Abstract
Recent studies have identified a new human dental derived progenitor cell population with multi-lineage differentiation potential referred to as human periapical cyst mesenchymal stem cells (hPCy-MSCs). In the present study, we compared two subpopulations of hPCy-MSCs characterised by the low or high expression of CD146 to establish whether this expression can regulate their stem cell properties. Using flow cytometry, we evaluated the stem cell marker profile of hPCy-MSCs during passaging. Furthermore, CD146Low and CD146High cells were sorted by magnetic beads and subsequently both cell populations were evaluated for differences in their proliferation, self-renewal, stem cell surface markers, stemness genes expression and osteogenic differentiation potential.
We found that hPCy-MSCs possessed a stable expression of several mesenchymal stem cell surface markers, whereas CD146 expression declined during passaging.
In addition, sorted CD146Low cells proliferated significantly faster, displayed higher colony-forming unit-fibroblast capacity and showed higher expression of Klf4 when compared to the CD146High subset. Significantly, the osteogenic potential of hPCy-MSCs was greater in the CD146Low than in CD146High population. These results demonstrate that CD146 is spontaneously downregulated with passaging at both mRNA and protein levels and that the high expression of CD146 reduces the proliferative, self-renewal and osteogenic differentiation potential of hPCy-MSCs. In conclusion, our study demonstrates that changes in the expression of CD146 can influence the stem cell properties of hPCy-MSCs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Today the use of mesenchymal stem cells (MSCs) to produce different cell types represents a new promising approach to treat or replace diseased, damaged or missing tissues [1]. In recent years, tissues of oral cavity have been successfully used in alternative to bone marrow as sources of MSCs for regenerative medicine due to their ease of procurement and availability [2]. MSCs have been identified in several tissues of the oral cavity including dental pulp [3], periodontal ligament [4], exfoliated deciduous teeth [5], dental follicle [6], apical papilla [7] and gingival tissues [8]. These dental-derived stem cells (DSC) are an attractive source not only for their use in regenerative dentistry but also for a broad range of potential therapeutic applications in all fields of regenerative medicine [1, 9]. For example, the transformation of DSC into chondrocytes can be used in cartilage-related diseases [10], whereas the differentiation of DSCs into cardiomyocytes has opened a new way to treat cardiovascular diseases [11]. Moreover, cell-based strategies with DSCs have been successfully used to regenerate bone [12], liver [13], muscular [14] as well as to treat spinal cord injuries [15], diabetes [16] and neurological diseases [17].
Recently, a new population of progenitor/multipotent stem cells named periapical cyst-derived mesenchymal stem cells (hPCy-MSCs) has been identified in human periapical cystic tissues by our group [18]. We reported that the hPCy-MSCs possess mesenchymal stem cell-like properties including self-renewal, high proliferative potential, stem cell markers expression and the ability to differentiate into osteogenic and adipogenic lineages [18, 19]. Moreover, their ability to differentiate into neural and glial cells was also recently shown, indicating that the hPCy-MSCs can also be considered for the replacement of neuronal cells that have been damaged by disease [2, 19]. Taken together, our previously published reports indicate that the hPCy-MSCs represent a new cell source with high application potential in the field of regenerative medicine [2, 18–20].
To enhance the potential of these newly discovered cells and given that previous studies have reported that CD146-based selection might be exploited to obtain multipotential MSCs fractions [21–24], we decided to explore the effect of CD146 expression in the hPCy-MSCs.
The immunoglobulin superfamily member CD146, also known as MUC-18 or melanoma cell adhesion molecule (Mel-CAM) has been originally identified as a melanoma marker [25]. It is primarily an endothelial cell marker that is also expressed in perivascular cells, smooth muscle, myofibroblasts cells, active T lymphocytes and MSCs isolated from multiple adult organs including those derived from dental tissues [26–28]. It has been shown that the high expression of CD146 on the surface of MSCs is linked with multipotency of these cells [29]. Although the specific function of CD146 is not entirely understood, it has been associated with several cellular processes regulated by transmembrane signalling including cell proliferation, adhesion, migration, cytoskeletal organisation and differentiation [26]. Gronthos et al. have shown that dental pulp stem cells (DPSCs) express the cell surface antigen CD146 and display phenotype consistent with perivascular cell populations. Furthermore, the same study showed the efficacy of using magnetic cell sorting (MACS) to enhance the clonogenic potential of DPSCs [26].
Therefore, the objective of this study was to evaluate the use of CD146 as a surface marker for the enrichment of a specific progenitor subpopulation from a heterogeneous hPCy-MSCs population. Here, we evaluated the expression of CD146 in the hPCy-MSCs during passaging and compared the biological characteristics of CD146High and CD146Low populations obtained after magnetic cell sorting. To that end, we compared proliferation, self-renewal, MSC surface markers and expression of stem cell transcription factors between sorted populations of hPCy-MSCs. In addition, the osteogenic potential of CD146High and CD146Low populations was evaluated to explore the role of CD146 in regulating the osteogenic differentiation of the hPCy-MSCs.
