Abstract
Purpose of Review
Acute respiratory infections caused by influenza virus are a major cause of viral respiratory diseases globally. Surveillance of circulating subtypes and estimation of disease burden is of utmost clinical importance. Molecular surveillance and proper disease burden estimates are scarce in India although clinical influenza infections are on the rise. Our study aims to delineate the prevalent influenza subtypes in a South Indian population from cases requiring hospital visits. Using real-time polymerase chain reaction (RT-PCR), 2154 throat/nasopharyngeal swabs from patients attending Government Medical College, Thiruvananthapuram, Kerala, India, with suspected influenza-like illness, were tested for the presence of different influenza subtypes.
Research Findings
Forty-three percent of specimens were positive for the influenza virus. Among these, prevalence of influenza A(H3N2), influenza B, and H1N1pdm09 was 26.7, 6.3, and 10%, respectively. Nominal co-infections were detected. An easy to use commercial kit was used for the majority of the study after proper evaluation for sensitivity and specificity against a gold standard protocol.
Summary
Specific diagnosis using molecular tools caters to the urgency, and a precise measure of the disease burden and management actions are needed, especially in developing countries like India. Infection rate estimation using a sensitive RT-PCR assay signified that influenza was highly prevalent in the region. The study data generated will help understand the epidemiology of influenza in India as well as generate information for global influenza surveillance and disease burden.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Influenza A and B viruses belong to the family Orthomyxoviridae with a genome makeup of a single-stranded, negative-sense RNA. They cause acute respiratory infections and form a major chunk of the global disease burden for viral respiratory diseases [1••, 2••, 3••]. Influenza A viruses are further categorized into subtypes H1N1 and H3N2 on the basis of surface glycoproteins, hemagglutinin (HA), and neuraminidase (NA). While antigenically distinct B strains are currently circulating globally, these strains are not too genetically different from the A subtype. Influenza B viruses are currently grouped into two lineages, Victoria and Yamagata, but are not subtyped any further. Influenza A viruses circulate among a diverse range of host species [3••]. The WHO Global Influenza Surveillance Network has greatly contributed to the knowledge about circulating influenza viruses, including the emergence of novel strains [4••, 5••, 6••]. A limited number of studies on public health surveillance of influenza had been reported from India, but credible data on influenza disease burden from the southern region is still missing. Several outbreaks in Pune, Himachal Pradesh, Delhi, Kashmir, and Kolkata have been investigated [7••, 8••, 9••, 10••]. To date, the requisite data to estimate influenza-associated disease burden is scant or absent in most developing countries (http://www.who.int/influenza/resources/publications/manual_burden_of_disease/en/). In this report, we summarize the surveillance data on influenza from 2010 to 2016, in hospital attending/admitted patients from Thiruvananthapuram in southern India.
Traditional methods to detect influenza virus from clinical samples include virus culture, isolation, and characterization by immunoassay, which usually take 3–7 days [11••, 12••, 13]. Later, developed rapid point-of-care tests are simple to use and enable rapid testing within a few minutes, but generally has low sensitivity compared to viral culture or molecular techniques like polymerase chain reaction [14••, 15•, 16••, 17•, 18••]. Other diagnostic methods employ real-time (RT) PCR and multiplex RT-PCR combined with chip-based detection [19••, 20•, 21••, 22••]. However, the drawback with established RT-PCR protocols is that individual influenza subtypes have to be detected in separate reactions increasing cost and effort in disease diagnosis [23••, 24••, 25••, 26••].
There is a need for reliable disease burden estimates especially from developing countries to provide a better understanding of the impact of influenza in vulnerable communities or subpopulations. Use of molecular tests like RT-PCR is important for surveillance in order to accurately identify which influenza subtypes are circulating and the rate of co-infections with other seasonally co-occurring viral/bacterial pathogens or other influenza subtypes. Molecular tests also help clinicians reliably confirm the highly virulent 2009 influenza H1N1 virus (H1N1pdm09) from patients without the need of prolonged exposure and also help in better and quick management of such patients.
