Abstract
Pluripotent stem cells have great potential for developmental biology and regenerative medicine. Embryonic stem cells, which are obtained from blastocysts, and induced pluripotent stem cells, which are generated by the reprogramming of somatic cells, are two main types of pluripotent cells. It is important to understand the regulatory network that controls the pluripotency state and reprogramming process. Various types of noncoding RNAs (ncRNAs) have emerged as substantial components of regulatory networks. The most studied class of ncRNAs in the context of pluripotency and reprogramming is microRNAs (miRNAs). In addition to canonical microRNAs, other types of small RNAs with miRNA-like function are expressed in PSCs. Another class of ncRNAs, long ncRNAs, are also involved in pluripotency and reprogramming regulation. Thousands of ncRNAs have been annotated to date, and a significant number of the molecules do not have known function. In this review, we briefly summarized recent advances in this field and described existing genome-editing approaches to study ncRNA functions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Protein-coding genes constitute a small part of most eukaryotic genomes. In humans, only 1,2% of DNA encode proteins. The other large part of the genome is referred to as “junk DNA”, except some housekeeping RNA molecules and regulatory and structural elements. Initially, the term “junk DNA” was applied by Ohno for pseudogenes, and subsequently, this term was extended for noncoding DNA without known function [1, 2]. The development of massive parallel sequencing technologies allows us to analyse transcriptomes at a deeper level. The studies revealed that most of the metazoan genome is transcribed [3]. Two major classes of noncoding RNA (ncRNA) are defined based on their length: long ncRNA (lncRNA), which is more than 200 nt in length, and small ncRNA, which is less than 200 nt in length. lncRNAs are usually classified according to the location in the genome [4]. However, information about the location does not provide the information about lncRNA function. Small RNAs, in turn, are classified based on their function in the most cases. Deep sequencing studies of small RNA fraction and subsequent analyses have discovered a high number of small RNA types. Among these diverse RNAs, small ncRNAs, which are involved in the regulation of gene expression, are of particular interest. Distinct classes, including microRNAs (miRNAs), piwi-interacting RNAs (piRNAs), sno-derived RNAs (sdRNAs), endogenous small interfering RNAs (siRNAs), and endogenous small hairpin RNAs (shRNAs), are referred to as small regulatory ncRNAs (Fig. 1) [5, 6]. Some of these molecules, such as piRNAs, are expressed in specific cell types and involved in the regulation of transposon expression in germ cells [7]. Other molecules, such as miRNAs, regulate gene expression in various cell types and define cell identity.
Biogenesis pathways of miRNA and miRNA-like molecules. a Canonical miRNA are grouped into clusters, transcribed by RNA-pol II and then processed by Drosha and Dicer ribonucleases. b Endogenous shRNAs are transcribed as small hairpins. Their biogenesis is Drosha independent. c The mirtron pathway. The miRNAs are encoded into short introns, spliced and debranched. Ldbr – lariat debranching enzyme. d snoRNA-derived pathway. e Endogenous siRNAs are formed by the transcription of inverted repeats. In mouse ESCs, siRNAs are predominantly formed from B1/Alu SINE elements [6]. f tRNA can form alternative hairpin structures that are cleaved by Dicer
Increasing evidence suggests that ncRNAs play important roles in different biological processes during development. Embryonic development starts from the totipotent zygote, which divides to form a blastocyst. The inner cell mass of the blastocyst consists of pluripotent stem cells (PSCs) that can self-renew and differentiate into any cell type in an organism. These embryonic stem cells (ESCs) can be isolated and indefinitely maintained under proper culture conditions [8, 9]. Another type of PSCs is reprogrammed from somatic cells by the overexpression of certain transcription factors (Oct4, Sox2, Klf-4, c-Myc) and referred to as induced pluripotent stem cells (iPSCs) [10]. PSCs have great potential for regenerative medicine and genetic therapy, especially iPSCs, which can be derived from the patient. Research in the field of pluripotency and self-renewal regulation is required for the efficient derivation and cultivation of PSCs. Significant progress has been made in the investigation of regulatory mechanisms using transcriptomic, epigenomic, proteomic, and metabolomic approaches. An important regulatory role was also established for many ncRNAs. In this review, we highlighted recent advances in the field of pluripotency and reprogramming regulation by ncRNAs.
MicroRNAs
miRNAs in Pluripotent Stem Cells
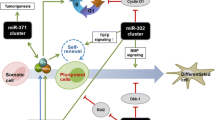
The most studied class of small ncRNAs is miRNA. Canonical miRNAs are 20–23 nt in length and transcribed by RNA-polymerase II into long pri-miRNA. RNase III Drosha with Dgcr8 forms a Microprocessor complex that cleaves pri-miRNAs to short hairpins termed pre-miRNAs [11]. These pre-miRNAs are exported to the cytoplasm and processed by RNase III Dicer to an RNA duplex. Then, the duplex is bound by Ago protein to form a RNA-induced silencing complex (RISC). One strand of this miRNA duplex is degraded, while the other strand functions as a guide for binding to the target mRNA. The biogenesis of miRNAs was excellently described in several previous reviews [12, 13]. miRNAs play important roles in the regulation of cell state and different processes, including self-renewal, differentiation, and reprogramming. Knockout or knockdown of miRNA processing machine components in human and mouse ESCs results in proliferation and differentiation defects [14,15,16,17]. Expression analysis revealed that the majority of miRNAs in mouse ESCs are related to six genomic loci [18]. The most abundant of these molecules are miRNAs from miR-290-295 (miR-371-373 in humans) and miR-17-92 clusters. Members of the miR-290-295 cluster contribute to approximately 70% of all miRNAs expressed in ESCs [19, 20]. These miRNAs predominantly share the same seed sequence AAGUGC and are named ESC-specific cell cycle-regulating (ESCC) miRNAs [17]. ESCC miRNAs are involved in the pluripotency regulation network, and their promoters are occupied by core pluripotency factors [20]. Core pluripotency factors also occupy the promoters of differentiation-related miRNAs to repress their expression and prevent differentiation. For example, miR-145 represses the pluripotency of human ESCs by directly targeting OCT4, SOX2 and KLF4 mRNAs, whereas the expression of miR-145 is repressed by OCT4 [21]. These links represent a double-negative feedback loop of pluripotency regulation. miRNAs from the miR-302-367 cluster share the same seed sequence as ESCC miRNAs, and this cluster is highly expressed in the mouse epiblast stem cells (EpiSCs) and human ESCs [22]. The molecular features of human ESCs are similar to those of mouse EpiSCs [23]. Another miRNA cluster with the AAGUGC seed sequence is located on human chromosome 19 and includes miRNAs of the miR-519/520 series. This cluster is primate-specific and expressed in human ESCs, placenta, and cancer cells [24,25,26,27].
Nucleotides 2–8 of mature miRNA represent a seed sequence that determines binding to the mRNA target [28]. Interestingly, different miRNA clusters with the same seed sequence characterize different cell types [29]. These miRNA clusters presumably may realize the same functions. However, this fact may be explained by various types of miRNA-mRNA interactions, except classic seed base pairing, which have previously been detected [30]. Additional base pairing of the 3′ miRNA end with target mRNA leads to the specification of targets of different miRNAs with the same seed [31]. These modes of miRNA actions may be responsible for different targets of miRNAs from miR-290-295 and miR-302-367 clusters. Currently, ESCC miRNAs have been implicated in the regulation of numerous processes, including the cell cycle, apoptosis, cellular metabolism, DNA methylation, and polycomb-mediated silencing of differentiation genes [17, 32,33,34,35].
