Abstract
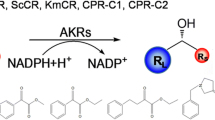
(R)-3, 5-Bis(trifluoromethyl)phenyl ethanol is a key chiral intermediate for the synthesis of aprepitant. Through a genome mining approach, an NADPH-dependent short-chain dehydrogenases derived from Burkholderia cenocepacia (Bc-SDR) was discovered with excellent anti-Prelog’s stereoselectivity of reducing 3, 5-bis(trifluoromethyl) acetophenone. The enzyme with 247 amino acids was successfully expressed in Escherichia coli and the molecular weight was about 26 kDa. Optimization of reaction conditions showed that the optimum temperature and pH of the enzyme was 25 °C and pH 7.0, respectively. Strong enhancement of enzyme activity was observed in the presence of 1 mM Mn2+. In addition, Bc-SDR exhibited (R)-selective enantioselectivity toward acetophenone derivatives, which makes it a potential catalyst for obtaining aromatic chiral alcohols as useful blocks in pharmaceutical applications.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Biocatalysis is well matched to chemical synthesis in the manufacture of pharmaceutical intermediates [1]. Through three waves of research and technological innovations [2], using enzyme catalysis is deemed as a key step in industrial pharmaceutical synthesis. In recent years, the number of biocatalysts used for the production of chiral compounds has rapidly increased. Chiral alcohols are among the most valuable key intermediates for the manufacture of pharmaceuticals. Carbonyl reductases, application in ketone reduction, are now often a preferred catalyst for the synthesis of chiral alcohols for its reliability chemoselectivity and stereoselectivity [3, 4].
3, 5-Bis(trifluoromethyl) acetophenone (3,5-BTAP) is a key precursor of (R)-3,5-bis (trifluoromethyl)phenyl ethanol ((R)-3,5-BTPE), which is an important chiral intermediate for the synthesis of aprepitant [5, 6]. Aprepitant as a neurokinin-1 receptor antagonist is widely used in the prevention of chemotherapy-induced nausea and vomiting [7, 8].
In recent years, asymmetric reduction of the prochiral ketone 3, 5-BTAP via microorganisms or enzyme catalysis has been studied to produce highly optically active (R)-3, 5-BTPE. It was reported that only five microorganisms, such as Leifsonia xyli, Lactobacillus kefir, Penicillium expansum, Microbacterium oxydans, and Trichoderma asperellum ZJPH0801, can enantioselectively reduce 3,5-BTAP to (R)-3,5-BTPE with > 99% ee, but with unsatisfied yields [5, 6, 9,10,11]. Wang et al. also did a series of investigation. They purified a carbonyl reductase from L. xyli HS0904 and determined its biochemical properties [12]. Then, they reported that a mutant form of carbonyl reductase from L. xyli HS0904 reduced 3, 5-BTAP to (R)-3, 5-BTPE at a high substrate concentration of 256 g/L in the presence of 20% (v/v) isopropanol, but with an unsatisfactory 82.5% product yield [13]. Except theses five natural “producers” of (R)-3,5-BTPE, few ketone/carbonyl reductases have been discovered with high production of (R)-3,5-BTPE from 3,5-BTAP. Ketone reductase ChKRED20 from Chryseobacterium sp. CA49 reduce 3, 5-BTAP to (R)-3, 5-BTPE with >99% conversion and 99.9% ee. But ChKRED20 requires 40% (v/v) isopropanol as a co-substrate and expensive nicotinamide adenine dinucleotide phosphate [NAD (P) +] in the oxidized form as a cofactor [14]. Ketoreductase P1B2, the commercially available ketoreductase from Codexis, reduced 3, 5-BTAP to (R)-3, 5-BTPE with 98% conversion and >99% ee, also requires 30% (v/v) isopropanol as co-substrate and expensive NAD(P) + as cofactor [15]. Recently, an NADPH-dependent carbonyl reductase was discovered from L. kefir and coexpressed with glucose dehydrogenase in E. coli to efficiently synthesize (R)-3, 5-BTPE by whole-cell biotransformation [16].
