Abstract
The propensity of induced pluripotent stem (iPS) cells to differentiate into specific lineages may be influenced by a number of factors, including the selection of the somatic cell type used for reprogramming. Herein we report the generation of new iPS cells, which we derived from human articular chondrocytes and from cord blood mononucleocytes via lentiviral-mediated delivery of Oct4, Klf4, Sox2, and cMyc. Molecular, cytochemical, and cytogenic analyses confirmed the acquisition of hallmark features of pluripotency, as well as the retention of normal karyotypes following reprogramming of both the human articular chondrocytes (AC) and the cord blood (CB) cells. In vitro and in vivo functional analyses formally established the pluripotent differentiation capacity of all cell lines. Chondrogenic differentiation assays comparing iPS cells derived from AC, CB, and a well established dermal fibroblast cell line (HDFa-Yk26) identified enhanced proteoglycan-rich matrix formation and cartilage-associated gene expression from AC-derived iPS cells. These findings suggest that the tissue of origin may impact the fate potential of iPS cells for differentiating into specialized cell types, such as chondrocytes. Thus, we generated new cellular tools for the identification of inherent features driving high chondrogenic potential of reprogrammed cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Induced pluripotent stem (iPS) cells are relevant cellular tools for modeling human musculoskeletal development and disease. Therefore, they hold tremendous promise for the development of specialized patient-specific therapies to restore damaged tissues with poor regenerative capacity, such as articular cartilage [1–3]. Several important aspects must, however, be addressed before human iPS cells can be used for the regeneration of cartilage, musculoskeletal disease modeling, or drug screening. Chief among these are our ability to activate the developmentally relevant signaling pathways and transcription programs to efficiently direct and restrict differentiation of pluripotent stem cells into functional chondrocytes. A wide range of developmental studies using various model systems have shed light on the inductive signal inputs and the myriad of transcription factors which orchestrate the formation of cartilage tissue from mesenchymal progenitor cells [4–6]. Applying this fundamental knowledge, we, and others, have previously demonstrated the progression of human cells from a pluripotent state to a prechondrogenic mesenchymal state within high density cultures treated with members of the TGFβ superfamily [7–12]. Within these differentiating pluripotent stem cell cultures, chondrogenic induction was associated with the expression of certain key transcription factors necessary for the formation of a cartilaginous, proteoglycan-rich extraceullalar matrix [7]. This initial work provided the framework for enriching differentiating human iPS cell cultures with a population of mesenchymal-like progenitors that exhibited high chondrogenic potential [7, 13, 14].
Traditionally, human dermal fibroblasts are recognized as a readily accessible somatic cell type with high proliferative capacity, making them well suited for iPS cell reprogramming. However, one potential limitation of dermal fibroblasts is that they also carry a higher likelihood of genetic mutations on account of their exposures to environmental insults such as ultra-violet radiation [15]. CD34+ blood cells, which can be obtained using minimally invasive procedures, and do not require the establishment of primary cultures, have gained appeal as a potential donor cell source for developing iPS cell reprogramming [16–18]. Moreover, an increasing number of reports have demonstrated that, depending on the tissue o origin, human iPS cells can harbor differences in developmental potency, transcriptional activities, and epigenetic regulation [2, 19–30]. Thus, retention of the parental cell type epigenome and transcriptome markers, even following reprogramming, can contribute to an enhanced capacity for differentiation to the cell type of origin both in vitro and in vivo [19]. Collectively, these studies raise the question as to whether the tissue of origin contributes to the future potential of iPS cells to generate chondrocytes for disease modeling, drug screening, or cell replacement therapies [23]. We generated iPS cells from human articular chondrocytes and further asked whether these cells constitute a source of iPS cells with high chondrogenic capacity. Here, we report the establishment and characterization of novel iPS cells from human articular chondrocytes as well as human cord blood cells in which we compared chondrogenicity in vitro.