In light of the above facts, the present report investigates the influenza virus infection as the cause of an influenza-like illness (ILI) and severe acute respiratory infection (SARI)-like illness, which is a clinical syndrome characterized by symptoms such respiratory distress, fever, etc. Respiratory samples were taken from southern India to perform diagnosis of influenza as the cause of respiratory syndrome using influenza A(H1N1pdm09 and H3N2) and B (Victoria and Yamagata) specific real-time PCR. We also compared an easy to use commercial kit with the established CDC (Centre for Disease Control, USA) RT-PCR-based assay to assess the efficacy of the commercial kit which does not require handling of multiple reagents thus preventing chances of cross-contamination in a routine diagnostic laboratory setting.
Material and Methods
Sample Collection and Transport
A total of 2154 acute phase throat/nasopharyngeal swab samples were collected from patients suspected with influenza virus infection as the cause of ILI- and/or SARI-like illness, presenting between 3 and 7 days of onset of fever (with case definition of sudden onset of fever > 38 °C, cough, or sore throat as per WHO) at the Government Medical College in Thiruvananthapuram, Kerala, India [5••]. Informed consent was taken from patients prior to enrollment. Samples were received at the laboratory from the hospital in viral transport medium (Himedia, India) at 4 °C and in standard triple packaging. All the samples were processed in a high-containment facility.
Samples from 2010 to 2012 were assayed using the CDC protocol during which the commercial kit was also simultaneously tested for sensitivity and specificity. Both influenza-positive and influenza-negative clinical samples pretested with CDC protocol were included.
Viral RNA Extraction
Viral RNA was extracted from throat and nasal secretions using the QIAamp Viral RNA Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions for use with the CDC-based protocol.
Spin Star nucleic acid kit (ADT Biotech, Malaysia) was used to extract total nucleic acid from the clinical samples for influenza typing using the commercial kit (Real Star Influenza RT-PCR Kit, Altona diagnostics GmbH, Germany). Viscous samples were pretreated with kit-provided mucolytic agent prior to extraction.
CDC-Based Assay
The assay employs a one-step approach for reverse transcribing the viral RNA and then utilizing a multiplex RT-PCR approach using a mixture of various primer-probe sets to detect various influenza subtypes. The primer-probe mixtures used were Inf A for universal detection of seasonal A influenza viruses A(H3N2), and primer/probe set specifically to detect highly virulent A(H1N1) pdm09. The fourth primer/probe set targeted the human RNase P gene and served as an internal control [25••].
The RT-PCR assays were performed using the AgPath-ID one-step RT-PCR kit (Applied Biosystems). Briefly, 5 μl of purified RNA was reverse transcribed and amplified in a 25 μl reaction mixture containing 12.5 μl of 2XRT-PCR buffer (Applied Biosystems), 1 μl of 25XRT-PCR enzyme mix (Applied Biosystems), 300 nM forward primer, 300 nM reverse primer, and 75 nM probe [26]. RT-PCR was performed in a ABI 7500 real-time PCR system (Applied Biosystems) and analyzed by SDS software v2.0.1 (Applied Biosystems). The thermal cycling conditions comprised a 10-min reverse transcription step at 45 °C followed by a 10-min initial PCR activation step at 95 °C, and 40 cycles of 95 °C for 15 s and 55 °C for 45 s each.
Commercial RT-PCR Kit
The RT-PCR assays for single tube detection and differentiation of seasonal human influenza A (A(H1N1)pdm09, A(H3N2)) and B (Victoria and Yamagata) strains from clinical samples was done using the Real Star Influenza RT-PCR Kit 3.0 (Altona diagnostics GmbH, Germany) following manufacturer’s instructions. Ten microliters of extracted eluent was used as a template for PCR amplification. All assays also detected the amplification of an internal control that did not interfere with target amplification, to check for possible PCR inhibitors. All RT-PCR reactions were performed on the Rotor gene 5 plex HRM real-time platform (QIAGEN, Germany) (Fig.1a, b, c, d).
Sensitivity and Specificity Testing of the Commercial Kit Compared to CDC Protocol
We compared the clinical sensitivity and specificity of the easy to use preassembled commercial kit as compared to those of the robust CDC protocol for detection of influenza virus using the RT-PCR assay. We randomly selected 10% of samples (from 2010 to 2012) diagnosed either positive or negative for influenza using the CDC protocol and retested the samples using the commercial kit and the CDC protocol simultaneously.