However, the carbonyl reductase that can produce (R)-3, 5-BTPE is still rare, because the overwhelming majority of enzymatic and microbial reductions follow the Prelog’s rule [17]. The enzyme prefers to transfer the pro-(R) hydrogen of NAD (P)H to the re-face of the ketone in the hydrogen transfer process to generate the (S)-configuration alcohols in nature [6]. Therefore, it is of great significance in searching for a carbonyl reductase with anti-Prelog’s specificity on 3, 5-BTAP.
B. cenocepacia was a promising catalyst for the reduction of ketones [18], and we found that it was able to reduce 3, 5-BTAP to the corresponding alcohol with considerable enantioselectivity. Therefore, in this study, to find a catalyst with excellent enantioselectivity toward 3, 5-BTAP, 20 known or putative carbonyl reductase genes from the genomic DNA of B. cenocepacia by a genome mining approach were selected. An NADPH-dependent short-chain dehydrogenases/ reductases (NCBI Reference Sequence: WP_012492403.1) from B. cenocepacia (Bc-SDR) was discovered as a practical catalyst to generate optically pure (R)-3,5-BTPE for synthesizing aprepitant (Fig. 1) and chosen for further studies.
Material and Methods
Chemicals, Microorganisms, and Plasmids
3, 5-BTAP was purchased from Beijing Ouhechem Company (China). (R)-3, 5-B TPE was prepared using the synthesis method through reduction of 3, 5-BTAP [19]. Acetophenone, the acetophenone derivatives, and other chemical reagents involved in this study were analytical grade and obtained from commercial sources. All organic solvents of high-performance liquid chromatography (HPLC) grade were purchased from Acros (USA).
B. cenocepacia was purchased from the China General Microbiological Culture collection Center. Restriction enzymes (NdeI and HindIII), rTaq DNA polymerase, T4 DNA ligase were purchased from Takara (Japan). The expression vector pET-22b (+) was purchased from invitrogen. E. coli DH5α and Rosetta (DE3) that were purchased from TransGen Biotech Company (China) were used as a cloning host and a expression host, respectively. The QIAquick PCR purification kit was purchased from QIAGEN (Germany). The genome DNA purification kit and plasmid extraction kit were obtained from Tiangen (China). All primers used in this study were synthesized by Sangon Biotech (China). Other materials were obtained from commercial sources.
Construction of the Recombinant Strains
All genes were searched from the NCBI database (http://www.ncbi.nlm.nih. gov). The non-redundant protein sequences of B. cenocepacia genomes were searched through a BLAST by using the carbonyl reductase sequence (GenBank: AGW01025.1) from L. xyli HS0904 as a template. Twenty putative carbonyl reductase genes (Fig. S1) were amplified through polymerase chain reaction and then inserted into expression vector pET-22b (+) by using the NdeI and HindIII restriction sites. Recombinant vector was transformed into E. coli DH5α for sequencing prior to introduce into E. coli Rosetta (DE3) for expression.
The recombinant strains were cultivated in 250 mL lysogeny broth (LB) medium (0.5% (w/v) yeast extract, 1% (w/v) peptone, and 1% (w/v) NaCl) including 50 μg/mL ampicillin at 37 °C with shaking at 220 rpm. When cells were grown to an optical density of 0.6–0.8 (OD) at 600 nm, isopropyl-β-d-thiogalactopyran-oside (IPTG) was added to the culture medium with a final concentration of 0.2 mM, and the temperature was decreased to 28 °C. After 16 h, the cells were harvested by centrifugation (10 min, 6000 rpm) at 4 °C.
