Abstract
The sweet potato whitefly, Bemisia tabaci (G.) biotype B (Hemiptera: Aleyrodidae), is one of the most economically important pest, by being a dreaded vector of Geminiviruses, and also causes direct damage to the crops by sucking phloem sap. Glutathione S-transferase (GST) is a large family of multifunctional enzymes that play pivotal roles in the detoxification of secondary allelochemical produced by the host plants and in insecticide resistance, thus regulates insect growth and development. The objective of this study is to show the potential of RNA interference (RNAi) in the management of B. tabaci. RNAi is a sequence-specific gene silencing mechanism induced by double-stranded RNA (dsRNA) which holds tremendous potential in pest management. In this regard, we sequenced the GST from B. tabaci and synthesized approximately 500-bp dsRNA from the above and delivered through diet to B. tabaci. Real-time quantitative PCR (RT-qPCR) showed that continuous application of dsGST at 1.0, 0.5, and 0.25 μg/μl reduced mRNA expression levels for BtGST by 77.43, 64.86, and 52.95 % which resulted in mortality by 77, 59, and 40 %, respectively, after 72 h of application. Disruption of BtGST expression will enable the development of novel strategies in pest management and functional analysis of vital genes in B. tabaci.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bemisia tabaci (Gennadius) (Hemiptera: Aleyrodidae) is a pest of global importance that causes significant crop loss as a direct pest and more importantly as a vector of Geminiviruses [1]. The emergence of novel genetic groups in B. tabaci increased the risk of transmission of Geminiviruses to wide varieties of crops worldwide. It is reported that B. tabaci infests more than 900 species of plants of agricultural, fiber, vegetables, and ornamentals and also a dreaded vector of approximately 111 Geminiviruses [2]. It is also the world’s number one invasive species among the 100 invasive ones according to the International Union for Conservation of Nature and Natural resources (IUCN, http://www.iucn.org/). In addition, B. tabaci excretes copious amount of honeydew that promotes the growth of saprophytic fungi affecting photosynthesis [3]. Chemical management of B. tabaci has become increasingly ineffective due to the development of high levels of resistance to insecticides that include organophosphates, carbamates, and pyrethroids [4, 5]. Therefore, there is an urgent need to harness modern molecular technique as an alternative, where ribonucleic acid interference (RNAi) holds immense potential in developing species-specific insecticides [6–8].
RNAi is the sequence-specific posttranscriptional gene silencing mechanism, discovered more than a decade ago in Caenorhabditis elegans [9]. RNAi has shown its potential in establishing the functions of genes as well as in pest and disease management [10]. RNAi is induced either by endogenous or exogenous double-stranded RNA (dsNA) molecule, which destroys the cognate mRNA, thus affecting the physiology of the organism [9, 11]. To achieve gene silencing, the cognate dsRNA is delivered through various methods such as through diet, soaking, microinjection, feeding bacteria-expressed dsRNA, and dsRNA-expressing transgenic plants [11, 12]. However, dsRNAs delivered either as a spray or through transgenic plants are more pertinent to the field level pest management practices.
RNAi has been successfully used in the management of insect pests by silencing a wide range of genes including gut proteins in tsetse fly [13], molting genes in Tribolium catstaneum [14], aquaporins in aphids [15], digestive and caste regulatory genes in Reticulitermes flavipes [16], olfactory receptors in Apolygus lucorum [17], and extensively in other insect pests [12]. In this study, we have silenced glutathione S-transferase (GST) of the sigma class and evaluated its potential as pest management candidate in B. tabaci. GST is a cytolytic enzyme found in all eukaryotic organisms that participates in a number of different reactions which are important in defense against oxidative damage both by oxygen and free radicals [18]. However, GST has also been implicated in insecticide resistance in whiteflies [19].
In this study, we assessed the feasibility of diet-delivered dsRNA in silencing and persistence of the gene knockdown in two different time intervals. Additionally, we employed less invasive method of dsRNA administration (parafilm feeding method) to increase the ease of experimentation with improved whitefly survival.
Materials and Methods
Insect Culture
B. tabaci (=MEAM-1 genetic group, biotype B , Bemisia argentifolli Bellows and Perring) was utilized from the continuous culture maintained at the Division of Entomology, IIHR, Bangalore, maintained on cotton, Gossypium hirsutam at 28 °C, 45 % RH, and 14:10 photoperiod [11].
Cloning of GST
Total RNA from ten adult flies was extracted employing the RNeasy Mini Kit (Qiagen, Netherlands) following the manufacturer’s protocol. DNA contaminant was removed by treating with RNase-free DNase I (Ferments Life Sciences, USA) and purified by Phenol/chloroform (25:24) extraction [20]. The RNA concentration was estimated using Nanodrop ND-1000 (Thermo Scientific, Belgium) and on 2 % agarose gel electrophoresis. First-strand complementary DNA (cDNA) was prepared from 1 μg RNA employing PrimeScript™ First-Strand cDNA Synthesis Kit (TaKaRa) following the manufacturer’s protocol. Subsequently, cDNA was diluted (1:5) for PCR amplification with gene-specific primers (Table 1). PCR was performed in a final volume of 25 μl containing Taq buffer, 2.5 mm dNTP, 10 pmol of forward and reverse primers, 1 U TaqDNA polymerase, 3 μl of diluted cDNA, and water to make up the volume. The reaction condition was 94 °C for 5 min, followed by 35 cycles of 94 °C for 30 s, 54 °C for 30 s, 72 °C for 45 s, and final extension of 72 °C for 5 min. A negative control containing no DNA template was also included.