Data Analysis
Statistical analysis was carried out using Epitools (Epitools epidemiological calculators, Aus Vet Animal Health Services; http://epitoolsausvet.com.au) and concordance between the two tests was determined using concordance and kappa of agreement.
Results
Viral Surveillance
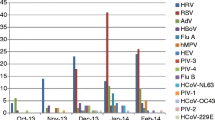
A total of 2154 throat and nasal swab samples were received with acute respiratory infections during October 2010 to December 2016. Using either of the real-time PCR-based methods, 43.2% were positive for influenza virus when tested. Among these, 136 (6.3%) samples were positive for influenza B (IFB), 576 (26.7%) samples were positive for A(H3N2), and 219 (10%) samples were positive for A(H1N1) pdm09. Among the positive cases, 652 (70%) were males and 279 (30%) were females. The clinical presentation in influenza-positive population included fever, cough, sore throat, nasal catarrh, headache, shortness of breath, and vomiting (Table 1).
The A(H3N2), IFB, and A(H1N1) pdm09 detection threshold cycle (Ct) mean and standard deviation values were 27.39 ± 3.61, 34.22 ± 2.67, and 28.87 ± 1.77, respectively (Table 2; Fig. 1a, b, c, d). Acceptable mean Ct (23.34 ± 0.45) was obtained from the kit internal controls of the two real-time PCR methods used in this study. Apparently, the viral load detected from throat/nasopharyngeal swab was clearly much lower for IFB compared to that for A(H3N2) and (A(H1N1) pdm09.
Analysis of the data revealed co-infections of A(H3N2) and A(H1N1) pdm09 in only eight cases (3.6%) and even lower co-infection of IFB with A(H1N1) pdm09 (3 cases; 1.3%). There were no observed co-infections for A(H3N2) and IFB (Table 1). This observation suggests that these pathogens probably do not co-infect frequently and are probably mutually exclusive of each other. Also, co-infections do not contribute significantly to the overall disease burden.
Comparison of the Two Real-time RT-PCR Assays
The concordance and kappa coefficients for detection of individual viral pathogens compared to those for CDC protocol were as follows: A(H3N2) 100.0%, 1.000; IFB 100.0%, 1.000; A(H1N1) pdm09 100.0%, 1.000, respectively (Table 3). Sensitivity and specificity of CDC protocol were overall 100.0 and 100%, respectively, and for individual pathogens were A(H3N2) 100.0, 100.0%; IFB 100.0, 100%; A(H1N1) pdm09 100.0, 100.0%, respectively, when commercial kit detection was considered as the gold standard (Table 3). On the other hand, sensitivity and specificity of commercial kit were overall 100 and 100.0%, respectively, when CDC protocol was taken as the gold standard. Individual pathogens then showed the following sensitivity and specificity, respectively: A(H3N2) 100.0, 100.0%; IFB 100%, 100.0%; A(H1N1) pdm09 100.0, 100.0%. Thus, the overall concordance between both the methods was 100% and kappa correlation was 1.000. Of note, we also observed that the Ct values for detection of each of the pathogens from the same samples done simultaneously by the two different methods were highly comparable, further reinstating that even at the quantitative levels, these two methods matched each other perfectly.
Discussion
In Asian countries, respiratory infections caused by the influenza virus have been generally ignored by the healthcare facilities. Information about influenza strains circulating in the Indian subcontinent stands majorly unknown due to lack of systemic studies. The available information on epidemiological and clinical features of influenza virus is entirely from research studies alone. Such studies are important to keep track of antigenic shifts in influenza virus which can lead to major endemics and economic losses [27, 28•]. With the rapid increase in number of influenza cases each year across the globe, and advent of more virulent strains and continued viral persistence in India and neighboring countries, there is an urgent need to systematically track the global dispersion of this virus in humans using molecular means.
A(H3N2) emerged as the etiology for the bulk of the cases positive for influenza. Contemporarily, this trend is also seen globally as reported from other studies reported from the USA and several other countries where A(H3N2) was detected in 26 to 30% of all influenza cases [29•].
Co-infections of different influenza viruses are rarely reported and reports focus solely on co-infections among different subtypes but not antigenic variants of the same subtype strains [30•, 31••, 32••]. The rate of co-infection determined in this study was 1.2% (n = 931) among all three tested pathogens. This rate is also agreeable with reported literature showing 0 to 3% co-infections among influenza subtypes [33•, 34•]. Although influenza co-infections are rare, we have shown that they occur during the first stage of a pandemic while seasonal strains co-circulated. This co-circulation poses a risk for further genetic reassortment in influenza strains, which could result in development and spread of new strains with pandemic potential.
