Abstract
Termites are well recognized for their thriving on recalcitrant lignocellulosic diets through nutritional symbioses with gut-dwelling microbiota; however, the effects of diet changes on termite gut microbiota are poorly understood, especially for the lower termites. In this study, we employed high-throughput 454 pyrosequencing of 16S V1–V3 amplicons to compare gut microbiotas of Tsaitermes ampliceps fed with lignin-rich and lignin-poor cellulose diets after a 2-week-feeding period. As a result, the majority of bacterial taxa were shared across the treatments with different diets, but their relative abundances were modified. In particular, the relative abundance was reduced for Spirochaetes and it was increased for Proteobacteria and Bacteroides by feeding the lignin-poor diet. The evenness of gut microbiota exhibited a significant difference in response to the diet type (filter paper diets < corn stover diets < wood diets), while their richness was constant, which may be related to the lower recalcitrance of this biomass to degradation. These results have important implications for sampling and analysis strategies to probe the lignocellulose degradation features of termite gut microbiota and suggest that the dietary lignocellulose composition could cause shifting rapidly in the termite gut microbiota.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Termites are extremely successful wood-degrading organisms distributed worldwide, which are important for carbon turnover in the environment and are potential sources of biochemical catalysts [1–4]. It is well known that termites form symbiotic association with diverse microorganisms in their gut, which as a result of evolution and adaptation, makes the termites capable of degrading the lignocelluloses in their diets, as well as improving their immunity, reproduction, and other physiological functions [3, 5–8]. The microsymbionts were mainly the bacteria in the so-called higher termites (family Termitidae), while the guts of lower termites (other families) harbored many protozoans as well as the prokaryotes [9]. Since uncultivable bacteria, like those in TG1 (termite gut 1) phylum, have been identified as endosymbionts of a variety of termite gut-flagellated protists [10], the bacteria, protists, and lower termite species might form a complex nutritional symbiosis [3, 11]. In addition, it has been evidenced that the lignocellulose degrading capability of termites are depending on their gut symbionts [3, 12–14], while certain symbiotic bacteria in the termite intestines act directly on cellulose and xylan hydrolysis [6, 7, 15].
Previously, it has been reported that the gut microbiota of termite was mainly determined by the termite phylogeny, but was also shaped by the diet habits of termites [16–18]. Recently, several studies have been realized on the relationships among the diets, the capabilities of lignocellulose digestion, and the structure of gut microbial community of termites [19–23]. Diet has been reported as the primary determinant of bacterial community structure in the guts of fungus, wood/grass, soil, and humus/litter-feeding higher termites [24]. For the lower termites, Boucias et al. [21] reported that diet-dependent shifts in the gut microbiota of Reticulitermes flavipes fed with lignin-rich and lignin-poor cellulose diets were not statistically significant; but Huang et al. [20] demonstrated that the gut bacterial communities were different among the R. flavipes populations fed with woody diet, corn stover, and sorghum. However, both of the reports have flaws: Boucias et al. [21] fed the termite for only 1 week that may not be sufficient to induce significant change in gut microbiota, while Huang et al. [20] did not include the comparison with no lignin diets. In addition, [9] indicated that “the changes in diet or overall host physiology affected the protist and bacterial populations in the hindgut” of six R. flavipes populations collected from different geographic locations or sources, but the diet features were not reported in their study [9]. Therefore, further study is needed to verify the effects of lignin contents in the diets on the gut microbiota of lower termites.
As a local species in Henan Province of China, the study on Tsaitermes ampliceps, a lower termite species, is rare up to date. Considering that clarifying the response of gut microorganisms to different kinds of diets is important for both the potential application of termite gut microbiota in biotransformation of lignocelluloses, hemicelluloses, and cellulose, and for understanding the termite adaptation in environment, we did the present study by feeding T. ampliceps populations with wood, corn stover, and filter paper separately in a laboratory environment. And then the gut bacterial communities in different treatments were comparatively analyzed by 454 high-throughput sequencing. The aims of our study were to facilitate a more comprehensive analysis of gut microbiota associated with termite fed with different diets and to find the effects of diet on the gut bacterial community. The results provide insights into the relationships among the intestinal bacteria and the diet lignin contents of the lower termites.
Materials and Methods
Termite Collection and Diet Manipulation Bioassays
T. ampliceps samples were collected from Shangcheng County, one of the two habitat regions (another one is Yuzhou County) for this termite, in Henan Province (N31°27′, E115°19′) [25, 26]. All collected locations are in wild fields not disturbed by human activities, while T. ampliceps is not an endangered species (Fig. 1). Termite populations were maintained together with logs and soils collected from the sampling site in complete darkness at room temperature with >70 % humidity until further processing. Termite species was identified based on morphology [25].
Feeding boxes and the gut sample of termite species Tsaitermes ampliceps used in this study. Plastic boxes with dimensions of 15 × 15 × 10 cm were filled with 2 cm thickness of 1 % (w/v) agarose gel as basal substrate incubated in darkness at 27 °C and 70 % relative humidity, supplied with: a Pine wood (18.5 % lignin and 66.2 % cellulose-hemicellulose). b Corn stover (11.8 % lignin and 62.3 % cellulose–hemicellulose). c Filter paper (100 % cellulose). d Entire gut compartments sampled from termite T. ampliceps
T. ampliceps individuals collected from the same population were divided into three groups (about 1,000 termite workers per group) and fed separately with the diets of pine wood (W, contained 18.5 % lignin and 66.2 % cellulose–hemicellulose), corn stover (C, contained 11.8 % lignin and 62.3 % cellulose–hemicellulose), and filter paper (F, contained cellulose alone) in darkness at 27 °C and 70 % relative humidity, in the plastic boxes (15 × 15 × 10 cm) filled with 2 cm thickness of 1 % (w/v) agarose gel as basal substrate. Dietary materials (strips of 0.5 cm in width) were saturated with deionized water and placed on the gel in boxes (Fig. 1). The dietary materials were regularly moistened with deionized water every day for 2 weeks.