Enzyme Activity Assay
Activity of recombinant strains were measured by reduction of 3,5-BTAP using whole cells. The reaction conditions contained 250 mg wet cell mass and 10 mM BTAP (dissolved in 50 μl DMSO) as substrate in 1 mL phosphate buffer (100 mM, pH 7.3). The mixture was incubated at 30 °C for 24 h and was extracted with 1 mL hexane by centrifugation. Then, 10 μL supernatant of organic solution was analyzed by HPLC. The analytical procedures of HPLC are as follows: column, Chiralcel OD-H (4.6 mm × 250 mm, 5 μm, Diacel Chemical industries, USA); eluent, 2-propanol/hexane 2:98; temp, 25 °C; flow rate, 0.75 mL/min; detection, 206 nm light; retention time of 3,5-BTAP, 5.76 min; retention time of 3,5-BTPE, t S = 10.96 min; t R = 12.55 min. One unit of enzyme activity was defined as the amount of enzyme that reduced 1 μM 3,5- BTAP per hour under above reaction conditions.
Optimization of Reaction Conditions
First, the optimum reaction time was determined. In a typical assay, 250 mg/mL wet cells were incubated with 10 mM 3, 5-BTAP in 1 mL phosphate buffer (100 mM, pH 7.3). The reactions were carried out at 30 °C at 220 rpm for different time ranging from 6 to 72 h. Then reaction mixtures were extracted with 1 mL hexane and analyzed by HPLC.
To test the optimal concentration of wet recombinant cells used in the reaction, different amounts of wet cells ranging from 50 to 750 mg were suspended in 1 mL phosphate buffer (100 mM, pH 7.3) containing 10 mM 3,5-BTAP. The assay mixtures were then incubated at 30 °C at 220 rpm for 24 h. After reaction, the mixtures were extracted and analyzed by HPLC.
The optimum reaction temperature of reducing 3, 5-BTAP by using Bc-SDR was investigated. To find the optimum temperature, experiment was carried out at different temperatures varying from 20 to 50 °C by adding 250 mg wet cells and 10 mM 3,5-BTAP (dissolved in 50 μl DMSO) to 1 mL phosphate buffer (100 mM, pH 7.3) for 24 h. The highest activity was set to 100%, and others can be calculated relatively.
To study the optimum pH, a different buffer system (100 mM) at the pH range of 5.0–10.0 (phosphate buffer (NaH2PO4–Na2HPO4) for pH 5.0–9.0 and NaHCO3–Na2CO3 for pH 9.0–10.0) was prepared. The experiment was performed at above standard condition, and the reaction temperature was 25 °C. The highest activity was set to 100%, and others can be calculated relatively.
The effect of metal ions on the activity of Bc-SDR was examined at a final concentration of 1 mM metal ions. Thirteen metal ions (Li+, K+, Na+, Ni2+, Zn2+, Mg2+, Sn2+, Co2+, Ca2+, Mn2+, Cu2+, Fe2+, Fe3+) were investigated under the optimum temperature and pH value at above standard condition for 24 h. Under this assay system, the activity of control group without metal ions was set to 100%, and others were expressed as a relative activity.
We also investigated the effects of organic solvent/water (v/v = 1/2) biphasic systems. Nine different kinds of organic solvents including ethyl acetate, methyl tert-butyl ether, isopropyl ether, dichloromethane, n-hexane, cyclohexane, n-hepane, toluene, and aether were tested as organic phases. The activity of control group without metal ions was set to 100%, and others were expressed as a relative activity.
Substrate Specificity
Substrate specificity of Bc-SDR was investigated in our study by bioreductions of potential substrates, including acetophenone and acetophenone derivatives. The reaction was performed with a 10 mM substrate concentration in above standard condition and then the activities toward different substrates were determined by HPLC. The specific activity of Bc-SDR toward 3, 5-BTAP was set to 100%, and others were expressed as a relative activity.
Results and Discussion
Identification of Bc-SDR from B. cenocepacia
Through a genome mining approach, 20 recombinant strains of putative carbonyl reductase were successfully constructed and their activities were tested. It turned out that only Bc-SDR was observed with excellent enantioselectivity of reducing 3,5-BTAP to (R)-3,5-BTPE, while others performed poor activity on the substrate. Bc-SDR is an NADPH-dependent short-chain dehydrogenase/reductase with a known structure (PDB; 4K6F_A), and 3D structure displays a typical sequence motifs, comprising Rossmann-fold elements for coenzyme-binding and the highly conserved catalytic tetrad of Asn-Ser-Tyr-Lys [20, 21]. Bc-SDR enzyme is a homotetramers with 247 resuides, which belongs to the “classical” SDRs family [22]. The sequence of Bc-SDR showed 34.1% identity and 53.6% similarity with the template sequence from L. xyli (Fig. S2). The bioreduction catalyzed by Bc-SDR was done with anti-Prelog’s specificity, which made Bc-SDR a potential biocatalyst for the preparation of aprepitant intermediate.