Amplified PCR product were eluted using gel extraction kit (Machery Nagel, Germany) and cloned in to general purpose cloning vector (PT257R/T, Thermo Scientific, USA), and three clones were sequenced in both the directions.
Sequence and Phylogenetic Analyses
Sequences assembly (both forward and reverse) were performed in BioEdit [21] and analyzed for homology with other sequences from insects in NCBI-BLAST, and the final sequences were submitted to NCBI-GenBank (Table 1). The annotated GST sequences from B. tabaci and 27 insect species were aligned using MUSCLE [22]. The alignment was used to construct the maximum likelihood (ML) trees using RaxML version 7.26 [23] under GTR + I + G model estimated by ProtTest [24]. To assess the branch support, 500 bootstrapping were performed using CIPRES interface [25].
dsRNA Synthesis
An off-target minimized region of 416 bp was identified for GST employing dsCheck (http:dscheck.RNAi.jp/) [26]. The T7 RNA polymerase promoter sequences (5′-TAATACGACTCACTATAGGG-3′) were tailed to the 5′ of the gene-specific primers that were employed for dsRNA synthesis. DNA templates for dsRNA were synthesized using PCR with similar reagents and PCR conditions, expect for DNA templates (1:100 diluted plasmid clones of the gene) and annealing temperature (65 °C for 45 s). The PCR products were resolved in 1.2 % agarose gel and purified using Nucleospin Extract II kit (Macherey Nagel, Germany). One microgram of the eluted DNA template was used for in vitro transcription using the T7 Ribomax™ express RNAi system (Promega, USA), precipitated with isopropanol and resuspended in nuclease-free water, followed by quantification in Nanodrop™ 1000 (ThermoScientific, USA). The integrity was analyzed by agarose gel electrophoresis (1.5 %) and kept at −70 °C until further use. dsRNA was diluted with artificial diet [27] to yield with three different concentrations, viz., 0.25, 0.50, and 1.0 μg/μl.
Insect Bioassay
A diet-based dsRNA delivery technique as previously reported [27–29] was followed in this study. The above diet was prepared in molecular biology grade water, filter-sterilized (0.22 μm) prior to the mixing of dsRNA at three concentrations, viz., 0.25, 0.50, and 1.0 μg/μl, and sandwiched between the two layers of UV-sterilized parafilm and one stretched on the inner surface of the cap. About 40 newly emerged adults of B. tabaci were taken in each tube, allowing them to feed on diet alone for 6 to 8 h, and then, healthy live 20 insects were transferred to a new tube with the diet containing dsRNA and the mortality was recorded daily. There were five treatments, such as diet alone (blank control), the diet containing dsRNA for Lac-Z (negative control), and the diets containing dsGSTs. All the treatments were replicated five times, and a total of 100 adults in each treatment were used.
RT-qPCR
Total RNA extracted from ten adults and converted to cDNA as described previously. cDNA was diluted tenfold of which 5 μl was used for PCR amplification employing gene-specific primers (Table 1). Real-time PCR was performed employing SYBR Green Jump Start™ Taq ReadyMix™ (Sigma-Aldrich, USA) according to the manufacturer’s protocol in a Light Cycler 480 II (Roche Applied Science, Switzerland). All the RT-qPCR assays were performed according to the MIQE guidelines [30]. Actin was used as an internal control gene for normalization, which was found to be the most suitable reference gene in a previous study (unpublished data). All the assays were triplicated, and also, no template controls were included. The relative gene expression data were analyzed using 2−ΔΔCT method as described by Livak and Schmittgen [31], and normalized values were expressed as percent silencing and compared to negative controls.
Data Analyses
The data is presented as mean ± SE and were analyzed by using t test employing GraphPad Prism v.5 (GraphPad Software, Inc., USA) at P < 0.05.
Results
Molecular Cloning and Sequence Analyses
A full-length GST (609 bp) was amplified employing the cDNA prepared from the total RNA from adult B. tabaci and submitted to NCBI-GenBank (accession number KJ913696). The BLAST results of the sequence indicated 100 % match to the previously reported GST sequences from B. tabaci. The deduced sequences of the cDNA consisted of 203 amino acid residues with a calculated molecular mass (MM) and isoelectric point (PI) of 50.09 kDa and 5.19, respectively, employing ExPASy Proteomics Website (http://web.expasy.org/). TMHMM Server v. 2.0 (http://www.cbs.dtu.dk/services/TMHMM-2.0/ [32] could not predict any transmembrane helix from our GST sequence. Similarly, no signal peptide motif was identified by SignalP 4.0 Server (http://www.cbs.dtu.dk/services/SignalP/) (Supplementary Fig. 1) as a result of no cleavage site evident.
