Abstract
Purpose of Review
This review aims to give an update on recent findings related to the cardiac splicing factor RNA-binding motif protein 20 (RBM20) and RBM20 cardiomyopathy, a form of dilated cardiomyopathy caused by mutations in RBM20.
Recent Findings
While most research on RBM20 splicing targets has focused on titin (TTN), multiple studies over the last years have shown that other splicing targets of RBM20 including Ca2+/calmodulin-dependent kinase IIδ (CAMK2D) might be critically involved in the development of RBM20 cardiomyopathy. In this regard, loss of RBM20 causes an abnormal intracellular calcium handling, which may relate to the arrhythmogenic presentation of RBM20 cardiomyopathy. In addition, RBM20 presents clinically in a highly gender-specific manner, with male patients suffering from an earlier disease onset and a more severe disease progression.
Summary
Further research on RBM20, and treatment of RBM20 cardiomyopathy, will need to consider both the multitude and relative contribution of the different splicing targets and related pathways, as well as gender differences.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Dilated cardiomyopathy (DCM), as defined by left ventricular or biventricular systolic dysfunction and dilatation that are not explained by abnormal loading conditions or coronary artery disease, is a leading cause of death worldwide, and one of the main reasons for heart transplantation [1]. Its prevalence has long been thought to be around 1:2500 based on a study in Olmsted county (USA) published in 1989, but newer studies estimate it up to 1:250 [2, 3]. The European Society of Cardiology divides DCM into familial (genetic) and nonfamilial (non-genetic) forms, with 30–50% of cases being familial in nature [4, 5]. In contrast to hypertrophic cardiomyopathy (HCM) and arrhythmogenic right ventricular cardiomyopathy (ARVC), where a small number of genes account for most of the genetic causes, DCM-causing mutations have been observed in a variety of genes of diverse ontology [2]. One of these genes is RBM20, of which mutations have been observed in 2–3% of familial DCM cases, and which interestingly has been shown to account for a number of DCM cases for which previously no genetic cause could be established [5,6,7,8]. The RBM20 gene encodes RNA binding motif protein 20, a trans-acting splicing factor [7]. RBM20 is highly expressed in striated muscle, especially the heart, and has been shown to affect splicing of a variety of genes important for cardiac function [7, 9, 10]. RBM20 cardiomyopathy manifests as a highly penetrant and aggressive disease which is correlated with high rates of heart failure, arrhythmias, and sudden cardiac death [7, 11•, 12, 13]. The most frequently investigated splicing target of RBM20 is TTN, which encodes the giant sarcomeric protein titin. It is important to note that mutations in the TTN gene itself are also frequent causes of DCM, accounting for ~ 20% of familial and ~ 15% of sporadic DCM cases [5, 14]. In 2012, Guo and colleagues described the first Rbm20-deficient animal model, namely a rat model with an unusually large titin protein, which they found to be due to a naturally occurring deletion in the Rbm20 gene [9]. Therefore, because TTN was among the first genes shown to be regulated by RBM20, its connection to DCM, and its importance in cardiac biology, the research focus on RBM20 cardiomyopathy mostly lay on TTN. However, recent findings suggest that RBM20 cardiomyopathy cannot solely be explained by changes in TTN splicing, and that missplicing of additional RBM20 target genes is likely to contribute to the phenotype [11•]. Currently, over 30 genes have been shown to be a target of RBM20 (Table 1). Among them are further genes important for sarcomeric function like LDB3 and TNNT2, as well as many genes crucial for calcium handling in cardiomyocytes. One example is the calcium/calmodulin-dependent protein kinase II (CaMKII), which modulates excitation-contraction coupling by phosphorylating targets like the ryanodine receptor (RyR), phospholamban (PLN), and the L-type Ca2+ channel, some of which are also targets of RBM20 [15]. Rbm20 deficiency has been shown to lead to altered cellular calcium handling, signified by a cellular calcium overload [11•]. This may be caused by the altered splicing of Camk2d towards the CaMKIIδA and CaMKIIδ9 isoforms, as other studies showed that an increase in CaMKIIδA is sufficient to cause similar calcium handling defects in mice [16]. Remarkably, in these models, gender differences were observed, with male mice being more affected [16]. Likewise, male patients with RBM20 cardiomyopathy also show a more severe disease phenotype [17•]. Taken together, these findings imply that misspliced CaMKIIδ might critically contribute to the RBM20 cardiomyopathy phenotype, which would explain pathological calcium handling, arrhythmias, and sudden cardiac death, and might also be responsible for the observed gender differences. In this review, we aim to summarize these recent insights in RBM20 cardiomyopathy, and provide possible new translational approaches based on these new insights.