In addition to mouse and human ESCs, miRNA expression was analysed in rat ESCs and iPSCs [36], pig iPSCs [37], and rabbit ES-like cells [38]. Rat ESCs and iPSCs are characterized by miR-290-295 cluster expression, and the overall expression pattern was similar to that of mouse ESCs. Pig miRNA expression was analysed in two types of iPSCs that differ by culture conditions. One iPSC type was cultured under conditions suitable for mouse ESCs, and the other iPSC type was cultured under human ESCs conditions. The miRNA expression pattern of the two types of pig iPSCs differs. However, both iPSC types express the miR-302 cluster. This miRNA cluster is also expressed in Rabbit ES-like cells, which are more similar to human than to mouse ESCs.
miRNA expression in mouse ESCs depends on the culture conditions. ESCs cultured with inhibitors of MEK1/2 (PD0325901) and GSK3 (CHIR99021) kinases (2i ES cells) represent a ground pluripotency state in contrast to serum-cultured ESCs [39,40,41,42]. ESCC miRNAs are expressed at similar levels in 2i and serum-cultured ESCs [43]. Notably, some differentiation-inducing miRNAs, such as miR-26a-5p, miR-99-5p, miR-218-5p and let-7 family miRNA, are upregulated in 2i ESCs. However, these miRNAs cannot silence self-renewal without the addition of serum. A comprehensive bioinformatics analysis of the miRNA-mRNA interaction network in serum-cultured ESCs revealed that one of the primary miRNA functions is the repression of differentiation signals [19, 44]. The inhibitors PD0325901 and CHIR99021 influence the maturation of miRNAs through the inhibition of Microprocessor complex [45, 46]. Particularly, the inhibition of GSK3 kinase induces the cytoplasmic localization of Drosha, and the inhibition of MEK1/2 decreases the stability of Dgcr8. In addition to miRNA processing, Dgcr8 is also involved in the splicing of Tcfl7 mRNA [47]. Tcfl7 is required for the activation of lineage-specific differentiation programmes. Despite impaired miRNA processing, the lack of differentiation signals from serum and the inhibition of MEK1/2 presumably lead to stable ground state ESC culture.
miRNAs and Reprogramming
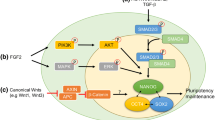
The classic reprogramming factors are Oct4, Sox2, Klf4, and c-Myc [10]. Different combinations of these and other pluripotency factors are used to obtain iPSCs [48]. Somatic cells can also be reprogrammed to a pluripotent state using small molecules [49]. The search for additional methods to obtain iPSC has not stopped. The first evidence of miRNA-based reprogramming was reported in 2008. Two human cancer cell lines were reprogrammed to an ES-like state using the expression of miR-302-367 cluster [50]. Subsequently, this miRNA cluster was used to reprogramme human and mouse fibroblasts to a pluripotent state with higher efficiency compared to Yamanaka factors [51]. Another group used miR-200c, members of the miR-302-367 cluster, and miR-369 mature miRNA mimics to reprogramme human and mouse adipose stromal cells and human dermal fibroblasts [52]. The advantage of the latter approach is the absence of lenti- or retroviral vectors that integrate into the genome. miRNA can also be used in combination with Yamanaka factors to increase the reprogramming efficiency and obtain a homogeneous population of iPSCs. The miRNAs miR-291, miR-294, and miR-295 substitute c-Myc in the reprogramming of mouse cells [53]. Moreover, the efficiency of such an approach is higher than that of the classic approach. Moreover, uniform populations of iPSCs are formed. Reprogramming to a pluripotent state is a dynamic process accompanying changes in gene expression, including ncRNAs, proteome, epigenome, and metabolome [54,55,56,57,58]. ESCC miRNAs and other miRNAs with the AAGUGC seed sequence increase reprogramming efficiency through different mechanisms, such as the regulation of the cell cycle, epigenetics, transcription factors, metabolic pathways, vesicular transport and the mesenchymal-to-epithelial transition (MET) [59,60,61,62]. MET is a key process of the initiation stage of reprogramming [63]. The inhibition of its reverse process, the epithelial-to-mesenchymal transition (EMT), enhances the reprogramming of somatic cells [64]. Various miRNAs promote MET by targeting different genes involved in this process. Some miRNAs, such as miR-106b, miR-93, members of the miR-302-367 cluster, and miR-372, inhibit EMT and promote MET by targeting TGFBR2 [60,61,62]. Members of the miR-200 family have been shown to target ZEB transcription factors, which are significant modulators of EMT [63, 65,66,67]. miRNAs belonging to the miR-181 family stimulate the reprogramming of mouse embryonic fibroblasts at the early stages by promoting the initiation phase [68]. This effect is achieved by the activation of Wnt and the repression of TGF-β signalling pathways.
The regulation of core pluripotency factors is a crucial process that influences the reprogramming efficiency. For example, transcription factor NR2F2 negatively regulates OCT4 expression in human cells, but during reprogramming, this factor is downregulated by miR-302 [69]. The miR-34 family of miRNAs provides a barrier for reprogramming by the repression of Nanog, Sox2, and Mycn expression [70]. The modulation of cellular physiology also affects reprogramming [71]. Cells change metabolism from oxidative phosphorylation to glycolysis during reprogramming [57, 72, 73]. Pyruvate kinase Pkm2 is involved in glycolysis and is highly expressed in pluripotent cells [33]. The expression of Pkm2 is regulated by the miR-290-295 cluster, which activates the Mbd2-Myc-Pkm2 axis through the downregulation of Mbd2. miR-369 stabilizes the translation of the splicing factor HNRNPA2B1, which is required for Pkm2 expression [74]. miR-31 plays a significant role in altering mitochondrial function by targeting succinate dehydrogenase complex subunit A [75].
A number of known miRNAs have been shown to mediate reprogramming. For example, miR-29b, miR-138, miR-19a/b, and miR-6539 increase reprogramming efficiency [76,77,78,79]. miR-34a, miR-29a, miR-21, let-7 family, miR-212, miR-132, miR-145, miR-27a, miR-24, miR-134 inhibit reprogramming [80,81,82,83,84,85]. We summarized the miRNAs implicated in reprogramming in the Table 1. However, iPSCs can be derived from Dgrc8-deficient mouse fibroblasts and neural stem cells, although with decreased efficiency compared to wild-type [86]. This finding suggests that canonical miRNAs are dispensable for reprogramming. Other small ncRNA or non-canonical miRNA species, such as mirtrons, for example, may compensate for the absence of canonical miRNAs. The miRNA-like molecules may be generated through different pathways, some of which are Drosha or Dicer independent [6, 87]. Further studies of individual miRNAs and small ncRNAs with miRNA-like functions may shed light on this problem.
IsomiRs
The pre-miRNA hairpins can be processed with some alterations, resulting in the addition or deletion of several nucleotides on the 5′ or 3′ ends of mature miRNA [88]. miRNAs may also be subjected to posttranscriptional modification by A-to-I editing [89]. A-to-I editing is performed by adenosine deaminases acting on RNA (ADAR) [90]. Inosines are recognized as guanosines, and the change appears as A-to-G. These mature molecules with 5′ or 3′ shifts or post-transcriptional edits constitute a minor fraction of miRNAs, called isomiRs. Evidence suggests that isomiRs are functional and important for the evolution of miRNAs [91, 92]. miRNA and corresponding isomiRs regulate genes that are enriched in the same functional pathway [93]. However, changes in the 5′ end of the miRNA may lead to a different pool of mRNA targets. For example, miR-302a-5p, which is expressed in human ESCs, has isomiR miR-302a-5p (+ 3) with three distinct 5′ end nucleotides shifted to the 3′ end [94, 95]. OTX2 is a miR-302a-5p target, whereas miR-302a-5p (+ 3) targets TSC1 expression and does not regulate OTX2. A total of 19 of the 24 pluripotency-associated miRNAs from miR-290-295 and miR-302-367 clusters are processed in different isomiRs in mouse ESCs [96]. These findings raise questions regarding which of the produced isomiRs are actually functional and how the pool of targets differs compared to classic sequences. Further studies may shed light on the answers to these questions and elucidate isomiR functions in PSCs.