Phylogenetic Analyses
NCBI-GenBank database search identified 27 sigma-class GSTs, and the retrieved sequences exhibited perfect match with GST sequence of B. tabaci from this study implemented in the MUSCLE program fixed in the MEGA 6.0 [22]. The alignment was used to construct a phylogenetic tree using MEGA 6.0. Neighbor-joining, maximum parsimony, ML, and minimum evolution methods generated trees with similar topology and bootstrap values; thus, only the ML tree is presented here (Fig. 1). The overall sequence homology shared between the BtGST and known sigma-class GSTs from other species of insects was about 49 to 65 % with highest homology to Acyrthosiphon pisum. The evolutionary relationship of GST proteins derived from B. tabaci and 28 other species of insects were evaluated. The ML trees for the GSTs of B. tabaci grouped with various aphid species viz., Acyrthosiphon pisum, Aphis citricidus, Aphis gossypii, and Kaburagia rhusicola that are nearest to the order Hemiptera.
Maximum likelihood (ML) tree showing phylogenetic relationship of the newly identified sigma-class GST from Bemisia tabaci (BtGST) with other corresponding sequences from different insects. Species name of insects with NCBI-GenBank accession numbers mentioned in the tree with indicated bootstrap values >50 %. Apart from sigma class, other classes of GSTs viz., theta, mu, zeta, omega, delta, and epsilon from Bombyx mori and Sarcoptes scabiei used as outgroup
Insect Bioassay
The administration of dsRNA (Fig. 2) for GST showed significant levels of mortality due to silencing. The mortality data recorded in different time intervals such as 24, 36, 48, 60, and 72 h for three concentrations of cognate dsRNA viz., 0.25, 0.50, and 1.0 μg/μl and are given in the Supplementary Table 1. In each treatment, 20 adult whiteflies were used and the mortality recorded at different time intervals as independent experiments. Diet-delivered dsRNA for GST produced mortality ranging from 3.0 to 77 % (F = 1.335, df = 4, 20, P < 0.05) (Fig. 3). Mortality was observed in treated (non-target gene, dsLacZ) and untreated (diet alone) controls, which was in the range of 3.0–20.00 and 2.0–21.00 %, respectively. However, our results indicated that diet-delivered dsRNA for LacZ did not show any impact on whitefly mortality, since the similar percent mortality was observed in the untreated control (Figs. 3 and 4).
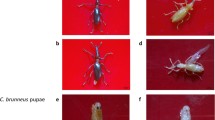
Collection of whiteflies and bioassays: diet pouch with diet were prepared using sterilized stretched parafilm on individual caps of specimen tubes (A1 and A2). Adult flies were collected on cabbage, Brassica oleracea plants and transferred into 30 ml specimen tubes (A3 and A4). Control live white flies feeding on the sandwiched diet (48 h). Diet pouch with dsRNA were prepared using sterilized stretched parafilm on individual caps of specimen tubes (B1 and B2). Dead white flies upon the treatment inside the tubes (48 h) (side view) (B3 and B4)
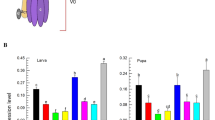
The percent mortality of whiteflies after diet-mediated dsGST ingestion. Mortality rate was counted in every 24, 36, 48, 60, and 72 h. The values are represented as the average of five biological replications. Error bars represent the standard deviation value. Asterisks above the bars indicate that the values were significantly different (P < 0.05, Tukey’s t test, n = 5)
Percent silencing of whiteflies after diet-mediated dsGST delivery. Expression was monitored for 2 days. Sampling was done on 24 and 48 h. The values are represented as the average of three biological replications. β-Actin was used as an internal reference gene. The results were analyzed using 2−∆∆CT method. Error bars represent the standard deviation of 2−∆∆CT value. Asterisks above the bars indicate the values were significantly different (P < 0.001, Tukey’s t test, n = 3)
Target Gene Silencing
Real-time PCR efficiency was calculated for both GST (target) and Actin (internal control) as 1.936 (96.8 %) and 1.942 (97.1 %), respectively. Melt curve analyses of the amplicons of these genes exhibited single peak that indicated single product, which was reconfirmed by cloning, sequencing, and BLAST analyses of the same.
Our results showed downregulation of BtGST transcript in comparison with both controls on different time intervals across various dsRNA concentrations (Fig. 4). The percent silencing of BtGST varied from 7.84 % (0.25 μg/μl–24 h) to 77.43 % (1.0 μg/μl–72 h) compared to the dsLacZ and untreated whiteflies (Fig. 4). Percent silencing is directly proportional to the dsRNA the concentration (Fig. 4) and mortality (Figs. 3 and 4).
