Abstract
The binding interaction of the cobalt(II) 1,10-phenanthroline complex (Co(phen) 2+3 , phen = 1,10-phenanthroline) with bovine serum albumin (BSA) was investigated by fluorescence spectroscopy combined with UV–Vis absorption and circular dichroism measurements under simulative physiological conditions. The experiment results showed that the fluorescence intensity of BSA was dramatically decreased owing to the formation of Co(phen) 2+3 –BSA complex. The corresponding association constants (K a) between Co(phen) 2+3 and BSA at four different temperatures were calculated according to the modified Stern–Volmer equation. The enthalpy change (ΔH°) and entropy change (ΔS°) were calculated to be −2.73 kJ mol−1 and 82.27 J mol−1 K−1, respectively, which suggested that electrostatic interaction and hydrophobic force played major roles in stabilizing the Co(phen) 2+3 –BSA complex. Site marker competitive experiments indicated that the binding of Co(phen) 2+3 to BSA primarily took place in site I of BSA. A value of 4.11 nm for the average distance r between Co(phen) 2+3 (acceptor) and tryptophan residues of BSA (donor) was derived from Förster’s energy transfer theory. The conformational investigation showed that the presence of Co(phen) 2+3 resulted in the change of BSA secondary structure and induced the slight unfolding of the polypeptides of protein, which confirmed the microenvironment and conformational changes of BSA molecules.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Serum albumin, as the most abundant protein in plasma, functions in the binding and transportation of various ligands such as fatty acids, hormones, and drugs [1]. The distribution, free concentration, and metabolism of these ligands strongly depend on their binding ability with serum albumin [2]. Strong binding decreases the concentrations of free ligands in plasma, whereas weak binding leads to a short lifetime or poor distribution of ligands [3]. Consequently, investigations on the affinity of ligands to serum albumins are of fundamental importance.
Cobalt is one of the indispensable microelements in human body. A characteristic feature of cobalt is the ability to form complexes with a variety of neutral molecules such as phenanthroline (phen) and bipyridine (bpy). These are widely used as a classical N,N-bidentate ligand in coordination chemistry due to their high affinity to metal ions. Generally, cobalt complexes show antitumor activities in vivo [4]. Jung et al. in their studies concerning the biological activity of several cobalt complexes have reported that these complexes can inhibit the growth of human melanoma and lung carcinoma cell lines [5]. In a recent report, Shimakoshi et al. have shown that a dicobalt complex was more efficient in cleaving DNA compared to an equivalent monomeric complex [6]. Some researches have been performed on investigation of the interaction between DNA and cobalt complexes containing 1, 10-phenanthroline ligand [7, 8]. However, the particular interaction mechanism between cobalt complexes and protein and the impact of these complexes on the conformation of protein are still unknown. In this manuscript, we report our studies on the interaction between Co(phen) 2+3 and bovine serum albumin (BSA). BSA was selected as our protein because of its medical importance, stability, and unusual ligand-binding properties [9, 10]. Studies which are directed toward the binding site- and conformation-specific reagents with protein enable chemists to extensively study the ability of these complexes to act as probes and thereby provide some scientific information at life science and clinical medicine fields.
In the present paper, the interaction between Co(phen) 2+3 and BSA was studied under physiological conditions by fluorescence spectroscopy, UV–Vis absorption, and circular dichroism (CD) spectroscopy. In order to determine the affinity of Co(phen) 2+3 to BSA and investigate the thermodynamics of their interaction, we planned to carry out detailed investigation of BSA–Co(phen) 2+3 association using fluorescence spectroscopy. CD techniques may provide information of the structural features that determine the biological effect of Co(phen) 2+3 . Particularly, the site marker competitive experiments were also carried out to determine the specific binding site of Co(phen) 2+3 to BSA. This study may provide valuable information to the great concerns regarding the biological effects of cobalt complexes on organisms.
Materials and Methods
Materials
BSA (fatty acid free) was purchased from Sigma Chemical Co. (USA) and used without further purification. Stock solution of BSA was prepared by dissolving it in Tris–HCl buffer solution (0.05 mol L−1 Tris base (2-amino-2-(hydroxymethyl)-1, 3-propanediol), 0.15 mol L−1 NaCl, pH 7.4) with the final concentration of 2.0 × 10−6 mol L−1. Co(phen) 2+3 was synthesized and characterized according to the method described in literature [11], and its molecular structure was presented in Fig. 1. The solution of Co(phen) 2+3 was prepared by dissolving it in doubly distilled water. Other chemicals were all of analytical grade, and doubly distilled water was used throughout the experiment.