Termite Gut Dissection and DNA Extraction
After 2 weeks of feeding, termite workers were used for extraction of metagenomic deoxyribonucleic acid (DNA). Briefly, after washing with 70 % ethanol and sterilized water (30 s for each), the whole intestinal tracts of 200 termites of the same treatment were dissected on ice using sterilized tweezers and insect needles under anatomical microscope (Fig. 1). The isolated guts were immersed in ice-cold 0.2 M phosphate-buffered saline containing (g/L) 8.0 NaCl, 0.2 KCl, 1.44 Na2HPO4, and 0.24 KH2PO4 with pH 7.4 [27] and were homogenized by Bullet Blender (Next Advance, America). After centrifugation at 3,000×g for 5 min at 4 °C, the supernatant was used for metagenomic DNA extraction using the E.Z.N. A® Tissue DNA Kit (OMEGA, USA), according to the manufacturer’s instruction. DNA quality was evaluated by electrophoresis in 1.0 % (w/v) agarose gels stained with SYBR Safe, visualized on a CCD compact image system (Major Science, USA). The DNA samples were stored at −20 °C before further processing. Triplicates for each feeding treatment were included as biological replicates for intestinal dissection, DNA extraction, and the subsequent PCR amplication and sequencing.
PCR Amplification of Microbial 16S rRNA Genes
The polymerase chain reaction (PCR) amplification of variable regions V1–V3 of 16S rRNA was performed by using the forward primer fP (5′-TGG AGA GTT TGA TCC TGG CTC AG-3′) and reverse primer rP (5′-TAC CGC GGC TGC TGG CAC-3′) [28] containing 10 pb multiplex identifiers (MID) with a GeneAmp® PCR Systems 9700 (Applied Biosystems, USA) using the corresponding protocol [28] in 50 μl of reaction mix. The PCR products were gel-purified using Agencourt AMPure XP beads (Beckman Coulter, USA) according to the manufacturer’s protocol and the concentrations were quantified using an Agilent BioAnalyzer 2100 (Invitrogen, USA).
Pyrosequencing and Bioinformatic Analysis of the 16S rRNA Amplicons
The purified DNA amplicons from metagenomic DNAs were analyzed by pyrosequencing with a 454 Life Sciences Genome Sequencer FLX Titanium (GS-Titanium; 454 Life Sciences, Branford, CT, USA), which produced 400-bp reads on average. The sequence preprocessing, identifying operational taxonomic units (OTUs), taxonomic assignment, and community structure comparison of the microbiota were analyzed with MOTHUR [29] (http://www.mothur.org/). To control the sequence quality, reads <150 bp, with an average quality score of <35 in each 50-bp window rolling along the whole read, with an ambiguous base call (N), with any homopolymers of >8 bases, or without the primer sequence were excluded [30]. The remaining reads were then sorted based on the tag sequences. A pre-clustering step using the “pre.cluster” script in MOTHUR was performed and all the chimerical sequences detected by UCHIME [31] were removed for reducing sequencing noise in the pyrosequencing data. These trimming processes were completed using RDP Initial Process (http://pyro.cme.msu.edu). To conduct termite gut microbiome analysis, the 16S rRNA gene reads that were obtained using the 454 GS-Titanium were further processed by customized computational pipelines. The 16S rRNA gene reads obtained in this study were deposited in the Sequence Read Archive (SRA) of NCBI under the accession number of SRP067996 and were further processed by computational pipelines that were customized for termite gut microbiome analysis.
For assessment of microbial diversity, the trimmed reads were clustered using UCLUST (http://www.drive5.com/uclust/) and the reads sharing identical sequences were defined as a phylotype. To determine the alpha diversity, an in-house Perl script was then used to convert the output from UCLUST into a format recognized by MOTHUR software package and the reads sharing similarities ≥97 % were assigned as the same OTU. All of the OTUs were aligned to the Greengenes database [2] using the “claasify.seqs” command in MOTHUR, with an 80 % confidence threshold to obtain their taxonomic assignments. Then, alpha diversity, including the analysis of Good’s coverage, diversity estimators (Shannon’s and Invsimpson), richness estimators (ACE and Chao1), and rarefaction curves were calculated based on the OTUs using the summary single command in MOTHUR (http://www.mothur.org/). The relative abundances of bacterial taxa were calculated at the phylum, class, order, family, and genus levels, as well as unclassified taxa if they were not clearly defined into any level. The microbial community structures were compared using principal coordinates analysis (PCoA) [32] based on the matrices of pairwise distances among all the microbiomes or using the heat map generated with the program at http://www.metagenassist.ca/METAGENassist/faces/UploadView.jsp2. For statistical analyses, the data were typically presented as the mean ± S.E.M. One-way analysis of variance (ANOVA) was used to detect statistically significant differences, and Tukey HSD was used for multiple comparisons with SPSS 17.0. All the statistical tests were two-sided with a significance level of p < 0.05. If appropriate, the p values were corrected based on Holm’s adjustment for multiple comparisons [33]. To probe the microbial metabolic and functional pathways in the microbiota, the OTUs were automatically taxonomy-to-phenotype mapped using approximately 20 different phenotypic categories in METAGENassist (http://www.metagenassist.ca/METAGENassist/faces/Home.jsp).
Results
Diversity of Gut Microbiota from T. ampliceps with Different Diets
After strict quality control and chimera checking, a total of 95,195 post-trimming 16S rRNA gene reads were obtained from the gut microbiota in the three treatments of T. ampliceps, with an average of 10,577 reads per sample (Table 1). The number of OTUs varied between 824 and 1,230 in the samples. The richness of total bacterial phylotypes ranged between 1,105 and 1,570 among the replicates in each sample. The diversity (Inv Simpson = 162.06) of the termite gut bacteria in treatment F was significantly lower than that in treatment C (181.94; p < 0.05) and was lower than that in W (189.69); however, the Shannon index showed a similar level (around 5.91) in all the three treatments (Table 1). The ratio of OTU/Chao1 was 0.719–0.748 and the rarefaction curves from the three biological replicate samples did not reach clear plateaus (Table 1, Online Resource Fig. S1), indicating that most of the OTUs have been detected, but more OTUs might be obtained if the number of reads was enlarged. Comparisons of the rarefaction curves showed that the three diets exhibited different levels of richness for OTU (Online Resource Fig. S1).