Materials and Methods
Isolation and Culture of hPCy-MSCs
Human periapical cyst mesenchymal stem cells (hPCy-MSCs) were isolated as described previously by Marrelli et al. [2, 18–20]. Briefly, human periapical cyst tissues were obtained from healthy volunteers of 25 ± 3 years. Written informed consents were obtained from all participants of this study using a specific form approved by the Calabrodental dental clinic (Crotone, Italy). The study followed the “Ethical principles for medical research involving human subjects” of the Helsinki Declaration. After extraction, the enucleated cyst tissues were immediately stored in a physiological solution and were transported to the research lab (Tecnologica SRL, Research Institute, Crotone, Italy). For digestion, the cyst tissues were washed three times with phosphate-buffered saline (PBS) containing antibiotics and subsequently placed in tissue culture dishes for mechanical disruption. Then, cyst tissues were minced into small pieces and digested with a solution containing 3 mg/ml type I collagenase (Gibco) and 4 mg/ml dispase (Gibco) for 1 h at 37 °C.
Next, the isolated cells were filtered through 70 μM falcon strainers (Becton Dickinson) and plated in α-MEM culture medium containing 10 % FBS (Gibco), 100 μM L-ascorbic acid-2-phosphate, 100 U/ml penicillin, 2 mM glutamine and 100 mg/ml streptomycin (Invitrogen). The obtained hPCy-MSCs were incubated at 37 °C with 5 % CO2, and the medium was changed every 3 days.
CD146 Magnetic Cell Sorting
The hPCy-MSCs (10 X 106 cells) were enzymatically dissociated into a single-cell suspension and incubated with FcR blocking reagent and then with a microbeads-conjugated anti-CD146 antibody (Miltenyi Biotech) at 4 °C for 15 min. Then, the cells were washed with PBS buffer containing 0.5 % BSA and 2 mM EDTA and centrifuged for 10 min. The cell pellets were resuspended in 500 μl of the same buffer and passed through an LS column placed in a magnetic field of a MACS separator, followed by three washes of the column with 3 ml MACS separation buffer. While the cells were passing through the column, the magnetically labelled hPCy-MSCs CD146+ were mainly retained on the column and then unlabelled cells (CD146Low) were eluted. Then, the column was removed from the MACS separator and the bound CD146+ cells (CD146High) were flushed out and collected in a separate tube. Finally, CD146High, CD146Low and the unsorted cells were cultured with the complete α-MEM medium. The purification of the sorted cells (CD146High and CD146Low) was confirmed by cytofluorimetric, immunofluorescence and qRT-PCR analyses.
Cell Proliferation and Cell Viability
Sorted cells were seeded onto 96-multiwell plates and cultured with complete growth medium at 37 °C and 5 % CO2. Cells viability was measured after 2 to 16 days by Prestoblue™ Cell Viability Reagent (Invitrogen) following manufacturer’s instructions. Briefly, growth medium was removed, and cells were washed and incubated with Prestoblue reagent for 2 h at 37 °C and 5 % CO2. Subsequently, the absorbance was read by a Multiskan GO microplate spectrophotometer. Cell viability of sorted hPCy-MSCs was analysed by the PI staining method combined with advanced image analysis (Accuchip cell counter, Adam, Nanoentek).
Flow Cytometry
The expression of MSC-specific cell-surface markers in sorted hPCy-MSCs was analysed as described previously by Paduano et al. [18, 30]. Briefly, cells were harvested using Trypsin-EDTA (Gibco) and aliquoted at 0.3 x 106 cells into FACS tubes for each antibody. Then, the hPCy-MSCs were stained using PE-conjugated anti-CD13 (CD13-PE), APC-conjugated anti-CD29 (CD29-APC), FITC-conjugated anti-CD44 (CD44-FITC), fluorescein isothiocyanate-conjugated anti-CD73 (CD73-FITC), phycoerythrin-conjugated anti-CD90 (CD90-PE), allophycocyanin-conjugated anti-CD105 (CD105-APC) and allophycocyanin-H7-conjugated anti-CD45 (CD45-APC-H7). All antibodies were obtained from BD (Becton Dickinson). At the end of the incubation period, cells were washed, and 500 μl of PBS were added to each sample. Fluorescein isothiocyanate-conjugated anti IgG1, phycoerythrin-conjugated anti-IgG1, allophycocyanin-H7-conjugated anti-IgG1 and allophycocyanin-H7-conjugated anti-IgG1 have been used as isotype controls. The fluorescence intensities were measured by flow cytometer (NAVIOS, Beckman Coulter) and data were analysed using Kaluza software (Beckman Coulter) and FlowJo software (Tree Star).
Immunofluorescence Analysis
Sorted hPCy-MSCs cultured in growth medium were rinsed with PBS, fixed in 4 % paraformaldehyde for 20 min at RT and blocked with 3 % BSA for 20 min at RT. Then, the hPCy-MSCs were incubated for 30 min with phycoerythrin-conjugated anti-CD146 antibody, washed twice with PBS and incubated with a solution containing 1 μg/ml of 4,6-diamidino-2-phenylindole (DAPI; Sigma) for 5 min. Then, cells were mounted with anti-fading medium (Invitrogen) and observed by confocal microscopy (Leica, TCS SP5). Fluorescein isothiocyanate-conjugated anti-IgG1 was used as negative control.