Fluorescence Measurements
All fluorescence spectra were measured on an LS-55 Spectrofluorometer (Perkin-Elmer Co., USA) equipped with a water bath and a quartz cell (1.0 × 1.0 cm). In a typical fluorescence measurement, 2.0-mL BSA solution with the concentration of 2.0 × 10−6 mol L−1 was added accurately into the quartz cell and then was titrated by successive additions of 1.0 × 10−3 mol L−1 Co(phen) 2+3 using a 2-μL trace syringe to attain a series of final concentrations. Titrations were operated manually and mixed moderately.
The fluorescence emission spectra were measured at 292, 298, 304, and 310 K using a thermostat bath to maintain the temperatures. The width of the excitation and emission slits was set at 15.0 and 4.0 nm, respectively. An excitation wavelength of 285 nm was chosen, and the emission wavelength was recorded from 300 to 450 nm. The results obtained were analyzed using the Stern–Volmer equation or modified Stern–Volmer equation to calculate the quenching constants.
The three-dimensional fluorescence spectra were performed under the following conditions: the emission wavelength was recorded between 200 and 500 nm; the initial excitation wavelength was set to 200 nm with increment of 5 nm; the number of scanning curves was 31; and other scanning parameters were just the same as those of the fluorescence emission spectra.
UV–Vis Absorption and Circular Dichroism Spectra
The UV–Vis absorption spectra were measured on a TU-1901 spectrophotometer (Puxi Analytic Instrument Ltd. of Beijing, China) with the wavelength range between 200 and 320 nm. Quartz cells with 1.0-cm optical path were used.
CD measurements were performed on a J-810 spectropolarimeter (Jasco, Tokyo, Japan) at room temperature under constant nitrogen flush over a wavelength range of 270 to 190 nm. The instrument was controlled by Jasco’s Spectra Manager™ software, and the scanning speed was set at 200 nm min−1. A quartz cell having a path length of 0.1 cm was used, and each spectrum was the average of three successive scans. Appropriate buffer solutions running under the same conditions were taken as blank, and their contributions were subtracted from the experimental spectra. The concentration of BSA was kept at 2.0 × 10−6 mol L−1, and the molar ratio of BSA to Co(phen) 2+3 was varied as 1:0, 1:1, 1:3, and 1:6.
Site Marker Competitive Experiments
Binding location studies between Co(phen)3 2+ and BSA in the presence of two site markers (warfarin and ibuprofen) were measured using the fluorescence titration methods. The concentrations of BSA and warfarin/ibuprofen were all stabilized at 2.0 × 10−6 mol L−1. Co(phen) 2+3 was then gradually added to the BSA–warfarin or BSA–ibuprofen mixtures. An excitation wavelength of 285 nm was selected and the fluorescence spectra were recorded in the range of 300 to 475 nm.
Results and Discussion
Fluorescence Quenching Mechanism and Quenching Constant
Fluorescence spectroscopy has been widely used to investigate the interaction between drugs and proteins owing to its unique sensitivity, selectivity, convenience, and abundant theoretical foundation [12]. Fluorescence quenching is the decrease of the quantum yield of fluorescence from a fluorophore induced by a variety of molecular interactions, such as excited-state reactions, energy transfer, ground-state complex formation, and collisional quenching [13]. The different mechanisms of quenching are usually classified as either dynamic quenching or static quenching, which can be distinguished by their different dependence on temperature and viscosity or preferably by lifetime measurements. Dynamic quenching and static quenching are caused by collisional encounters and ground-state complex formation between fluorophores and quenchers, respectively. As higher temperatures result in larger diffusion coefficients, dynamic quenching constants are expected to be higher with increasing temperature. In contrast, the increase of temperature is likely to result in decreased stability of complexes; thus, the values of the static quenching constants are expected to be smaller [13]. In this paper, we measured the fluorescence quenching spectra of BSA in the presence of different concentrations of Co(phen) 2+3 at four different temperatures (the temperatures used were 293, 298, 303, and 308 K) to elucidate the quenching mechanism.