Materials and Methods
iPS cell Induction and Culture
STEMCAA lentiviruses [31] pseudotyped with the vesicular somatitis virus G glycoprotein were produced by transient transfection of the packaging plasmid (pCAG-HIVgp), the VSV-G glycoprotein- and Rev-expressing plasmid (pCMV-VSV-G-RSV-Rev), and the lentiviral vector plasmid into 293 T cells. Normal human articular chondrocytes (AC) (#CC-2250, NHAC-kn, Lonza, Walkersville Inc) or cord blood mononucleocytes (#2C-150A Poietics cord blood MNC, Lonza, Walkersville Inc) were tranduced for 24 h with STEMCAA lentivirus expressing the reprogramming factors Oct3/4, Sox2, Klf4, and cMyc. Cells were cultured at 37 °C in a humidified atmosphere with 5 % CO2. Transduced cells initially cultured in chondrocyte growth media (DMEM/F12, 10 % fetal bovine serum, 1 % penicillin/streptomycin) were then passaged onto a feeder layer of irradiated mouse embryonic fibroblasts (MEFs) (20,000 cells per cm2) in the presence of embryonic stem cell-supportive media consisting of DMEM:F12, 20 % Knock-out Serum Replacement (Hyclone), 10 ng/mL basic fibrobast growth factor, β-mercaptoethanol, 1 % glutamax, 1 % nonessential amino acids, and penicillin/streptomycin. For clonal expansion of candidate iPS cells, individual colonies were manually picked by dissociation using 1 mg/mL type II collagenase (Sigma Aldrich) and replated onto irradiated MEFs on 0.1 % gelatin-coated six-well culture plates. iPS cell lines derived from either human dermal fibroblasts (HDFa-YK26) [32], cord blood (CBLR.1), or normal human articular chondrocytes (AC-iPSCs) were routinely grown as bulk cultures on irradiated MEFs and maintained in embryonic stem cell-supporting media described above and cultured for less than twenty passages.
Embryoid bodies were formed by detaching undifferentiated iPS colonies from monolayer culture using dispase (1 mg/mL). Cell clumps were washed several times with 1X PBS, then transferred to low attachment culture flasks (Corning). Cells were maintained in suspension in differentiation media consisting of DMEM high glucose supplemented with 10 % defined fetal bovine serum, 10 % serum replacement, 1 % nonessential amino acids, 1 % sodium pyruvate, 1 % glutamax, and pencillin/streptomycin for up to 21 days. Differentiation media was exchanged every 2 to 3 days.
Immunocytochemistry and Histological Staining
iPS cells were seeded onto a feeder layer of irradiated MEFs on 0.1 % gelatin coated glass coverslips, cultured for 5 days, and then fixed with 4 % paraformaldehyde. Immunostaining of nuclear proteins (Naong. Oct4) was performed by permeabilizing the cells for 10 min at room temperature in PBS buffer containing 0.4 % Triton-X-100. For immunodetection of surface proteins (SSEA3, TRA-60), cells were permeabilized with 0.1 % Triton-X-100 in PBS containing 5 % normal goat serum, and 1 % bovine serum albumin. Following blocking of nonspecific binding with 5 % normal goat serum and 0.2 % Tween-20, cells were incubated at 37 °C for 1 h with primary antibodies against SSEA3 (Santa Cruz), TRAI-60 (Santa Cruz), Nanog, and Oct4 (Stemgent). Cells were washed in PBS, then incubated with the appropriate AlexaFluor 488-conjugated secondary antibodies (Molecular Probes) for 1 h at 37 °C. Cells were mounted onto coverslips using VectaShield mounting media with DAPI (Vector Laboratories). Fluorescent images were acquired using Northern Eclipse software (Empix Imaging, Cheektowaga, NY) on an Olympus IX71 inverted microscope. Alkaline phosphatase (ALP) staining (Sigma) was performed using formalin-fixed iPS cells, as per the manufacturere’s recommendations.
Karyotype Analysis
Normal chromosome constitution of the newly-derived iPS cells was verified by conventional karyotyping. iPS cell lines were arrested in metaphase via treatment with 0.1 μg/ml colcemid for 1–2 h. Cells were trypsinized and swelled with 0.075 M KCl at 37 °C for 15–20 min. The preparation was fixed with a 3:1 methanol:glacial acetic acid solution. Cells were dropped onto acetone-cleaned slides, air-dried overnight, dehydrated, and stained by standard G-banding methods. Briefly, G-banding was performed by partial digestion of the chromosome slide preparations in a 0.005 % trypsin solution in phosphate buffered saline, pH 7.0 followed by staining in 5 % Giemsa in Gurr buffer pH 6.8 buffer. Chromosomes were analyzed for at least 20 mitotic cells per sample, and representative images were captured using an Olympus AX-71 equipped with the Genus imaging software (Applied Imaging).
Teratoma Formation
All animal procedures were carried out in accordance with protocol approval by the Institutional Animal Care and Use Committee (IACUC) at the University of Connecticut Health Center. Undifferentiated iPS cells derived from normal human articular chondrocytes (AC-iPS cells #14, passage 11) were harvested by accutase (5 min, 37 °C). 1×106 cells suspended in 10 uL DMEM/F12 media was injected intramuscularly into the right hind limb of three eight week old Scid/Nod mice (SCID, NOD.CB17-Prkdc-scid/J; Jackson Laboratory, Bar Harbor, ME) using a 28.5 gauge needle. 8 weeks after cell injections, mice were sacrificed and the tumors explanted. Tumor specimens from each of the three mice were fixed overnight in 10 % formalin, dehydrated through sequential ethanol and xylene washes, then embedded in paraffin. Paraffin-embedded tissue blocks were sectioned (6um) and stained with hematoxylin and esoin.