MicroRNA-offset-RNAs
Another type of small RNA molecules produced from pre-miRNA hairpins is microRNA-offset-RNA (moRNA). moRNAs are located adjacent to mature miRNAs in conserved regions across species. First, moRNAs was identified in human small RNA sequencing data in 2009 [97]. Further, 326 moRNAs were found in human ESCs, whereas a significantly lower number of moRNAs were found in fibroblasts [98]. Transfection of the moRNA-103a-2-3p mimic results in the downregulation of 538 genes, and a substantial part of the genes have seed matches in the 3′UTR of moRNA-103a-2-3p. Another study showed that moRNA-21 is functional, and its function depends on the seed sequence and requires Ago2 for gene repression [99]. These studies suggest that this type of small ncRNA regulates gene expression through a miRNA-like pathway.
snoRNAs and sno-derived RNAs
snoRNAs are small ncRNAs of 60–140 nt in length. Two classes of snoRNAs exist: H/ACA box and C/D box. snoRNAs function as guides for ribosomal RNA modifications. H/ACA box snoRNAs promote 2′-O-ribose methylation in complex with fibrillarin, and C/D box snoRNAs promote pseudouridylation in complex with dyskerin [100]. Fibrillarin is essential for early development, and homozygous mutations lead to death prior to implantation [101]. A decreased level of fibrillarin affects the expression of snoRNAs encoded in introns. Indeed, the disruption of one snoRNA may lead to significant changes in cellular homeostasis. For example, the inhibition of H/ACA box snoRNA 7A by antisense oligonucleotide suppresses the proliferation and self-renewal of umbilical cord blood-derived mesenchymal stem cells [102]. Dyskerin ribonucleoprotein complex (DKC1) is involved in the transcriptional regulation of core pluripotency genes as an OCT4/SOX2 coactivator [103]. Several snoRNAs are differentially expressed between mouse ESCs and their differentiated derivatives [104]. ESC snoRNAs may guide DKC1 to gene enhancers, and the disruption of the complex presumably leads to changes in the transcriptional gene network. Additional studies will help to understand the functions of snoRNAs in self-renewal and pluripotency regulation.
In addition to isomiRs and moRNAs, which are produced from the pre-miRNA hairpin, functional miRNA-like molecules can be derived from snoRNAs [105,106,107]. These small ncRNAs are called sno-derived RNAs (sdRNAs). snoRNAs can be processed into sdRNAs that perform posttranscriptional gene silencing. However, the function of sdRNAs in pluripotency has not yet been investigated, and future studies are required to analyse the sdRNA expression in PSCs and identify pathways that are regulated by this type of small ncRNAs.
lncRNAs
lncRNAs Expression and Function in PSCs
Another enormous class of noncoding RNAs is lncRNAs. lncRNAs are more than 200 nt in length, transcribed by RNA pol II, polyadenylated, capped, and often spliced from pre-lncRNA. lncRNAs are involved in gene expression regulation in different biological processes in embryonic development [108]. Progress in massively parallel sequencing technologies enabled the identification of thousands of lncRNAs. Guttman et al. identified approximately 1600 large intergenic ncRNAs (lincRNAs) by analysis of ChIP-seq data of trimethylated lysines 4 and 36 of histone H3 in four cell types [109]. Further, this group developed the algorithm of transcriptome reconstruction, Scripture, and identified 1140 novel lincRNAs using RNA-seq data [110]. Approximately 1500 very large intergenic ncRNAs (vlincRNAs) were identified in human cells [111]. To date, more than 14,000 human lncRNAs transcripts were manually annotated by the GENCODE consortium [112].
A loss-of-function study revealed that 137 lincRNAs are involved in the transcriptional network of mouse ESCs, and 26 of these molecules regulate the maintenance of the pluripotent state [113]. In turn, the expression of these lincRNAs is regulated by core pluripotency factors Oct4, Sox2, Nanog, c-Myc, and Klf4. In another study, 20 lincRNAs were functionally verified as pluripotency maintaining players [114]. Some lncRNA molecules expressed in ESCs are required for the repression of differentiation-related and lineage-specific genes [113, 115,116,117,118,119]. Knockdown of such lncRNAs is accompanied by the loss of pluripotency state and activation of lineage-specific markers. For example, the knockdown of AK028326 (Gomafu/Miat) results in the upregulation of trophoblast-specific transcripts [117]. Loss of Panct1 activates expression of endoderm and ectoderm markers [115]. In contrast to miRNAs, lncRNAs can regulate gene expression at the transcriptional level. The inactivation of the entire X-chromosome is initiated by the well-studied lincRNA Xist, which interacts with polycomb repressive complex 2 (PRC2) [120, 121]. The hypothesis that other lincRNAs can represent a scaffold for chromatin modifying enzymes was confirmed by a number of studies. A significant part of human lincRNAs in various cell types is physically associated with the PRC2 complex [122]. Further, the immunoprecipitation of RNA–protein complexes showed that ESC lincRNAs interact with chromatin complexes, which are involved in the “reading”, “writing”, and “erasing” of epigenetic marks [113]. Some examples of lincRNAs function in ESCs were investigated in the details. tsRMST lncRNA interacts with the pluripotency factor NANOG and the component of PRC2 repressive complex SUZ12. This interaction leads to the repression of differentiation-related transcription factors and non-canonical Wnt ligand WNT5A [118, 119]. lncRNA-ES1 and lncRNA-ES3 contribute to the suppression of differentiation through interactions with SOX2 and SUZ12 [116]. A number of lncRNAs found in ESCs, bind to WDR5 [123]. WDR5 is a subunit of the MLL complex, which implements histone H3 lysine 4 trimethylation and promotes a pluripotency state.
At the post-transcriptional level, lncRNAs can act as a competitive endogenous RNA or “sponges” for miRNAs to block its function. In mouse ESCs, lncRNA AK048794 binds to miR-592 and negatively modulates the expression of Oct4, Sox2 and Nanog through targeting FAM91A1 mRNA [124]. lincRNA-ROR (Regulator of Reprogramming) promotes pluripotency by binding to miR-145, the negative regulator of OCT4, SOX2, KLF4, and NANOG expression [21, 125]. Human lincRNA HPAT5 modulates the expression of the let-7 miRNA family and prevents differentiation [126].
The term ncRNA implies that RNA molecules do not encode proteins. Nevertheless, a great number of small open reading frames (ORF) were found in lncRNAs using a bioinformatics approach [127]. Verification of the translation and function of the small ORF is a difficult process. The high-throughput analysis of ribosome-protected RNA fragments (ribosome profiling or Ribo-seq) will help to identify novel and confirm the translation of some predicted small regulatory peptides encoded by lncRNAs [128]. In mouse ESCs, approximately half of lncRNAs display ribosome profiling signals [129]. Additional studies suggest that the part of lncRNAs are actually translated [130], and other studies have shown that the majority of lncRNAs do not have coding potential [131, 132]. In human cancer cells, 510 of the 1189 expressed lncRNAs have ORFs according to ribosome profiling analysis, yet their translated peptides are likely not functional [133]. Nevertheless, several studies have demonstrated the translation of small polypeptides from lncRNAs and analysed their functions [134, 135]. One of these small polypeptides is CRNDEP, which is translated from human lncRNA CRNDE in highly proliferative tissues. This small peptide is presumably involved in the regulation of cell proliferation [135]. Notably, the mouse orthologue of CRNDE linc1399 is also associated with the maintenance of a pluripotent state [113, 135]. Evolutionary conservation supports the hypothesis of the functionality of this lncRNA-encoded polypeptide. Thus, combining transcriptome, translatome, and proteomic studies may reveal lncRNAs that encode proteins, and shed light on the question of ncRNA translation.
lncRNAs and Reprogramming
The involvement of the lncRNAs in the reprogramming process has been established in several studies. The expression of lncRNAs was analysed during the reprogramming of mouse fibroblasts to a pluripotent state [136, 137]. Approximately 1200 lncRNAs were differentially expressed at different stages during the process [136]. Another analysis showed that approximately 300 lncRNAs were activated in transitional cells and/or iPSCs, and some of these molecules are involved in the suppression of lineage-specific genes and the regulation of the metabolic gene expression [137]. However, the first functional example of lncRNA involved in the reprogramming process, called lincRNA-ROR, was previously established [138]. Knockdown of lincRNA-ROR decreased reprogramming efficiency, and its overexpression resulted in increased iPSC colony formation. One of the established functions of lincRNA-ROR is the inhibition of the p53-mediated cell cycle arrest and apoptosis [139]. iPSCs show properties similar to ESCs. Fully pluripotent iPSC lines can contribute to the development of tetraploid blastocysts and generation of full-term mice [140]. However, some iPSC clones failed to pass this test. RNA-seq analysis of genetically identical iPSC and ESC lines revealed that the aberrant silencing of the lncRNAs Gtl2 and Rian distinguishes iPSC clones from ESCs and restricts their developmental potential [141]. Notably, the expression of a few transcripts located in the imprinted Dlk1-Dio3 cluster determines cell potential.