Gene Expression Analysis
Total RNA was extracted from sorted hPCy-MSCs using the Purelink™ RNA mini kit (Applied Biosystems) and the extracted RNA was quantified using a Multiskan Go spectrophotometer (Thermo Scientific). Gene expression was analysed as previously described [18, 20, 30]. Briefly, 200 ng of RNA were reverse-transcribed using M-MuLV reverse transcriptase (Applied Biosystems) in a 10 μL volume containing buffer with 400 μmol/L dNTPs and 0.2 μg Oligo(dT). Subsequently, 0.5 μL of cDNA were amplified by real-time PCR (qPCR) using primers (5 pmol) and power SYBR green Master Mix (Applied Biosystems). The qPCR reactions were performed using a Pikoreal 96 system (Thermo Scientific) with the following parameters: an initial denaturation step at 95 °C for 10 min; 40 cycles of 10 s at 95 °C and 1 min at 60 °C. Efficiencies of qPCR were calculated from the cycle threshold (Ct) curves obtained from the amplification of the ten-fold serial dilutions of cDNA. Moreover, melting curve analyses were performed to verify the specificity of the PCR products. After amplification, mRNA expressions of specific genes were obtained by the 2−ΔΔCt method with the levels of gene expression normalised to the housekeeping gene hypoxanthine phosphoribosyltransferase (HRPT). The primers for the CD146, kruppel-like factor 4 (Klf4), octamer-binding transcription factor 4 (OCT4), homeobox transcription factor Nanog (Nanog), v-myc avian myelocytomatosis viral oncogene homolog (c-myc), SRY-related HMG-box 2 (Sox2), runt-related transcription factor 2 (Runx-2), osteopontin (OPN), hypoxanthine phosphoribosyltransferase (HPRT) are listed in Appendix Table 1.
Osteogenic Differentiation
To obtain osteogenic differentiation the sorted hPCy-MSCs were plated on plate and cultured with osteogenic medium containing α-MEM (Sigma), 20 % FBS (Invitrogen), 0.2 mM L-ascorbic acid-2-phosphate (Sigma), 100 nM dexamethasone (Sigma), 10 mM β-glicerophosphate (Sigma), 100 U/mL penicillin, 0.1 mg/mL streptomycin, and 0.25 mg/mL amphotericin B. Subsequently, the sorted hPCy-MSCs were fixed with 4 % paraformaldehyde (Sigma) for 20 min at RT and washed twice with PBS. Then, a solution of 5 mg/mL Alizarin Red S (Sigma) was added to the cells for 30 min. The sorted hPCy-MSCs were washed three times with H2O and observed under microscopy. Alizarin Red precipitates were quantified as previously described [18]. Briefly, samples were incubated with 800 μl acetic acid (10 %) for 30 min and subsequently the supernatant was transferred into a tube and boiled for 10 min at 85 °C. After centrifugation (15 min, 15.000×g), 150 μl of samples were transferred into a 96-well plate, and the optical density was evaluated at 405 nm using a Multiskan Go Spectrophotometer (Thermo Scientific).
Statistical Analysis
Results are shown as means ± standard deviations (SD) from three independent experiments. The differences between groups were analysed with an unpaired two-tailed Student’s t-test using the GraphPad Prism software package (Graphpad Software Inc). Differences were considered statistically significant when P < 0.05. *P < 0.05; **P < 0.01; and ***P < 0.001.
Results
Isolation and Sorting of hPCy-MSCs
Mesenchymal stem cells were isolated from the periapical cyst and separated by magnetic cell sorting (MACS) according to their expression of CD146. A schematic representation of experimental approach is shown in Fig. 1. Briefly, MSCs were obtained from the inflammatory periapical cyst by enzymatic digestion, cultured and analysed during passaging for the expression of CD146 at both mRNA and protein levels. Subsequently, cells were sorted by magnetic beads into two populations named CD146High and CD146Low and analysed for MSC surface markers expression, proliferation, CFU-F, stemness genes expression and osteogenic differentiation potential (Fig. 1). The objective was to identify the role of CD146 expression in the regulation of the stem cell properties of hPCy-MSCs.
Schematic overview of the experimental setup. Mesenchymal stem cells were isolated from the human periapical cyst (hPCy-MSCs). After culture, cells were analysed for surface marker expression and for CD146 mRNA expression during different in vitro passaging (p2, p4, p6, p8). Subsequently, hPCy-MSCs were sorted by magnetic beads for CD146. Sorted cells were analysed as described
hPCy-MSCs Possess a Stable Expression of Several Mesenchymal Stem Cell Surface Markers, whereas CD146 Declines during Passaging
First, we investigated the expression of CD146 on hPCy-MSCs by flow cytometry at four different passages (p2, p4, p6 and p8). Phenotypic characterization showed that MSC-specific markers CD13, CD29, CD44, CD73 and CD105 were uniformly positive during passaging, indicating a stable expression profile throughout the expansion (Fig. 2a). On the contrary, CD146 expression declined gradually during passaging in hPCy-MSCs. The expression of CD146 decreased along passages also in three different oral-derived mesenchymal stem cells such as dental pulp stem cells (DPSCs), periodontal ligament stem cells (PDLSCs) and dental follicle stem cells (DFPCs, Supplementary Fig. 1). Furthermore, we also observed that the mRNA expression levels of CD146 in hPCy-MSCs were significantly downregulated during the expansion in vitro (Fig. 2b). These data indicate that hPCy-MSCs possess a stable expression of several mesenchymal stem cell surface markers during expansion, whereas CD146 is spontaneously downregulated at both mRNA and protein levels.