Pluritest/Microarray Analysis
RNA was isolated from cord blood-derived iPS cells (CB-iPS) at passage 12 using the RNeasy mini kit (Qiagen). The quality of the isolated RNA was evaluated using a BioAnalyzer. High quality RNA samples were submitted to Expression Analysis Inc for gene expression profiling using the Illumina HumanHT-12 Expression BeadChip. The PluriTest was applied for the evaluation of pluripotency based on gene expression profiles, as described in [33].
Chondrogenic Differentiation
Chondrogenic differentiation was induced by culturing iPS cells in micromass (2×105 cell/10uL drop) in defined, serum-free media as previously described [7]. On day two of micromass formation and for the duration of the differentiation assay, iPS cell micromasses were treated with human recombinant BMP-2 (100 ng/mL; R&D Systems, Minneapolis, MN). Media and growth factor were replenished every other day. Micromass cultures were harvested for RNA extraction (days 7, 14 and 21), or fixed in formalin for staining with 1 % Alcian Blue solution in acetic acid, pH 2.5 (Polysciences Inc). Semi-quantititave analyses of proteoglycan content was evaluated in day twenty-one chondrogenic micromasses generated using AC-iPS cells (#14, #15), CB-iPS cells, and HDFa-YK26 skin fibroblast (SF) iPS cells. Micromasses harvested from each iPS cell lines were fixed in formalin, embedded in paraffin and sectioned for Alcian blue staining. The relative amount of Alcian blue staining was determined as a percentage of total micromass area for 3–6 micromasses from AC-iPS cells, CB-iPS cells, and SF-iPS cells using the Image Pro software area algorithm.
Gene Expression Analyses
Total RNA was extracted from undifferentiated and differentiated iPS cell cultures with TRIzol reagent (Invitrogen). RNA concentrations were determined by measuring the absorbance at 260 nn. Samples of one microgram were treated with DNaseI (Biorad) to eliminate contamination with genomic DNA, then reverse transcribed to single stranded cDNA using iScript Reverse Transcriptase (Qiagen). 20 ng of cDNA was used for quantitative PCR analyses. Quantitative PCR was performed using SYBR Green I Mastermix (Roche) and analyzed using a 7600 Real Time qPCR machine (Applied Biosystems). Gene expression values are expressed as 2^DeltaDeltaCt, with DeltaDeltaCt defined as the difference in crossing threshold (Ct) values between undifferentiated and differentiated iPS cell samples, using Gapdh as an internal standard. Oligonucleotide primer sequences were as previously described in [7].
Statistical Analyses
Experiments were repeated at least three times using a minimum of three biological replicates per condition. Data are presented as mean of three separate experiments ± standard error. A one-way analysis of variance was performed using the Tukey’s posthoc test. For all tests, p < 0.05 was considered significant.
Results
Generation of iPS Cells from Normal Articular Chondrocytes
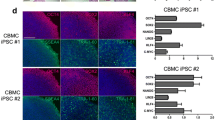
Monolayer cultures of normal human articular chondrocytes shown in Fig. 1a were transduced with the STEMCAA-lentiviral vector expressing the human reprogramming transcription factors Oct, Klf4, Sox2, and Myc [31]. Following passage onto a feeder layer of irradiated mouse embyronic fibroblasts (MEFs), transduced cells gave rise to colonies displaying morphological features characteristic of human ES cells. Several colonies were selected on the basis of morphology, then expanded onto irradiated MEFs for further characterizations. The selected articular chondrocyte-derived iPS cell clones (designated AC-iPSC #7, #14, and #15) exhibited alkaline phosphatase (ALP) staining (Fig. 1a). By real time PCR, we assessed the relative expression of canonical stemness genes including Sox2, Klf4, ALP, Oct4, and Nanog in each of the three AC-iPS cell clones as well as the parental articular chondrocytes. Little to no expression of Sox2, ALP, Oct4, or Nanog was detected in the donor articular chondrocytes, whereas, the expressions of the stemness-associated genes were significantly increased in all AC-iPS cell lines (Fig. 1b). In contrast, Klf4 expression was detected in both the parental articular chondrocytes as well as the derived AC-iPS cells. Immunofluorescent stainings for cell surface molecules SSEA3, SSEA4, TRAI60, and TRAI80, as well as nuclear factors Oct4 and Nanog, further validated the induction of pluripotency in human articular chondrocytes (Fig. 1c). Retention of the normal karyotype in AC-iPS cells was demonatrated by G-banding chromosome analyses shown in Fig. 1d.