Properties of RBM20
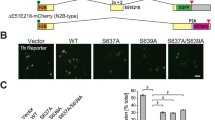
The RBM20 gene is located on chromosome 10, contains 14 exons, and encodes a protein of 1227 amino acids. The protein contains two zinc finger domains, an RNA recognition motif (RRM), a serine- and arginine-rich region (RS region), a leucine-rich region, and a glutamate-rich region (Fig. 1) [18]. These regions, especially the RS region, are highly conserved among species and mutations in these domains often cause a loss of RBM20 function [7, 12, 19]. RBM20 is predominantly located in the nucleus, and its nuclear localization is required for its function [10, 20]. The most frequent disease-causing RBM20 mutations occur in the RS region [19]. The serine and arginine residues in the RS region (RSRSP stretch) can be phosphorylated, which is necessary for the nuclear localization of Rbm20 [20]. The loss of phosphorylation in these mutated residues translocates RBM20 out of the nucleus, and this likely negates RBM20’s function [20]. The RS region also mediates protein-protein interactions with other splicing factors such as U2AF65 and U2AF35, potentially explaining the critical function of this domain for splicing activity [21]. The RS region, however, might not be the only region of RBM20 important for nuclear localization, as a truncated RBM20 peptide (including the RRM but not the RS region) is sufficient for nuclear localization [10]. This suggests that not the presence, but the phosphorylation status, of the RS domain determines nuclear localization. It would be interesting to see if a truncated peptide that includes the RRM, but also a mutated (and phospho-resistant) RS domain would localize to the cytoplasm.
Schematic visualization of RBM20 protein structure, including functional domains. Known mutations shown above the corresponding domains. The RRM and RS region, especially the RSRSP stretch from amino acids 634–638 (marked in orange), are both important for RBM20’s nuclear localization. The RRM is required to bind to the “UCUU” RNA sequence found on targets of RBM20. RBM20’s regulatory function is likely achieved by the combined function of its functional domains. (ZnF, zinc finger domain; RRM, RNA recognition motif; RS region, serine-/arginine-rich region) (A single asterisk indicates a nonsense mutation) [6, 7, 12, 13, 17, 23, 68, 102,103,104,105,106,107,108,109,110,111]
Of a set of RBM20 mutants selectively lacking one of the zinc finger domains, the leucine-rich region, the RRM or the glutamate-rich region, only the loss of the glutamate-rich region led to a significant decrease in splicing activity [20]. Truncating RBM20 on the N-terminal part of the RS region, and thereby losing the glutamate-rich region, completely stopped splicing activity, although this cannot conclusively be explained by the loss of the glutamate-rich region [22]. Mutations of the glutamate-rich region, while not as frequent as mutations in the RS region, are also known to be associated with RBM20 cardiomyopathy [6, 23]. In one case, a mutation at amino acid 913 exchanging glutamate for lysine leads to a substantial reduction in RBM20 protein amount, which is hypothesized to be caused by misfolding of RBM20 due to the mutation [23]. It might be the case that the glutamate-rich region is more resistant to mutations affecting single amino acids compared with the RS region, as there is perhaps a greater degree of redundancy in the glutamate-rich region. The RRM also has functional relevance, as truncated RBM20 peptides without the RRM and the C-terminus of the protein lost all splicing function [22]. Selectively removing only the RRM however did not significantly reduce splicing function of RBM20, indicating that different regions of RBM20 affect its regulatory function [20]. Taken together, these data suggest that the different domains of Rbm20 have different functions, and that proper function of Rbm20 is an orchestrated process of most, if not all, of these domains. RBM20’s RRM binds to an RNA recognition element defined by a UCUU core, which most commonly is located up to 50 nucleotides upstream or 100 nucleotides downstream in the introns flanking an alternatively spliced exon [21, 24]. In the TTN pre-mRNA, RBM20 represses splicing of large stretches of exons, prohibiting the removal of flanking