Abstract
MicroRNAs (miRNAs) are small noncoding RNAs that are involved in key biological processes, including development, differentiation, and regeneration. The global miRNA expression profile that regulates the regenerative potential of the neonatal mouse heart has not been reported. We performed deep sequencing to determine the genome-wide miRNA expression profile of the neonatal mouse heart at three key ages (1, 6, and 7 days). The miRNAs at least 1.4-fold differentially expressed between the three time points were selected for further analysis. Two miRNAs (mmu-miR-22-5p and mmu-miR-338-3p) were significantly upregulated, and nine miRNAs (mmu-miR-324-5p, mmu-miR-337-5p, mmu-miR-339-5p, mmu-miR-365-1-5p, mmu-miR-500-3p, mmu-miR-505-5p, mmu-miR-542-5p, mmu-miR-668-3p, and mmu-miR-92a-1-5p) were significantly downregulated in cardiac tissue of 7-day-old mice compared to 1- and 6-day-old mice. The expression patterns of five significantly different miRNAs were verified by quantitative real-time PCR. Furthermore, the potential targets of these putative miRNAs were suggested using miRNA target prediction tools. The candidate target genes are involved in the myocardial regenerative process, with a prominent role for the Notch signaling pathway. Our study provides a valuable resource for future investigation of the biological function of miRNAs in heart regeneration.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
MicroRNAs (miRNAs) are an abundant class of 21–24 nucleotide noncoding RNAs that negatively regulate gene expression at the post-transcriptional level [1, 2]. At least 30 % of genes are predicted to be regulated by miRNAs. MiRNAs have been shown to play important roles in diverse biological processes, including proliferation, development, differentiation, and cell death [3]. Emerging evidence indicates that multiple miRNAs are dynamically regulated during tissue regeneration, including heart regeneration [4]. We sought to test whether miRNAs are differentially regulated during the process of regeneration in the neonatal mouse heart. Our results identify the expression profiles of miRNAs that are differentially regulated during heart regeneration. Several potential miRNA target genes involved in cardiac differentiation and the Notch signaling pathway were predicted, suggesting a role for miRNAs in heart regeneration.
Urodele amphibians and zebrafish are known to possess the capacity for cardiac regeneration [5, 6]. Poss et al. [5] reported complete cardiac regeneration in adult zebrafish within 2 months after resecting 20 % of the ventricle. However, mature mammalian cardiac myocytes have long been believed to be terminally differentiated, as they are replaced by fibrous scar tissue without regenerative capability. A recent study shows that the neonatal mouse also retains a regenerative capacity over a small window of time (ages 1–6 days), but this capacity disappears by day 7 [7]. This suggests that the mammalian heart also possesses a limited capacity to regenerate. However, it is unknown whether miRNAs are involved in this process. In this study, to identify miRNAs that regulate gene expression during cardiac regeneration, we selected three key ages (1, 6, and 7 days) to perform miRNA screening by high-throughput deep sequencing in C57BL/6 mice. We validated five differentially regulated miRNAs using quantitative real-time PCR (qRT-PCR). Furthermore, we performed a comprehensive analysis of the differentially expressed miRNA target genes and pathways based on bioinformatic analysis.
Materials and Methods
Experimental Animals and Tissue Collection
Our study had been approved by the Animal Care and Use Committee of Nanjing Medical University, China. C57BL/6J male and female mice were housed under pathogen-free conditions. Animals were kept in individual cases in a temperature-controlled room (18–24 °C) with a 12-h light/dark cycle. C57BL/6J females were mated with males at the age of 6 months. The left ventricular apex was removed from 1-day-old, 6-day-old to 7-day-old mice [7]. Specimens (five mice per group) were snap frozen in liquid nitrogen and subsequently stored at −80 °C. Total RNA was extracted using TRIzol reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions.