A substantial amount of human lincRNAs is enriched in transposable elements (TE), particularly endogenous retroviruses (ERVs) [142]. Distinct classes of ERVs are expressed at different stages in human preimplantation embryos [143]. HERVH was established as the most highly expressed class of endogenous retroviruses in human PSCs [144]. Further, the expression of HERVH was linked to a naïve pluripotency state [145]. Currently, the functions of many individual TE-derived lincRNAs remain elusive due to their repetitive structures. Nevertheless, recently, three human lincRNAs were functionally studied. HPAT2, 3, and 5 were implicated in the pluripotency regulation network and reprogramming of human fibroblasts [126]. We summarized the lncRNAs implicated in pluripotency regulation and reprogramming in the Table 2.
Genome-Editing Technologies for Studying ncRNA
Thousands of ncRNAs that belongs to different classes have been annotated to date, but few of these molecules have been functionally studied, particularly in the context of pluripotency. In the case of miRNA and miRNA-like molecules, the crucial step is to find target mRNAs. A large number of the ESCC miRNA targets have been established to date. However, additional studies are required to elucidate the entire pool of the ESCC miRNAs targets, discriminate the functions of different miRNAs with similar sequences, and find targets of other miRNAs expressed in ESCs and involved in reprogramming. One miRNA can regulate hundreds of genes, thus identifying target genes and conducting functional analyses are challenging [28]. The existing computational prediction algorithms generate many false positive and false-negative results [146]. Large-scale studies using HITS-CLIP or PAR-CLIP approaches with the overexpression of certain miRNAs will help to identify targets of non-abundant miRNAs [147, 148]. However, miRNA overexpression may introduce biases in the results. To confirm miRNA-mRNA interactions, luciferase reporter constructs are typically used [149]. The main drawback of this method is the non-physiological conditions caused by transfection of exogenous reporters and miRNA mimics. In addition, this approach is often performed in a heterogeneous system, such as HEK293 cells, which are easy to transfect. These drawbacks limit the use of the luciferase approach to understand the function of this miRNA binding site or not in certain cell types. Recently, a genome-editing approach was utilized to analyse the functionality of miRNA target sites [150]. Candidate target sites can be replaced with molecular barcodes using homology-directed repair induced by the CRISPR/Cas9 system. Then, the effect of the microRNA target site mutation can be analysed by quantitative PCR.
Genetic knockdown or knockout approaches may be utilized to understand miRNA functions. The widespread method of miRNA knockdown is the delivery of antisense inhibitors. miRNA antisense inhibitors may be chemically synthesized or expressed from transgenes, which contain tandem repeats of the miRNA complementary sequence [151, 152]. Using chemically synthesized inhibitors is expensive and suitable for short-term studies. The knockout of miRNA may be realized using genome-editing methods, such as TALENs or CRISPR/Cas9 systems [153,154,155]. The expression of miRNA may be disrupted by introducing indels in the processing sites, which impede miRNA biogenesis, or by deleting the region encoding target miRNA genes or whole clusters (Fig. 2a). The expression of Cas9 and two sgRNA results in the deletion of regions up to several megabases between sgRNA sites [156]. This approach allows the stable knockout of protein-coding genes and various ncRNAs, including lncRNAs, both in vitro and in vivo (Fig. 2b). For example, a locus of approximately 6,5 kb, encoding the HPAT5 lincRNA, was deleted using the CRISPR/Cas9 system to obtain a knockout cell line and investigate HPAT5 function in reprogramming [126]. Mice with a 23-kb deletion of the Rian lncRNA gene were obtained by the injection of Cas9 protein and two single guide RNAs into one-cell stage mouse embryos [157]. The CRISPR/Cas9 system was also used to study ncRNAs, such as miR-21, miR-29a, lncRNA-21A, UCA1, and AK023948, in human cancer cell lines [158]. Moreover, the method of ncRNA knockout was adapted for high-throughput screening with lentiviral paired-guide RNA library [159]. However, the removal of large regions in the genome has potential pitfalls. These regions may contain some regulatory elements, or genes of small ncRNAs, such as miRNAs, snoRNAs, etc. Additionally, the homozygous deletion of large regions remains challenging [155]. Another way to eliminate ncRNA expression is the removal of the promoter region, which is typically shorter than the entire lncRNA (Fig. 2c).
ncRNA knockout and knockdown using the CRISPR/Cas9 system. a miRNA cluster, single miRNA or other small RNA genes can be deleted using CRISPR/Cas9 with two sgRNAs, which flank the target region. b Knockout of lncRNAs can be achieved using the same approach. However, the deletion of a larger region is more difficult than the deletion of a smaller region. c Alternatively, promoters containing the transcription start site can be deleted to decrease lncRNA expression. d Catalytically inactive dCas9 fused with the KRAB repressor domain can be used to knockdown ncRNAs
Several methods have been used to avoid genetic alterations in the targeted region to obtain knockdown. For example, transcriptional repressors based on TALE proteins fused with the KRAB domain are utilized [154, 160]. The CRISPR interference system can also be used to decrease ncRNA expression. In this method, sgRNA targeting the 5′ region or gene promoter is coexpressed with catalytically inactive dCas9 protein [161]. The binding of this complex to the nontemplate DNA strand blocks the elongation of transcription or prevents initiation. To increase the efficiency of gene knockdown, dCas9 can be fused with the KRAB domain (Fig. 2d) [162].
Conclusion
PSCs are a unique model for studying early development, modelling hereditary diseases, and drug screening. PSCs differentiate into many cell types of the human body, which makes these cells indispensable for regenerative medicine. Knowledge of pluripotency regulation mechanisms is required for proper culture conditions and the efficient derivation of patient-specific iPSCs, which are suitable for subsequent differentiation and transplantation. High-throughput studies revealed that the pluripotency regulation network is complex, and ncRNAs play important roles in its maintenance. The first step is identifying ncRNA expression patterns. Large-scale loss-of-function studies are the next step after massive parallel sequencing to reveal substantial ncRNAs in pluripotency regulation. Finally, functional studies are required to understand the functions of individual ncRNAs. Currently, various classes of ncRNAs have been identified, and most part of these molecules are “dark matter” of the genome. Future studies will shed light on the world of ncRNAs and their roles in pluripotency regulation.
References
Comings, D. E. (1972). The structure and function of chromatin. Advances in Human Genetics, 3, 237–431.
Ohno, S. (1972). So much “junk” DNA in our genome. Brookhaven Symposia in Biology, 23, 366–370.
Djebali, S., Davis, C. A., Merkel, A., et al. (2012). Landscape of transcription in human cells. Nature, 489, 101–108.
Kung, J. T., Colognori, D., & Lee, J. T. (2013). Long noncoding RNAs: past, present, and future. Genetics, 193, 651–669.
Mattick, J. S., & Makunin, I. V. (2005). Small regulatory RNAs in mammals. Human molecular genetics, 14 Spec No 1, R121-132.
Babiarz, J. E., Ruby, J. G., Wang, Y., Bartel, D. P., & Blelloch, R. (2008). Mouse ES cells express endogenous shRNAs, siRNAs, and other Microprocessor-independent, Dicer-dependent small RNAs. Genes & Development, 22, 2773–2785.
Aravin, A. A., Hannon, G. J., & Brennecke, J. (2007). The Piwi-piRNA pathway provides an adaptive defense in the transposon arms race. Science, 318, 761–764.
Evans, M. J., & Kaufman, M. H. (1981). Establishment in culture of pluripotential cells from mouse embryos. Nature, 292, 154–156.
Thomson, J. A., Itskovitz-Eldor, J., Shapiro, S. S., et al. (1998). Embryonic stem cell lines derived from human blastocysts. Science, 282, 1145–1147.
Takahashi, K., & Yamanaka, S. (2006). Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell, 126, 663–676.
Gregory, R. I., Yan, K. P., Amuthan, G., et al. (2004). The Microprocessor complex mediates the genesis of microRNAs. Nature, 432, 235–240.