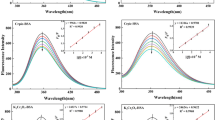
Figure 2 shows the fluorescence emission spectra of BSA in the presence of various concentrations of Co(phen) 2+3 at 298 K. When a different amount of Co(phen) 2+3 was titrated into a fixed concentration of BSA, the fluorescence intensity of BSA at around 350 nm decreased regularly, but the emission maximum did not move to shorter or longer wavelength. These results indicated that the interaction between Co(phen) 2+3 and BSA occurred and the intrinsic fluorescence of BSA has been quenched by Co(phen) 2+3 . Curve L (baseline) corresponds to the emission spectrum obtained for the blank solution containing Co(phen) 2+3 without BSA. It indicates that Co(phen) 2+3 has no effect on the fluorescence intensity of BSA at around 350 nm. The fluorescence quenching was usually analyzed using the classical Stern–Volmer equation [14]:
where F 0 and F represent the fluorescence intensities in the absence and presence of the quencher, respectively; K SV is the Stern–Volmer quenching constant; [Q] is the concentration of the quencher; τ 0 is the average fluorescence lifetime of biomolecular complex and equal to 10−8 s [15]; k q, which is equal to K SV/τ 0, is the apparent bimolecular quenching rate constant. The evaluation of k q can be useful since it reflects the efficiency of quenching or the accessibility of the fluorophores to the quencher. Diffusion-controlled quenching typically results in values of k q near 1 × 1010 M−1 S−1. The value of k q smaller than the diffusion-controlled value can result from steric shielding of the fluorophore or a low quenching efficiency. Apparent values of k q larger than the diffusion-controlled limit usually indicate some type of binding reaction [16]. Figure 3a shows the Stern–Volmer plots of F 0/F versus [Q] at the four temperatures; the calculated K SV and k q values were presented in Table 1. The results show that K SV is inversely correlated with temperature and the average value of k q is around 9.70 × 1012 M−1 S−1, which is larger than the diffusion-controlled limit. It suggests that the fluorescence quenching of BSA was caused by complex formation and the quenching mechanism may be a static quenching [17].
Emission spectra of BSA in the presence of Co(phen) 2+3 at different concentrations (T = 298 K, λ ex = 285 nm). c(BSA) = 2.0 × 10−6 mol L−1; c(Co(phen) 2+3 )/(10−6 mol L−1), A–K 0, 1.0, 2.0, 3.0, 4.0, 5.0, 6.0, 7.0, 8.0, 9.0, 10.0, respectively. Curve L shows the emission spectrum of Co(phen) 2+3 only, c(Co(phen) 2+3 ) = 2.0 × 10−6 mol L−1
One additional method to distinguish static and dynamic quenching is by careful examination of the absorption spectra of the fluorophore. Collisional quenching only affects the excited states of the fluorophores, and thus no changes in the absorption spectra are expected. In contrast, ground-state complex formation will frequently result in perturbation of the absorption spectrum of the fluorophore [16]. The UV–Vis absorption spectra of Co(phen) 2+3 , BSA, and BSA–Co(phen) 2+3 (subtracting the corresponding spectrum of Co(phen) 2+3 in the buffer) solutions were measured and shown in Fig. 4. As is depicted in Fig. 4, the absorption intensity of BSA decreased obviously at around 223 nm with the addition of Co(phen) 2+3 , which further demonstrated that the fluorescence quenching of BSA was mainly caused by complex formation between BSA and Co(phen) 2+3 . Therefore, the fluorescence quenching of BSA by Co(phen) 2+3 should be analyzed using the modified Stern–Volmer equation [18]:
In this case, ΔF is the difference in fluorescence intensity between the absence and presence of quencher at concentration [Q]; ƒ a is the fraction of accessible fluorescence, and K a is the modified Stern–Volmer association constant for the accessible fluorophores. F 0/ΔF is linear with the reciprocal value of the quencher concentration [Q], with slope equal to the value of (f a K a)−1. Figure 3b displays the modified Stern–Volmer plots, and the corresponding values of K a at different temperatures are presented in Table 2. The decreasing trend of K a with increasing temperature is in accordance with K SV's dependence on temperature, which coincides with the static type of quenching mechanism [16].