Molecular methods, including one-step PCR and RT-PCR, have provided a convenient and sensitive approach for the diagnosis of influenza virus within a reasonable turnaround time [18••, 35••, 36••]. Our study reinstates the importance of clear diagnosis and overall surveillance of the highly adaptive and evolving influenza virus using molecular methods to aid healthcare providers to adapt to the ever-changing clinical manifestations of antigenically mutated strains.
The molecular testing of influenza patients helped the clinicians in timely diagnosis and treatment of these patients during the study. The RT-PCR test has higher sensitivity and specificity; hence, it is considered to be the gold standard for true estimation of disease burden, as compared to the any other commercial antigen-based or other tests. Therefore, continued surveillance of the circulating influenza viruses in India will help implement influenza control and also serve as cues for determining priority populations for possible vaccination.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
•• Neumann G, Noda T, Kawaoka Y. Emergence and pandemic potential of swine-origin H1N1 influenza virus. Nature. 2009;459(7249):931–9. Influenza viruses especially in the context of overcrowded regions like India where infectious diseases can spread like wildfire and management of the disease can be a nightmare. To understand the recent emergence of swine-origin H1N1 viruses, this publication was significantly useful to control these outbreaks and for real-time monitoring of the evolution of this virus.
•• Prachayangprecha S, Makkoch J, Suwannakarn K, Vichaiwattana P, Korkong S, Theamboonlers A, et al. Epidemiology of seasonal influenza in Bangkok between 2009 and 2012. J Infect Dev Ctries. 2013;7(10):734–40. A study showing a prevalence and seasonal pattern of influenza viruses between 2009 and 2012.
•• Wright PFNG, Kawaoka Y. Orthomyxoviruses. In: Knipe DMHP, editor. Fields virology. 5th ed. Philadelphia: Lippincott Williams & Wilkins; 2007. p. 1691–740. This book help us to understand the basic molecularbiology of virus and diagnostic methodos used for detection.
•• Ortiz JR, Sotomayor V, Uez OC, Oliva O, Bettels D, McCarron M, et al. Strategy to enhance influenza surveillance worldwide. Emerg Infect Dis. 2009;15(8):1271–8. This study describe a sentinel surveillance system that could enhance the quality of influenza epidemiologic and laboratory.
•• Rambaut A, Pybus OG, Nelson MI, Viboud C, Taubenberger JK, Holmes EC. The genomic and epidemiological dynamics of human influenza A virus. Nature. 2008;453(7195):615–9. A study suggest a sink-source model of viral ecology in which new lineages are seeded from a persistent influenza reservoir, which we hypothesize to be located in the tropics, to sink populations in temperate regions.
•• Chadha MS, Potdar VA, Saha S, Koul PA, Broor S, Dar L, et al. Dynamics of influenza seasonality at sub-regional levels in India and implications for vaccination timing. PLoS One. 2015;10(5):e0124122. This study reveals to understand regional differences in influenza seasonality at regional and sub-regional level, especially in countries with large latitude span.
•• Garten RJ, Davis CT, Russell CA, Shu B, Lindstrom S, Balish A, et al. Antigenic and genetic characteristics of swine-origin 2009 A(H1N1) influenza viruses circulating in humans. Science. 2009;325(5937):197–201. A study shows worldwide monitoring of the antigenic and genetic properties of the 2009 A(H1N1) viruses continues for, among other reasons, detecting any changes and thus any necessity for selecting further vaccine candidates or changes in antiviral recommendations.
•• Rao BL. Epidemiology and control of influenza. The National medical journal of India. 2003;16(3):143–9. This study describes there is a urgent need to expand influenza surveillance in developing coutries like india.
•• Koul PA, Mir MA, Bali NK, Chawla-Sarkar M, Sarkar M, Kaushik S, et al. Pandemic and seasonal influenza viruses among patients with acute respiratory illness in Kashmir (India). Influenza Other Respir Viruses. 2011;5(6):e521–7. A study reveals that the 2009A/H1N1 in Srinagar is genetically similar to globally circulating clade 7 strains, with unique signature sequences in the HA gene.