MiRNA Sequencing and Sequence Analysis
The small RNAs were isolated using the previously described methods [8]. RNA fragments of 18–30 bases were collected using a Novex 15 % TBE-urea gel. Next, 5′ and 3′ adaptors were ligated to the ends of the small RNAs, and the RNA was converted to DNA by reverse transcription-PCR (RT-PCR). The RT-PCR products were purified on a 6 % TBE PAGE gel. The purified cDNA library was used directly for cluster generation and high-throughput sequencing with a Solexa sequencer (Huada Genomics Institute Co. Ltd, China) according to the manufacturer’s protocol. The 50-nt sequence tags from high-throughput sequencing were screened for known miRNAs, novel miRNAs, and potential base edits of known miRNAs by the following procedure: (1) data cleaning was performed to remove low-quality reads and several varieties of contaminants, and the length distribution of these clean reads was then calculated. The length of small RNA is between 18 and 30 nt; (2) samples with common and specific sequences were grouped; (3) the expression and genomic distribution of small RNA tags were mapped by SOAP (Short Oligonucleotide Alignment Program); (4) alignments to known miRNA were performed; (5) an expression profile was generated for the known miRNAs; (6) small RNA tags were aligned to repeat-associated RNA to find matched tags in the samples; (7) the small RNA tags were screened for matches with rRNA, scRNA, snoRNA, snRNA and tRNA from GenBank, and matches to unannotated tags were eliminated; (8) the small RNA tags were screened for matches to sequences within Rfam, and matches to unannotated tags were eliminated; (9) small RNA tags were aligned to exons and introns of mRNA sequences to eliminate degraded fragments of mRNA in the small RNA tags; and (10) differential expression of known miRNAs was examined by Cluster analysis to annotate the clean tags into categories and to predict their mRNA targets.
Real-Time qPCR Validation of Differentially Expressed miRNAs
Real-time qRT-PCR was performed to confirm the differential expression of miRNAs by Solexa sequencing. Total RNA was isolated from neonatal mouse heart tissue using TRIzol (Invitrogen). A reverse transcription mixture was assembled containing 1 μg total RNA, 0.5 μl miRNA reverse primer (1 μM; Table 1), 0.3 μl RNase inhibitor (40 U/μl), 2 μl of 10× buffer, 2 μl RNasin (10 U/μl), and RNAse-free H2O to a total volume of 20 μl. The reaction mixture was incubated at 16 °C for 30 min and then 42 °C for 40 min, followed by heat inactivation at 85 °C for 5 min. Real-time qRT-PCR was performed on an ABI 7300 real-time PCR system (Applied Biosystems, Carlsbad, CA, USA) with 2.5 μl reverse transcription product, 0.5 μl forward and reverse primers each (Table 2), and 12.5 μl SYBR-Green real-time PCR master mix. The PCR cycling parameters were as follows: 95 °C for 5 min; 40 cycles of 95 °C for 10 s, 60 °C for 20 s, 72 °C for 20 s; and 78 °C for 20 s. The amount of mmu-miRNA was calculated relative to U6 small nuclear RNA (internal control) using the 2−ΔΔCt method. All reactions were run at least in triplicate.
Target Gene Prediction
Predicted targets for mmu-miR-532-5p, mmu-miR-22-5p, mmu-miR-338-3p, mmu-miR-542-5p, mmu-miR-339-5p, and mmu-miR-668-3p were analyzed using miRanda (http://microrna.sanger.ac.uk/sequences/index.shtml) and DIANA Lab (http://diana.cslab.ece.ntua.gr).
Statistical Analysis
Putative miRNA candidates were selected according to the following criteria: (1) fold-change was calculated based on the ratio of normalized counts between different time points; (2) statistical analyses were performed using one-way analysis of variance (ANOVA), and t tests or Student’s-tests were performed with a correction for multiple comparisons; and (3) values were expressed as mean ± SEM. P < 0.05 was considered to indicate a statistically significant result.
Results
Overview of the Solexa Sequencing Data
To identify miRNA expression profiles in the regenerative heart, deep sequencing of small RNA was performed using Solexa technology, and raw reads were obtained. After eliminating low-quality tags, contaminants, and trimming adaptors, we obtained 10784964, 10993620, and 10833043 clean reads from 1-, 6-, and 7-day-old groups, respectively. The length distribution of small RNA was used to analyze the composition of small RNA (Fig. 1). Most of the mature miRNAs were between 19 and 23 nucleotides (nt) for all three groups. The majority of small RNAs from the libraries were 22 nt long. The common and specific tags for the libraries, including a summary of the numbers of unique tags and total tags, are listed in Tables 3, 4, and 5. The results indicate that most of the miRNA sequences were conserved between all of the groups, with the number of group-specific sequences being much smaller.