Kim, V. N., Han, J., & Siomi, M. C. (2009). Biogenesis of small RNAs in animals. Nature Reviews Molecular Cell Biology, 10, 126–139.
Winter, J., Jung, S., Keller, S., Gregory, R. I., & Diederichs, S. (2009). Many roads to maturity: microRNA biogenesis pathways and their regulation. Nature Cell Biology, 11, 228–234.
Kanellopoulou, C., Muljo, S. A., Kung, A. L., et al. (2005). Dicer-deficient mouse embryonic stem cells are defective in differentiation and centromeric silencing. Genes & Development, 19, 489–501.
Murchison, E. P., Partridge, J. F., Tam, O. H., Cheloufi, S., & Hannon, G. J. (2005). Characterization of Dicer-deficient murine embryonic stem cells. Proceedings of the National Academy of Sciences of the United States of America, 102, 12135–12140.
Qi, J., Yu, J. Y., Shcherbata, H. R., et al. (2009). microRNAs regulate human embryonic stem cell division. Cell cycle (Georgetown, Tex.), 8, 3729–3741.
Wang, Y., Baskerville, S., Shenoy, A., Babiarz, J. E., Baehner, L., & Blelloch, R. (2008). Embryonic stem cell-specific microRNAs regulate the G1-S transition and promote rapid proliferation. Nature Genetics, 40, 1478–1483.
Calabrese, J. M., Seila, A. C., Yeo, G. W., & Sharp, P. A. (2007). RNA sequence analysis defines Dicer’s role in mouse embryonic stem cells. Proceedings of the National Academy of Sciences of the United States of America, 104, 18097–18102.
Leung, A. K., Young, A. G., Bhutkar, A., et al. (2011). Genome-wide identification of Ago2 binding sites from mouse embryonic stem cells with and without mature microRNAs. Nature Structural & Molecular Biology, 18, 237–244.
Marson, A., Levine, S. S., Cole, M. F., et al. (2008). Connecting microRNA genes to the core transcriptional regulatory circuitry of embryonic stem cells. Cell, 134, 521–533.
Xu, N., Papagiannakopoulos, T., Pan, G., Thomson, J. A., & Kosik, K. S. (2009). MicroRNA-145 regulates OCT4, SOX2, and KLF4 and represses pluripotency in human embryonic stem cells. Cell, 137, 647–658.
Jouneau, A., Ciaudo, C., Sismeiro, O., et al. (2012). Naive and primed murine pluripotent stem cells have distinct miRNA expression profiles. Rna, 18, 253–264.
Tesar, P. J., Chenoweth, J. G., Brook, F. A., et al. (2007). New cell lines from mouse epiblast share defining features with human embryonic stem cells. Nature, 448, 196–199.
Bar, M., Wyman, S. K., Fritz, B. R., et al. (2008). MicroRNA discovery and profiling in human embryonic stem cells by deep sequencing of small RNA libraries. Stem cells (Dayton, Ohio), 26, 2496–2505.
Laurent, L. C., Chen, J., Ulitsky, I., et al. (2008). Comprehensive microRNA profiling reveals a unique human embryonic stem cell signature dominated by a single seed sequence. Stem cells (Dayton, Ohio), 26, 1506–1516.
Lin, S., Cheung, W. K., Chen, S., et al. (2010). Computational identification and characterization of primate-specific microRNAs in human genome. Computational Biology and Chemistry, 34, 232–241.
Nguyen, P. N., Huang, C. J., Sugii, S., Cheong, S. K., & Choo, K. B. (2017). Selective activation of miRNAs of the primate-specific chromosome 19 miRNA cluster (C19MC) in cancer and stem cells and possible contribution to regulation of apoptosis. Journal of Biomedical Science, 24, 20.
Bartel, D. P. (2009). MicroRNAs: target recognition and regulatory functions. Cell, 136, 215–233.
Lewis, B. P., Burge, C. B., & Bartel, D. P. (2005). Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell, 120, 15–20.
Cloonan, N. (2015). Re-thinking miRNA-mRNA interactions: intertwining issues confound target discovery. BioEssays : News and Reviews in Molecular, Cellular and Developmental Biology, 37, 379–388.
Broughton, J. P., Lovci, M. T., Huang, J. L., Yeo, G. W., & Pasquinelli, A. E. (2016). Pairing beyond the Seed Supports MicroRNA Targeting Specificity. Molecular Cell, 64, 320–333.
Zheng, G. X., Ravi, A., Calabrese, J. M., et al. (2011). A latent pro-survival function for the mir-290-295 cluster in mouse embryonic stem cells. PLoS Genetics, 7, e1002054.
Cao, Y., Guo, W. T., Tian, S., et al. (2015). miR-290/371-Mbd2-Myc circuit regulates glycolytic metabolism to promote pluripotency. The EMBO Journal, 34, 609–623.
Sinkkonen, L., Hugenschmidt, T., Berninger, P., et al. (2008). MicroRNAs control de novo DNA methylation through regulation of transcriptional repressors in mouse embryonic stem cells. Nature Structural & Molecular Biology, 15, 259–267.
Kanellopoulou, C., Gilpatrick, T., Kilaru, G., et al. (2015). Reprogramming of Polycomb-Mediated Gene Silencing in Embryonic Stem Cells by the miR-290 Family and the Methyltransferase Ash1l. Stem Cell Reports, 5, 971–978.
Sherstyuk, V. V., Medvedev, S. P., Elisaphenko, E. A., et al. (2017). Genome-wide profiling and differential expression of microRNA in rat pluripotent stem cells. Scientific Reports, 7, 2787.
Zhang, W., Zhong, L., Wang, J., & Han, J. (2016). Distinct MicroRNA Expression Signatures of Porcine Induced Pluripotent Stem Cells under Mouse and Human ESC Culture Conditions. PloS One, 11, e0158655.
Maraghechi, P., Hiripi, L., Toth, G., Bontovics, B., Bosze, Z., & Gocza, E. (2013). Discovery of pluripotency-associated microRNAs in rabbit preimplantation embryos and embryonic stem-like cells. Reproduction (Cambridge, England), 145, 421–437.
Nichols, J., Silva, J., Roode, M., & Smith, A. (2009). Suppression of Erk signalling promotes ground state pluripotency in the mouse embryo. Development (Cambridge, England), 136, 3215–3222.
Silva, J., Barrandon, O., Nichols, J., Kawaguchi, J., Theunissen, T. W., & Smith, A. (2008). Promotion of reprogramming to ground state pluripotency by signal inhibition. PLoS Biology, 6, e253.
Wray, J., Kalkan, T., Gomez-Lopez, S., et al. (2011). Inhibition of glycogen synthase kinase-3 alleviates Tcf3 repression of the pluripotency network and increases embryonic stem cell resistance to differentiation. Nature Cell Biology, 13, 838–845.
Ying, Q. L., Wray, J., Nichols, J., et al. (2008). The ground state of embryonic stem cell self-renewal. Nature, 453, 519–523.
Yan, Y., Yang, X., Li, T. T., et al. (2017). Significant differences of function and expression of microRNAs between ground state and serum-cultured pluripotent stem cells. Journal of Genetics and Genomics = Yi chuan xue bao, 44, 179–189.
Wang, A., He, Q., & Zhong, Y. (2015). Systematically dissecting the global mechanism of miRNA functions in mouse pluripotent stem cells. BMC genomics, 16, 490.
Ai, Z., Shao, J., Shi, X., et al. (2016). Maintenance of Self-Renewal and Pluripotency in J1 Mouse Embryonic Stem Cells through Regulating Transcription Factor and MicroRNA Expression Induced by PD0325901. Stem Cells International, 2016, 1792573.
Wu, Y., Liu, F., Liu, Y., et al. (2015). GSK3 inhibitors CHIR99021 and 6-bromoindirubin-3′-oxime inhibit microRNA maturation in mouse embryonic stem cells. Scientific Reports, 5, 8666.
Cirera-Salinas, D., Yu, J., Bodak, M., Ngondo, R. P., Herbert, K. M., & Ciaudo, C. (2017). Noncanonical function of DGCR8 controls mESC exit from pluripotency. The Journal of Cell Biology, 216, 355–366.