•• Hampson AW. Epidemiological data on influenza in Asian countries. Vaccine. 1999;17(Suppl 1):S19–23. A study shows Epidemiological data on pneumococcal infections in Asian countries.
•• Wang W, Ren P, Mardi S, Hou L, Tsai C, Chan KH, et al. Design of multiplexed detection assays for identification of avian influenza a virus subtypes pathogenic to humans by SmartCycler real-time reverse transcription-PCR. J Clin Microbiol. 2009;47(1):86–92. This study publication helped diagnosis using four real-time protocols developed to screen influenza viruses from nasopharyngeal swab specimens from children with acute respiratory infections. This study helped designing real-time protocols for remote hospitals in the field.
•• Ellis JS, Zambon MC. Molecular diagnosis of influenza. Rev Med Virol. 2002;12(6):375–89. This publication gave an idea on how to use molecular techniques to understand the epidemiology of influenza viruses.
Storch GA. Essentials of diagnostic virology. New York: Churchill Livingstone Inc.; 2000.
•• Eggers M, Roth B, Schweiger B, Schmid M, Gregersen JP, Enders M. Comparison of the novel ResPlex III assay and existing techniques for the detection and subtyping of influenza virus during the influenza season 2006-2007. Eur J Clin Microbiol Infect Dis. 2012;31(6):1257–65. A study shows a comparison of in-house real-time RT-PCR (RRT-PCR), and ResPlex III (Qiagen, Hilden, Germany). ResPlex IIIhad the highest sensitivity for detecting influenza virus in clinical specimens, followed by in-house RRT-PCR (96% compared with ResPlexIII). Conventional cell culture in MDCK cells, rapid culture, and quick test assays were substantially less sensitive (55%, 72%, and 39%, respectively).
• Espy MJ, Smith TF, Harmon MW, Kendal AP. Rapid detection of influenza virus by shell vial assay with monoclonal antibodies. J Clin Microbiol. 1986;24(4):677–9. A study shows that the shell vial assay using these recently developed monoclonal antibodies is readily adaptable to laboratories that wish to use immunofluorescence for the diagnosis of influenza virus infections.
•• Rawlinson WD, Waliuzzaman ZM, Fennell M, Appleman JR, Shimasaki CD, Carter IW. New point of care test is highly specific but less sensitive for influenza virus A and B in children and adults. J Med Virol. 2004;74(1):127–31. A study describes the modified point of care diagnostic test was assessed on 469 nasopharyngeal aspirates and 260 nose/throat swabs taken from children and adults. The test was specific (77-98%) for all specimen types for influenza virus A and B, depending upon incubation conditions.
• Stockton J, Ellis JS, Saville M, Clewley JP, Zambon MC. Multiplex PCR for typing and subtyping influenza and respiratory syncytial viruses. J Clin Microbiol. 1998;36(10):2990–5. A study shows a typing and subtyping of influenza A and B viruses and RSV types A and B, the multiplex RT-PCR gave an excellent (100%) correlation with the results of conventional typing and subtyping with specific antisera. Also be used to accurately detect more than one viral template in the same reaction mixture, allowing viral coinfections to be identified with the same respiratory specimen.
•• Mahony JB. Detection of respiratory viruses by molecular methods. Clin Microbiol Rev. 2008;21(4):716–47. A study review describes the molecular methods used to detect respiratory viruses and discusses the contribution that molecular testing, especially multiplex PCR, has made to our ability to detect respiratory viruses and to increase our understanding of the roles of various viral agents in acute respiratory disease.
•• Fan J, Cui D, Lau S, Xie G, Guo X, Zheng S, et al. Detection of a novel avian influenza A (H7N9) virus in humans by multiplex one-step real-time RT-PCR assay. BMC Infect Dis. 2014;14:541. In the present study, we have enlightened that the multiplex PCR is sensitive in detecting novel influenza strains.
• Kang X, Wu W, Zhang C, Liu L, Feng H, Xu L, et al. Detection of avian influenza A/H7N9/2013 virus by real-time reverse transcription-polymerase chain reaction. J Virol Methods. 2014;206:140–3. A study developed the sensitive and specific assay for detecting influenza A virus H7N9 and it could efficiently differentiate the H7N9 infection from other infection with Influenza A virus.