In the PCoA analysis, the gut microbiotas of T. ampliceps fed on different diets were separated (p < 0.01) from each other and those in C and W were more similar compared with that in F (Figs. 2 and 3). The Jclass-based PCoA, a thetayc scheme PCoA analysis, produced further consistent results (Fig. 3). In our study, the majority of gut bacteria (41.6 % of OTU, accounting for 94.8 % of sequence reads; Fig. 2b) were shared across the three treatments. Combined with the analysis of diversity index, it seems that the composition of OTUs in core microbial community had no significant difference (according to Shannon index), while their evenness was significantly different in the three treatments (according to Inv Simpson index, p = 0.017). Therefore, the gut microbiota community structure of T. ampliceps was affected by the diet shifting mainly in the relative abundances of different taxa, and the filter paper diet presented greater influence than the corn stover diets.
Principal component analysis (PCA). a The microbiota grouping of replicates for the same treatment and separation of mcrobiotas in different diet treatments is showed. b Venn diagram demonstrated the distribution of gut bacterial OTUs in the Tsaitermes ampliceps feeding with three diets with different lignocellulosic contents. In a, each point corresponds to a microbial community and the triplicate samples of the same treatment were presented in the same color. The percentages of variation explained by the plotted principal coordinates are indicated on the axes. T*-F shows T. ampliceps feeding on filter paper, T*-C shows T. ampliceps feeding on corn stover, T*-W shows T. ampliceps feeding on wood. In b, 784 OTUs were shared by the three treatments and the others shared by two treatments or found in single treatments
Principal coordinates analysis (PCoA) showing that the gut bacterial communities in Tsaitermes ampliceps populations feeding on the three diets with different lignocellulosic contents were significantly different (**p < 0.01; ***p < 0.001 according to Kruskal–Wallis test). a The community structures analyzed by PCoA based on thetayc distance matrix. b The community structures analyzed by PCoA based on Jclass distance matrix. Each point corresponds to a microbial community where the color indicates its treatment. PCO1 and PCO2 are shown with the percentage variation explained for each axis. T*-F shows T. ampliceps feeding on filter paper, T*-C shows T. ampliceps feeding on corn stover, and T*-W shows T. ampliceps feeding on wood
Community Composition of T. ampliceps Gut Microbiota with Three Diets
The T. ampliceps gut microbiota of the three diet treatments were all covered 22 bacterial phyla and some unclassified members (Fig. 4a and Online Resource Table S1). However, some significant differences in the relative abundances were observed for eight phyla, including Spirochaetes, Proteobacteria, Bacteroidetes, Firmicutes, Planctomycetes, Acidobacteria, Thermi, and Fusobacteria (p < 0.05; Fig. 4a and Online Resource Table S1). The relative abundance of Spirochaetes was the highest in all the three treatments, and significant differences were observed in the order of W > C > F. The second and third abundant phyla in all the treatments were Proteobacteria and Bacteroides, and both were higher in C and F than in W. The forth abundant phylum was Firmicutes and its relative abundances in the three treatments were significantly different, in the order of F > W > C. The relative abundance of Planctomycetes was lower in C than in the other treatments. No difference was detected in the other five abundant phyla (Fig. 4a and Online Resource Table S1). Generally, the diversity of gut bacteria in F was significantly lower than those in C and W treatments (Inv Simpson index being 162.06, 181.69, and 181.94, respectively).
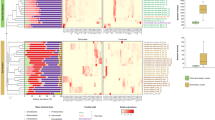
Variations of bacterial communities in the gut of Tsaitermes ampliceps feeding on three diets with different lignocellulosic contents. a The variation of core phyla of the gut bacteria. The bars marked with different letters for the same bacterial phylum are significant different in the abundance based upon the statistical analysis (p < 0.5). b Heat map presenting the variation of core genera of gut bacteria in different treatments
Among the three diet treatments of T. ampliceps, a total of 122 genera were identified, and about one third (3,100/9,147) of the sequences were unclassified. Generally, 81, 56, and 80 genera were identified within the samples of C, W, and F, respectively (Fig. 4b, Online Resource Table S2). The main genera were similar among the three treatments, but their relative abundances varied. The genus Treponema was the most abundant one, occupying 54.57, 50.77, and 38.32 % of all the OTUs, followed by Candidatus genus Azobacteroides (3.07, 3.00, and 2.55 %), Desulfovibrio (1.62, 1.92, and 1.63 %), and Dysgonomonas (1.10, 1.80, and 1.30 %) for W, C, and F treatments, respectively. The other genera presented at relative abundance ≤0.87 % (Fig. 4b, Online Resource Table S2). The main differences among the three treatments were that higher abundance of Treponema (54.57 %) and lower abundance of Dysgonomonas (1.10 %) were found in W than in the other two. In addition, 39 minor genera were found only in C or F, but not in the wood-fed populations (Fig. 4b, Online Resource Table S2). Lower abundances of Treponema (38.32 %) and Dysgonomonas (1.30 %) differentiated the F treatment from C (50.77 and 1.80 %, respectively). The other differences were found in the minor genera, such as rare Pseudomonas (0.07 %) and Acinetobacter (0.03 %) in W, but little bit abundant in C (0.73 and 0.38 %, respectively) and F (0.63 and 0.54 %, respectively); absence of za29 and presence of Chitinophaga in C; absence of Flavobacterium and Stenotrophomonas in W. These differences demonstrated that the diet characters might have regulated the gut core bacterial community structure of the termite species after 2 weeks of feeding.