Generation of iPS cells from normal human articular chondrocytes. a Morphology of the parental human articular chondrocytes AC in monolayer culture (i). Typical iPS cell colony following the transduction of human articular chondrocytes with STEMCAA lentiviruses is shown in (ii). Representative TRAI-81 live staining of a colony derived from AC-iPS cell clone #15 (iii). Alkaline phosphatase staining of undifferentiated AC-iPS cell clones #7, #14, and #15 are shown in (iv-vi). Scale bar, 100 μm. b Q-PCR analyses showed the induced expressions of canonical stemness genes (Nanog, Oct4, ALP) in AC-iPS cells, with little to no expression detected in the parental articular chondrocytes. Klf4 expression was detected in both the parental articular chondrocytes (AC) and AC-iPS cell clones. Gapdh served as the housekeeping gene and internal control. Gene expression data is represented as expression relative to undifferentiated human H9 ES cells, set at 1.0. d Immunofluorescence staining of pluripotency markers showed expression of nuclear Oct4 and Nanog proteins, as well as surface TRAI60 and SSEA4 antigens in AC-iPS cell clones #7, #14, and #15. DAPI nuclear counterstain is shown in blue. Scale bar, 100 μm. d G banding chromosome analyses of AC-iPS cells (#14, #15) confirmed a normal karyotype
Functional Characerization of Human Articular Chondrocyte-Derived iPS Cells
Undirected differentiation assays were performed in vitro and in vivo to confirm that AC-iPS cells exhibited the standard criteria for reprogrammed pluripotent cells. Hematoxylin and eosin (H&E) staining of three week old embryoid bodies (EBs) formed from AC-iPS cells (clones #7, #14, and #15, respectively) showed their spontaneous differentiation into multiple cell derivatives of the three germ layers (Fig. 2a). Teratoma assays were performed in three mice for in vivo assessments of the pluripotential differentiation capability of iPS cells derived from human articular chondrocytes (AC-iPS cell clone #14, passage 11). Tumors formed in all mice within 6 weeks of intra-muscular injection of the undifferentiated AC-iPS cells. As shown in Fig. 2b, discernible differentiation into tissue derivatives of each germ layer, including cartilage, bone, skeletal muscle, blood, gut-like epithelium, and neural tissue was identifed by conventional H&E staining. Thus, our collective molecular, phenotypic, and functional analyses revealed induction of pluripotency in human articular chondrocytes.
AC-iPS cells exhibit pluripotency. a Analysis of Hematoxylin-eosin (H&E) stains of 21 day embryoid bodies (EBs) formed from normal articular chondrocye-derived iPS cells (AC-iPS cells, clones #7, #14, #15). Each AC-iPS cell clone exhibited spontaneous differentiation into derivatives of ectoderm, endoderm, and mesoderm. b Histological evidence of teratoma formation 8 weeks following intramuscular injection of AC-iPS cells (clone 14, passage 11) into NOD/SCID mice. H&E staining of tumor samples showed formation of diverse tissue types derived from mesoderm (cartilage, bone, muscle, blood), endoderm (gut epithelium), and ectoderm (neural tissue). Scale bar, 100 μm
Generation of iPS Cells from Cord Blood Cells
Cord blood-derived iPS cells (CBLR.1, designated as CB-iPSCs) were generated using the same STEMCAA lentiviral-mediated reprogramming approach described above for AC-iPS cell generation. iPS cells derived from CD34-positive cord blood-derived mononucleocytes exhibited typical human ES cell morphology and expressed the canoncial pluripotency markers (Oct4, Nanog, Klf4, ALP) by mRNA analyses (Fig. 3a). To confirm that CB-iPS cells fulfilled the established criteria for induced pluripotency, we performed standardized bioninformatic analyses of genome-wide transcriptional profiles using PluriTest [33, 34]. Assessment of the expression profiles in undifferentiated CB-iPS cells confirmed Pluripotency scores that were consistent with normal, well-established human iPS cells (Fig. 3b). As shown in Fig. 3c, CB-iPS cells maintained a normal karyotype. Based on these established criteria, we successfully generated a new iPS cell line from human cord blood.