•• Templeton KE, Scheltinga SA, Beersma MF, Kroes AC, Claas EC. Rapid and sensitive method using multiplex real-time PCR for diagnosis of infections by influenza A and influenza B viruses, respiratory syncytial virus, and parainfluenza viruses 1, 2, 3, and 4. J Clin Microbiol. 2004;42(4):1564–9. This paper describes the application of real-time PCR to clinical samples increases the sensitivity for respiratory viral diagnosis and also would improve patient management and infection control.
•• Kuo RL, Yang SL, Liu YC, Chen LT, Mok CK, Kuo SM, et al. Influenza A/B virus detection and influenza A virus subtyping with emphasis on the novel H7N9 virus by using multiplex real-time RT-PCR. J Virol Methods. 2014;208:41–6. A study shows a multiplex or singular real time RT-PCR platforms were established for differentially detecting the recent highly pathogenic H7N9, 2009 pandemic H1N1, seasonal H3, and avian H5 genomes.The platforms effectively exclude the false positive cases from other respiratory infection agents. The sensitivity for detecting the H7 and N9 genes was 5 copies per reaction, and analyzing clinical specimens by using this platform required only 3 h.
•• Li Y, Wu T, Qi X, Ge Y, Guo X, Wu B, et al. Simultaneous detection of hemagglutinin and neuraminidase genes of novel influenza A (H7N9) by duplex real-time reverse transcription polymerase chain reaction. J Virol Methods. 2013;194(1–2):194–6. The paper reports that the assay is suitable for large-scale screening due to short turnaround times and high specificity, sensitivity, and reproducibility.
•• Wang D, Coscoy L, Zylberberg M, Avila PC, Boushey HA, Ganem D, et al. Microarray-based detection and genotyping of viral pathogens. Proc Natl Acad Sci U S A. 2002;99(24):15687–92. This paper describes micro array based method is versatile and greatly expands the spectrum of detectable viruses in a single assay while simultaneously providing the capability to discriminate among viral subtypes.
•• Liao RS, Landt O, Hill JT. Comparison of a laboratory-developed RT-PCR and the CDC RT-PCR protocol with rapid immunodiagnostic testing during the 2009 H1N1 influenza A pandemic. Diagn Microbiol Infect Dis. 2011;70(2):236–9. This paper is our source to design a work plan to compare two different protocols and do statstically significant analysis.
•• Whiley DM, Bialasiewicz S, Bletchly C, Faux CE, Harrower B, Gould AR, et al. Detection of novel influenza A(H1N1) virus by real-time RT-PCR. J Clin Virol. 2009;45(3):203–4. The paper describes the results showed that the RT-PCR methods were suitable for sensitive and specific detection of novel influenza A(H1N1) RNA in human samples.
York I, Donis RO. The 2009 pandemic influenza virus: where did it come from, where is it now, and where is it going? Curr Top Microbiol Immunol. 2013;370:241–57.
• Centers for Disease C, Prevention. Influenza activity—United States, 2012–13 season and composition of the 2013–14 influenza vaccine. MMWR Morb Mortal Wkly Rep. 2013;62(23):473–9. This report summarizes influenza activity in the United States during the 2012-13 influenza season and reports the recommendations for the components of the 2013-14 Northern Hemisphere influenza vaccine.
• Toda S, Okamoto R, Nishida T, Nakao T, Yoshikawa M, Suzuki E, et al. Isolation of influenza A/H3 and B viruses from an influenza patient: confirmation of co-infection by two influenza viruses. Jpn J Infect Dis. 2006;59(2):142–3. This study shows a influenza A/H3 and B viruses from an influenza patient: confirmation of co-infection by two influenza viruses in 2004-2005 epidemic season, Japan.
• Falchi A, Arena C, Andreoletti L, Jacques J, Leveque N, Blanchon T, et al. Dual infections by influenza A/H3N2 and B viruses and by influenza A/H3N2 and A/H1N1 viruses during winter 2007, Corsica Island, France. J Clin Virol. 2008;41(2):148–51. A study describes the Influenza viruses were detected in 93 (69.4%) of 134 patients with influenza-like illness using the combination of classical and molecular assays. Dual respiratory infections by influenza viruses were detected in 3.2%.