Discussion
The GST is a large family of multifunctional enzymes that play a vital role in the detoxification of both endogenous and xenobiotic compounds [33]. GSTs primarily catalyze the conjugation of the thiol group of reduced glutathione to the electrophilic centers of lipophilic compounds rendering them more water soluble and can be rapidly excreted [34, 35]. Majority of studies on insect GSTs have been focused on their role in detoxification of insecticides and allelochemicals and oxidative stress response [35–37]. GSTs are primarily classified into two major groups based on their location within the cell, i.e., microsomal and cytosolic. A third group of GSTs, the kappa class, is located in mammalian mitochondria and peroxisomes [38, 39]. Later, phylogenetic analyses of insect GST genes showed that it can be broadly classified into different classes, viz., sigma, delta, and epsilon, and majority of the remaining cytosolic insect GSTs are members of the zeta, theta, and omega classes [40–42]. Specifically, GSTs have been shown their potential in developing insecticide resistance in B. tabaci [5], ticks [43, 44], mites [45], etc. In this study, we sequenced BtGST and showed that it is phylogenetically more closely related to sigma class (insect class II) and more divergent from other classes of GSTs such as mu, epsilon, delta, zeta, and theta (Fig. 1).
In this study, we also demonstrated that diet-mediated delivery of dsGST can successfully inhibit the translation of the same in B. tabaci. Oral delivery of dsRNA has enormous potential applications in the management of agricultural insect pests. Diet-mediated dsRNA-induced gene silencing has been previously evidenced in a wide variety of agricultural insect pests of the order Lepidoptera [46–49], Diptera [13, 50–53], Orthoptera [54], Hymenoptera [55], Hemiptera [15, 56–65], Coleoptera [63, 66–69], and Isoptera [16]. So far, dsRNA delivery to various insect has been accomplished through oral feeding, microinjection, soaking, and transgenics [12, 70, 71, 46]. Among which, microinjection is the most commonly used method, but the disadvantages are (i) laborious, (ii) requirement of expertise and equipments, and (iii) mortality rates will be higher due to the invasive nature. Hence, we employed alternate less invasive and easy method of diet-mediated dsRNA delivery to assess the effect of RNAi in B. tabaci. In this study, we demonstrated that the ingestion of dsRNA can bring about the gene silencing of target genes in B. tabaci. A reduction in the BtGST gene transcripts resulted in mortality of B. tabaci during the bioassay. Thus, our study showed that BtGST downregulation mediated by dsRNA ingestion was lethal to B. tabaci and revealed the role of GST in cell signaling for cell proliferation required for growth and development of this hemimetabolous insect.
The effect of diet-delivered dsRNA on BtGST transcript levels in B. tabaci has been directly proportional to the dose (Fig. 4). Similar results were observed in Sogatella furcifera [72], Nilapravata lugens [73], Aedes aegypti [51], etc. However, certain other studies have also observed that increasing dsRNA concentration beyond an optimal dose did not improve extent of gene silencing, although the optimal concentration of dsRNA may vary for the insect species in question, mode of cognate dsRNA delivery, and the developmental stages of insects [15, 12]. Our percent silencing ranged 7.84–21.43, 16.67–30.85, 25.15–42.65, 37.77–57.94, and 52.95–77.43 % for BtGST in 24, 36, 48, 60, and 72 h, respectively. Thus, our data is comparable to that of the previous studies conducted in many other insects, where the percent silencing typically ranged from 30 to 80 % with either mortality or observable phenotypic changes [13, 16, 48, 51, 52, 57, 64, 74].
It is known that few factors are critical for exogenous dsRNA-mediated gene silencing in insects such as physiological importance of the target genes, concentration of cognate dsRNA, frequency of administration, and presence or absence of key core components for systemic RNAi machinery such as systemic interference defective (sid-1) and RNA-dependent RNA polymerase (RdRp), which are less understood in insects [75, 76]). As far as target genes are concerned, our results showed that BtGSTs could potentially serve as a target for developing novel insect pest management strategy. Different target genes resulted in various levels of silencing with either mortality or phenotypes [12, 14]. Our results showed that after 72 h of ingestion of dsGST, it resulted in 77 % mortality, while 21 and 20 % mortality was observed in untreated and dsLacZ, respectively. It has been previously reported that about 70 % silencing was evident after hemocoelic injection of dsRNA in B. tabaci [11] as against 29–97 % mortality [29], which supports the results obtained in the present study.
Depletion of 7.84 % (0.25 μg/μl–24 h) to 77.43 % (1.0 μg/μl–72 h) of BtGST transcripts though was not high as compared to the previous studies on Spodoptera litura, where 95 % reduction of amino peptidase transcript was evident. It could be due to the pooled samples of B. tabaci that were used to construct the overall efficiency of dsGST. Both mortality data and percent silencing of BtGST indicated that the extent of mortality and silencing was directly proportional to the concentration of dsRNA administered. Thus, gene knockdown by diet-mediated delivery of dsGST in B. tabaci will be undoubtedly an invaluable tool in the management of this invasive insect pest.