Expression of Bc-SDR in E. coli
The Bc-SDR enzyme gene was successfully amplified through PCR reactions and inserted into expression vector pET-22b (+). Then, a Bc-SDR recombinant strain was successfully constructed. After induced overnight with the addition of 0.2 mM IPTG, the expression of Bc-SDR enzyme was monitored by SDS-PAGE with harvest recombinant strains of 200 μL bacteria liquid. SDS-PAGE analysis showed that Bc-SDR was successfully expressed with a high expression level. The majority of the target protein existed in the soluble with a single band between 25 and 30 kDa, which is corresponding to the predicted molecular mass of 26 kDa (Fig. 2). The expression vector pET-22b (+) has a C-terminal His-Tag sequence that can purify enzyme by using immobilized Ni-NTA affinity chromatography. However, the purified carbonyl reductase directly precipitated after purification and completely lost their bioactivity. To solve this problem, purified factors like buffer, pH, and different fusion tags were optimized, but none of them worked (data not shown). It seems that Bc-SDR is very sensitive outside the cells, and thus, the whole cell of recombinant strains was selected as the biocatalyst for reduction of 3, 5-BTAP. Compared with the isolated carbonyl reductase, whole-cell catalysts exhibit higher degrees of stability without addition of expensive coenzyme. The biotransformation with Bc-SDR recombinant strain was measured by chiral HPLC. As shown in Fig. 3, the other enantiomer (S)-3, 5-BTPE was not detected, which showed that the enantiomeric excess was more than 99.9%. We calculated that the activity toward 3, 5-BTAP was 1.258 U/mg.
SDS-PAGE analysis of the expression of Bc-SDR in E.coli. Lane M, the marker proteins with molecular mass from 14 to 100 kDa; lane 1, crude protein of extract of E. coli Rosetta (DE3)/ pET-22b(+)-Bc-SDR before induction; lane 2, crude protein of extract of E. coli Rosetta (DE3)/ pET-22b(+)-Bc-SDR after induction
Effect of Incubation Time and Wet Cell Weight in Aqueous System
We tried to optimize the reaction condition for biotransformation process. A time course study was first performed. As shown in Fig. 4a, the substrate conversion increased with the extension of reaction time, but after 36 h, the difference of the conversion rate is slight. Thus, the optimal incubation time was determined as 36 h. We then turned to invest the effect of wet cell weight on the biotransformation process. Under the reaction of 10 mM substrate, the conversion increased almost linerly with the increment of wet cells within the range of 50–450 mg (Fig. 4b). Conversion reached 99% when 750 mg/mL of wet cell was used.
Characterizations of the Bc-SDR. a The effect of reaction time (range from 6 to 72 h) on the biocatalytic reaction. Two hundred fifty milligrams per milliliter wet cells was incubated with 10 mM 3,5-BTAP in 1 mL phosphate buffer (100 mM, pH 7.3). Bc-SDR activity was monitored at 30 °C at 220 rpm for different time. b The effect of wet cell weight on the biocatalytic reaction. Different amount of wet cells were used in 1 mL phosphate buffer containing 10 mM substrates for 24 h
Effect of Reaction Temperature/pH
Normally, temperature and pH have a tremendous impact on the activity of short-chain dehydrogenases, which influence the production titer of biotransformation. In our study, the optimum temperature and pH were investigated to optimize the reaction conditions of reducing 3, 5-BTAP by Bc-SDR. The results (Fig. 5a) showed that the activity increased sharply over a range of 20–25 °C, and the optimum reaction temperature was 25 °C. When the temperature was above 25 °C, the activity decreased gradually, 56% of which still remained at 50 °C.