Generation of iPS cells from human cord blood. a Quantitative PCR analyses showed expression of stemness genes (Nanog, Oct4, ALP, Klf4) in cord blood-derived iPS (CB-iPS) cells. b PluriTest plot depicting the Pluripotency scores of established human iPS cells and the newly-derived CB-iPS cells. Scores indicate that undifferentiated CB-iPS cells fall within the empirically set thresholds for pluripotent cells. c G banding chromosome analysis confirmed a normal karyotype in CB-iPS cells
Enhanced Proteoglycan Accumulation in iPS Cells Derived from Articular Chondrocytes
Together with the previously established skin fibroblast-derived iPS cell line (HDFa-YK26, designated as SF-iPS), we used our newly generated human AC-iPS cells and CB-iPS cells [7, 32] to examine whether iPS cells from different tissue sources exhibit distinctive chondrogenic differentiation potentials in vitro. We previously demonstrated that culturing SF-iPS cells in micromass in serum-free media alone can induce chondrogenic differentiation [7], suggesting that cell-cell interactions are sufficient to initiate chondrogenic induction in naïve iPS cells. Alcian blue staining of day twenty-one micromass cultures formed from differentiating iPS cells enabled us to visualize the relative enrichment of cartilaginous, proteoglycan-rich matrix within iPS cells derived from skin fibroblasts (SF-iPSCs), CB-iPS cells, and AC-iPS cells (Fig. 4a). By qualitative assessment, the Alcian blue staining patterns appeared uniform within day twenty-one chondrogenic micromasses generated from AC-iPS cells (Fig. 4a). By contrast extensive cell type heterogeneity, inconsistent with the histological features of cartilage tissue, was observed in the day twenty-one micromasses generated from SF-iPS cells and CB-iPS cells (Fig. 4a). Semi-quantitiative analyses of the Alcian blue staining area was used to evaluate the relative proteoglycan content in micromasses formed from AC-iPS cells, CB-iPS cells, and SF-iPS cells. As shown in Fig. 4b, chondrogenic micromasses from AC-iPS cells (clones #14 and #15) displayed a significantly higher percentage of Alcian blue staining area (41.1 % ± 14.0, and 51.4 % ± 14.7 respectively) compared to micromasses from SF-iPS cells (21.8 % ± 8.143 SD).
Increased proteoglycan content in AC-iPS cells. a Alcian blue staining of day twenty-one micromasses harvested from SF-iPS cells, CB-iPS cells, and SF-iPS cells. Micromasses from each iPS cell line were paraffin embedded and sectioned at 6 μm, then stained with Alcian blue to visualize proteoglycan deposition. Scale bar, 100 um. b Semi-quantitative assessment of the percent Alcian blue stained micromass area showed increased proteoglycan content in 21 days chondrogenic micromasses from AC-iPS cells as compared to the SF-iPS cells. Error bars indicate standard error. * denotes significance versus SF-iPS cells, P < 0.05
Cell-Type Specific iPS Cells Display Differential Capaciites for Chondrogenic Differentiation in Vitro
Previously, we reported that stimulation of high-density iPS cell micomasses with human recombinant BMP-2 enhanced the condensation of differentiating iPS cells in micromass culture and led to temporal accumulation of Alcian-blue stained extracellular matrix [7]. At the transcriptional level, stimulation with human recombinant BMP-2 induced a significant, temporal induction of genes indicative of chondroprogenitors (Sox9) and differentiated chondrocytes (Col2a1, Aggrecan, Runx2) [7]. Thus, we used this directed differentation assay to compare the relative in vitro chondrogenic potential of AC-iPS cells, SF-iPS cells, and CB-iPS cells over a 21 days differentiation time course. In the presence of BMP-2 containing chondrogenic media, micromass cultures formed from SF-iPS cells, CB-iPS cells, and AC-iPS cells showed temporal accumulation of Alcian blue-stained proteoglycan matix (Fig. 5a).
AC-iPS cells exhibit high chondrogenic differentiation potential. a Alcian blue staining of BMP-2 treated micromasses from SF-iPS cells, CB-iPS cells, and AC-iPS cells over a time course of 7, 14, and 21 days. b-f Chondrocyte marker gene expression analyses by quantitative PCR. The relative expressions of cartilage-assoicated genes (Sox9, Col2a1, Col2B, Runx2) and the fibrocartilage marker, Col1a1 were evaluated over a time course of 21 days in BMP-2 stimulated micromasses from SF-iPS cells, CB-iPS cells, and AC-iPS cells (clone #7, #14, #15). The values represent mean fold induction over undifferentiated cells ± standard error (SE). * denotes significance versus SF-iPS cells; @ denotes significance versus CB-iPS cells, # denotes significance versus AC-iPS cell clones #15; % denotes significance versus AC-iPS cell clones #14. & denotes significance versus AC-iPS cell clones at P < 0.05, respectively
By real-time PCR analyses, we assessed the relative expressions of key chondrogenic genes in differentiating micromasses from SF-iPS cells, CB-iPS cells, and three separate AC-iPS cell clones following 7, 14 and 21 days of BMP-2 treatment. At multiple time points during the differentiation process, we observed increased Sox9 induction in certain AC-iPS cell clones compared to either SF-iPS cells or CB-iPS cells (Fig. 5b). Next we assessed the relative expression of Col2a1, a downstream transcriptional target of Sox9, and the primary extracellular consitutent of articular cartilage.. At an early stage of differentiation (day 7), the induction of Col2a1 transcripts was elevated in three AC-iPS cell clones as compared to either the SF-iPS cell line or the CB-iPS cell line. By 14 days of differentiation, Col2a1 induction was consistently higher in the AC-iPSC clones, and this elevated expression persisted in two of the AC-iPS cell clones at day 21. (Fig. 5c). Consistent with these observations, the relative expression of Col2B transcripts were significantly elevated in AC-iPS cells (clone #7, #14) compared to either SF-iPS cells or CB-iPS cells (Fig. 5d) by 21 days of differentiation. However, variable induction of aggrecan was observed among the three AC-iPS cell clones at day twenty one of differentiation (Fig. 5e). Among the different iPS cell lines examined, BMP-2 treated SF-iPS cell micromasses exhibited a significantly higher induction of the prehypertrophic/hypertrophic chondrocyte marker Runx2 as compared to the AC-iPS cell clones (Fig. 5f). We also monitored the expression of Col1a1, a marker of fibrocartilage formation, in the late stage BMP-2 treated micromass cultures from SF-iPS cells, CB-iPS cells and AC-iPS cells. Afte 21 days of differentiation, the level of Col1a1 mRNA induction was highest in SF-iPS cells, and significantly reduced in the three AC-iPS cells (Fig. 5f).