introns [25]. This allows alternative splice sites at the 3′ or 5′ end of RBM20-repressed regions to splice together, which can skip large numbers of exons and create different splice products depending on which alternate splice sites splice together [22, 25]. RBM20 also acts as a splicing repressor for many of its other splicing targets [21]. Interestingly, the interaction between RBM20 and TTN also affects the regulation of other RBM20 targets. In the nucleus, RBM20 is mainly localized in two speckles, which are the two transcriptional sites of the TTN gene [25]. During the transcriptional process, RBM20 colocalizes with the TTN pre-mRNA, which is simultaneously spliced [25, 26]. These clusters of TTN pre-mRNA and RBM20 are able to spatially attract the gene loci of other RBM20 targets like CACNA1C and CAMK2D [27•]. Remarkably, this colocalization leads to an increased splicing activity of RBM20 on these targets, and loss of TTN pre-mRNA leads to similar splicing changes in CACNA1C and CAMK2D as loss of RBM20 itself [27•]. These data suggest that the TTN pre-mRNA creates a RBM20 splicing factory, which is necessary for correct splicing of other RBM20 targets [27•]. RBM20 seems also able to generate circular RNA transcripts (circRNAs) from TTN and several of its other targets [28, 29]. In the case of TTN, RBM20-dependent exon skipping provides the substrate for back-splicing these skipped exons together to produce circRNAs. However, this accounts only for a small amount of cardiac circRNAs, while the vast majority is created in non-RBM20-dependent manner from constitutively spliced exons, competitively with the generation of the linear transcript [28]. Interestingly, RBM20 not only affects splicing but can also affect transcription of its targets, which might be related to its ability to generate circRNAs [30].
Upstream Regulators of RBM20
In mice, Rbm20 is induced during early cardiac development (Carnegie stage E8.5), and loss of Rbm20 leads to a dysregulation of cardiogenic genes [31]. The main pathway implicated in Rbm20 regulation so far is the PI3K-Akt-mTOR axis, which was found to upregulate Rbm20 expression [32, 33]. Activation of PI3K (phosphoinositide 3-kinase) causes Akt (also known as PKB; protein kinase B) to become phosphorylated, which in turn activates mTORC1. mTORC1 phosphorylates S6KI and 4E-BP1, which promote expression of Rbm20 [32]. The PI3K-Akt-mTOR axis is activated by exercise and various growth factors, including insulin, insulin-like growth factor 1, and neuregulin, and its activation is cardioprotective in DCM [36, 37]. Akt especially has been shown to regulate other SR proteins in a growth factor–dependent manner [38]. From these growth factors, currently only insulin and the thyroid hormone T3 have been shown to have an effect on Rbm20, both leading to an upregulation of Rbm20 expression [32, 33]. The connection to the thyroid hormones is interesting, since hypothyroidism is known to have adverse effects on heart function and correlates with disease severity in heart failure patients [39, 40]. In that regard, hypothyroidism could contribute to disease severity by regulating RBM20 expression and missplicing of downstream splicing targets, as it is known that RBM20 levels correlate with the amount of missplicing of its targets [21]. Another pathway connected to Rbm20 is the mitogen-activated protein kinase (MAPK) pathway. Like the PI3K axis, it can be activated by growth factors like epidermal growth factor [41]. Angiotensin II has also been shown to activate this pathway, triggering its downstream target ETS transcription factor (ELK1), which binds to the Rbm20 promotor and upregulates Rbm20 expression [42]. The methylation status of Rbm20 also affects its expression. In a study of rats treated with doxorubicin, which decreased the global amount of methylated DNA and specifically altered methylation in 7 different Rbm20 nucleotide residues, Rbm20 expression was increased almost twofold [43]. Interestingly, RBM20 methylation correlates with fasting insulin levels, although it is unknown if there is a functional relevance to this [44]. RBM20 expression was also found to be positively correlated with BMI, and together these findings suggest an important influence of the body’s metabolic state on RBM20 [45].