UV–Vis absorption spectra of BSA in the absence and presence of Co(phen) 2+3 . A: The absorption spectrum of Co(phen) 2+3 only, c(Co(phen) 2+3 ) = 2 × 10−6 mol L−1; B: The absorption spectrum of BSA only, c(BSA) = 2 × 10−6 mol L−1; C: The difference absorption spectra between BSA–Co(phen) 2+3 and Co(phen) 2+3 at the same concentration, c(BSA) = c(Co(phen) 2+3 ) = 2 × 10−6 mol L−1
Determination of the Force Acting Between Co(phen) 2+3 and BSA
In general, the interaction forces between endogenous or exogenous ligands and biological macromolecules may include hydrophobic force, multiple hydrogen bond, van der Waals force, electrostatic interactions, etc. [19]. The signs and magnitudes of the thermodynamic parameters (ΔH° and ΔS°) can account for the main forces involved in the binding process. To elucidate the interaction between Co(phen) 2+3 and BSA, the thermodynamic parameters were calculated from the van't Hoff plot. If the enthalpy change (ΔH°) does not vary significantly in the temperature range studied, both the enthalpy change (ΔH°) and entropy change (ΔS°) can be evaluated from the van't Hoff equation:
where K is analogous to the modified Stern–Volmer association constant K a at the corresponding temperature and R is the gas constant. The temperatures used were 292, 298, 304, and 310 K. It can be seen from Fig. 5 that there was a good linear relationship between lnK and 1/T. The enthalpy change (ΔH°) could be calculated from the slope of the van't Hoff plot. The free energy change (ΔG°) was then estimated from the following relationship:
Table 2 shows the values of ΔH° and ΔS° obtained from the slopes and ordinates at the origin of the fitted lines. The negative values of free energy (ΔG°) support the assertion that the binding process was spontaneous. According to viewpoints of Ross and Subramanian [20], the small negative enthalpy change value (−2.73 kJ mol−1, almost zero) and the positive entropy change value (82.27 J mol−1 K) of the interaction between Co(phen) 2+3 and BSA indicated that electrostatic interactions played a major role in the binding reaction. While from the point of view of water structure, a positive ΔS° value is frequently taken as evidence for hydrophobic force [21]. Moreover, as described in the literature [2], the isoelectric point of BSA is about pH 4.7, and acidity has some influences on the binding of ligands to BSA. When the value of pH is larger than 4.7, BSA has net negative charge because of the ionization of amino acid residues. In the present case, the pH value of the BSA solution is 7.4 and the Co(phen) 2+3 complexes in solution have a positive charge, which further supports the involvement of electrostatic interactions in the binding of Co(phen) 2+3 to BSA. Therefore, electrostatic interactions most likely played a major role in the binding of Co(phen) 2+3 to BSA, but hydrophobic force could not be excluded.
Number of Binding Sites and Identification of the Binding Location of Co(phen) 2+3 on BSA
When small molecules bind independently to a set of equivalent sites on a macromolecule, the equilibrium between free and bound molecules is given by the following equation [22]:
where K b and n are the binding constant and the number of binding sites, respectively. Thus, a plot of log (F 0 − F)/F versus log [Q] yields the K b and n values and the results at the four different temperatures are listed in Table 3. The linear correlation coefficient is larger than 0.999, and the standard deviation is no more than 0.010, indicating that the assumptions underlying the derivation of Eq. 5 are satisfied. As can be seen from Table 3, K b decreased with the increasing of temperature, which indicated the formation of an unstable Co(phen) 2+3 –BSA complex in the binding reaction, and it would possibly be partly disassociated when the temperature increased. The values of n at the experimental temperatures are all approximately equal to 1, suggesting that there was one class of binding sites to Co(phen) 2+3 in BSA.
Crystal structure of BSA shows that BSA is a heart-shaped helical monomer composed of three homologous domains named I, II, III, and each domain includes two subdomains called A and B to form a cylinder. The principal regions of ligand-binding sites on albumin are located in hydrophobic cavities in subdomains IIA and IIIA, which exhibit similar chemistry properties [23]. The binding cavities associated with subdomains IIA and IIIA are also referred to as site I and site II according to the terminology proposed by Sudlow et al. [24]. As the data in the preceding discussion did not allow us to give the precise binding location of Co(phen) 2+3 on BSA, the site marker competitive experiments were then carried out, using drugs which specifically bind to a known site or region on BSA. As described in the literature, warfarin has been demonstrated to bind to subdomain IIA, while ibuprofen is considered as subdomain IIIA binder [25]. Then, information about the Co(phen) 2+3 binding site can be gained by monitoring the changes in the fluorescence intensity of Co(phen) 2+3 -bound albumin that was brought about by site I (warfarin) and site II (ibuprofen) markers (Fig. 6).