In conclusion, we have demonstrated the utility of the exogenously administered cognate dsRNA for GST of B. tabaci, a notorious vector of Geminiviruses and direct pests of wide varieties crops worldwide. We have also designed a simple, inexpensive, high-throughput method of dsRNA administration, which resulted in mortality due to downregulation of BtGST. Mortality was evaluated across three concentrations of diet-delivered dsGST of sigma class with highest mortality of 77.43 % for 1.0 μg/μl after 72 h, while mortality in control was 21 % under the same conditions. Based on the present study, we conclude that oral delivery of cognate dsRNA through diet or transgenic crops expressing dsRNA has potential applications for whitefly management and deserve further research.
References
Byrne, D. N., & Bellows, T. S. (1991). Whitefly biology. Annual Review of Entomology, 36, 431–457.
Jones, D. R. (2003). Plant viruses transmitted by whiteflies. European Journal of Plant Pathology, 109, 195–219.
Brown, J. K., & Czosnek, H. (2002). Whitefly transmission of plant viruses. In R. T. Plumb (Ed.), Advances in botanical research (Plant virus vector interactions, Vol. 36, pp. 65–100). New York: Academic.
Horowitz, A. R., Kontsedalov, S., Denholm, I., & Ishaaya, I. (2002). Dynamics of insecticide resistance in Bemisia tabaci: a case study with the insect growth regulator pyriproxyfen. Pest Management Science, 58, 100–112.
Foster, S. P., Devine, G., & Devonshire, A. L. (2007). Insecticide resistance. In H. F. van Emden & R. Harrington (Eds.), Aphids as crop pests (pp. 261–285). UK: CABI.
Caplen, N. J., Fleenor, J., Fire, A., & Morgan, R. A. (2000). DsRNA-mediated gene silencing in culture Drosophila Cells: a tissue culture model for analysis of RNA interference. Gene, 252, 95–105.
Price, D. R., & Gatehouse, J. A. (2008). RNAi-mediated crop protection against insects. Trends in Biotechnology, 26, 393–400.
Mao, Y. B., Tao, X. Y., Xue, X. Y., Wang, L. J., & Chen, X. Y. (2011). Cotton plants expressing CYP6AE14 double-stranded RNA show enhanced resistance to bollworms. Transgenic Research, 20, 665–673.
Fire, A., Xu, S., Montgomery, M. K., Kostas, S. A., Driver, S. E., & Mello, C. C. (1998). Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature, 391, 806–811.
Christiaens, O., Swevers, L, & Smagghe, G. (2014). DsRNA degradation in the pea aphid (Acyrthosiphon pisum) associated with lack of response in RNAi feeding and injection. Peptides, 53, 307–314.
Ghanim, M., Kontsedalov, S., & Czosnek, H. (2007). Tissue-specific gene silencing by RNA interference in the whitefly Bemisia tabaci (Gennadius). Insect Biochemistry and Molecular Biology, 37, 732–738.
Huvenne, H., & Smagghe, G. (2010). Mechanisms of dsRNA uptake in insects and potential of RNAi for pest control: a Review. Journal of Insect Physiology, 56, 227–235.
Walshe, D. P., Lehane, S. M., Lehane, M. J., & Haines, L. R. (2009). Prolonged gene knockdown in the tsetse fly Glossina by feeding double stranded RNA. Insect Molecular Biology, 18, 11–19.
Konopova, B., & Jindra, M. (2007). Juvenile hormone resistance gene methoprene-tolerant controls entry into metamorphosis in the beetle, Tribolium catstaneum. Proceedings of the National Academy of Science, 104, 10488–10493.
Shakesby, A. J., Wallace, I. S., Isaacs, H. V., Pritchard, J., Roberts, D. M., et al. (2009). A water-specific aquaporin involved in aphid osmoregulation. Insect Biochemistry and Molecular Biology, 39, 1–10.
Zhou, X., Wheeler, M. M., Oi, F. M., & Scharf, M. E. (2008). RNA interference in the termite Reticulitermes flavipes through ingestion of double-stranded RNA. Insect Biochemistry and Molecular Biology, 38, 805–815.
Zhou, Y. L., Zhu, X. Q., Gu, S. H., Cui, H. H., Guo, Y. Y., Zhou, J. J., & Zhang, Y. J. (2014). Silencing in Apolygus lucorum of the olfactory coreceptor Orco gene by RNA interference induces EAG response declining to two putative semiochemicals. Journal of Insect Physiology, 60, 31–39.
Sharp, P. J., Smith, D. R., Bach, W., Wagland, B. M., & Cobon, G. S. (1991). Purified glutathione S-transferases from parasites as candidate protective antigens. International Journal for Parasitology, 21, 839–846.
Yang, N., Xie, W., Yang, X., Wang, S., Wu, Q., et al. (2013). Transcriptomic and proteomic responses of sweet potato whitefly, Bemisia tabaci, to thiamethoxam. PLoS ONE, 8(5), e61820. doi:10.1371/journal.pone.0061820.
Sambrook, J., & Russell, D.W. (2001). Molecular cloning. Cold Spring Harbor Laboratory Press, Cold Spring Harbor.
Hall, T. A. (1999). BioEdit: a user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Nucleic Acids Symposium Series, 41, 95–98.