Functional Implications of the Different Diets
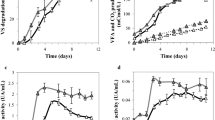
For T. ampliceps fed with three kinds of lignocellulosic diets, 27 types of bacterial metabolic activities (phenotypic category of metabolism) were identified (Fig. 5). The abundances for different metabolic types, especially the nitrogen cycle, sulfur cycle, and degrading bacteria, varied among different treatments (Fig. 5). In general, the nitrogen-fixing bacteria were the most abundant group (with relative abundance of 20–44 %), followed by ammonia oxidizer and dehalogenation (about 12 %). Sulfate reducer, nitrite reducer, sulfide oxidizer, and aromatic hydrocarbons degrader represented the third level of abundance (4–7 %), whereas those of denitrifying, dinitrogen-fixing, and streptomycin producer were low (approximately 0.1 %; Fig. 5). Comparing the three treatments, the microbiota in F were characterized by the lower nitrogen-fixing and the greater sulfate reducer, nitrite reducer, sulfide oxidizer, aromatic hydrocarbons, xylan degrader, etc., that was significantly different from those in the C and W treatments.
Relative abundance of putative metabolic groups in termite gut microbiota of Tsaitermes ampliceps feeding on three diets with different lignocellulosic contents. To probe the microbial metabolic and functional pathways, the OTUs were automatically taxonomy-to-phenotype mapped using approximately 20 different phenotypic categories in METAGENassist (http://www.metagenassist.ca/METAGENassist/faces/Home.jsp). The comparison was conducted at phylum level. The significantly varied metabolic groups among the three treatments were marked with * (p < 0.05) or *** (p < 0.001). T*-F shows T. ampliceps feeding on filter paper, T*-C shows T. ampliceps feeding on corn stover, and T*-W shows T. ampliceps feeding on wood
Discussion
According to the previous studies [3, 10, 34], the synergistic association with protozoans and bacteria is essential for the lower termites to digest the ligniocellulos. Although flagellates have been believed to play an important role in the lignocellulose degradation based upon their dense colonization and production of multiple enzymes in the gut of lower termites [34, 35], the endophytic bacteria in the gut are also important in order to maintain the gut environment, such as producing acetate by recycling H2 and CO2 produced by the flagellates, consuming O2, and fixing nitrogen [36]. Considering that the different diets may change the gut environments, such as C/N ratio, composition, and intermediate metabolites, the shifting in bacterial community is expected corresponding to the diet change. In the present study, we fed the termite 2 weeks and analyzed for the first time the response of gut bacterial communities in T. ampliceps to three different lignocellulosic diets. The high coverage values (0.94–0.98; Table 1) and the well clustering of bacterial communities from the three subsamples for each diet in PCA (Fig. 2) demonstrated that most of the gut bacteria associated with the sampled termites have been detected and the methods in this study were reproducible.
In general, the structure composition of gut microbiota characterized by predominant phyla Spirochaetes (56.2 %), Proteobacteria (12.6 %), Firmicutes (8.2 %), and Bacteroidetes (6.3 %) in T. ampliceps gut (Fig. 4a) was similar to that reported previously for another wood-feeding lower termite R. flavipes [9, 20, 21]. However, the dominance of Proteobacteria (12.6 %) and the relative abundances of the other dominant phyla in T. ampliceps were differed from the gut microbiota of R. flavipes [9, 20, 21]. These variations may be related to the differences of the diet habits, environmental factors, or genetics of their hosts, as reported previously [16, 17, 21]. The dominance of Spirochaetes (mainly Treponema) was common in the gut microbiota for both the lower and the higher termites [7, 9, 20, 21, 37], implying that bacteria in this phylum may play important role in the life of termites. In addition, the absence of Fibrobacteres in the lower termite but as the second abundant phylum in higher termite (13 %) [7, 37] could be a microbial marker to differentiate the lower and higher termites.
In the present study, we focused on the effects of diet lignocellulosic composition on the gut microflora. The significant variation in 58.4 % of OTUs in the gut bacteria (Fig. 2b) is an evidence that 2 weeks of feeding was sufficient to cause the detectable shifting in gut microbiota of T. ampliceps. Our results confirmed the previous reports that microbial communities are considered sensitive to environmental perturbation, including diets [3, 4, 20], and that the conclusion of Boucias et al. [21] was a feeding time-depending event. Thus, we can consider that altering diet composition would impact gut microbiota composition. Interestingly, the significantly lower diversity in gut bacteria of T. ampliceps fed with filter paper than those in corn stover and wood fed populations and the clustering results (Table 1, Fig. 2) were consistent with the previous studies that the diets of corn stover and filter paper reduced gut bacterial richness and diversity in R. flavipe colonies compared to wood diets [20, 21]. Combining our results with the previous reports, the significantly reduced diversity level of termite gut microbiota associated with diet without lignin (filter paper) may demonstrate that more complex synergistic interactions among the gut microorganisms are needed for degrading the high-lignin-containing materials. Since previous studies have revealed that the gut microbiota plays an essential role in the life of termites by digesting the food, fixing nitrogen, and producing amino acids/vitamins etc. [7, 37], the shifting in the gut microbiota responding to the diet will be discussed in relation to the possible function of the bacteria.
In a previous research, core microbiome was defined as the common bacterial taxa presented in the guts of different termite species or the same termite fed with different diets, but the relative abundance of each taxon in the microbiome varied [17, 20, 38]. In the present study, the majority of detected phyla were shared across all the treatments (60.8 % of phyla, accounting for 94.9 % of OTUs; Fig. 2; Online Resource Table S2) and the core bacterial taxa belonged to the phyla Spirochetes (51.19 %), Proteobacteria (15.74 %), Bacteroidetes (10.28 %), and Firmicutes (6.62 %) (Online Resource Table S2).
In particular, the relative abundance of Spirochaetes was reduced while Proteobacteria and Bacteroides increased in the gut microbiota when the lignin content in the diet decreased (Figs. 3 and 4). Our data reinforce previous findings that phylum Spirochetes played an important role in plant lignocelluloses degradation in the lower termite guts [17, 20, 38]. However, no known gene directly involved in the lignin degradation was detected in metagenomic analyses of several higher termites, and the release of cellulose and hemicelluloses from the lignocellulose was estimated to be performed by the termites in their alkaline parts of gut [3, 37]. Therefore, the lignocellulose-degrading ability of gut bacteria in the lower termites needs further study.