Influence of selected site markers on the fluorescence of Co(phen) 2+3 bound to BSA (T = 298 K, λ ex = 285 nm). a c(BSA) = c(warfarin) = 2.0 × 10−6 mol L−1; b c(BSA) = c(ibuprofen) = 2.0 × 10−6 mol L−1; c(Co(phen) 2+3 )/(10−6 mol L−1), A–K: 0, 1.0, 2.0, 3.0, 4.0, 5.0, 6.0, 7.0, 8.0, 9.0, 10.0, respectively. The inserts correspond to the molecular structures of site markets
During the experiment, Co(phen) 2+3 was gradually added to the solution of BSA with site markers held in equimolar concentrations (2.0 × 10−6 mol L−1). As shown in Fig. 6a, with addition of warfarin into the BSA solution, the maximum emission wavelength of BSA had a slight red shift, and the fluorescence intensity was significantly lower than that of the solution without warfarin. Then, with the continuing addition of Co(phen) 2+3 into the above system, the fluorescence intensity of the BSA solution with warfarin held in equimolar concentrations decreased gradually, and the intensity was much lower than that of the solution without warfarin (Fig. 2), indicating that the binding of Co(phen) 2+3 to BSA was affected by adding warfarin. By contrast, in the presence of ibuprofen, the fluorescence intensity of the Co(phen) 2+3 –BSA complex was almost the same as that of the solution without ibuprofen (Figs. 2 and 6b), indicating that site II marker ibuprofen did not prevent the binding of Co(phen) 2+3 in its usual binding location. In order to facilitate the comparison of the influence of warfarin and ibuprofen on the binding of Co(phen) 2+3 to BSA, the fluorescence quenching data of the Co(phen) 2+3 –BSA system with the presence of site markers were also analyzed using the modified Stern–Volmer equation, as shown in Fig. 7. The modified Stern–Volmer association constants of the systems, which can be calculated from the slope values of the plots, were listed in Table 4. The association constant of the system with warfarin was approximately 60% of that without warfarin, while the constant of the system with and without ibuprofen had only a small difference. It indicated that warfarin could significantly affect the binding of Co(phen) 2+3 to BSA while ibuprofen had only a small influence on the binding of Co(phen) 2+3 to BSA. The above experimental results and analysis demonstrated that the binding of Co(phen) 2+3 to BSA is mainly located within site I (subdomain IIA).
Binding Distance Between Co(phen) 2+3 and BSA
Energy transfer phenomena have wide applications in energy conversion process [26]. According to Förster theory of nonradioactive energy transfer [27], a transfer of energy could take place through direct electrodynamic interaction between the primarily excited molecule and its neighbors, which will happen under the following conditions: (1) the donor can produce fluorescence light; (2) fluorescence emission spectrum of the donor and UV–Vis absorbance spectrum of the acceptor have overlap; and (3) the distance between the donor and the acceptor approaches and is lower than 8 nm. The energy transfer efficiency is related not only to the distance between the acceptor and donor but also to the critical energy transfer distance, R 0, as described in Eq. 6:
where r represents the distance between the donor and acceptor. R 0, the critical distance at which transfer efficiency equals to 50%, is given by the following equation [28]:
where K 2 is the orientation factor related to the geometry of the donor–acceptor dipole; n is the refractive index of medium; ϕ is the fluorescence quantum yield of the donor, and J expresses the degree of spectral overlap between the donor emission and the acceptor absorption, which could be calculated by the following equation [28]:
where F(λ) is the fluorescence intensity of the donor at wavelength range λ; ε(λ) is the molar absorption coefficient of the acceptor at wavelength λ. By assumption that most of the fluorescence quenching of BSA is from Förster energy transfer process, the binding distance between Co(phen) 2+3 and BSA can be obtained. The overlap of the absorption spectrum of Co(phen) 2+3 and the fluorescence emission spectrum of BSA is shown in Fig. 8. The overlap integral, J, can be evaluated by integrating the spectra in Fig. 8 according to Eq. 8. In the present case, K 2 = 2/3, n = 1.336, and \( \varphi \) = 0.15 [29]. From Eqs. 6–8, we would be able to calculate that J = 3.57 × 10−14 cm3 L mol−1, R 0 = 3.12 nm, and E = 0.16. Then, the binding distance r between Co(phen) 2+3 and BSA may be as close as 4.11 nm. The donor-to-acceptor distance r is less than 8 nm, indicating that the energy transfer from BSA to Co(phen) 2+3 occurred with high possibility [30].
Conformational Investigation of BSA After the Co(phen) 2+3 Binding
We had ascertained that it was the complex formation between Co(phen) 2+3 and BSA that caused the fluorescence quenching of BSA, but it is still a puzzle about whether the binding affects the conformation and/or microenvironment of BSA. To further investigate BSA structure after Co(phen) 2+3 binding, the methods of synchronous fluorescence, CD and three-dimensional fluorescence spectroscopy were utilized.