Characterizations of the Bc-SDR. a The effect of reaction temperature (range from 20 to 50 °C) on reducing 3,5-BTAP by Bc-SDR. Reaction was performed in phosphate buffer. b The effect of reaction pH (pH 5.0–9.0, NaH2PO4–Na2HPO4 buffer; pH 9.0–10.0, NaHCO3–Na2CO3 buffer) on reducing 3,5-BTAP by Bc-SDR. Temperature was 25 °C
The enzyme’s activity and stability can be significantly influenced by pH. From the results (Fig. 5b), we can see that the optimum pH of the enzyme was 7.0 and the highly acidic or alkaline conditions may cause the activity of enzyme to decrease dramatically. With a narrow fluctuation around pH 7.0, the production titer was significantly dropped of the biotransformation. When pH > 10.0, the Bc-SDR lost all its activity. Thus, the optimum reaction temperature and pH used for reducing 3, 5-BTAP by Bc-SDR was 25 °C and 7.0.
Effects of Metal Ions and Organic Solvents
We have investigated the influence of various metal ions toward reaction. The experiment was performed at a final concentration of 1 mM metal ions under the optimum temperature and pH. Results (Fig. 6a) showed that 11 metal ions facilitated the reduction of 3, 5-BTAP by using the whole cell of Bc-SDR recombinant strain. Among the tested metal ions, Li+, K+, Na+, Mg2+, Mn2+, and Fe3+ showed a stronger enhancement of the enzyme activity than Zn2+, Sn2+, Co2+, Ca2+, and Fe2+, and the highest relative activity (128%) was achieved when the ions group was observed in the presence of 1 mM Mn2+. On the contrary, Ni2+ and Cu2+ have apparent negative effect on enzyme activity, which inhibited the biotransformation of 3, 5-BTAP.
Characterizations of the Bc-SDR. a The effect of metal ions on reducing 3,5-BTAP by Bc-SDR. The activity of Bc-SDR was examined at a final concentration of 1 mM metal ions under the optimum temperature and pH value. b The effect of organic solvents on the activity of the Bc-SDR. Bc-SDR activity was monitored at 30°C for 24 h with each organic solvent/water (v/v = 1/2) biphasic systems
We also investigated the influence of the various organic solvent/water (v/v = 1/2) biphasic systems. As shown in Fig. 6b, ethyl acetate, methyl tert-butyl ether, isopropyl ether, dichloromethane, toluene, and ether could greatly inhibit Bc-SDR activity at the concentration of 1/2 (v/v). When n-hexane, cyclohexane, and n-hepane were used as co-solvent, enzyme activities were partially reduced. None of biphasic systems had a improvement to Bc-SDR activity.
Substrate Specificity
Substrate specificity of Bc-SDR was studied. The specific activity of Bc-SDR toward 3, 5-BTAP was set as 100%, and the data (Table 1) showed that the relative activity of Bc-SDR toward 2′-fluoroacetophenone and acetophenone were 200.1 and 119.5%, respectively. Others showed a relative activity between 42.7 and 70.5%. As a result, with increased size of substitutions at meta-position and ortho-position on the aromatic ring of acetophenone, the relative activity of Bc-SDR was decreased gradually. However, a higher relative activity of Bc-SDR toward 2′-fluoroacetophenone than acetophenone was observed, on account that fluorine and hydrogen may form a hydrogen bond to strengthen the interaction between substrate and enzyme. From the table, we can see that Bc-SDR exhibited excellent anti-Prelog’s specificity toward acetophenone and acetophenone derivatives to generate (R)-selective product. Compared with 2′-chloroacetophenone and 2′-bromoacetophenone, the meta-substituted of acetophenone, acetophenone, and 2′-fluoroacetophenone exhibited the highest enantioselectivity, and the ee remained > 99.9%. The bromine and chlorine atoms at ortho-position may increase the pro-S orientation, resulting in a stereoselective deterioration of (R)-selective product. In summary, the Bc-SDR had broad substrate specificity for acetophenone derivatives, and could be a promising biocatalyst with excellent anti-Prelog’s specificity.