Gonzalez, F., Boue, S., & Izpisua Belmonte, J. C. (2011). Methods for making induced pluripotent stem cells: reprogramming a la carte. Nature Reviews Genetics, 12, 231–242.
Hou, P., Li, Y., Zhang, X., et al. (2013). Pluripotent stem cells induced from mouse somatic cells by small-molecule compounds. Science, 341, 651–654.
Lin, S. L., Chang, D. C., Chang-Lin, S., et al. (2008). Mir-302 reprograms human skin cancer cells into a pluripotent ES-cell-like state. Rna, 14, 2115–2124.
Anokye-Danso, F., Trivedi, C. M., Juhr, D., et al. (2011). Highly efficient miRNA-mediated reprogramming of mouse and human somatic cells to pluripotency. Cell Stem Cell, 8, 376–388.
Miyoshi, N., Ishii, H., Nagano, H., et al. (2011). Reprogramming of mouse and human cells to pluripotency using mature microRNAs. Cell Stem Cell, 8, 633–638.
Judson, R. L., Babiarz, J. E., Venere, M., & Blelloch, R. (2009). Embryonic stem cell-specific microRNAs promote induced pluripotency. Nature Biotechnology, 27, 459–461.
Bernstein, B. E., Mikkelsen, T. S., Xie, X., et al. (2006). A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell, 125, 315–326.
Chin, M. H., Mason, M. J., Xie, W., et al. (2009). Induced pluripotent stem cells and embryonic stem cells are distinguished by gene expression signatures. Cell Stem Cell, 5, 111–123.
Meissner, A., Mikkelsen, T. S., Gu, H., et al. (2008). Genome-scale DNA methylation maps of pluripotent and differentiated cells. Nature, 454, 766–770.
Panopoulos, A. D., Yanes, O., Ruiz, S., et al. (2012). The metabolome of induced pluripotent stem cells reveals metabolic changes occurring in somatic cell reprogramming. Cell Research, 22, 168–177.
Phanstiel, D. H., Brumbaugh, J., Wenger, C. D., et al. (2011). Proteomic and phosphoproteomic comparison of human ES and iPS cells. Nature Methods, 8, 821–827.
Gruber, A. J., Grandy, W. A., Balwierz, P. J., et al. (2014). Embryonic stem cell-specific microRNAs contribute to pluripotency by inhibiting regulators of multiple differentiation pathways. Nucleic Acids Research, 42, 9313–9326.
Li, Z., Yang, C. S., Nakashima, K., & Rana, T. M. (2011). Small RNA-mediated regulation of iPS cell generation. The EMBO Journal, 30, 823–834.
Liao, B., Bao, X., Liu, L., et al. (2011). MicroRNA cluster 302–367 enhances somatic cell reprogramming by accelerating a mesenchymal-to-epithelial transition. The Journal of Biological Chemistry, 286, 17359–17364.
Subramanyam, D., Lamouille, S., Judson, R. L., et al. (2011). Multiple targets of miR-302 and miR-372 promote reprogramming of human fibroblasts to induced pluripotent stem cells. Nature Biotechnology, 29, 443–448.
Samavarchi-Tehrani, P., Golipour, A., David, L., et al. (2010). Functional genomics reveals a BMP-driven mesenchymal-to-epithelial transition in the initiation of somatic cell reprogramming. Cell Stem Cell, 7, 64–77.
Li, R., Liang, J., Ni, S., et al. (2010). A mesenchymal-to-epithelial transition initiates and is required for the nuclear reprogramming of mouse fibroblasts. Cell Stem Cell, 7, 51–63.
Gregory, P. A., Bert, A. G., Paterson, E. L., et al. (2008). The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nature Cell Biology, 10, 593–601.
Korpal, M., Lee, E. S., Hu, G., & Kang, Y. (2008). The miR-200 family inhibits epithelial-mesenchymal transition and cancer cell migration by direct targeting of E-cadherin transcriptional repressors ZEB1 and ZEB2. The Journal of Biological Chemistry, 283, 14910–14914.
Wang, G., Guo, X., Hong, W., et al. (2013). Critical regulation of miR-200/ZEB2 pathway in Oct4/Sox2-induced mesenchymal-to-epithelial transition and induced pluripotent stem cell generation. Proceedings of the National Academy of Sciences of the United States of America, 110, 2858–2863.
Judson, R. L., Greve, T. S., Parchem, R. J., & Blelloch, R. (2013). MicroRNA-based discovery of barriers to dedifferentiation of fibroblasts to pluripotent stem cells. Nature Structural & Molecular Biology, 20, 1227–1235.
Hu, S., Wilson, K. D., Ghosh, Z., et al. (2013). MicroRNA-302 increases reprogramming efficiency via repression of NR2F2. Stem cells (Dayton, Ohio), 31, 259–268.
Choi, Y. J., Lin, C. P., Ho, J. J., et al. (2011). miR-34 miRNAs provide a barrier for somatic cell reprogramming. Nature Cell Biology, 13, 1353–1360.
Prigione, A., Rohwer, N., Hoffmann, S., et al. (2014). HIF1alpha modulates cell fate reprogramming through early glycolytic shift and upregulation of PDK1-3 and PKM2. Stem cells (Dayton, Ohio), 32, 364–376.
Folmes, C. D., Martinez-Fernandez, A., Faustino, R. S., et al. (2013). Nuclear reprogramming with c-Myc potentiates glycolytic capacity of derived induced pluripotent stem cells. Journal of Cardiovascular Translational Research, 6, 10–21.
Zhang, J., Nuebel, E., Daley, G. Q., Koehler, C. M., & Teitell, M. A. (2012). Metabolic regulation in pluripotent stem cells during reprogramming and self-renewal. Cell Stem Cell, 11, 589–595.
Konno, M., Koseki, J., Kawamoto, K., et al. (2015). Embryonic MicroRNA-369 Controls Metabolic Splicing Factors and Urges Cellular Reprograming. PloS One, 10, e0132789.
Lee, M. R., Mantel, C., Lee, S. A., Moon, S. H., & Broxmeyer, H. E. (2016). MiR-31/SDHA Axis Regulates Reprogramming Efficiency through Mitochondrial Metabolism. Stem Cell Reports, 7, 1–10.
Guo, X., Liu, Q., Wang, G., et al. (2013). microRNA-29b is a novel mediator of Sox2 function in the regulation of somatic cell reprogramming. Cell Research, 23, 142–156.
He, X., Cao, Y., Wang, L., et al. (2014). Human fibroblast reprogramming to pluripotent stem cells regulated by the miR19a/b-PTEN axis. PloS One, 9, e95213.
Wu, F., Tao, L., Gao, S., et al. (2017). miR-6539 is a novel mediator of somatic cell reprogramming that represses the translation of Dnmt3b. The Journal of Reproduction and Development, 63, 415–423.
Ye, D., Wang, G., Liu, Y., et al. (2012). MiR-138 promotes induced pluripotent stem cell generation through the regulation of the p53 signaling. Stem cells (Dayton, Ohio), 30, 1645–1654.
Yang, C. S., Li, Z., & Rana, T. M. (2011). microRNAs modulate iPS cell generation. Rna, 17, 1451–1460.
Worringer, K. A., Rand, T. A., Hayashi, Y., et al. (2014). The let-7/LIN-41 pathway regulates reprogramming to human induced pluripotent stem cells by controlling expression of prodifferentiation genes. Cell Stem Cell, 14, 40–52.
Pfaff, N., Liebhaber, S., Mobus, S., et al. (2017). Inhibition of miRNA-212/132 improves the reprogramming of fibroblasts into induced pluripotent stem cells by de-repressing important epigenetic remodelling factors. Stem Cell Research, 20, 70–75.
Barta, T., Peskova, L., Collin, J., et al. (2016). Brief Report: Inhibition of miR-145 Enhances Reprogramming of Human Dermal Fibroblasts to Induced Pluripotent Stem Cells. Stem cells (Dayton, Ohio), 34, 246–251.
Ma, Y., Yao, N., Liu, G., et al. (2015). Functional screen reveals essential roles of miR-27a/24 in differentiation of embryonic stem cells. The EMBO Journal, 34, 361–378.