MSC surface markers and CD146 mRNA expression in expanded hPCy-MSCs. a Surface marker expression levels of CD146, CD13, CD29, CD44, CD73, CD90, CD105 and CD45 in hPCy-MSCs during different passages (p2, p4, p6, p8). b Expression of CD146 mRNA levels in hPCy-MSCs during passaging (p2, p4, p6, p8). The results are presented as fold increase (2-(ΔΔCT)) with respect to the level expressed in hPCy-MSCs at p2. The data shown are the means ± SDs of three independent experiments, *P < 0.05, *** P < 0.001 compared with hPCy-MSCs at p2. p: passage
Analysis of CD146Low and CD146High Sorted Populations of hPCy-MSCs
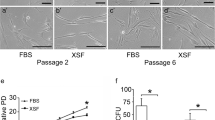
An early passage of hPCy-MSCs was separated by the magnetic cell sorting (MACS) to generate CD146Low and CD146High fractions (Fig. 3a). The efficacy of the separation was confirmed by flow cytometry immediately after cell sorting and by immunofluorescence and qRT-PCR after expansion in culture.
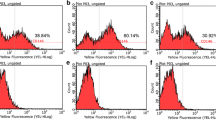
Analysis of CD146Low and CD146High sorted populations of hPCy-MSCs. a Morphological aspects of the hPCy-MSCs CD146High and CD146Low subpopulations compared to unsorted cells, scale bar, 100 μm. b Flow cytometry analysis of the unsorted hPCy-MSCs, CD146High and CD146Low sorted fractions. Isotype control is indicated by red and positive signals by blue and the percentages of positively labelled cells are indicated. c Immunofluorescence analysis of CD146 expression in sorted cells, scale bar, 100 μm. d Expression of CD146 mRNA levels in hPCy-MSCs CD146High and CD146Low subpopulations compared to unsorted cells evaluated by qRT-PCR. The results are presented as fold increase (2-(ΔΔCT)) with respect to the level expressed in unsorted hPCy-MSCs. *P < 0.05, *** P < 0.001
Cytofluorimetric analyses indicate that CD146Low and CD146High populations of hPCy-MSCs possess low and high expression of CD146 (10.4 % and 94.2 %, respectively) compared to 39.5 % of the unsorted cells (Fig. 3b). These results were confirmed by immunofluorescence analysis, which showed that CD146 protein was highly expressed in the CD146High fraction and lowly expressed in the CD146Low fraction in comparison with unsorted cells (Fig. 3c). Moreover, the mRNA expression levels of CD146 were reduced 0.5-fold in CD146Low population and increased 20-fold in the CD146High population with respect to unsorted hPCy-MSCs (Fig. 3d). Collectively, these data indicate that MACS technology can enrich the CD146High and CD146Low fractions from hPCy-MSCs populations with good efficacy.
Sorted CD146Low Cells Proliferate significantly faster, Have Higher Colony-Forming Unit-Fibroblast and Show Higher Expression of Klf4 than CD146High Cells
To analyse the role of CD146 expression on stem cell properties of hPCy-MSCs, we evaluated the expression of several mesenchymal surface markers including CD13, CD29, CD44, CD73, CD90, CD105 and CD45. Characterization of sorted cells revealed that the expression of MSC markers was similar in both the CD146Low and CD146High fractions compared to that of the unsorted cells (Fig. 4a). Having shown that both sorted hPCy-MSCs populations possessed a phenotype identical to those of originally derived cells, we next analysed the proliferation rate of CD146Low and CD146High populations. Importantly, we observed that the proliferation of the CD146Low cells was slightly higher than control cells whereas was significantly greater with respect to CD146High cells (Fig. 4b). Furthermore, primary cultures of sorted hPCy-MSCs were established at clonal density in order to obtain discrete colonies (CFU-F). Results show that CD146Low cells produce a similar number of CFU-F than unsorted cells. On the contrary, the self-renewal capability is significantly lower in CD146High cells with respect to control (Fig. 4c). Having seen that the low expression of CD146 is required for the optimal proliferation and CFU-F efficiency of hPCy-MSCs, a viability assay was performed to evaluate the percentage of live/dead cells in all populations described above. Results show that CD146High and CD146Low populations do not clearly differ in the viability of the cells compared to unsorted population (Fig. 4d).
Characterization of CD146Low and CD146High sorted populations of hPCy-MSCs. a Flow cytometry analysis of MSC cell surface marker expression in sorted hPCy-MSCs. Isotype control is indicated by the dotted line and antibody staining by a solid line. b Prestoblue assay for the proliferation of CD146Low and CD146High sorted populations compared to unsorted cells from day 2 to 16. c Colony forming unit fibroblast (CFU-F) in CD146Low and CD146High cell populations. Results are mean ± SDs. d Percentage of cell viability of sorted fractions of hPCy-MSCs. e mRNA expression of stemness genes Klf4, OCT4, Nanog, c-myc and Sox-2 in CD146Low and CD146High sorted populations evaluated by qRT-PCR. Results are represented as fold increase compared to the level expressed in unsorted hPCy-MSCs. *P < 0.05, **P < 0.01, ***P < 0.001, N.S., not significant
Next, we questioned if the CD146 expression could modify the expression of stemness genes in hPCy-MSCs. We observed that unsorted hPCy-MSCs expressed several transcripts of the embryonic stem cells such as Klf4, Oct4, Nanog, Sox2 and c-myc. Importantly, the expression of Klf4 was significantly higher in the CD146Low population whereas was lower in CD146High compared to control (Fig. 4e).