Conclusion
This study aims to find a catalyst with excellent anti-Prelog’s enantioselectivity of reducing 3, 5-BTAP to the corresponding alcohol, and consequently an NADPH-dependent short-chain dehydrogenases/ reductase was discovered from B. cenocepacia by genome mining. A Bc-SDR recombinant E. coli was successfully constructed, and Bc-SDR enzyme was successfully induced with a high expression level at molecular mass of 26 kDa. As shown in Table 2, three kinds of catalysis were used in the biotransformation of 3, 5-BTAP to (R)-3, 5-BTPE including dry cells, wet cells, and lyophilized enzyme. Bc-SDR catalyzes the reduction of 3, 5-BTAP to (R)-3, 5-BTPE with high enantiopurity (> 99.9% ee). Besides, Bc-SDR exhibited (R)-enantioselectivity toward acetophenone and 2′-fluoroacetophenone which showed potential to be used in the bioreduction of acetophenone derivatives. The discovery of Bc-SDR enriches and develops the biotransformation of 3, 5-BTAP to produce (R)-3, 5-BTPE. As an anti-Prelog’s biocatalyst, it may play an important role in pharmaceutical industry and is worth of further investigation.
References
Moore, J. C., Pollard, D. J., & Kosjek, B. (2007). Advances in the enzymatic reduction of ketones. Accounts of Chemical Research, 40, 1412–1419.
Bornscheuer, U. T., Huisman, G. W., & Kazlauskas, R. J. (2012). Engineering the third wave of biocatalysis. Nature, 485, 185–194.
Huisman, G. W., Liang, J., & Krebber, A. (2010). Practical chiral alcohol manufacture using ketoreductases. Current Opinion in Chemical Biology, 14, 122–129.
Noey, E. L., Tibrewal, N., & Jiménez-Osés, G. (2015). Origins of stereoselectivity in evolved ketoreductases. Proceedings of the National Academy of Sciences, 112, E7065–E7072.
Wang, P., Cai, J. B., & Ouyang, Q. (2011). Asymmetric biocatalytic reduction of 3,5-bis (trifluoromethyl) acetophenone to (1R)-[3,5-bis (trifluoromethyl) phenyl] ethanol using whole cells of newly isolated Leifsonia xyli HS0904. Applied Microbiology and Biotechnology, 90, 1897–1904.
Gelo-Pujic, M., Le Guyader, F., & Schlama, T. (2006). Microbial and homogenous asymmetric catalysis in the reduction of 1-[3,5-bis (trifluoromethyl) phenyl] ethanone. Tetrahedron: Asymmetry, 17, 2000–2005.
Brands, K. M. J., Payack, J. F., & Rosen, J. D. (2003). Efficient synthesis of NK1 receptor antagonist aprepitant using a crystallization-induced diastereoselective transformation. Journal of the American Chemical Society, 125, 2129–2135.
Nakade, S., Ohno, T., & Kitagawa, J. (2008). Population pharmacokinetics of aprepitant and dexamethasone in the prevention of chemotherapy-induced nausea and vomiting. Cancer Chemotherapy and Pharmacology, 63, 75–83.
Kurbanoglu, E. B., Zilbeyaz, K., & Taskin, M. (2009). Total production of (R)-3, 5-bistrifluoromethylphenyl ethanol by asymmetric reduction of 3,5-bis (trifluoromethyl)-acetophenone in the submerged culture of Penicillium expansum isolate. Tetrahedron: Asymmetry, 20, 2759–2763.
Gai, P., Tang, C., Liu, J., Liu, Y., Zhang, C., & Wu, Z. (2013). Asymmetric anti-Prelog reduction of 3,5-bis(trifluoromethyl)-acetophenone by microbacterium oxydans C3. Chinese Journal of Applied & Environmental Biology, 19, 37–42.
Li, J., Wang, P., He, J. Y., Huang, J., & Tang, J. (2013). Efficient biocatalytic synthesis of (R)-[3,5-bis (trifluoromethyl) phenyl] ethanol by a newly isolated Trichoderma asperellum ZJPH0810 using dual cosubstrate: ethanol and glycerol. Applied Microbiology and Biotechnology, 97, 6685–6692.