Zhang, L., Zheng, Y., Sun, Y., et al. (2016). MiR-134-Mbd3 axis regulates the induction of pluripotency. Journal of Cellular and Molecular Medicine, 20, 1150–1158.
Liu, Z., Skamagki, M., Kim, K., & Zhao, R. (2015). Canonical MicroRNA Activity Facilitates but May Be Dispensable for Transcription Factor-Mediated Reprogramming. Stem Cell Reports, 5, 1119–1127.
Miyoshi, K., Miyoshi, T., & Siomi, H. (2010). Many ways to generate microRNA-like small RNAs: non-canonical pathways for microRNA production. Molecular Genetics and Genomics : MGG, 284, 95–103.
Morin, R. D., O’Connor, M. D., Griffith, M., et al. (2008). Application of massively parallel sequencing to microRNA profiling and discovery in human embryonic stem cells. Genome Research, 18, 610–621.
Luciano, D. J., Mirsky, H., Vendetti, N. J., & Maas, S. (2004). RNA editing of a miRNA precursor. Rna, 10, 1174–1177.
Yang, W., Chendrimada, T. P., Wang, Q., et al. (2006). Modulation of microRNA processing and expression through RNA editing by ADAR deaminases. Nature Structural & Molecular Biology, 13, 13–21.
Pantano, L., Estivill, X., & Marti, E. (2010). SeqBuster, a bioinformatic tool for the processing and analysis of small RNAs datasets, reveals ubiquitous miRNA modifications in human embryonic cells. Nucleic Acids Research, 38, e34.
Tan, G. C., Chan, E., Molnar, A., et al. (2014). 5′ isomiR variation is of functional and evolutionary importance. Nucleic Acids Research, 42, 9424–9435.
Cloonan, N., Wani, S., Xu, Q., et al. (2011). MicroRNAs and their isomiRs function cooperatively to target common biological pathways. Genome Biology, 12, R126.
Hinton, A., Hunter, S. E., Afrikanova, I., et al. (2014). sRNA-seq analysis of human embryonic stem cells and definitive endoderm reveals differentially expressed microRNAs and novel IsomiRs with distinct targets. Stem cells (Dayton, Ohio), 32, 2360–2372.
Yuan, Z., Ding, S., Yan, M., et al. (2015). Variability of miRNA expression during the differentiation of human embryonic stem cells into retinal pigment epithelial cells. Gene, 569, 239–249.
Clancy, J. L., Patel, H. R., Hussein, S. M., et al. (2014). Small RNA changes en route to distinct cellular states of induced pluripotency. Nature Communications, 5, 5522.
Langenberger, D., Bermudez-Santana, C., Hertel, J., Hoffmann, S., Khaitovich, P., & Stadler, P. F. (2009). Evidence for human microRNA-offset RNAs in small RNA sequencing data. Bioinformatics (Oxford, England), 25, 2298–2301.
Asikainen, S., Heikkinen, L., Juhila, J., et al. (2015). Selective microRNA-Offset RNA expression in human embryonic stem cells. PloS One, 10, e0116668.
Zhao, J., Schnitzler, G. R., Iyer, L. K., Aronovitz, M. J., Baur, W. E., & Karas, R. H. (2016). MicroRNA-Offset RNA Alters Gene Expression and Cell Proliferation. PloS One, 11, e0156772.
Dupuis-Sandoval, F., Poirier, M., & Scott, M. S. (2015). The emerging landscape of small nucleolar RNAs in cell biology. Wiley Interdisciplinary Reviews RNA, 6, 381–397.
Newton, K., Petfalski, E., Tollervey, D., & Caceres, J. F. (2003). Fibrillarin is essential for early development and required for accumulation of an intron-encoded small nucleolar RNA in the mouse. Molecular and Cellular Biology, 23, 8519–8527.
Zhang, Y., Xu, C., Gu, D., et al. (2017). H/ACA Box Small Nucleolar RNA 7A Promotes the Self-Renewal of Human Umbilical Cord Mesenchymal Stem Cells. Stem cells (Dayton, Ohio), 35, 222–235.
Fong, Y. W., Ho, J. J., Inouye, C., & Tjian, R. (2014). The dyskerin ribonucleoprotein complex as an OCT4/SOX2 coactivator in embryonic stem cells. eLife, 3.
Skreka, K., Schafferer, S., Nat, I. R., et al. (2012). Identification of differentially expressed non-coding RNAs in embryonic stem cell neural differentiation. Nucleic Acids Research, 40, 6001–6015.
Brameier, M., Herwig, A., Reinhardt, R., Walter, L., & Gruber, J. (2011). Human box C/D snoRNAs with miRNA like functions: expanding the range of regulatory RNAs. Nucleic Acids Research, 39, 675–686.
Ender, C., Krek, A., Friedlander, M. R., et al. (2008). A human snoRNA with microRNA-like functions. Molecular Cell, 32, 519–528.
Taft, R. J., Glazov, E. A., Lassmann, T., Hayashizaki, Y., Carninci, P., & Mattick, J. S. (2009). Small RNAs derived from snoRNAs. Rna, 15, 1233–1240.
Mattick, J. S. (2007). A new paradigm for developmental biology. The Journal of Experimental Biology, 210, 1526–1547.
Guttman, M., Amit, I., Garber, M., et al. (2009). Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature, 458, 223–227.
Guttman, M., Garber, M., Levin, J. Z., et al. (2010). Ab initio reconstruction of cell type-specific transcriptomes in mouse reveals the conserved multi-exonic structure of lincRNAs. Nature Biotechnology, 28, 503–510.
St Laurent, G., Vyatkin, Y., Antonets, D., et al. (2016). Functional annotation of the vlinc class of non-coding RNAs using systems biology approach. Nucleic Acids Research, 44, 3233–3252.
Derrien, T., Johnson, R., Bussotti, G., et al. (2012). The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Research, 22, 1775–1789.
Guttman, M., Donaghey, J., Carey, B. W., et al. (2011). lincRNAs act in the circuitry controlling pluripotency and differentiation. Nature, 477, 295–300.
Lin, N., Chang, K. Y., Li, Z., et al. (2014). An evolutionarily conserved long noncoding RNA TUNA controls pluripotency and neural lineage commitment. Molecular Cell, 53, 1005–1019.
Chakraborty, D., Kappei, D., Theis, M., et al. (2012). Combined RNAi and localization for functionally dissecting long noncoding RNAs. Nature Methods, 9, 360–362.
Ng, S. Y., Johnson, R., & Stanton, L. W. (2012). Human long non-coding RNAs promote pluripotency and neuronal differentiation by association with chromatin modifiers and transcription factors. The EMBO Journal, 31, 522–533.
Sheik Mohamed, J., Gaughwin, P. M., Lim, B., Robson, P., & Lipovich, L. (2010). Conserved long noncoding RNAs transcriptionally regulated by Oct4 and Nanog modulate pluripotency in mouse embryonic stem cells. Rna, 16, 324–337.
Wu, C. S., Yu, C. Y., Chuang, C. Y., et al. (2014). Integrative transcriptome sequencing identifies trans-splicing events with important roles in human embryonic stem cell pluripotency. Genome Research, 24, 25–36.
Yu, C. Y., & Kuo, H. C. (2016). The Trans-Spliced Long Noncoding RNA tsRMST Impedes Human Embryonic Stem Cell Differentiation Through WNT5A-Mediated Inhibition of the Epithelial-to-Mesenchymal Transition. Stem cells (Dayton, Ohio), 34, 2052–2062.
Shevchenko, A. I., Zakharova, I. S., & Zakian, S. M. (2013). The evolutionary pathway of x chromosome inactivation in mammals. Acta Naturae, 5, 40–53.
Zhao, J., Sun, B. K., Erwin, J. A., Song, J. J., & Lee, J. T. (2008). Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science, 322, 750–756.
Khalil, A. M., Guttman, M., Huarte, M., et al. (2009). Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proceedings of the National Academy of Sciences of the United States of America, 106, 11667–11672.
Yang, Y. W., Flynn, R. A., Chen, Y., et al. (2014). Essential role of lncRNA binding for WDR5 maintenance of active chromatin and embryonic stem cell pluripotency. eLife, 3, e02046.