Posttranslational Regulation of RBM20
Little is known about posttranslational regulation of RBM20, but it is likely that RBM20 is additionally regulated by posttranslational modifications (PTMs) such as phosphorylation, as other SR proteins have been shown to alter their localization, conformation, and splicing function if phosphorylated, and the phosphorylation of RBM20’s RS region has already been shown to be functionally relevant for splicing regulation [20, 34, 35]. Another PTM worth investigating may be ubiquitination of RBM20. In a study of a family with RBM20 cardiomyopathy caused by an E913K mutation in RBM20, protein levels of RBM20 were strongly reduced, while mRNA levels and stability were not, which may suggest that the new lysine functions as a substrate for ubiquitination, and leads to degradation by the proteasomal complex [23]. Further identification of PTMs of RBM20, and what enzymes/kinases are involved, will be valuable to unravel the mechanism(s) through which RBM20 functions, and could underlie new therapeutic approaches. One possible approach here would be the use of in silico predictions of the interactions of potential kinases with both wild-type and mutated forms of RBM20. Furthermore, RBM20-dependent splicing is regulated by polypyrimidine tract-binding protein 1 (PTBP1), also known as PTB4 or hnRNPI. PTBP1 belongs to a larger family of genes regulating mRNA and can also alter splicing [46,47,48]. The RRM of PTBP1 shares similarities in structure with RBM20’s RRM, and PTBP1 is able to bind simultaneously with RBM20 to the UCUU-binding motif shared by RBM20’s targets [22, 24]. Binding of PTBP1 counteracts RBM20’s effect on TTN splicing in a manner dependent on the ratio of PTBP1 to RBM20 [22]. Antagonistic mRNA regulation is also seen in other RNA-binding proteins; for example, the CELF and MNBL proteins regulate over 200 exons in an antagonistic manner [49]. However, PTBP1 and RBM20 do not only oppose each other, as splicing of formin homology 2 domain-containing 3 (FHOD3), a protein relevant for the assembly of the sarcomere, was changed towards an increase of FHOD3 isoform F both by addition of RBM20 and PTBP1 in a dose-dependent manner [50]. Examining the effect of PTBP1 on other known RBM20 targets and vice versa might lead to a better understanding of the interactions between these two factors.
RBM20 Regulates Splicing of Sarcomeric Genes Including TTN
Many of RBM20’s known targets can be categorized as either sarcomeric, like TTN, LDB3, TNNT2 (cardiac muscle troponin T), MYH6, and MYH7 (myosin heavy chain alpha and beta isoform), or related to cellular ion handling, like CAMK2D, RYR2, SLC8A1 (the sodium-calcium exchanger; also NCX), and CACNA1C (encoding the alpha 1C sub-unit of the L-type calcium channel; also CaV1.2). The most frequently studied splicing target of RBM20 is TTN, but it should be noted that evidence that TTN is the only critical target of RBM20 is currently lacking. The TTN gene contains 363 exons, and its translational product (titin, also known as connectin) is the largest human protein, with a size of over 3 MDa. Titin connects the sarcomeric Z-disk and thick filaments in striated muscle, where it is among the most common proteins [51, 52]. It functions as a molecular spring, and together with collagen, titin is the main determinant of passive tension (as measured in demembranated cellular systems) in the sarcomere and thereby the muscle [51]. Cardiac titin mainly occurs in two isoforms which are co-expressed [53]. The smaller isoform is named N2B titin after the N2B element, an additional elastic segment found only in cardiac titin [54, 55]. The larger N2BA isoform additionally contains an elastic N2A element, as well as multiple elastic Ig-like and PEVK domains [54, 55]. This increase in size and the number of elastic elements in the N2BA isoform reduces the stiffness of this isoform compared to N2B titin [54, 55]. The ratio of these isoforms in a myocyte defines the cellular stiffness, with larger amounts of the larger N2BA isoform leading to myocytes with a reduced stiffness [55, 56]. Functionally, this results in an increased whole-heart compliance, as the cardiomyocytes react less rigidly to the force applied by the blood flow [55, 56]. In DCM and ischemic heart disease, the N2BA:N2B ratio is