•• Eshaghi A, Blair J, Burton L, Choi KW, De Lima C, Duncan C, et al. Characterization of an influenza A and influenza B co-infection of a patient in a long-term care facility with co-circulating influenza a and influenza B. Int J Infect Dis. 2009;13(3):e127–8. Present study clearly stated that molecular techniques are powerful tools that enable laboratories to extensively characterize circulating strains of influenza, including co-infections, without the use of culture-based methods. Our study also support this data.
•• Chidlow G, Harnett G, Williams S, Levy A, Speers D, Smith DW. Duplex real-time reverse transcriptase PCR assays for rapid detection and identification of pandemic (H1N1) 2009 and seasonal influenza A/H1, A/H3, and B viruses. J Clin Microbiol. 2010;48(3):862–6. A study describes a total 11,000 clinical samples were then tested for influenza A and B matrix gene targets and specific hemagglutinin gene targets for seasonal influenza A/H1, A/H3, and pandemic A (H1N1) 2009. Minimum sensitivities and specificities were 98.8% and 100%, respectively, for pandemic (H1N1) 2009, 81.5% and 98.9% for seasonal A/H1, and 96.3% and 99.6% for A/H3.
• Palacios G, Hornig M, Cisterna D, Savji N, Bussetti AV, Kapoor V, et al. Streptococcus pneumoniae coinfection is correlated with the severity of H1N1 pandemic influenza. PLoS One. 2009;4(12):e8540. A study shows the association of Streptococcus. pneumoniae with morbidity and mortality is established in the current and previous influenza pandemics. However, this study is the first to demonstrate the prognostic significance of non-invasive antemortem diagnosis of S. pneumoniae infection and may provide insights into clinical management.
• Ghedin E, Fitch A, Boyne A, Griesemer S, DePasse J, Bera J, et al. Mixed infection and the genesis of influenza virus diversity. J Virol. 2009;83(17):8832–41. A study reveal that individuals can harbor influenza viruses that differ in major phenotypic properties, including those that are antigenically distinct and those that differ in their sensitivity to antiviral agents.
•• Nguyen Van JC, Camelena F, Dahoun M, Pilmis B, Mizrahi A, Lourtet J, et al. Prospective evaluation of the Alere i influenza a&B nucleic acid amplification versus Xpert Flu/RSV. Diagn Microbiol Infect Dis. 2016;85(1):19–22. This study indicate that the Alere i Influenza A&B assay has a good overall analytical performance and a high degree of concordance with the PCR-based Xpert Flu/RSV assay. The Alere i Influenza A&B isothermal nucleic acid amplification test is a powerful tool for influenza detection due to its high sensitivity and specificity as well as its ability to generate results within 15min.
•• Chiarella FC, Culebras E, Fuentes-Ferrer ME, Picazo JJ. Evaluation of the Alere i influenza A&B assay for rapid identification of influenza A and influenza B viruses. J Med Microbiol. 2016;65(6):456–61. This technology is very much useful in remote limited settings (RLS) to detect influenza A and B. Isothermal nucleic acid amplification test is a powerful tool for influenza detection due to its high sensitivity and specificity as well as its ability to generate results within 15 min. Our study’s turnaround time, sensitivity, and specificity were comparable.–61.
Acknowledgments
The authors are thankful to the Indian Council for Medical Research (ICMR), Government of India for providing necessary infrastructure and facilities at the Viral Disease Biology Program Network Grade 1 Lab at RGCB for carrying out this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Seetha Dayaker, Heera R. Pillai, Vineetha P. Thulasi, Devakikutty Jayalekshmi, and Radhakrishnan R. Nair declare that they have no conflict of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors. Informed consent was taken from patients prior to enrollment.
Additional information
This article is part of the Topical Collection on Respiratory Infections
Rights and permissions
About this article
Cite this article
Dayakar, S., Pillai, H.R., Thulasi, V.P. et al. Comparative Study of Molecular Approaches for the Detection of Influenza Virus from Patient Samples Using Real-time PCR: Prospective Disease Burden Study in Kerala (India) from 2010 to 2016. Curr Infect Dis Rep 20, 24 (2018). https://doi.org/10.1007/s11908-018-0632-y
Published:
DOI: https://doi.org/10.1007/s11908-018-0632-y