The graduate decrease of Spirochetes (mainly Treponema) in C and F might be indirect evidence that these bacteria were important in both the hemicelluloses and lignin degradation (Fig. 3). According to the metagenomic studies [7, 37], Treponema belonging to the most abundant phylum Spirochetes is responsible for transformation of H2 and CO2 into acetate, that is the main source of carbon and energy for termites. The majority of glycoside hydrolase (GH) catalytic domains in a termite were encoded by treponemes [37]. Therefore, it is reasonable that Treponema is detected as one of the most abundant bacterial groups in the guts, despite the phylogeny of termites. In the present study, another main genus was Za29 that has been classified as Spirochaeta and presented greater abundance in treatment F than in W and C. Although its physiological features are not known, a related strain SPN1 presented several enzyme activities involved in degradation of cellulose [39].
In the phylum Bacteroidetes, three genera, Candidatus genus Azobacteroides, Dysgonomonas, and Sphingobacterium, were found in the top 11 genera in the gut microbiota (Fig. 4). Azobacteroides is a bacterium coupled N2 fixation to cellulolysis within protist cells in termite gut [40]. The abundance of Azobacteroides was constant in all the three treatments, implying that it was not directly participated in the polymer degradation. The genome analysis of this bacterium demonstrates that it is able to fix nitrogen, to synthesize amino acids/cofactors, to recycle the waste of protist (urea and ammonia), to ferment the monomers (glucose, xylose, and hexouronates) produced from lignocellulose degradation by protist, and to uptake H2 produced from lignocellulose degradation and nitrogen fixation [40]. In contrast to Azobacteroides, genus Dysgonomonas increased its relative abundance in treatments C and F than that in W (Fig. 4b). Two species in this genus, D. macrotermitis [41] and D. termitidis [42], have been described for isolates from termite guts and they are able to ferment soluble starch, xylan, pectin, CM-cellulose, and aesculin. So, the enhanced abundance of this genus or total Bacteroidetes might be a response to the increase of cellulose in the diet of termite. Abundance of Sphingobacterium was significantly greater in treatment F than in W and C, and it has been isolated as cellulolytic bacterium from gastrointestinal tract of snail [43].
Candidatus genus Tammella and TG5 (a group specific to termite gut) are members of recently recognized phylum Synergistetes that harbors some anaerobic, Gram-negative rod/vibrioid bacterial groups. The physiology of Tammella is not known, but its function as the motility ectosymbiont of the termite gut flagellate Caduceia versatilis has been confirmed [44]. Since the relative abundance of Tammella is constant in all the three treatments, it may not be related to the degradation. Different from Tammella, the relative abundance of TG5 was gradually increased in treatments W, C, and F (from 0.46 to 0.59 and 0.87 %), which might be related to the degradation of cellulose, but it is only an estimation since no phenotypic features is available for this genus [9].
As a member of Proteobacteria, the detection of greater abundances of Desulfovibrio, Pseudomonas and Acinetobacter in treatments C and F than in W might related to their degradation ability of hemicellulose and cellulose. Pseudomonas as a dominant cellulolytic group has been isolated from snail gastrointestinal tract [43] and degradation of various compounds has also been reported for them [45]. Similar to Pseudomonas, Acinetobacter strains are also frequently isolated as crude oil and/or lignocellulose degrading bacteria [46, 47]. Desulfovibrio is a well-known anaerobic degrader and it has been isolated from lignocellulose-degrading consortia [48]. Based upon it physiological features, these bacteria may not directly degrade the lignocelluloses, but it can remove the inhibition of organic acids to the lignocelluloses degradation by anaerobic respiration. Therefore, it is possible that these genera also participate directly or indirectly in the degradation of lignocelluloses, especially the hemicelluloses and cellulose, inside the gut of termite, since Proteobacteria is an abundant phylum and were enriched by the low-lignocellulose diets in treatments C and F.
Belonging to Tenericutes, Mycoplasma has been reported mainly as pathogen for animals and human. Because of the small size of the genome, its degradation ability is limited and its function in the termite gut is unknown. The constant abundance of Tenericutes in the three treatments of the present study implied that this group may not be related to the degradation.
In addition to the mentioned phyla and genera above, the other bacteria detected in the termite gut may also play some important roles in maintaining the efficiency of degradation and transformation in the gut system. No abundant genus was detected in Firmicutes, although it was the fourth abundant phylum. The decrease in treatment C and increase in treatment F demonstrated the possibility that this group was inhibited by hemicellulose, but enriched by cellulose. In the anaerobic conditions, the main Firmicutes is Clostridia, which contained some lignocellulosic [47] and cellulolytic [49] hydrolyzing members. In higher termites, most of the xylanase (hemicelluloses-degrading) genes (GH11 family) were assigned to Firmicutes [37]. The higher abundance of Firmicutes detected in treatment F than in treatments W and C demonstrated the possibility that Firmicutes may not be the main hemicelluloses degrader in T. ampliceps gut. In the higher termites, some cellulose-degrading genes (cohesins and dockerins) were affiliated to Clostridia [37], but these bacteria were decreased in the C treatment and were not presented as the main genera (Fig. 4). In addition, no Clostridium-related cellulase gene GH5 was detected in the metagenomic study [7]. Therefore, further study is needed to confirm the cellulose degrading activity of clostridia in termite gut.
Because of the absence of information about the metabolic functions of the bacterial genus or OTUs in the gut of lower termites, the shifting in functional groups of termite gut microbiota associated with the dietary lignocellulose composition was estimated using the 20 different phenotypic categories in METAGENassist (http://www.metagenassist.ca/METAGE382Nassist/faces/Home.jsp) as references.
The variation of microbial functional groups in T. ampliceps was highly correlated with the termite diet types (Fig. 5). As the cellulose content increased in the diets, the relative abundances of phyla with cellulose degradation and xylan degradation functions also increased in the gut of T. ampliceps. Thus, the diets tune the functions of the microbes in the termite gut and correspondingly their biomass degradation capability. The finding of changes in enzyme activities responding to the diet features [19, 22] could be evidence for this estimation. Recent studies showed that bacteria from the phyla Firmicutes [1, 50], Proteobacteria [50], Armatimonadetes [51], and Spirochaetes [52] may have important roles in lignin degradation, and they were also the abundant ones in the wood treatment in the present study. This may facilitate the identification of strains related to lignocellulose degradation, i.e., the diet-specific bacterial OTUs may represent organisms with the ability to degrade lignin. These diet-dependent taxa may have unique selective effects on each biomass type processed by termite gut bacterial communities [1].