The synchronous fluorescence spectra can provide information about the molecular microenvironment in a vicinity of the chromophore molecules [31]. Yuan et al. [32] suggested a useful method to study the environment of amino acid residues by measuring the possible shift in wavelength emission maximum, λ max, which correspond to the changes of the polarity around the chromophore molecule. When the D value (Δλ) between excitation and emission wavelength are stabilized at 60 nm, the synchronous fluorescence gives the characteristic information of tryptophan residues [33]. As shown in Fig. 9, the maximum emission wavelength had a slight red shift (from 286 to 289 nm) at the investigated concentrations range. It indicates that the polarity around the tryptophan residues was increased and the hydrophobicity was decreased [34].
CD spectra, a sensitive technique to monitor the secondary structural change of protein [35], have been employed to investigate the structural changes of BSA after the binding of Co(phen) 2+3 . The CD spectra of BSA with various concentrations of Co(phen) 2+3 at pH 7.4 and room temperature are shown in Fig. 10. As is shown in Fig. 10, BSA exhibited two negative bands at around 208 and 222 nm in the ultraviolet region, characteristic of α-helix structure in protein. The band at 222 nm is contributed to the n→π* transition for the peptide bond of α-helix [36]. The CD spectra of BSA in the presence and absence of Co(phen) 2+3 were similar in shape, indicating that the structure of BSA was still predominantly α-helix.
In order to quantify the different types of secondary structure content, the CD spectra have been analyzed by the algorithm SELCON3, with 43 mode proteins with known precise secondary structures used as the reference set [37, 38]. The fraction contents of different secondary structures for BSA in the absence and presence of Co(phen) 2+3 are presented in Table 5. A decreased percentage of the α-helical content and an increased percentage of β-strands, turn, and unordered structure contents were observed with the increasing concentration of Co(phen) 2+3 (Table 5). As the secondary structure contents are related close to the ligand binding and transport capability of BSA, the secondary structural changes here meant the loss of the biological activity of BSA upon interaction with high concentration of Co(phen) 2+3 . The conformational changes here meant that Co(phen) 2+3 bound with the amino acid residues of the main polypeptide chain of BSA and destroyed their hydrogen bonding networks, making the serum albumin adopt a more incompact conformation state [39].
Three-dimensional fluorescence spectra, which can comprehensively exhibit the fluorescence information and conformational change of protein [3], were also employed in the present study. The three-dimensional fluorescence spectra of BSA and Co(phen) 2+3 –BSA complex are shown in Fig. 11; the corresponding characteristic parameters are presented in Table 6. By comparing the spectral changes of BSA in the absence and presence of Co(phen) 2+3 , the conformational and microenvironmental changes of BSA can be obtained. As Fig. 11 showed, peak a is the Rayleigh scattering peak (λ ex = λ em); peak b is the second-ordered scattering peak (λ em = 2λ ex) [17], and the fluorescence intensity of peak a increased with the addition of Co(phen) 2+3 . The reasonable explanation is that a BSA–Co(phen) 2+3 complex came into being after the addition of Co(phen) 2+3 , making the diameter of the macromolecule increased, which in turn resulted in the enhanced scattering effect [40]. Peak 1 (λ ex = 280.0 nm, λ em = 351.0 nm), which mainly reveals the spectral behavior of tryptophan and tyrosine residues, is the primary fluorescence peak we studied. Besides peak 1, there is another strong fluorescence peak 2 (λ ex = 230.0 nm, λ em = 352.0 nm) that mainly exhibits the fluorescence spectral behavior of polypeptide backbone structures, and the fluorescence intensity of this peak is correlated with the secondary structure of protein [3]. As shown in Fig. 11, the fluorescence intensity of peak 2 decreased obviously after the addition of Co(phen) 2+3 , which means that the secondary structure of BSA has been changed. This result was in accordance with what we got from the CD spectra. Analyzing from the fluorescence intensity changes of peak 1 and peak 2 (the intensity values were listed in Table 6), they both decreased obviously but to different degrees: the fluorescence intensity of peak 1 has been quenched to 17.2% while peak 2 to 19.7%. The decrease of the fluorescence intensity of the two peaks in combination with the synchronous fluorescence and CD spectra results indicated that the interaction of Co(phen) 2+3 with BSA induced the slight unfolding of the polypeptides of protein, which resulted in a conformational change of the protein to increase the exposure of some hydrophobic regions that had been buried [41]. All these phenomenon and analysis revealed that the binding of Co(phen) 2+3 to BSA induced some microenvironmental and conformational changes in BSA.