increased; however, it is unclear if this is a causative or compensatory effect [57,58,59]. A high percentage of DCM cases are caused by mutations leading to a truncated form of titin [5, 14]. However, in comparison to other forms of DCM, TTN cardiomyopathy represents a mild entity of DCM that becomes symptomatic at a higher age, has a better treatability, and relatively benign outcomes [60]. TTN cardiomyopathy is also diagnosed at a higher age when compared directly to RBM20 cardiomyopathy [61]. The loss of RBM20 function does not lead to truncation of the titin protein. Instead, the average titin protein becomes even larger—the amount of the smaller N2B isoform is reduced, while the larger N2BA isoform is upregulated and another, giant and even more compliant 3.9 MDa titin isoform called N2BA-G appears, which reduces the myocardial stiffness (as measured in intact cardiomyocytes/myocardial tissue) and increases resting sarcomere length [9, 23, 62,63,64]. Functionally, compliance of the heart increases, and passive tension of the sarcomere is reduced [23, 63]. Additionally, the Frank-Starling mechanism is impaired due to a diminished length-dependent activation (a reduction of the force generated by an increase in muscle length), which was reduced in both heterozygous and homozygous Rbm20-deficient mice [63, 65]. This results in a reduced fractional shortening (FS), and an even further reduced FS when the afterload is increased (for example through application of noradrenergic stress) [63]. Furthermore, the length-dependent Ca2+ sensitivity of the sarcomere is also affected by loss of RBM20 [23, 65]. A study in human RBM20-deficient cardiomyocytes found an increase in Ca2+ sensitivity, which could be counteracted by application of protein kinase A (PKA), while control cardiomyocytes showed no change after PKA application [23]. It is also known from other forms of heart failure that a reduction of PKA-mediated phosphorylation of sarcomeric proteins caused by a reduction of β-adrenergic signaling leads to an increase in Ca2+ sensitivity [112]. However, this is notably only found in human tissues, as in animal models the opposite (i.e., a reduction of Ca2+ sensitivity) is observed [65]. Some sarcomeric proteins are also phosphorylated by targets of RBM20; for example, CaMKII-dependent phosphorylation of titin is able to reduce the passive tension of the sarcomere [66]. The interplay of titin stiffness and calcium sensitivity also affects cross-bridge cycling, as loss of RBM20 causes a slowed cross-bridge detachment rate [65, 67]. Interestingly, a slowdown of cross-bridge detachment normally leads to an increased Ca2+ sensitivity, while the opposite is the case when RBM20 is lost [65, 67]. This may be explained by the altered titin, as titin plays a large part in modulation of the sarcomeric complex and interacts both with the thin and thick filament [65, 114]. Interestingly, heightened levels of collagen are observed in the RBM20-deficient heart, which may be a compensatory effect working against the reduced titin stiffness [63]. Together, these physiological changes likely contribute to the disease presentation of RBM20 cardiomyopathy. However, RBM20 cardiomyopathy shows a higher penetrance when compared with dilated cardiomyopathy in general, and with TTN cardiomyopathy specifically, with over 70% having a familial history of DCM and over 50% having a familial history of sudden cardiac death (SCD) [61, 11•]. Strikingly, RBM20 cardiomyopathy presents with a high prevalence of arrhythmias, especially sustained ventricular arrhythmia (VA) and tachycardia (VT), but also non-sustained VT and SCD, as well as high rates of atrial fibrillation [61, 11•]. This is in stark contrast to TTN cardiomyopathy [11•, 60]. Consequently, over 60% of RBM20 cardiomyopathy patients require implantation of an internal cardioverter defibrillator (ICD) compared with 27% of general DCM patients and < 10% of TTN cardiomyopathy patients [11•, 5]. RBM20 cardiomyopathy patients are also more likely to need a heart transplant compared to other DCM patients, and need it at a significantly younger age, with a mean of 28.5 years [8]. Together, these data suggest that the RBM20 cardiomyopathy phenotype is more severe and arrhythmogenic, and cannot be solely explained by changes in titin.