Discussion
While an improved understanding of the molecular cues driving chondrogenesis has heightened the translational appeal of iPS cell-derived progenitors, the selection of the donor cell source for reprogramming has emerged as an important consideration. Findings from previous studies utilizing both mouse and human cells supported the hypothesis that iPS cells are not created equal and that differences in their differentiation tendencies may be attributed, at least in part, to the cell type of origin [2, 19–30]. We generated new sources of iPS cells from human articular chondrocytes as well as cord blood cells using the STEMCAA lentiviral-mediated delivery of the four Yamanaka reprogramming transcription factors [31]. Through molecular, cytochemical, and cytogenetic analyses, we confirmed that all analyzed iPS cell lines derived from human articular chondrocytes and cord blood cells acquired hallmark features of pluripotent cells and retained a stable karyotype. In vitro differentiation to derivatives of the three germ layers coupled with teratoma formation in vivo validated the pluripotent potential of these new sources of cell type-specific iPS cells.
With the generation of these new lines, we compared the chondrogenic differentiation potential of the AC-iPS cells to that of an established skin fibroblast-derived iPS cell line [7, 32] and our newly derived CB-iPS cells. Our histological analyses showed a higher content proteoglycan-rich cartilaginous matrix within AC-iPS cells, suggestive of enhanced recruitment of AC-iPS cells to the chondrogenic lineage. When exposed to BMP-2, the induction of key cartilage-associated genes (Sox9, Col2a1, Col2B) was observed in micromass cultures formed from iPS cell lines derived from either skin, cartilage and blood. Most notably, induction of Col2a1, the primary extracellular matrix component of cartilage, was significantly enhanced in all three AC-iPS cell clones as compared to either SF-iPS cell line or CB-iPS celll line at early stages of the chondrogenic assay. In contrast, only a single AC-iPS cell clone (clone #15) exhibited significantly higher induction of aggrecan. We postulate that aggrecan expression may serve as a valid read-out for enhanced chondrogenic differentiation capacity in iPS cells, whereas the reduced levels of Col1a1 and Runx2 induction in each of the three AC-iPS cell clones may reflect higher quality cartilage formation. We further note that it may be challenging to identify an iPS cell line with better chondrogenic differentiation capacity on the basis of a single gene, and raise the importance of screening multiple cartilage genes over a time course of differentiation.
Previous studies have demonstrated superior differentiation of iPS cells to the cell type of origin and analyzed the importance of somatic cell type selection for the derivation of specialized cell types [2, 19–30]. Our data suggest that reprogramming surplus articular chondrocytes may be a viable approach for generating iPS cells with high chondrogenic potential. Nevertheless, we acknowledge that a more extensive evaluation of the chondrogenic differentiation capacity of multiple sources and clones of cell type-specific iPS cells will be informative. Moreover, the variability we observed in the chondrogenic responses of AC-iPS cell clones highlight the importance of clonal screening for selection of human iPS cells with high chondrogenic potential. Future studies will evaluate whether in vitro observations of enhanced chondrogenic differentiation potential among humna cell type-specific iPS cells are indeed predictive of better cartilage formation and repair in vivo.