MiRNA Expression Profiles
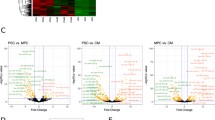
We used Solexa deep sequencing technology to evaluate the expression profiles of miRNAs from the cardiac tissue of the three groups. Comparison analysis showed the differential expression of microRNAs in myocardial tissues of 1-, 6-, and 7-day-old mice. Although these groups expressed many of the same miRNAs (Tables 3, 4, 5), the miRNA expression pattern within each group was found to vary significantly (Fig. 2). The miRNAs with at least 1.4-fold difference between all three time points were selected for further analysis. Specifically, we identified four miRNAs with increased expression and 22 with decreased expression in cardiac tissue of 6-day-old mice compared to 1-day-old mice (Table 6), eight miRNAs with increased expression and 40 miRNAs with decreased expression in cardiac tissue of 7-day-old mice compared to 1-day-old mice (Table 7), and nine miRNAs with increased expression and 10 miRNAs with decreased expression in cardiac tissue of 7-day-old mice compared to 6-day-old mice (Table 8).
Hierarchical miRNA clustering of the 1-, 6-, and 7-day-old groups (P1 represents 1-day-old group; P2 represents 6-day-old group; P3 represents 7-day-old group). Distinguishable miRNA expression profiles are observed between the three groups. Red indicates high relative expression, whereas green indicates low relative expression (Color figure online)
Validation of Differently Expressed miRNAs Using Real-Time qRT-PCR
To validate the Solexa deep sequencing, real-time qRT-PCR analysis was performed. Two upregulated miRNAs (mmu-miR-22-5p and mmu-miR-338-3p) and three downregulated miRNAs (mmu-miR-542-5p, mmu-miR-339-5p, and mmu-miR-668-3p) were selected, and expression was examined in cardiac tissue of 7-day-old mice compared to 1- and 6-day-old mice. The expression data obtained by real-time qRT-PCR analysis were consistent with the high-throughput sequencing results (Fig. 3).
Prediction of miRNA Target Genes
To better understand the biological functions of differently expressed miRNAs, the putative target sites of these miRNAs were predicted using online miRNA target prediction tools (http://microrna.sanger.ac.uk/sequences/index.shtml and http://diana.cslab.ece.ntua.gr). Among the miRNA target genes, we found that GATA1, TBX5, Baf60c, DLK1, Dvl1, and LFNG were targets of mmu-miR-532-5p, mmu-miR-22-5p, mmu-miR-338-3p, mmu-miR-542-5p, mmu-miR-668-3p, and mmu-miR-339-5p, respectively.
Discussion
It is well established that urodele amphibians and teleost fish have life-long myocardial regeneration capacity [6, 9]. A recent study found that the neonatal mouse also retains capacity for cardiac regeneration, but that this ability persists only in the first week of life [7]. The molecular mechanisms underlying this phenomenon have not been elucidated. MiRNAs have recently been shown to play a vital role in cardiac regeneration [10, 11]. We collected myocardial tissue from 1-, 6- (turning point) and 7-day-old mice, and used deep sequencing technology to determine the comprehensive miRNA expression profiles in myocardial tissue of these three groups. mmu-miR-532-5p expression gradually reduced during the period of neonatal heart development. Based on our findings from miRNA target prediction software, GATA1 was identified as a target of mmu-miR-532-5p. GATA1 is a member of the GATA family of zinc finger transcription factors, which is essential for red blood cell and megakaryocyte maturation [12]. Caprioli et al. [13] showed that the differentiation of stem cells into hematopoietic cells is suppressed at low levels of GATA1 expression, but that this transcriptional regulator may also promote the differentiation of embryonic stem cells into myocardial cells. We hypothesize that due to reduction in mmu-miR-532-5p expression, the expression of GATA1 in 7-day-old mice cardiac tissue gradually increases and thereby inhibits the regeneration of myocardial cells.