The diet-driven changes in termite gut bacterial communities and the decrease of diversity along with the decreased content of lignin in diets may be explained as the recalcitrance of diets relate to gut bacterial richness and diversity by impacting the potential for niche differentiation. The recalcitrance of plant cell walls to efficient deconstruction to simple sugars could be efficiently performed by enzymatic decay of biomass of microbial communities existing in soils, swamps, and the guts of wood-eating worms and termites [53]. For instance, the recalcitrance index (RI) values decreased in the order of softwood (0.87), hardwood (0.56), and corn stover (0.26) due to their different lignin content (the order of 25–35, 18–25, and 17–21) [54]. Among the substrates tested here, filter paper is only composed of cellulose, with a bundle of linear poly β-1,4-glucan chains organized by H-bonding networks. The recalcitrance of wood and corn stover extending far beyond filter paper, with cross-linking of these matrixing polymers (cellulose, hemicelluloses, pectins, and lignins), could result in the elimination of water from the wall and that such dehydration could alter the structure of the cellulose fibrils [53]. Filter paper diets with low recalcitrance may lead to quick exclusion of some bacteria competing for the same homogenous substrates; so, rapid degradation of biomass could be accomplished by a decreased number of microbial taxa. This may explain the lower gut bacterial richness and diversity for termites fed with filter paper compared to corn stover and wood diets. In addition, plant secondary metabolites, including alkaloids, phenolic, and essential oils compounds, could act as inhibitors against some members of the gut microbiome [55]; the diet-specific bacterial OTUs may represent organisms with the capacity to detoxify some of these secondary metabolites, including degrades aromatic hydrocarbons, sulfate reducer and sulfur reducer, and so on (Fig. 5). However, microbial functions do not offer a clear explanation for the varied richness and diversity of termite gut bacterial communities.
In conclusion, experiments on T. ampliceps using different lignocellulosic diets showed that the dietary lignocellulose composition partially shaped the termite gut microbiota, and the gut microbial diversity of T. ampliceps feeding with filter paper diets was significantly lower than that in T. ampliceps fed with corn stover diets and wood diets. The majority of bacterial taxa were shared across all diets, but each diet significantly enriched certain taxa. These results have important implications for sampling and analysis strategies to probe the temporal and epidemiological features of termite gut microbiota and to unveil the intimate co-evolution of insect hosts and their gut microbiota. These findings contribute important new knowledge enhancing our understanding of termite symbiosis and symbiont-assisted lignocellulose digestion.
References
Bugg, T. D., Ahmad, M., Hardiman, E. M., & Singh, R. (2011). The emerging role for bacteria in lignin degradation and bio-product formation. Current Opinion in Biotechnology, 22, 394–400.
McDonald, J. E., Rooks, D. J., & McCarthy, A. J. (2012). Methods for the isolation of cellulose-degrading microorganisms. Methods in Enzymology, 510, 349–374.
Brune, A. (2014). Symbiotic digestion of lignocellulose in termite guts. Nature Reviews Microbiology, 12, 168–180.
Scharf, M. E. (2015). Omic research in termites: an overview and a roadmap. Frontiers in Genetics, 6, 76.
Fraune, S., & Bosch, T. C. G. (2010). Why bacteria matter in animal development and evolution. BioEssays, 32, 571–580.
Scharf, M. E. (2015). Termites as targets and models for biotechnology. Annual Review of Entomology, 60, 77–102.
Warnecke, F., Luginbühl, P., Ivanova, N., Ghassemian, M., Richardson, T. H., Stege, J. T., Cayouette, M., McHardy, A. C., Djordjevic, G., Aboushadi, N., Sorek, R., Tringe, S. G., Podar, M., Martin, H. G., Kunin, V., Dalevi, D., Madejska, J., Kirton, E., Platt, D., Szeto, E., Salamov, A., Barry, K., Mikhailova, N., Kyrpides, N. C., Matson, E. G., Ottesen, E. A., Zhang, X., Hernández, M., Murillo, C., Acosta, L. G., Rigoutsos, I., Tamayo, G., Green, B. D., Chang, C., Rubin, E. M., Mathur, E. J., Robertson, D. E., Hugenholtz, P., & Leadbetter, J. R. (2007). Metagenomic and functional analysis of hindgut microbiota of a wood-feeding higher termite. Nature, 450, 560–565.
Werren, J. H., Baldo, L., & Clark, M. E. (2008). Wolbachia: master manipulators of invertebrate biology. Nature Reviews Microbiology, 6, 740–751.
Benjamino, J., & Graf, J. (2016). Characterization of the core and caste-specific microbiota in the termite, Reticulitermes flavipes. Frontiers in Microbiology, 7, 171.
Ohkuma, M., Sato, T., Noda, S., Ui, S., Kudo, T., & Hongoh, Y. (2007). The candidate phylum ‘termite group 1’ of bacteria: phylogenetic diversity, distribution, and endosymbiont members of various gut flagellated protists. FEMS Microbiology Ecology, 60, 467–476.
Brugerolle, G., & Radek, R. (2006). Symbiotic protozoa of termites. In H. König & A. Varma (Eds.), Intestinal microorganisms of soil invertebrates. Soil biology. Vol. 6 (pp. 243–269). Berlin: Springer.
Berlanga, M., Pasteur, B. J., Grandcolas, P., & Guerrero, R. (2011). Comparison of the gut microbiota from soldier and worker castes of the termite Reticulitermes grassei. International Microbiology, 14, 83–93.
Scharf, M. E., Karl, Z. J., Sethi, A., & Boucias, D. G. (2011). Multiple levels of synergistic collaboration in termite lignocellulose digestion. PloS One, 6, e21709.
Scharf, M. E., Karl, Z. J., Sethi, A., Sen, R., Raychoudhury, R., & Boucias, D. G. (2011). Defining host-symbiont collaboration in termite lignocellulose digestion, “the view from the tip of the iceberg. Communicative & Integrative Biology, 4, 761–763.