Zhou, Y., Dai, Q. S., Zhu, S. C., et al. (2016). AK048794 maintains the mouse embryonic stem cell pluripotency by functioning as an miRNA sponge for miR-592. The Biochemical Journal, 473, 3639–3654.
Wang, Y., Xu, Z., Jiang, J., et al. (2013). Endogenous miRNA sponge lincRNA-RoR regulates Oct4, Nanog, and Sox2 in human embryonic stem cell self-renewal. Developmental Cell, 25, 69–80.
Durruthy-Durruthy, J., Sebastiano, V., Wossidlo, M., et al. (2016). The primate-specific noncoding RNA HPAT5 regulates pluripotency during human preimplantation development and nuclear reprogramming. Nature Genetics, 48, 44–52.
Niazi, F., & Valadkhan, S. (2012). Computational analysis of functional long noncoding RNAs reveals lack of peptide-coding capacity and parallels with 3′ UTRs. Rna, 18, 825–843.
Mumtaz, M. A., & Couso, J. P. (2015). Ribosomal profiling adds new coding sequences to the proteome. Biochemical Society Transactions, 43, 1271–1276.
Ingolia, N. T., Lareau, L. F., & Weissman, J. S. (2011). Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell, 147, 789–802.
Ingolia, N. T., Brar, G. A., Stern-Ginossar, N., et al. (2014). Ribosome profiling reveals pervasive translation outside of annotated protein-coding genes. Cell Reports, 8, 1365–1379.
Banfai, B., Jia, H., Khatun, J., et al. (2012). Long noncoding RNAs are rarely translated in two human cell lines. Genome Research, 22, 1646–1657.
Guttman, M., Russell, P., Ingolia, N. T., Weissman, J. S., & Lander, E. S. (2013). Ribosome profiling provides evidence that large noncoding RNAs do not encode proteins. Cell, 154, 240–251.
Ji, Z., Song, R., Regev, A., & Struhl, K. (2015). Many lncRNAs, 5′UTRs, and pseudogenes are translated and some are likely to express functional proteins. eLife, 4, e08890.
Nelson, B. R., Makarewich, C. A., Anderson, D. M., et al. (2016). A peptide encoded by a transcript annotated as long noncoding RNA enhances SERCA activity in muscle. Science, 351, 271–275.
Szafron, L. M., Balcerak, A., Grzybowska, E. A., et al. (2015). The Novel Gene CRNDE Encodes a Nuclear Peptide (CRNDEP) Which Is Overexpressed in Highly Proliferating Tissues. PloS One, 10, e0127475.
Hussein, S. M., Puri, M. C., Tonge, P. D., et al. (2014). Genome-wide characterization of the routes to pluripotency. Nature, 516, 198–206.
Kim, D. H., Marinov, G. K., Pepke, S., et al. (2015). Single-cell transcriptome analysis reveals dynamic changes in lncRNA expression during reprogramming. Cell Stem Cell, 16, 88–101.
Loewer, S., Cabili, M. N., Guttman, M., et al. (2010). Large intergenic non-coding RNA-RoR modulates reprogramming of human induced pluripotent stem cells. Nature Genetics, 42, 1113–1117.
Zhang, A., Zhou, N., Huang, J., et al. (2013). The human long non-coding RNA-RoR is a p53 repressor in response to DNA damage. Cell Research, 23, 340–350.
Kang, L., Wang, J., Zhang, Y., Kou, Z., & Gao, S. (2009). iPS cells can support full-term development of tetraploid blastocyst-complemented embryos. Cell Stem Cell, 5, 135–138.
Stadtfeld, M., Apostolou, E., Akutsu, H., et al. (2010). Aberrant silencing of imprinted genes on chromosome 12qF1 in mouse induced pluripotent stem cells. Nature, 465, 175–181.
Sverdlov, E. D. (2000). Retroviruses and primate evolution. BioEssays : News and Reviews in Molecular, Cellular and Developmental Biology, 22, 161–171.
Goke, J., Lu, X., Chan, Y. S., et al. (2015). Dynamic transcription of distinct classes of endogenous retroviral elements marks specific populations of early human embryonic cells. Cell Stem Cell, 16, 135–141.
Santoni, F. A., Guerra, J., & Luban, J. (2012). HERV-H RNA is abundant in human embryonic stem cells and a precise marker for pluripotency. Retrovirology, 9, 111.
Wang, J., Xie, G., Singh, M., et al. (2014). Primate-specific endogenous retrovirus-driven transcription defines naive-like stem cells. Nature, 516, 405–409.
Yue, D., Liu, H., & Huang, Y. (2009). Survey of Computational Algorithms for MicroRNA Target Prediction. Current Genomics, 10, 478–492.
Chi, S. W., Zang, J. B., Mele, A., & Darnell, R. B. (2009). Argonaute HITS-CLIP decodes microRNA-mRNA interaction maps. Nature, 460, 479–486.
Hafner, M., Landthaler, M., Burger, L., et al. (2010). Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell, 141, 129–141.
Kuhn, D. E., Martin, M. M., Feldman, D. S., Terry, A. V. Jr., Nuovo, G. J., & Elton, T. S. (2008). Experimental validation of miRNA targets. Methods (San Diego, Calif.), 44, 47–54.
Bassett, A. R., Azzam, G., Wheatley, L., et al. (2014). Understanding functional miRNA-target interactions in vivo by site-specific genome engineering. Nature Communications, 5, 4640.
Esau, C. C. (2008). Inhibition of microRNA with antisense oligonucleotides. Methods (San Diego, Calif.), 44, 55–60.
Tay, F. C., Lim, J. K., Zhu, H., Hin, L. C., & Wang, S. (2015). Using artificial microRNA sponges to achieve microRNA loss-of-function in cancer cells. Advanced Drug Delivery Reviews, 81, 117–127.
Chang, H., Yi, B., Ma, R., Zhang, X., Zhao, H., & Xi, Y. (2016). CRISPR/cas9, a novel genomic tool to knock down microRNA in vitro and in vivo. Scientific Reports, 6, 22312.
Zhang, Z., Xiang, D., Heriyanto, F., Gao, Y., Qian, Z., & Wu, W. S. (2013). Dissecting the roles of miR-302/367 cluster in cellular reprogramming using TALE-based repressor and TALEN. Stem Cell Reports, 1, 218–225.
Liu, Z., Hui, Y., Shi, L., et al. (2016). Efficient CRISPR/Cas9-Mediated Versatile, Predictable, and Donor-Free Gene Knockout in Human Pluripotent Stem Cells. Stem Cell Reports, 7, 496–507.
Essletzbichler, P., Konopka, T., Santoro, F., et al. (2014). Megabase-scale deletion using CRISPR/Cas9 to generate a fully haploid human cell line. Genome Research, 24, 2059–2065.
Han, J., Zhang, J., Chen, L., et al. (2014). Efficient in vivo deletion of a large imprinted lncRNA by CRISPR/Cas9. RNA Biology, 11, 829–835.
Ho, T. T., Zhou, N., Huang, J., et al. (2015). Targeting non-coding RNAs with the CRISPR/Cas9 system in human cell lines. Nucleic Acids Research, 43, e17.
Zhu, S., Li, W., Liu, J., et al. (2016). Genome-scale deletion screening of human long non-coding RNAs using a paired-guide RNA CRISPR-Cas9 library. Nature Biotechnology, 34, 1279–1286.
Cong, L., Zhou, R., Kuo, Y. C., Cunniff, M., & Zhang, F. (2012). Comprehensive interrogation of natural TALE DNA-binding modules and transcriptional repressor domains. Nature Communications, 3, 968.
Qi, L. S., Larson, M. H., Gilbert, L. A., et al. (2013). Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell, 152, 1173–1183.
Gilbert, L. A., Larson, M. H., Morsut, L., et al. (2013). CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell, 154, 442–451.
Acknowledgements
This work was supported by the Russian Science Foundation (project №16-14-10084).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no conflicts of interest.
Rights and permissions
About this article
Cite this article
Sherstyuk, V.V., Medvedev, S.P. & Zakian, S.M. Noncoding RNAs in the Regulation of Pluripotency and Reprogramming. Stem Cell Rev and Rep 14, 58–70 (2018). https://doi.org/10.1007/s12015-017-9782-9
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12015-017-9782-9