Taken together, our newly generated AC-iPS cells clones exhibited histological features of cartilage matrix formation, with limited heterogeneity, and enhanced expression of type II collagen when cultured under chondrogenic-promoting conditions. The derivation of these new iPS cell lines and our initial studies demostrating differential chondrogenic responses in vitro, provide valuable tools for the assessment of genetic and epigenetic mechanisms governing stem cell commitment and differentiation to the chondrogenic lineage, as well as the identification of inherent features influencing the chondrogenicity of cell type-specific iPS cells.
Abbreviations
- AC:
-
articular chondrocytes
- ALP:
-
alkaline phosphatase
- BMP-2:
-
bone morphogenetic protein-2
- CB:
-
cord blood
- DMEM:
-
Dulbecco’s modified Eagle’s medium
- EB:
-
embryoid bodies
- ESC:
-
embryonic stem cell
- iPS cells:
-
induced pluripotent stem cells
- MEFs:
-
mouse embryonic fibroblasts
- SF:
-
skin fibroblasts
- SR:
-
serum replacement
References
Nakayama N. & Umeda K. (2011) From Pluripotent Stem Cells to Lineage-Specific Chondrocytes: Essential Signaling and Cellular Intermediates. Embryonic Stem Cells: The Hormonal Regulation of Pluripotency and Embryogenesis, ed Atwood C (InTech).
Robinton, D. A., & Daley, G. Q. (2012). The promise of induced pluripotent stem cells in research and therapy. Nature, 481, 295–305.
Wu, S. M., & Hochedlinger, K. (2011). Harnessing the potential of induced pluripotent stem cells for regenerative medicine. Nature Cell Biology, 13, 497–505.
Long, F., & Ornitz, D. M. (2013). Development of the endochondral skeleton. Cold Spring Harbor Perspectives in Biology, 5, a008334.
Lefebvre, V., & Smits, P. (2005). Transcriptional control of chondrocyte fate and differentiation. Birth Defects Research. Part C, Embryo Today, 75(3), 200–212.
Wuelling, M., & Vortkamp, A. (2010). Transcriptional networks controlling chondrocyte proliferation and differentiation during endochondral ossification. Pediatric Nephrology (Berlin, Germany), 25, 625–631.
Guzzo, R. M., Gibson, J., Xu, R. H., Lee, F. Y., & Drissi, H. (2013). Efficient differentiation of human iPSC-derived mesenchymal stem cells to chondroprogenitor cells. Journal of Cellular Biochemistry, 114, 480–490.
Toh, W. S., Lee, E. H., Richards, M., & Cao, T. (2010). In vitro derivation of chondrogenic cells from human embryonic stem cells. Methods in Molecular Biology, 584, 317–331.
Craft, A. M., Ahmed, N., Rockel, J. S., et al. (2013). Specification of chondrocytes and cartilage tissues from embryonic stem cells. Development (Cambridge, England), 140, 2597–2610.
Oldershaw, R. A., Baxter, M. A., Lowe, E. T., et al. (2010). Directed differentiation of human embryonic stem cells toward chondrocytes. Nature Biotechnology, 28, 1187–1194.
Diekman, B. O., Christoforou, N., Willard, V. P., et al. (2012). Cartilage tissue engineering using differentiated and purified induced pluripotent stem cells. Proceedings of the National Academy of Sciences of the United States of America, 109, 19172–19177.
Onyekwelu, I., Goldring, M. B., & Hidaka, C. (2009). Chondrogenesis, joint formation, and articular cartilage regeneration. Journal of Cellular Biochemistry, 107, 383–392.
Koyama, N., Miura, M., Nakao, K., et al. (2013). Human induced pluripotent stem cells differentiated into chondrogenic lineage via generation of mesenchymal progenitor cells. Stem Cells and Development, 22, 102–113.
Liu, Y., Goldberg, A. J., Dennis, J. E., Gronowicz, G. A., & Kuhn, L. T. (2012). One-step derivation of mesenchymal stem cell (MSC)-like cells from human pluripotent stem cells on a fibrillar collagen coating. PLoS One, 7(3), e33225.
Abyzov, A., Mariani, J., Palejev, D., et al. (2012). Somatic copy number mosaicism in human skin revealed by induced pluripotent stem cells. Nature, 492, 438–442.
Zhang, X. B. (2013). Cellular reprogramming of human peripheral blood cells. Genomics, Proteomics & Bioinformatics, 11, 264–274.
Meng, X., Neises, A., Su, R. J., et al. (2009). Efficient reprogramming of human cord blood CD34+ cells into induced pluripotent stem cells with OCT4 and SOX2 alone. Molecular Therapy, 20, 408–416.
Haase, A., Olmer, R., Schwanke, K., et al. (2009). Generation of induced pluripotent stem cells from human cord blood. Cell Stem Cell, 5, 434–441.