Two upregulated miRNAs (mmu-miR-22-5p and mmu-miR-338-3p) were differentially expressed in 7-day-old mice, compared to 1- and 6-day-old mice. A recent study in LO2/HBx-d382 cells indicated that miR-338-3p expression inhibits cell proliferation [14]. Given that the cardiac regenerative capacity is lost in 7-day-old mice, where mmu-miR-338-3p was expressed at the highest levels, upregulation of mmu-miR-338-3p could inhibit myocardial proliferation. Furthermore, the predicted target genes of mmu-miR-22-5p and mmu-miR-338-3p are TBX5 and Baf60c, respectively. TBX5 and Baf60c can direct ectopic differentiation of mouse mesoderm into beating cardiomyocytes, and TBX5 is essential for differentiation into contracting cardiomyocytes [15]. Ieda et al. [16] reported that GATA4, Mef2c, and TBX5 efficiently reprogrammed fibroblasts directly into functional cardiomyocytes. Based on our data and the reported role of these genes in cardiomyocyte differentiation, we hypothesize that mmu-miR-22-5p and mmu-miR-338-3p may reduce the expression of TBX5 and Baf60c, preventing the ability to regenerate the myocardium in 7-day-old mice.
A number of previous studies have identified the signaling pathways critical for cardiomyocyte proliferation, apoptosis, and regeneration [17–19]. In the present study, the target genes of the significantly downregulated miRNAs (mmu-miR-542-5p, mmu-miR-339-5p, and mmu-miR-668-3p) in cardiac tissue of 7-day-old mice included DLK1, LFNG, and Dvl1. These genes play an important role in regulating the Notch signaling pathway [20–22], providing support for a potential role for the Notch signaling pathway in the activation of heart regeneration [23].
In summary, using high-throughput Solexa sequencing, we classified a large number of miRNAs in the 1- to 7-day-old neonatal mouse heart. Importantly, we identified a series of miRNAs that are differentially expressed at these three time points. Cardiac regeneration is a complex biological process. More work will be needed to determine the functions, relationships, and mechanisms of action of miRNAs in cardiac regeneration.
References
Bartel, D. P. (2009). MicroRNAs: Target recognition and regulatory functions. Cell, 136, 215–233.
Lagos-Quintana, M., Rauhut, R., Lendeckel, W., & Tuschl, T. (2001). Identification of novel genes coding for small expressed RNAs. Science, 294, 853–858.
Mendell, J. T. (2005). MicroRNAs: Critical regulators of development, cellular physiology and malignancy. Cell Cycle, 4, 1179–1184.
Yin, V. P., Lepilina, A., Smith, A., & Poss, K. D. (2012). Regulation of zebrafish heart regeneration by miR-133. Development Biology, 365(2), 319–327.
Poss, K. D., Wilson, L. G., & Keating, M. T. (2002). Heart regeneration in zebrafish. Science, 298(5601), 2188–2190.
Oberpriller, J. O., & Oberpriller, J. C. (1974). Response of the adult newt ventricle to injury. Journal of Experimental Zoology, 187(2), 249–259.
Porrello, E. R., Mahmoud, A. I., Simpson, E., Hill, J. A., Richardson, J. A., Olson, E. N., et al. (2011). Transient regenerative potential of the neonatal mouse heart. Science, 331(6020), 1078–1080.
Lau, N. C., Lim, L. P., Weinstein, E. G., & Bartel, D. P. (2001). An abundant class of tiny RNAs with probable regulatory roles in Caenorhabditis elegans. Science, 294, 858–862.
Jopling, C., Sleep, E., Raya, M., Martí, M., Raya, A., & Belmonte, J. C. I. (2010). Zebrafish heart regeneration occurs by cardiomyocyte dedifferentiation and proliferation. Nature, 464(7288), 606–609.
Thatcher, E. J., & Patton, J. G. (2010). Small RNAs have a big impact on regeneration. RNA Biology, 7(3), 333–338.