Sinma, K., Khucharoenphaisan, K., Kitpreechavanich, V., & Tokuyama, S. (2011). Purification and characterization of a thermostable xylanase from Saccharopolyspora pathumthaniensis S582 isolated from the gut of a termite. Bioscience, Biotechnology, and Biochemistry, 75, 1957–1963.
Hongoh, Y. (2010). Diversity and genomes of uncultured microbial symbionts in the termite gut. Bioscience, Biotechnology, and Biochemistry, 74, 1145–1151.
Tai, V., James, E. R., Nalepa, C. A., Scheffrahn, R. H., Perlman, S. J., & Keeling, P. J. (2015). The role of host phylogeny varies in shaping microbial diversity in the hindguts of lower termites. Applied and Environmental Microbiology, 81, 1059–1070.
Abdul Rahman, N., Parks, D. H., Willner, D. L., Engelbrektson, A. L., Goffredi, S. K., Warnecke, F., Scheffrahn, R. H., & Hugenholtz, P. (2015). A molecular survey of Australian and north American termite genera indicates that vertical inheritance is the primary force shaping termite gut microbiomes. Microbiologica, 3, 5.
Karl, Z. J., & Scharf, M. E. (2015). Effects of five diverse lignocellulosic diets on digestive enzyme biochemistry in the termite Reticulitermes flavipes. Archives of Insect Biochemistry and Physiology, 90, 89–103.
Huang, X. F., Bakker, M. G., Judd, T. M., Reardon, K. F., & Vivanco, J. M. (2013). Variations in diversity and richness of gut bacterial communities of termites (Reticulitermes flavipes) fed with grassy and woody plant substrates. Microbial Ecology, 65, 531–536.
Boucias, D. G., Cai, Y., Sun, Y., Lietze, V. U., Sen, R., Raychoudhury, R., & Scharf, M. E. (2013). The hindgut lumen prokaryotic microbiota of the termite Reticulitermes flavipes and its responses to dietary lignocellulose composition. Molecular Ecology, 22, 1836–1853.
Raychoudhury, R., Sen, R., Cai, Y., Sun, Y., Lietze, V. U., Boucias, D. G., & Scharf, M. E. (2013). Comparative metatranscriptomic signatures of wood and paper feeding in the gut of the termite Reticulitermes flavipes (Isoptera: Rhinotermitidae). Insect Molecular Biology, 22, 155–171.
Brauman, A., Doré, J., Eggleton, P., Bignell, D., Breznak, J. A., & Kane, M. D. (2001). Molecular phylogenetic profiling of prokaryotic communities in guts of termites with different feeding habits. FEMS Microbiology Ecology, 35, 27–36.
Mikaelyan, A., Dietrich, C., Köhler, T., Poulsen, M., Sillam-Dussès, D., & Brune, A. (2015). Diet is the primary determinant of bacterial community structure in the guts of higher termites. Molecular Ecology, 24, 5284–5295.
Huang, F. S., Zhu, S. M., Ping, Z. M., He, X. S., & Li, G. X. (2000). Fauna Sinica Insecta, vol. 17 Isoptera (pp. 430–865). Beijing: Science Press.
Su, L. J., Liu, Y. Q., Liu, H., Wang, Y., Li, Y., Lin, H. M., Wang, F. Q., & Song, A. D. (2015). Linking lignocellulosic dietary patterns with gut microbial Enterotypes of Tsaitermes ampliceps and comparison with Mironasutitermes shangchengensis. Genetics and Molecular Research, 14, 13954–13967.
Liu, N., Yan, X., Zhang, M., Xie, L., Wang, Q., Huang, Y., Zhou, X., Wang, S., & Zhou, Z. (2011). Microbiome of fungus-growing termites: a new res ervoir for lignocellulase genes. Applied and Environmental Microbiology, 77, 48–56.
Schloss, P. D., Gevers, D., & Westcott, S. L. (2011). Reducing the effects of PCR amplification and sequencing artifacts on 16S rRNA-based studies. PloS One, 6, e27310.
Schloss, P. D., Westcott, S. L., Ryabin, T., et al. (2009). Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Applied and Environmental Microbiology, 75, 7537–7541.
Kunin, V., Engelbrektson, A., Ochman, H., & Hugenholtz, P. (2010). Wrinkles in the rare biosphere: pyrosequencing errors can lead to artificial inflation of diversity estimates. Environmental Microbiology, 12, 118–123.
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C., & Knight, R. U. (2011). CHIME improves sensitivity and speed of chimera detection. Bioinformatics, 27, 2194–2200.
Kuhnigk, T., & König, H. (1997). Degradation of dimeric lignin model compounds by aerobic bacteria isolated from the hindgut of xylophagous termites. Journal of Basic Microbiology, 37, 205–211.
Aickin, M., & Gensler, H. (1996). Adjusting for multiple testing when reporting research results: the Bonferroni vs Holm methods. American Journal of Public Health, 86, 726–728.
Berchtold, M., Chatzinotas, A., Schönhuber, W., Brune, A., Amann, R., Hahn, D., & König, H. (1999). Differential enumeration and in situ localization of microorganisms in the hindgut of the lower termite Mastotermes darwiniensis by hybridization with rRNA-targeted probes. Archives of Microbiology, 172, 407–416.
Ni, J., & Tokuda, G. (2013). Lignocellulose-degrading enzymes from termites and their symbiotic microbiota. Biotechnology Advances, 31, 838–850.
Brune, A. (2006). Symbiotic associations between termites and prokaryotes. In M. Dworkin, S. Falkow, E. Rosenberg, K.-H. Schleifer, & E. Stackebrandt (Eds.), The prokaryotes, vol. 1. Symbiotic associations, biotechnology, applied microbiology (pp. 439–474). New York: Springer.
He, S., Ivanova, N., Kirton, E., Allgaier, M., Bergin, C., Scheffrahn, R. H., Kyrpides, N. C., Warnecke, F., Tringe, S. G., & Hugenholtz, P. ((2013)). Comparative metagenomic and metatranscriptomic analysis of hindgut paunch microbiota in wood- and dung-feeding higher termites. PloS One, 8, e61126.