In addition, CD146High and CD146Low populations showed a weaker expression of the stemness genes OCT4, Nanog, Sox2 and c-myc compared to unsorted cells (Fig. 4e). However, the expression of OCT4 was slightly higher for CD146Low than CD146High populations whereas the expression of Sox2 was slightly lower for CD146Low than CD146High populations. Collectively, these results clearly indicate that CD146 expression can influence several stem cell properties of hPCy-MSCs.
Osteogenic Potential of hPCy-MSCs Is Higher in the CD146Low than in CD146High Cell Population
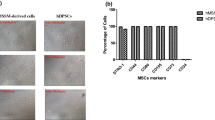
To test whether the CD146Low and CD146High populations exhibited differences in osteogenic capacity, we analysed the ability of these sorted populations to differentiate into osteoblast-like cells in vitro. Osteogenic differentiation of CD146Low and CD146High was detected by alizarin red staining, in which differentiation of unsorted cells was used as control. Overall, there was no significant osteogenic differentiation between CD146Low and unsorted cells. However, the osteogenic differentiation potential of the CD146High population was significantly lower with respect to CD146Low and control (Fig. 5a). These results were confirmed by spectrophotometric quantification, with significant less absorbance noted in the CD146High population (Fig. 5b).
Osteogenic differentiation potential of hPCy-MSCs CD146Low and CD146High sorted fractions. a Osteogenic differentiation of hPCy-MSCs CD146High and CD146Low subpopulations stained with alizarin red. b Alizarin red quantification of mineralized deposits in sorted fractions of hPCy-MSCs compared to unsorted cells. *P < 0.05. c mRNA expression of osteoblast marker genes RUNX-2 and OPN in sorted cell populations. hPCy-MSCs (CD146Low or High) undifferentiated and after 21 days of differentiation (Osteo). Results are represented as fold increase compared to the level expressed in unsorted hPCy-MSCs. *P < 0.05, **P < 0.01, ***P < 0.001
The osteogenic potential of sorted populations was further confirmed by gene expression after three weeks in osteogenic medium, examining the expression levels of osteopontin (OPN) and runt-related transcription factor 2 (Runx-2). CD146High and CD146Low populations cultured in osteogenic medium showed significant downregulation of Runx-2 than unsorted cells (Fig. 5c). In contrast, the expression levels of OPN were significantly upregulated in CD146Low cells and were downregulated in CD146High cells compared to that of control. qRT-PCR analyses were consistent with the alizarin red staining, demonstrating a decrease in the transcript levels of Runx-2 and OPN (3-fold and 22-fold, respectively) in CD146High with respect to the CD146Low population.
Discussion
Mesenchymal stem cells derived from the human periapical cyst (hPCy-MSCs), which possess multipotent differentiation capacity, are a useful cell source for regenerative medicine [18–20]. Importantly, compared to BMSCs and other dental-derived stem cells, hPCy-MSCs can be simply isolated from patients with minimal discomfort [2, 18, 20].
Therefore, a better knowledge of the functional markers able to improve stem cell properties of hPCy-MSCs should be investigated to potentiate their use in regenerative medicine.
Using dental-derived MSCs, Huang et al. have shown that STRO-1+ /CD146+ DPSCs exhibit higher MSC properties with respect to the negative counterpart [31], whereas Zhu et al. have observed that periodontal ligament stem cells (PDLSCs) CD146+ possess a higher proliferative and osteogenic potential compared to CD146 [24].
CD146 functions in melanoma, breast and prostate cancers have been well characterised and comprehended in great detail [25, 32]. However, the precise role of the adhesion molecule CD146 in MSCs and maintenance of dental-derived stem cells during in vitro expansion have not been yet understood. Gronthos et al. in 2000 demonstrated that dental-derived MSCs, such as those derived from dental pulp shared the expression of CD146 with BMSCs [3]. In this study, by evaluating the expression of stem cells surface markers during ex vivo expansion, we observed that the MSCs derived from dental tissues including periapical cyst, dental pulp, periodontal ligament and dental follicle exhibit a progressively decreased expression of the pericyte marker CD146 along passages. In accord with our results, previous studies with MSCs derived from human bone marrow and adipose tissue have shown a decrease in the percentage of CD146 positive cells during in vitro passages [33–36]. Furthermore, the decrease in CD146 expression from passage one to five was observed in two dental-derived stem cells such as human exfoliated deciduous teeth (SHED) and dental pulp stem cells (DPSCs) [27]. The spontaneous downregulation of CD146 with passaging seems to be shared by MSCs of different origin. However, it is not yet clear if the decrease in CD146 expression is intrinsic or reflects the culture conditions. For example, Russel et al. have demonstrated that cellular confluence is sufficient to downregulate the expression of CD146 in BMSCs [37], whereas Tormin et al. have observed that CD146 expression in BMSCs is downregulated under hypoxic conditions [38]. Similar to our results, Sharma et al. have shown that placenta-derived MSCs were initially CD146+ but then became negative during in vitro culturing on hydrogel-based matrix [39].
It has been well observed that CD146 expression in MSCs is correlated with their proliferative potential and trilineage differentiation capacity [40–42]. Starting from the concept that CD146 is an indicator of stemness properties, several researchers have used CD146 as a positive maker for sorting native MSCs [22, 38, 43, 44].