Wang, N., Huang, J., & Luo, H. (2013). Purification and characterization of a new carbonyl reductase from Leifsonia xyli HS0904 involved in stereoselective reduction of 3, 5-bis (trifluoromethyl) acetophenone. Journal of Molecular Catalysis B: Enzymatic, 92, 1–6.
Wang, N. Q., Sun, J., & Huang, J. (2014). Cloning, expression, and directed evolution of carbonyl reductase from Leifsonia xyli HS0904 with enhanced catalytic efficiency. Applied Microbiology and Biotechnology, 98, 8591–8601.
Liu, Y., Tang, T. X., & Pei, X. Q. (2014). Identification of ketonereductase ChKRED20 from the genome of Chryseobacterium sp. CA49 for highly efficient anti-Prelog reduction of 3,5-bis(trifluoromethyl)acetophenone. Journal of Molecular Catalysis B: Enzymatic, 102, 1–8.
Li, H. M., Moncecchi, J., & Truppo, M. D. (2015). Development of an immobilized ketoreductase for enzymatic (R)-1-(3,5-Bis(trifluoromethyl)phenyl)ethanol production. Organic Process Research & Development, 19, 695–700.
Chen, K., Li, K., & Deng, J. (2016). Carbonyl reductase identification and development of whole-cell biotransformation for highly efficient synthesis of (R)-[3,5-bis (trifluoromethyl) phenyl] ethanol. Microbial Cell Factories, 15, 191.
Prelog, V. (1964). Specification of the stereospecificity of some oxido-reductases by diamond lattice sections. Pure and Applied Chemistry, 9, 119–130.
Singh, A., Basit, A., & Banerjee, U. C. (2009). Burkholderia cenocepacia: a new biocatalyst for efficient bioreduction of ezetimibe intermediate. Journal of Industrial Microbiology & Biotechnology, 36, 1369.
Yıldız, T. (2015). An oxazaborolidine-based catalytic method for the asymmetric synthesis of chiral allylic alcohols. Tetrahedron: Asymmetry, 26, 497–504.
Hoffmann, F., & Maser, E. (2007). Carbonyl reductases and pluripotent hydroxysteroid dehydrogenases of the short-chain dehydrogenase/reductase superfamily. Drug Metabolism Reviews, 39, 87–144.
Kallberg, Y., Oppermann, U., & Jörnvall, H. (2002). Short-chain dehydrogenase/ reductase (SDR) relationships: a large family with eight clusters common to human, animal, and plant genomes. Protein Science, 11, 636–641.
Oppermann, U., Filling, C., & Hult, M. (2003). Short-chain dehydrogenases/reductases (SDR): the 2002 update. Chemico-Biological Interactions, 143, 247–253.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
ESM 1
Fig. S1 Multiple sequence alignment of the target sequence (NCBI Reference Sequence: ANZ75577.1, ANZ77886.1, ANZ77893.1, XP_002489352.1, XP_002492297.1, XP_002493760.1, XP_002494195.1, WP_006481546.1, WP_006481604.1, WP_006481801.1, WP_006483065.1, WP_006485075.1, WP_006485158.1, WP_006488202.1, WP_012336650.1, WP_012492403.1, WP_050013528.1, WP_058903664.1, WP_060214413.1, WP_077189930.1) with the template sequence (Leifsonia xyli HS0904) Fig. S2 Sequence alignment of the target sequence (Burkholderia cenocepacia) with the template sequence (Leifsonia xyli HS0904). (DOCX 9094 kb)
Rights and permissions
About this article
Cite this article
Yu, S., Li, H., Lu, Y. et al. A Catalyst from Burkholderia cenocepacia for Efficient Anti-Prelog’s Bioreduction of 3,5-Bis(Trifluoromethyl) Acetophenone. Appl Biochem Biotechnol 184, 1319–1331 (2018). https://doi.org/10.1007/s12010-017-2628-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-017-2628-8