Tamura, K., Stecher, G., Peterson, D., Filipski, A., & Kumar, S. (2013). MEGA6: molecular evolutionary genetics analysis version 6.0. Molecular Biology and Evolution, 30, 2725–2729.
Stamatakis, A. (2006). RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics, 22, 2688–2690.
Darriba, D., Taboada, G. L., Doallo, R., & Posada, D. (2011). ProtTest 3: fast selection of best-fit models of protein evolution. Bioinformatics, 27, 1164–1165.
Stamatakis, A. (2008). The RAxML 7.0.4 manual. http://icwww.epfl.ch/~stamatak/index-Dateien/countManual7.0.4.php. Accessed 12 Sept 2014.
Naito, Y., Tamuda, T., Mastumiya, T., Kumoko, U. T., Saigo, K., & Morihita, S. (2005). dsChech: highly sensitive off-target search software for double-stranded RNA-mediated RNA interference. Nucleic Acids Research, 33, 589–591.
Blackburn, M. B., Domek, J. M., Gelman, D. B., & Hu, J. S. (2005). The broadly insecticidal Photorhabdus luminescens toxin complex a (TCA): activity against the Colorado potato beetle and sweet potato whitefly. Journal of Insect Science, 5, 32.
Febvay, G., Delobel, B., & Rahbe, Y. (1988). Influence of the amino-acid balance on the improvement of an artificial diet for a biotype of Acyrthosiphon pisum (Homoptera, Aphididae). Canadian Journal of Zoology, 66, 2449–2453.
Upadhyay, S. K., Chandrashekar, K., Thakur, N., Singh, P. K., Tuli, R., et al. (2011). RNA interference for the control of whiteflies (Bemisia tabaci) by oral route. Journal of Biological Sciences, 36, 153–161.
Bustin, S. A., Benes, V., Garson, J. A., Hellemans, J., Huggett, J., & Kubista, M. (2009). The MIQE guidelines—minimum information for publication of quantitative real-time PCR experiments. Clinical Chemistry, 55, 611–622.
Livak, K. J., & Schmittgen, T. D. (2001). Real-time quantitative PCR. Methods, 25, 383–385.
Krogh, A., Larsson, B., Heijne, G., & Sonnhammer, E. L. (2001). Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. Journal of Molecular Biology, 305, 567–580.
Salinas, A. E., & Wong, M. G. (1999). Glutathione S-transferases—a review. Current Medicinal Chemistry, 6(4), 279–309.
Habig, W. H., Pabst, M. J., & Jakoby, W. B. (1974). Glutathione S-transferases. The first enzymatic step in mercapturic acid formation. Journal of Biological Chemistry, 249, 7130–7139.
Enayati, A. A., Ranson, H., & Hemingway, J. (2005). Insect glutathione transferases and insecticide resistance. Insect Molecular Biology, 14(1), 3–8. doi:10.1111/j.1365-2583.2004.00529.x.
Vontas, J. G., Small, G. J., & Hemingway, J. (2001). Glutathione S-transferases as antioxidant defence agents confer pyrethroid resistance in Nilaparvata lugens. Biochemistry Journal, 357, 65–72.
Sawicki, R., Singh, S. P., Mondal, A. K., Benes, H., & Zimniak, P. (2003). Cloning, expression and biochemical characterization of one Epsilon class (GST-3) and ten Delta class (GST-1) Glutathione-S-Transferases from Drosophila melanogaster, and identification of additional nine members of the Epsilon class. Biochemistry Journal, 370, 661–669.
Morel, F., Rauch, C., Petit, E., Piton, A. L., Theret, N., Coles, B., & Guillouzo, A. (2004). Gene and protein characterization of the human glutathione S-transferase kappa and evidence for a peroxisomal localization. The Journal of Biological Chemistry, 279, 16246–16253.
Lander, J. E., Parsons, J. F., Rife, C. L., Gilliand, G. L., & Armstrong, R. N. (2004). Parallel evolutionary pathways for glutathione transferases: structure and mechanisms of the mitochondrial class Kappa enzyme rGSTK1-1. Biochemistry, 43, 252–261.
Board, P. G., Baker, R. T., Chelvanayagam, G., & Jermiin, L. S. (1997). Zeta, a novel class of glutathione transferases in a range of species from plants to humans. Biochemical Journal, 328, 929–935.
Board, P. G., Coggan, M., Chelvanayagam, G., Easteal, S., Jermiin, L. S., et al. (2000). Identification, characterization, and crystal structure of the Omega class glutathione transferases. Journal of Biological Chemistry, 275(32), 24798–24806. doi:10.1074/jbc.m001706200.
Ranson, H., Claudianos, C., Ortelli, F., Abgrall, C., Hemingway, J., Sharakhova, M. V., Unger, M. F., Collins, F. H., & Feyereisen, R. (2002). Evolution of supergene families associated with insecticide resistance. Science, 298, 179–181.