To determine if the CD146 antigen is able to increase stem cell properties of hPCy-MSCs, we successfully enriched the CD146High and CD146Low cell fractions from a heterogeneous population by using a magnetic cell sorting (MACS). Of note, the MACS technology did not completely separate CD146+ from CD146- cells, but significantly induced an enrichment of both cell fractions.
Firstly, we observed that CD146Low and CD146High hPCy-MSCs did not differ in the expression of other MSC markers such as CD13, CD29, CD44, CD73 and CD90. As previously showed for BMSCs [43], these results indicate that CD146High and CD146Low enrichments do not alter the expression of other surface markers of the hPCy-MSCs. Furthermore, we found that proliferation and CFU-F capacity were significantly higher in CD146Low compared to CD146High populations of hPCy-MSCs. Recently, other have reported that CD146− clones isolated from BMSCs proliferate significantly greater than the corresponding CD146+ counterparts [44]. In contrast to our results, two reports have indicated that the CFU-efficiency of dental derived stem cells such as DPSCs and PDLSCs CD146+ are significantly higher than those of CD146− [24, 31].
Importantly, we observed that CD146Low cells expressed a high level of Klf4, indicating that this subpopulation displays a key characteristic of pluripotent cells. Furthermore, CD146Low and CD146High populations showed weaker expression of the stem cell transcription factors OCT4, Nanog, c-myc and Sox2 as compared to native populations. Similar results have been obtained by Espagnolle et al. for human BMSCs CD146+ and CD146− [44].
In this study, we observed a significant difference between the osteogenic potential of CD146Low and CD146High populations of hPCy-MSCs. In fact, the osteogenic differentiation capacity of the CD146Low population was significantly higher with respect to the CD146High counterpart. The osteogenic commitment of the CD146− MSCs was also observed in a previous study that indicated that BMSCs CD146− formed higher heterotopic bone in vivo with respect to CD146+ cells [43].
On the contrary, it has been recently reported that BMSCs CD146Low and CD146High do not differ in the osteogenic differentiation potential [38, 44]. To our knowledge, only one report on dental-derived MSCs has indicated that CD146+ PDLSCs possess higher osteogenic potential than CD146− [24]. The differences between Zhu et al. and our results regarding the osteogenic potential of sorted MSCs are probably due to the different tissue types from which cells were derived.
In this work, we sorted CD146Low and CD146High cells from a heterogeneous population of hPCy-MSCs. Interestingly, our results show that CD146 has a crucial role in regulating stem cell properties of hPCy-MSCs. We therefore conclude that CD146 could be used to select the most potent population of hPCy-MSCs for regenerative medical applications.
References
Egusa, H., Sonoyama, W., Nishimura, M., Atsuta, I., & Akiyama, K. (2012). Stem cells in dentistry--part I: stem cell sources. Journal of Prosthodontic Research, 56, 151–165.
Tatullo, M., Marrelli, M., & Paduano, F. (2015). The regenerative medicine in oral and maxillofacial surgery: the most important innovations in the clinical application of mesenchymal stem cells. International Journal of Medical Sciences, 12, 72–77.
Gronthos, S., Mankani, M., Brahim, J., Robey, P. G., & Shi, S. (2000). Postnatal human dental pulp stem cells (DPSCs) in vitro and in vivo. Proceedings of the National Academy of Sciences of the United States of America, 97, 13625–13630.
Seo, B. M., Miura, M., Gronthos, S., Bartold, P. M., Batouli, S., Brahim, J., et al. (2004). Investigation of multipotent postnatal stem cells from human periodontal ligament. Lancet (London, England), 364, 149–155.
Miura, M., Gronthos, S., Zhao, M., Lu, B., Fisher, L. W., Robey, P. G., et al. (2003). SHED: stem cells from human exfoliated deciduous teeth. Proceedings of the National Academy of Sciences of the United States of America, 100, 5807–5812.
Morsczeck, C., Gotz, W., Schierholz, J., Zeilhofer, F., Kuhn, U., Mohl, C., et al. (2005). Isolation of precursor cells (PCs) from human dental follicle of wisdom teeth. Matrix Biology , 24, 155–165.journal of the International Society for Matrix Biology
Sonoyama, W., Liu, Y., Fang, D., Yamaza, T., Seo, B. M., Zhang, C., et al. (2006). Mesenchymal stem cell-mediated functional tooth regeneration in swine. PloS One, 1, e79.
Huang, G. T., Gronthos, S., & Shi, S. (2009). Mesenchymal stem cells derived from dental tissues vs. those from other sources: their biology and role in regenerative medicine. Journal of Dental Research, 88, 792–806.
Kabir, R., Gupta, M., Aggarwal, A., Sharma, D., Sarin, A., & Kola, M. Z. (2014). Imperative role of dental pulp stem cells in regenerative therapies: a systematic review. Nigerian journal of surgery , 20, 1–8.official publication of the Nigerian Surgical Research Society
Rizk, A., & Rabie, A. B. (2013). Human dental pulp stem cells expressing transforming growth factor beta3 transgene for cartilage-like tissue engineering. Cytotherapy, 15, 712–725.
Xin, L. Z., Govindasamy, V., Musa, S., & Abu Kasim, N. H. (2013). Dental stem cells as an alternative source for cardiac regeneration. Medical Hypotheses, 81, 704–706.
Graziano, A., d'Aquino, R., Laino, G., & Papaccio, G. (2008). Dental pulp stem cells: a promising tool for bone regeneration. Stem Cell Reviews, 4, 21–26.