Conclusions
This paper provided an approach for studying the binding of Co(phen) 2+3 to BSA by employing different optical techniques. The studies presented here demonstrated that the fluorescence quenching of BSA resulted mainly from static mechanism and electrostatic interactions, and hydrophobic force played major roles in stabilizing the Co(phen) 2+3 –BSA complex. Experimental results showed that Co(phen) 2+3 was a strong quencher of the fluorescence of BSA and bound to the protein with high affinity. From site marker competitive experiments, it appeared that the binding site of Co(phen) 2+3 on the protein is around subdomain IIA. Results from the conformational investigation suggested that BSA underwent some conformational changes at secondary structure levels, which further indicated that the microenvironments of BSA were changed and its biological activity was weakened in the presence of Co(phen) 2+3 .
References
Zolese G, Falcioni G, Bertoli E, Galeazzi R, Wozniak M, Wypych Z, Gratton E, Ambrosini A (2000) Steady-state and time resolved fluorescence of albumins interacting with N-oleoylethanolamine, a component of the endogenous N-acylethanolamines. Proteins Struct Funct Genet 40:39–48
Kragh-Hansen U (1981) Molecular aspects of ligand binding to serum albumin. Pharmacol Rev 33:17–53
Zhang YZ, Zhou B, Liu YX, Zhou CX, Ding XL, Liu Y (2008) Fluorescence study on the interaction of bovine serum albumin with P-aminoazobenzene. J Fluoresc 18:109–118
Osinsky SP, Levitin IY, Sigan AL, Bubnovskaya LN, Ganusevich II, Campanella L, Wardman P (2003) Redox-active cobalt complexes as promising antitumor agents. Russ Chem Bull 52:2636–2645
Jung M, Kerr DE, Senter PD (1997) Bioorganometallic chemistry—synthesis and antitumor activity of cobalt carbonyl complexes. Arch Pharm Pharm Med Chem 330:173–176
Shimakoshi H, Kaieda T, Matsuo T, Sato H, Hisaeda Y (2003) Syntheses of new water-soluble dicobalt complexes having two cobalt-carbon bonds and their ability for DNA cleavage. Tetrahedron Lett 44:5197–5199
Peng B, Chao H, Sun B, Li H, Gao F, Ji LN (2007) Synthesis, DNA-binding and photocleavage studies of cobalt(III) mixed-polypyridyl complexes: [Co(phen)2(dpta)]3+ and [Co(phen)2(amtp)]3+. J Inorg Biochem 101:404–411
Kumar RS, Arunachalam S, Periasamy VS, Preethy CP, Riyasdeen A, Akbarsha MA (2008) Synthesis, DNA binding and antitumor activities of some novel polymer–cobalt(III) complexes containing 1, 10-phenanthroline ligand. Polyhedron 27:1111–1120
Guharay J, Sengupta B, Sengupta PK (2001) Protein–flavonol interaction: fluorescence spectroscopic study. Proteins 43:75–81
Zhang YZ, Zhang XP, Hou HN, Dai J, Liu Y (2008) Study on the interaction between Cu(phen) 2+3 and bovine serum albumin by spectroscopic methods. Biol Trace Elem Res 121:276–287
Inskeep RG (1962) The spectra of the tris complexes of 1, 10-phenanthroline and 2, 2-bipyridine with the transition metals iron(II) through zinc(II). J Inorg Nucl Chem 24:763–776
Cui FL, Wang JL, Cui YR, Li JP, Yao XJ, Lu Y, Fan J (2007) The binding of 3-(p-bromophenyl)-5-methyl-thiohydantoin with human serum albumin: investigation by fluorescence spectroscopy and molecular model. J Luminesc 127:409–415
Hu YJ, Liu Y, Zhang LX, Zhao RM, Qu SS (2005) Studies of interaction between colchicine and bovine serum albumin by fluorescence quenching method. J Mol Struct 750:174–178
Gelamo EL, Silva CHTP, Imasato H, Tabak M (2002) Interaction of bovine and human serum albumins with ionic surfactants: spectroscopy and modeling. Biochim Biophys Acta 1594:84–99
Lakowicz JR, Weber G (1973) Quenching of fluorescence by oxygen, a probe for structural fluctuations in macromolecules. Biochemistry 12:4161–4170
Lakowicz JR (2006) Principles of fluorescence spectroscopy, 3rd edn. Plenum, New York, pp 277–285
Mei P, Zhang YZ, Zhang XP, Yan CX, Zhang H, Liu Y (2008) Spectroscopic investigation of the interaction between copper (II) 2-oxo-propionic acid salicyloyl hydrazone complex and bovine serum albumin. Biol Trace Elem Res 124:269–282