Liu, N., & Olson, E. N. (2010). MicroRNA regulatory networks in cardiovascular development. Developmental Cell, 18(4), 510–525.
Kuhl, C., Atzberger, A., Iborra, F., Nieswandt, B., Porcher, C., & Vyas, P. (2005). GATA1-mediated megakaryocyte differentiation and growth control can be uncoupled and mapped to different domains in GATA1. Molecular and Cellular Biology, 25(19), 8592–8606.
Caprioli, A., Koyano-Nakagawa, N., Iacovino, M., Shi, X., Ferdous, A., Harvey, R. P., et al. (2011). Nk2–5 represses Gata1 gene expression and modulates the cellular fate of cardiac progenitors during embryogenesis. Circulation, 123(15), 1633–1641.
Fu, X., Tan, D., Hou, Z., Hu, Z., Liu, G., Ouyang, Y., et al. (2012). The effect of miR-338-3p on HBx deletion-mutant (HBx-d382) mediated liver-cell proliferation through cyclinD1 regulation. PLoS One, 7(8), e43204.
Takeuchi, J. K., & Bruneau, B. G. (2009). Directed transdifferentiation of mouse mesoderm to heart tissue by defined factors. Nature, 459(7247), 708–711.
Ieda, M., Fu, J. D., Delgado-Olguin, P., Vedantham, V., Hayashi, Y., Bruneau, B. G., et al. (2010). Direct reprogramming of fibroblasts into functional cardiomyocytes by defined factors. Cell, 142(3), 375–386.
Malekar, P., Hagenmueller, M., Anyanwu, A., Buss, S., Streit, M. R., Weiss, C. S., et al. (2010). Wnt signaling is critical for maladaptive cardiac hypertrophy and accelerates myocardial remodeling. Hypertension, 55, 939–945.
Kemi, O. J., Ceci, M., Wisloff, U., Grimaldi, S., Gallo, P., Smith, G. L., et al. (2008). Activation or inactivation of cardiac Akt/mTOR signaling diverges physiological from pathological hypertrophy. Journal of Cellular Physiology, 214, 316–321.
Stoick-Cooper, C. L., Weidinger, G., Riehle, K. J., Hubbert, C., Major, M. B., Fausto, N., et al. (2007). Distinct Wnt signaling pathways have opposing roles in appendage regeneration. Development, 134(3), 479–489.
Falix, F. A., Aronson, D. C., Lamers, W. H., & Gaemers, I. C. (2012). Possible roles of DLK1 in the notch pathway during development and disease. Biochimica et Biophysica Acta, 1822(6), 988–995.
Xu, K., Usary, J., Kousis, P. C., Prat, A., Wang, D. Y., Adams, J. R., et al. (2012). Lunatic fringe deficiency cooperates with the Met/Caveolin gene amplicon to induce basal-like breast cancer. Cancer Cell, 21(5), 626–641.
De Lange, R. P., Burr, K., Clark, J. S., Negrin, C. D., Brosnan, M. J., St Clair, D. M., et al. (2001). Mapping and sequencing rat dishevelled-1: A candidate gene for cerebral ischaemic insult in a rat model of stroke. Neurogenetics, 3(2), 99–106.
Raya, A., Koth, C. M., Büscher, D., Kawakami, Y., Itoh, T., Raya, R. M., et al. (2003). Activation of notch signaling pathway precedes heart regeneration in zebrafish. Proc Natl Acad Sci USA, 30(100 Suppl 1), 11889–11895.
Acknowledgments
This study was supported by Grants from the National Natural Science Foundation of China (Nos. 81070138 and 81200126) and the National Natural Science Foundation of Jiangsu Province of China (No. BK2010582).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Liu HL and Zhu JG contributed equally to this work.
Rights and permissions
About this article
Cite this article
Liu, H.L., Zhu, J.G., Liu, Y.Q. et al. Identification of the microRNA Expression Profile in the Regenerative Neonatal Mouse Heart by Deep Sequencing. Cell Biochem Biophys 70, 635–642 (2014). https://doi.org/10.1007/s12013-014-9967-7
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12013-014-9967-7