Ishkitiev, N., Yaegaki, K., Imai, T., Tanaka, T., Fushimi, N., Mitev, V., et al. (2015). Novel management of acute or secondary biliary liver conditions using hepatically differentiated human dental pulp cells. Tissue Engineering Part A, 21, 586–593.
Yang, R., Chen, M., Lee, C. H., Yoon, R., Lal, S., & Mao, J. J. (2010). Clones of ectopic stem cells in the regeneration of muscle defects in vivo. PloS One, 5, e13547.
Bianco, J., De Berdt, P., Deumens, R., & des Rieux, A. (2016). Taking a bite out of spinal cord injury: Do dental stem cells have the teeth for it? Cellular and Molecular Life Sciences , 73, 1413–1437.CMLS
Guimaraes, E. T., Cruz Gda, S., Almeida, T. F., Souza, B. S., Kaneto, C. M., Vasconcelos, J. F., et al. (2013). Transplantation of stem cells obtained from murine dental pulp improves pancreatic damage, renal function, and painful diabetic neuropathy in diabetic type 1 mouse model. Cell Transplantation, 22, 2345–2354.
Arthur, A., Rychkov, G., Shi, S., Koblar, S. A., & Gronthos, S. (2008). Adult human dental pulp stem cells differentiate toward functionally active neurons under appropriate environmental cues. Stem Cells (Dayton, Ohio), 26, 1787–1795.
Marrelli, M., Paduano, F., & Tatullo, M. (2013). Cells isolated from human periapical cysts express mesenchymal stem cell-like properties. International Journal of Biological Sciences, 9, 1070–1078.
Tatullo M, Falisi G, Amantea M, Rastelli C, Paduano F, Marrelli M. Dental pulp stem cells and human periapical cyst mesenchymal stem cells in bone tissue regeneration: comparison of basal and osteogenic differentiated gene expression of a newly discovered mesenchymal stem cell lineage. Journal of Biological Regulators and Homeostatic Agents 2015; 29:713–718.
Marrelli, M., Paduano, F., & Tatullo, M. (2015). Human periapical cyst-mesenchymal stem cells differentiate into neuronal cells. Journal of Dental Research, 94, 843–852.
Kouroupis, D., Churchman, S. M., McGonagle, D., & Jones, E. A. (2014). The assessment of CD146-based cell sorting and telomere length analysis for establishing the identity of mesenchymal stem cells in human umbilical cord. F1000Research, 3, 126.
Ulrich, C., Abruzzese, T., Maerz, J. K., Ruh, M., Amend, B., Benz, K., et al. (2015). Human Placenta-Derived CD146-Positive Mesenchymal Stromal Cells Display a Distinct Osteogenic Differentiation Potential. Stem Cells and Development, 24, 1558–1569.
Hagmann, S., Frank, S., Gotterbarm, T., Dreher, T., Eckstein, V., & Moradi, B. (2014). Fluorescence activated enrichment of CD146+ cells during expansion of human bone-marrow derived mesenchymal stromal cells augments proliferation and GAG/DNA content in chondrogenic media. BMC Musculoskeletal Disorders, 15, 322.
Zhu, W., Tan, Y., Qiu, Q., Li, X., Huang, Z., Fu, Y., et al. (2013). Comparison of the properties of human CD146+ and CD146- periodontal ligament cells in response to stimulation with tumour necrosis factor alpha. Archives of Oral Biology, 58, 1791–1803.
Luo, Y., Zheng, C., Zhang, J., Lu, D., Zhuang, J., Xing, S., et al. (2012). Recognition of CD146 as an ERM-binding protein offers novel mechanisms for melanoma cell migration. Oncogene, 31, 306–321.
Shi, S., & Gronthos, S. (2003). Perivascular niche of postnatal mesenchymal stem cells in human bone marrow and dental pulp. Journal of Bone and Mineral Research , 18, 696–704.the official journal of the American Society for Bone and Mineral Research
Sivasankar, V., & Ranganathan, K. (2015). Growth characteristics and expression of CD73 and CD146 in cells cultured from dental pulp. Journal of Investigative and Clinical Dentistry. doi:10.1111/jicd.12155.
Shi, S., Bartold, P. M., Miura, M., Seo, B. M., Robey, P. G., & Gronthos, S. (2005). The efficacy of mesenchymal stem cells to regenerate and repair dental structures. Orthodontics & craniofacial research, 8, 191–199.
Covas, D. T., Panepucci, R. A., Fontes, A. M., Silva Jr., W. A., Orellana, M. D., Freitas, M. C., et al. (2008). Multipotent mesenchymal stromal cells obtained from diverse human tissues share functional properties and gene-expression profile with CD146+ perivascular cells and fibroblasts. Experimental Hematology, 36, 642–654.
Paduano, F., Marrelli, M., White, L. J., Shakesheff, K. M., & Tatullo, M. (2016). Odontogenic differentiation of human dental pulp stem cells on hydrogel scaffolds derived from Decellularized bone extracellular matrix and Collagen Type I. PloS One, 11, e0148225.
Huang, C. E., FW, H., CH, Y., Tsai, L. L., Lee, T. H., Chou, M. Y., et al. (2014). Concurrent expression of Oct4 and Nanog maintains mesenchymal stem-like property of human dental pulp cells. International Journal of Molecular Sciences, 15, 18623–18639.
