:747–753
Anaya JM (2012) Common mechanisms of autoimmune diseases (the autoimmune tautology) Autoimmun Rev Feb 12. doi.10.1016/j.autrev.2012.02.002
Acknowledgments
We thank all patients and participants. This study was financed by Colciencias (122254531722), the School of Medicine and Health Sciences at the Universidad del Rosario, Bogota, Colombia, and Marshfield Foundation, WI, USA.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Anaya, JM., Castiblanco, J., Rojas-Villarraga, A. et al. The Multiple Autoimmune Syndromes. A Clue for the Autoimmune Tautology. Clinic Rev Allerg Immunol 43, 256–264 (2012). https://doi.org/10.1007/s12016-012-8317-z
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12016-012-8317-z