Enayati, A., Asgarian, A. A., Sharif, M., Boujhmehrani, H., Amouei, A., Vahedi, N., Boudaghi, B., Piazak, N., & Hemingway, J. (2009). Propetamphos resistance in Rhipicephalus bursa (Acari, Ixodidae). Veterinary Parasitology, 162, 135–141.
Enayati, A., & Hemingway, J. (2010). Malaria management: past, present, and future. Annual Review of Entomology, 55, 569–591. doi:10.1146/annurev-ento-112408-085423.
Molin, E. U., & Mattsson, J. G. (2008). Effect of acaricides on the activity of glutathione transferases from the parasitic mite Sarcoptes scabiei. Parasitology, 135, 115–123.
Turner, C. T., Davy, M. W., MacDiarmid, R. M., Plummer, K. M., Birch, N. P., & Newcomb, R. D. (2006). RNA interference in the light brown apple moth, Epiphyas postvittana (Walker) induced by double-stranded RNA feeding. Insect Molecular Biology, 15, 383–391.
Mao, Y. B., Cai, W. J., Wang, J. W., Hong, G. J., Tao, X. Y., et al. (2007). Silencing a cotton bollworm P450 monooxygenase gene by plant-mediated RNAi impairs larval tolerance of gossypol. Nature Biotechnology, 25, 1307–1313.
Bautista, M. A., Anita, M., Miyata, T., Miura, K., & Tanaka, T. (2009). RNA interference mediated knockdown of a cytochrome P450, CYP6BG1, from the diamondback moth, Plutella xylostella, reduces larval resistance to permethrin. Insect Biochemistry and Molecular Biology, 39, 38–46.
Ren, X. L., Chen, R. R., Zhang, Y., Ma, Y., Cui, J. J., et al. (2013). A Spodoptera exigua cadherin serves as a putative receptor for Bacillus thuringiensis Cry1Ca toxin and shows differential enhancement to Cry1Ca and Cry1Ac toxicity. Applied and Environmental Microbiology, 79, 5576–5583.
Li, J., Chen, Q. H., Lin, Y. J., Jiang, T. R., Wu, G., et al. (2011). RNA interference in Nilaparvata lugens (Homoptera: Delphacidae) based on dsRNA ingestion. Pest Management Science, 67, 852–859.
Singh, A. D., Wong, S., Ryan, C. P., & Whyard, S. (2013). Oral delivery of double stranded RNA in larvae of the yellow fever mosquito, Aedes aegypti: implications for pest mosquito control. Journal of Insect Science, 13, 69.
Zhang, X., Zhang, J., & Zhu, K. Y. (2010). Chitosan/double-stranded RNA nanoparticle-mediated RNA interference to silence chitin synthase genes through larval feeding in the African malaria mosquito (Anopheles gambiae). Insect Molecular Biology, 19, 683–693.
Cancino-Rodezno, A., Alexander, C., Villasenor, R., Pacheco, S., Porta, H., et al. (2010). The mitogen-activated protein kinase p38 is involved in insect defense against Cry toxins from Bacillus thuringiensis. Insect Biochemistry and Molecular Biology, 40, 58–63.
Meyering-Vos, M., & Muller, A. (2007). RNA interference suggests sulfakinins as satiety effectors in the cricket Gryllus bimaculatus. Journal of Insect Physiology, 53, 840–848.
Maori, E., Paldi, N., Shafir, S., Kalev, H., Tsur, E., et al. (2009). IAPV, a bee-affecting virus associated with colony collapse disorder can be silenced by dsRNA ingestion. Insect Molecular Biology, 18, 55–60.
Chen, J., Zhang, D., Yao, Q., Zhang, J., Dong, X., et al. (2010). Feeding-based RNA interference of a trehalose phosphate synthase gene in the brown plant hopper, Nilaparvata lugens. Insect Molecular Biology, 19, 777–786.
Li, X., Zhang, M., & Zhang, H. (2011). RNA interference of four genes in adult Bactrocera dorsalis by feeding their dsRNAs. PloS One, 6, e17788.
Zha, W., Peng, X., Chen, R., Du, B., Zhu, L., et al. (2011). Knockdown of midgut genes by dsRNA-transgenic plant-mediated RNA interference in the hemipteran insect Nilaparvata lugens. PLoS ONE, 6, e20504.
Zhang, Y., Li, S., Wang, L., Zi, J., He, P., et al. (2013). Over expression of carboxylesterase-1 and mutation (F439H) of acetylcholinesterase-1 are associated with chlorpyrifos resistance in Laodelphax striatellus. Pesticide Biochemistry and Physiology, 106, 8–13.
Sun, X. Q., Zhang, M. X., Yu, J. Y., Jin, Y., Ling, B., et al. (2013). Glutathione S-Transferase of brown plant hoppers (Nilaparvata lugens) is essential for their adaptation to gramine-containing host plants. PloS One, 8, e64026.
He, P., Zhang, J., Liu, N. Y., Zhang, Y. N., Yang, K., et al. (2011). Distinct expression profiles and different functions of odorant binding proteins in Nilaparvata lugens Stal. PloS One, 6, e28921.