Imbert, A. M., Garulli, C., Choquet, E., Koubi, M., & Aurrand-Lions, M. (2012). Chabannon C. CD146 expression in human breast cancer cell lines induces phenotypic and functional changes observed in Epithelial to Mesenchymal Transition. PloS One, 7, e43752.
Wagner, W., Bork, S., Horn, P., Krunic, D., Walenda, T., Diehlmann, A., et al. (2009). Aging and replicative senescence have related effects on human stem and progenitor cells. PloS One, 4, e5846.
Mitchell, J. B., McIntosh, K., Zvonic, S., Garrett, S., Floyd, Z. E., Kloster, A., et al. (2006). Immunophenotype of human adipose-derived cells: temporal changes in stromal-associated and stem cell-associated markers. Stem Cells (Dayton, Ohio), 24, 376–385.
Halfon, S., Abramov, N., Grinblat, B., & Ginis, I. (2011). Markers distinguishing mesenchymal stem cells from fibroblasts are downregulated with passaging. Stem Cells and Development, 20, 53–66.
Gharibi, B., & Hughes, F. J. (2012). Effects of medium supplements on proliferation, differentiation potential, and in vitro expansion of mesenchymal stem cells. Stem cells translational medicine, 1, 771–782.
Russell, K. C., Tucker, H. A., Bunnell, B. A., Andreeff, M., Schober, W., Gaynor, A. S., et al. (2013). Cell-surface expression of neuron-glial antigen 2 (NG2) and melanoma cell adhesion molecule (CD146) in heterogeneous cultures of marrow-derived mesenchymal stem cells. Tissue Engineering Part A, 19, 2253–2266.
Tormin, A., Li, O., Brune, J. C., Walsh, S., Schutz, B., Ehinger, M., et al. (2011). CD146 expression on primary nonhematopoietic bone marrow stem cells is correlated with in situ localization. Blood, 117, 5067–5077.
Sharma, M. B., Limaye, L. S., & Kale, V. P. (2012). Mimicking the functional hematopoietic stem cell niche in vitro: recapitulation of marrow physiology by hydrogel-based three-dimensional cultures of mesenchymal stromal cells. Haematologica, 97, 651–660.
Russell, K. C., Phinney, D. G., Lacey, M. R., Barrilleaux, B. L., Meyertholen, K. E., & O'Connor, K. C. (2010). In vitro high-capacity assay to quantify the clonal heterogeneity in trilineage potential of mesenchymal stem cells reveals a complex hierarchy of lineage commitment. Stem Cells (Dayton, Ohio), 28, 788–798.
Sorrentino, A., Ferracin, M., Castelli, G., Biffoni, M., Tomaselli, G., Baiocchi, M., et al. (2008). Isolation and characterization of CD146+ multipotent mesenchymal stromal cells. Experimental Hematology, 36, 1035–1046.
Baksh, D., Yao, R., & Tuan, R. S. (2007). Comparison of proliferative and multilineage differentiation potential of human mesenchymal stem cells derived from umbilical cord and bone marrow. Stem Cells (Dayton, Ohio), 25, 1384–1392.
Harkness L, Zaher W, Ditzel N, Isa A, Kassem M. CD146/MCAM defines functionality of human bone marrow stromal stem cell populations. Stem cell research & therapy 2016; 7:4.
Espagnolle, N., Guilloton, F., Deschaseaux, F., Gadelorge, M., Sensebe, L., & Bourin, P. (2014). CD146 expression on mesenchymal stem cells is associated with their vascular smooth muscle commitment. Journal of Cellular and Molecular Medicine, 18, 104–114.
Acknowledgments
The present study was supported by “ICARE Project - Infrastruttura Calabrese per la medicina Rigenerativa: “Generazione di biobanche per la criopreservazione di cellule staminali umane e di tessuto osseo per uso clinico e design e sviluppo di bioscaffold innovativi”. PON03PE_00009_2.2. The authors are grateful to Dr. Raghavendra Vasudeva Murthy and Dr. Pasquale Marrazzo for editing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interests
The authors declare that they have no conflicts of interest.
Additional information
Francesco Paduano; Massimo Marrelli and Marco Tatullo equally contributed to this work.
Electronic Supplementary Material
Supplementary Fig. 1
MSC surface marker expression on dental stem cells (DSCs) from various passages. Surface marker expression levels of CD146, CD13, CD29, CD44, CD73, CD90, CD105 and CD45 in a dental pulp stem cells (DPSCs), b dental follicle stem cells (DFPCs) and c periodontal ligament stem cells (PDLSCs) during various passages (p2, p4, p6, p8). (GIF 48 kb)
Supplementary Fig. 2
Growth kinetics and the osteogenic potential of hPCy-MSCs during expansion in vitro. a Long term growth curves of hPCy-MSCs, each obtained from an individual donor (n = 8). b Osteogenic differentiation of hPCy-MSCs at passage 0 and passage 10. Cells were exposed to osteogenic induction medium for 3 weeks, followed by staining with Alizarin Red. (GIF 176 kb)
Appendix
Appendix
Rights and permissions
About this article
Cite this article
Paduano, F., Marrelli, M., Palmieri, F. et al. CD146 Expression Influences Periapical Cyst Mesenchymal Stem Cell Properties. Stem Cell Rev and Rep 12, 592–603 (2016). https://doi.org/10.1007/s12015-016-9674-4
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12015-016-9674-4