Bar-Nur, O., Russ, H. A., Efrat, S., & Benvenisty, N. (2011). Epigenetic memory and preferential lineage-specific differentiation in induced pluripotent stem cells derived from human pancreatic islet Beta cells. Cell Stem Cell, 9, 17–23.
Kim, K., Doi, A., Wen, B., et al. (2010). Epigenetic memory in induced pluripotent stem cells. Nature, 467, 285–290.
Kim, K., Zhao, R., Doi, A., et al. (2011). Donor cell type can influence the epigenome and differentiation potential of human induced pluripotent stem cells. Nature Biotechnology, 29, 1117–1119.
Polo, J. M., Liu, S., Figueroa, M. E., et al. (2010). Cell type of origin influences the molecular and functional properties of mouse induced pluripotent stem cells. Nature Biotechnology, 28, 848–855.
Sullivan, G. J., Bai, Y., Fletcher, J., & Wilmut, I. (2010). Induced pluripotent stem cells: epigenetic memories and practical implications. Molecular Human Reproduction, 16, 880–885.
Ohi, Y., Qin, H., Hong, C., et al. (2011). Incomplete DNA methylation underlies a transcriptional memory of somatic cells in human iPS cells. Nature Cell Biology, 13, 541–549.
Xu, H., Yi, B. A., Wu, H., et al. (2011). Highly efficient derivation of ventricular cardiomyocytes from induced pluripotent stem cells with a distinct epigenetic signature. Cell Research, 22, 142–154.
Shao, K., Koch, C., Gupta, M. K., et al. (2012). Induced pluripotent mesenchymal stromal cell clones retain donor-derived differences in DNA methylation profiles. Molecular Therapy, 21, 240–250.
Rizzi, R., Di Pasquale, E., Portararo, P., et al. (2012). Post-natal cardiomyocytes can generate iPS cells with an enhanced capacity toward cardiomyogenic re-differentation. Cell Death and Differentiation, 19, 1162–1174.
Pfaff, N., Lachmann, N., Kohlscheen, S., et al. (2012). Efficient hematopoietic redifferentiation of induced pluripotent stem cells derived from primitive murine bone marrow cells. Stem Cells and Development, 21, 689–701.
Ghosh, Z., Wilson, K. D., Wu, Y., Hu, S., Quertermous, T., & Wu, J. C. (2010). Persistent donor cell gene expression among human induced pluripotent stem cells contributes to differences with human embryonic stem cells. PLoS One, 5, e8975.
Lee, S. B., Seo, D., Choi, D., et al. (2012). Contribution of hepatic lineage stage-specific donor memory to the differential potential of induced mouse pluripotent stem cells. Stem Cells, 30, 997–1007.
Sommer, C. A., Stadtfeld, M., Murphy, G. J., Hochedlinger, K., Kotton, D. N., & Mostoslavsky, G. (2009). Induced pluripotent stem cell generation using a single lentiviral stem cell cassette. Stem Cells, 27, 543–549.
Martins-Taylor, K., Nisler, B. S., Taapken, S. M., et al. (2011). Recurrent copy number variations in human induced pluripotent stem cells. Nature Biotechnology, 29, 488–491.
Muller, F. J., Schuldt, B. M., Williams, R., et al. (2011). A bioinformatic assay for pluripotency in human cells. Nature Methods, 8, 315–317.
Muller F.J., Brandl B., Loring J.F. (2008) Assessment of human pluripotent stem cells with PluriTest. In: StemBook. Cambridge, MA; 2008.
Acknowledgments
This work was funded by the State of Connecticut Stem Cell Seed Grants (#10SCA36 and #13-SCA-UCHC-11 to RMG) and the State of Connecticut Established Investigator Grant (#11SCB08 to HD). The authors are grateful for the stem cell services and technical support provided by Leann Crandall, Tiwanna Johnson and Jung Park from the University of Connecticut Stem Cell Core and Chromosome Facility We also acknowledge Dr. Judy Brown and Dr. Rachel O’Neil from the University of Connecticut Induced Pluripotent Stem Cell Core and Chromosome Facility for their expertise with chromosome analyses. The authors have no conflict of interest to declare.
Conflicts of Interest
None.
Author information
Authors and Affiliations
Corresponding author
Additional information
This study was funded by the State of Connecticut Stem Cell Program/Department of Public Health. Contract grant numbers: #10SCA36 and #13-SCA-UCHC-11 (to RMG), #11SCB08 (to HD)
Rights and permissions
About this article
Cite this article
Guzzo, R.M., Scanlon, V., Sanjay, A. et al. Establishment of Human cell Type-Specific iPS cells with Enhanced Chondrogenic Potential. Stem Cell Rev and Rep 10, 820–829 (2014). https://doi.org/10.1007/s12015-014-9538-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12015-014-9538-8