Lu, D. H., Wu, M., Pu, J., Zhang, Q., & Han, Z. J. (2013). A functional study of two dsRNA binding protein genes in Laodelphax striatellus. Pest Management Science, 69, 1034–1039.
Whyard, S., Singh, A. D., & Wong, S. (2009). Ingested double-stranded RNAs can act as species-specific insecticides. Insect Biochemistry and Molecular Biology, 39, 824–832.
Araujo, R., Santos, A., Pinto, F., Gontijo, N., Lehane, M., et al. (2006). RNA interference of the salivary gland nitrophorin 2 in the triatomine bug, Rhodnius prolixus (Hemiptera: Reduviidae) by dsRNA ingestion or injection. Insect Biochemistry and Molecular Biology, 36, 683–693.
Yao, J., Rotenberg, D., Afsharifar, A., Barandoc-Alviar, K., & Whitfield, A. E. (2013). Development of RNAi methods for Peregrinus maidis, the corn plant hopper. PLoS ONE, 8, e70243.
Baum, J. A., Bogaert, T., Clinton, W., Heck, G. R., Feldmann, P., et al. (2007). Control of coleopteran insect pests through RNA interference. Nature Biotechnology, 25, 1322–1326.
Zhu, F., Xu, J., Palli, R., Ferguson, J., & Palli, S. R. (2011). Ingested RNA interference for managing the populations of the Colorado potato beetle, Leptinotarsa decemlineata. Pest Management Science, 67, 175–182.
Wan, P. J., Lu, D., Guo, W. C., Ahmat, T., Yang, L., et al. (2013). Molecular cloning and characterization of a putative proline dehydrogenase gene in the Colorado potato beetle, Leptinotarsa decemlineata (Say). Insect Science. doi:10.1111/1744- 7917.12034.
Zhou, L. T., Jia, S., Wan, P. J., Kong, Y., Guo, W. C., et al. (2013). RNA interference ofa putative S-adenosyl-L-homocysteine hydrolase gene affects larval performance in Leptinotarsa decemlineata (Say). Journal of Insect Physiology, 59, 1049–1056.
Blandin, S., Moita, L. F., Kocher, T., Wilm, M., Kafatos, F. C., & Levashina, E. A. (2002). Reverse genetics in the mosquito Anopheles gambiae: targeted disruption of the Defensin gene. EMBO Reports, 3, 852–856.
Pitino, M., Coleman, A. D., Maffei, M. E., Ridout, C. J., & Hogenhout, S. A. (2011). Silencing of aphid genes by dsRNA feeding from plants. PloS One, 6, e25709.
Wan, P. J., Jia, S., Li, N., Fan, J. M., & Li, G. Q. (2014). RNA interference depletion of the Halloween gene disembodied implies its potential application for management of plant hopper Sogatella furcifera and Laodelphax striatellus. PLoS ONE, 9(1), e86675. doi:10.1371/journal.pone.0086675.
Liu, S., Ding, Z., Zhang, C., Yang, B., & Liu, Z. (2010). Gene knockdown by introthoracic injection of double-stranded RNA in the brown plant hopper, Nilaparvata lugens. Insect Biochemistry and Molecular Biology, 40, 666–671.
Campbell, E. M., Budge, G. E., & Bowman, A. S. (2010). Gene-knockdown in the honey bee mite, Varroa destructor by a non-invasive approach: studies on a glutathione S-transferase. Parasites & Vectors, 3, 73–83.
Garbutt, J. S., Belles, X., Richards, E. H., & Reynolds, S. E. (2012). Persistence of double stranded RNA in insect hemolymph as a potential determiner of RNA interference success: evidence from Manduca sexta and Blattella germanica. Journal of Insect Physiology. doi:10.1016/j.jinsphys.2012.05.013.
Belles, X. (2010). Beyond drosophila: RNAi in vitro and functional genomics in insects. Annual Review of Entomology, 55, 111–128.
Acknowledgments
We thank ICAR, New Delhi, for funding the project on 12th Plan Out Reach Program on Sucking Pests (ORP-SP). This work was supported by the Out Reach Program on Sucking Pests (ORP-SP) of Indian Council for Agricultural Research (ICAR), India. Our sincere thanks are due to the Director, IIHR, Bangalore, for encouragement and the necessary facilities. Finally, we are very grateful to Dr. V. V. Ramamurthy (Indian Agricultural Research Institute (IARI), New Delhi) for his continuous support and suggestions over the course of this work.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Figure 1
SignalP prediction for peptide motif of sigma class GST sequences from Bemisia tabaci. No signal peptide motif was identified, which was evident from the absence of cleavage site. (GIF 178 kb)
Supplementary Table 1
(XLSX 9 kb)
Rights and permissions
About this article
Cite this article
Asokan, R., Rebijith, K.B., Roopa, H.K. et al. Non-Invasive Delivery of dsGST Is Lethal to the Sweet Potato Whitefly, Bemisia tabaci (G.) (Hemiptera: Aleyrodidae). Appl Biochem Biotechnol 175, 2288–2299 (2015). https://doi.org/10.1007/s12010-014-1437-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-014-1437-6