Dietrich, C., Köhler, T., & Brune, A. (2014). The cockroach origin of the termite gut microbiota: patterns in bacterial community structure reflect major evolutionary events. Applied and Environmental Microbiology, 80, 2261–2269.
Dröge, S., Fröhlich, J., Radek, R., & König, H. (2006). Spirochaeta coccoides sp. nov., a novel coccoid spirochete from the hindgut of the termite Neotermes castaneus. Applied and Environmental Microbiology, 72, 392–397.
Hongoh, Y., Sharma, V. K., Prakash, T., Noda, S., Toh, H., Taylor, T. D., Kudo, T., Sakaki, Y., Toyoda, A., Hattori, M., & Ohkuma, M. (2008). Genome of an endosymbiont coupling N2 fixation to cellulolysis within protist cells in termite gut. Science, 322, 1108–1109.
Yang, Y. J., Zhang, N., Ji, S. Q., Lan, X., Zhang, K. D., Shen, Y. L., Li, F. L., & Ni, J. F. (2014). Dysgonomonas macrotermitis sp. nov., isolated from the hindgut of a fungus-growing termite. International Journal of Systematic and Evolutionary Microbiology, 64, 2956–2961.
Pramono, A. K., Sakamoto, M., Iino, T., Hongoh, Y., & Ohkuma, M. (2015). Dysgonomonas termitidis sp. nov., isolated from the gut of the subterranean termite Reticulitermes speratus. International Journal of Systematic and Evolutionary Microbiology, 65, 681–685.
Pinheiro, G. L., Correa, R. F., Cunha, R. S., Cardoso, A. M., Chaia, C., Clementino, M. M., Garcia, E. S., de Souza, W., & Frasés, S. (2015). Isolation of aerobic cultivable cellulolytic bacteria from different regions of the gastrointestinal tract of giant land snail Achatina fulica. Frontiers in Microbiology, 6, 860.
Hongoh, Y., Sato, T., Dolan, M. F., Noda, S., Ui, S., Kudo, T., & Ohkuma, M. (2007). The motility symbiont of the termite gut flagellate Caduceia versatilis is a member of the “Synergistes” group. Applied and Environmental Microbiology, 73, 6270–6276.
Wasi, S., Tabrez, S., & Ahmad, M. (2013). Use of Pseudomonas spp. for the bioremediation of environmental pollutants: a review. Environmental Monitoring and Assessment, 185, 8147–8155.
Jiménez, D. J., Korenblum, E., & van Elsas, J. D. (2014). Novel multispecies microbial consortia involved in lignocellulose and 5-hydroxymethylfurfural bioconversion. Applied Microbiology and Biotechnology, 98, 2789–2803.
Kannisto, M. S., Mangayil, R. K., Shrivastava-Bhattacharya, A., Pletschke, B. I., Karp, M. T., & Santala, V. P. (2015). Metabolic engineering of Acinetobacter Baylyi ADP1 for removal of clostridium butyricum growth inhibitors produced from lignocellulosic hydrolysates. Biotechnology for Biofuels, 8, 198.
Oosterkamp, M. J., Méndez-García, C., Kim, C.-H., Bauer, S., Ibáñez, A. B., Zimmerman, S., Hong, P.-Y., Cann, I. K., & Mackie, R. I. (2016). Lignocellulose-derived thin stillage composition and efficient biological treatment with a high-rate hybrid anaerobic bioreactor system. Biotechnology for Biofuels, 9, 120.
He, Y., Ding, Y., & Long, Y. (1991). Two cellulolytic clostridium species: Clostridium cellulosi sp. nov. and Clostridium cellulofermentans sp. nov. International Journal of Systematic Bacteriology, 41, 306–309.
Huang, X. F., Santhanam, N., Badri, D. V., Hunter, W. J., Manter, D. K., Decker, S. R., Vivanco, J. M., & Reardon, K. F. (2013). Isolation and characterization of lignin-degrading bacteria from rainforest soils. Biotechnology and Bioengineering, 110, 1616–1626.
Geib, S. M., Filley, T. R., Hatcher, P. G., et al. (2008). Lignin degradation in wood-feeding insects. Proceedings of the National Academy of Sciences of the United States of America, 105, 12932–12937.
Chaffron, S., & von Mering, C. (2007). Termites in the woodwork. Genome Biology, 8, 229.
Wei, H., Xu, Q., Taylor, L. E., Baker, J. O., Tucker, M. P., & Ding, S. Y. (2009). Natural paradigms of plant cell wall degradation. Current Opinion in Biotechnology, 20, 330–338.
Vikman, M., Karjomaa, S., Kapanen, A., Wallenius, K., & Itavaara, M. (2002). The influence of lignin content and temperature on the biodegradation of lignocellulose in composting conditions. Applied Microbiology and Biotechnology, 59, 591–598.
Reichling, J. (2010). Plant–microbe interactions and secondary metabolites with antibacterial, antifungal and antiviral properties. In M. Wink (Ed.), Annual plant reviews volume 39: functions and biotechnology of plant secondary metabolites (pp. 214–347). Oxford: Wiley.
Acknowledgments
This research was supported by National Science Foundation of China Grant 31170350.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
We declare that we have no financial and personal relationships with other people or organizations that can inappropriately influence our work. There is no professional or other personal interest of any nature or kind in any product, service, and/or company that could be construed as influencing the position presented in the manuscript entitled, “Variation in the gut microbiota and sensitivity to dietary changes in termite hosts”.
Additional information
Lijuan Su and Lele Yang contributed equally to this work.
Electronic supplementary material
Supplementary Fig. S1
(DOCX 156 kb)
Supplementary Table S1
(DOCX 29 kb)
Supplementary Table S2
(DOCX 68 kb)
Rights and permissions
About this article
Cite this article
Su, L., Yang, L., Huang, S. et al. Variation in the Gut Microbiota of Termites (Tsaitermes ampliceps) Against Different Diets. Appl Biochem Biotechnol 181, 32–47 (2017). https://doi.org/10.1007/s12010-016-2197-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-016-2197-2