Lehrer SS (1971) The quenching of the tryptophyl fluorescence of model compounds and of lysozyme by iodide ion. Biochemistry 10:3254–3263
Leckband D (2000) Measuring the forces that control protein interactions. Annu Rev Biophys Biomol Struct 29:1–26
Ross PD, Subramanian S (1981) Thermodynamic of protein association reactions: forces contributing to stability. Biochemistry 20:3096–3102
Wang YQ, Zhang HM, Zhang GC, Tao WH, Fei ZH (2007) Spectroscopic studies on the interaction between silicotungstic acid and bovine serum albumin. J Pharm Biomed Anal 43:1869–1875
Jin J, Zhang X (2008) Spectrophotometric studies on the interaction between pazufloxacin mesylate and human serum albumin or lysozyme. J Luminesc 128:81–86
Zhang GW, Que QM, Pan JH, Guo JB (2008) Study of the interaction between icariin and human serum albumin by fluorescence spectroscopy. J Mol Struct 881:132–138
Sudlow G, Birkett DJ, Wade DN (1976) Further characterization of specific drug binding sites on human serum albumin. Mol Pharmacol 12:1052–1061
He XM, Carter DC (1992) Atomic structure and chemistry of human serum albumin. Nature 358:209–215
Naik DB, Moorty PN, Priyadarsini KI (1990) Nonradiative energy transfer from 7-amino coumarin dyes to thiazine dyes in methanolic solutions. Chem Phys Lett 168:533–538
Förster T (1965) Delocalized excitation and excitation transfer. In: Sinanoglu O (ed) Modern quantum chemistry, vol 3. Academic, New York, pp 93–137
Sklar LA, Hudson BS, Simoni RD (1977) Conjugated polyene fatty acids as fluorescent probes: synthetic phospholipid membrane studies. Biochemistry 16:819–828
Mahammed A, Gray HB, Weaver JJ, Sorasaenee K, Gross Z (2004) Amphiphilic corroles bind tightly to human serum albumin. Bioconjugate Chem 15:738–746
Li DJ, Zhu JF, Jin J, Yao XJ (2007) Studies on the binding of nevadensin to human serum albumin by molecular spectroscopy and modeling. J Mol Struct 846:34–41
Zhang Y, Qi ZD, Zheng D, Li CH, Liu Y (2009) Interactions of chromium (III) and chromium (VI) with bovine serum albumin studied by UV spectroscopy, circular dichroism, and fluorometry. Biol Trace Elem Res 130:172–184
Yuan T, Weljie AM, Vogel HJ (1998) Tryptophan fluorescence quenching by methionine and selenomethionine residues of calmodulin: orientation of peptide and protein binding. Biochemistry 37:3187–3195
Miller JN (1979) Recent advances in molecular luminescence analysis. Proc Anal Div Chem Soc 16:203–209
Klajnert B, Bryszewska M (2002) Fluorescence studies on PAMAM dendrimers interactions with bovine serum albumin. Bioelectrochemistry 55:33–35
Yue YY, Zhang YH, Li Y, Zhu JH, Qin J, Chen XG (2008) Interaction of nobiletin with human serum albumin studied using optical spectroscopy and molecular modeling methods. J Luminesc 128:513–520
Yang P, Gao F (2002) The principle of bioinorganic chemistry. Science, Beijing, p 349
Sreerama N, Woody RW (1993) A self-consistent method for the analysis of protein secondary structure from circular dichroism. Anal Biochem 209:32–44
Whitmore L, Wallace BA (2004) An online server for protein secondary structure analyses from circular dichroism spectroscopic data. Nucleic Acids Res 32:668–673
Cui FT, Fan J, Hu ZD (2004) Interactions between 1-benzoyl-4-p-chlorophenyl thiosemicarbazide and serum albumin: investigation by fluorescence spectroscopy. Bioorg Med Chem 12:151–157
Zhang YZ, Dai J, Zhang XP, Yang X, Liu Y (2008) Studies of the interaction between Sudan I and bovine serum albumin by spectroscopic methods. J Mol Struct 888:152–159
Tian JN, Liu JQ, Hu ZD, Chen XG (2005) Interaction of wogonin with bovine serum albumin. Bioorg Med Chem 13:4124–4129
Acknowledgements
We gratefully acknowledge the financial support of Chinese 863 Program (2007AA06Z407); National Natural Science Foundation of China (grant nos. 30570015, 20621502).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, YZ., Li, HR., Dai, J. et al. Spectroscopic Studies on the Binding of Cobalt(II) 1,10-Phenanthroline Complex to Bovine Serum Albumin. Biol Trace Elem Res 135, 136–152 (2010). https://doi.org/10.1007/s12011-009-8502-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12011-009-8502-y