RBM20 Regulates Splicing of Ion- and Calcium-handling Genes Including CAMK2D
Several studies have now investigated if and how loss of RBM20 can impact calcium handling. In a stem cell RBM20 knockdown model, the peak amplitude of calcium transients highly increased and the area under the curve (AUC) more than doubled, while the beating frequency went down significantly [31]. Two models of patient-derived pluripotent stem cell cardiomyocytes with mutations in the RS domain (S635A and R636S, respectively) showed similarly altered calcium handling [68,69,70]. Interestingly, application of norepinephrine stress increased these differences in peak amplitude and AUC, and additional application of either the calcium channel antagonist verapamil or the beta-blocker carvedilol could return these values to those of WT controls [70]. In a RBM20–KO mouse model, isolated cardiomyocytes showed an altered L-type calcium current (ICa,L) density, which increased in amplitude almost twofold [11•]. In addition, these cardiomyocytes showed an intracellular calcium overload, signified by heightened diastolic Ca2+, heightened peak transient, and heightened SR calcium load, and they presented with an increased amount of pro-arrhythmic spontaneous calcium releases from the SR [11•]. The addition of norepinephrine increased the amount of spontaneous calcium releases, but they could be reduced to WT levels through treatment with verapamil [11•]. Verapamil is an inhibitor of the L-type calcium channel, so it seems likely that the increased L-type calcium current density is causative for the intracellular calcium overload and spontaneous calcium releases. The RBM20–KO mouse model also shows splicing differences in various ion handling genes, and it has been hypothesized that (some of) these underlie the calcium handling abnormalities [11•]. CaMKIIδ, for example, undergoes an almost complete switch from the normal δB and δC isoforms towards the δA and δ9 isoforms [11•]. CaMKIIδA was first described as a neurological isoform, but is also expressed during cardiac development up to 30 days after birth [16, 71]. Interestingly, its downregulation during cardiac development is dependent on RBM20 [31]. It is locally associated with the T-tubules and the intercalated discs, which are also the locations where calcium channels are highly concentrated [16]. In addition, a transgenic mouse model overexpressing specifically CaMKIIδA showed a similar phenotype to the one observed in RBM20 loss-of-function models, with increased SR calcium load and increased peak calcium transients [16]. Therefore, it has been hypothesized that CaMKIIδA could interact with the LTCC, and increase L-type calcium current [16, 11•]. CaMKIIδ9 is currently not well studied, but was recently found to be among the most abundant isoforms in the heart [72•]. Cardiac stress upregulates CaMKIIδ9 expression and hyper-activates it through phosphorylation and oxidation [72•]. CaMKIIδ9 promotes cardiomyopathy and cell death by phosphorylation of ubiquitin-conjugating enzyme E2T (UBE2T), an enzyme in the Fanconi anemia DNA repair pathway, which leads to degradation of UBE2T and thereby accumulation of DNA damage [72•]. Together, these results show that RBM20 cardiomyopathy is accompanied with an altered cellular calcium handling, which could be the reason for the heightened number of arrhythmias observed in patients. A possible mechanism for this could be increased activation of the LTCC through increased levels of CaMKIIδA, leading to a higher ICa,L and a cellular calcium overload; however, further research is required to validate this.
Sex and Gender Differences in RBM20 Cardiomyopathy
Sex and gender differences in cardiac disease have gained attention over the last years. Different prevalence of risk factors, socioeconomic factors leading to differing utilization of health resources, and genetic differences between men and women lead to pronounced differences both in prevalence and outcome of many cardiac diseases [73]. However, much of our knowledge on cardiac disease is still based on studies done in males, and even today much experimental work in animal studies neglects females. The prevalence of dilated cardiomyopathy is higher in men, with both historic and current studies showing a ratio of 25–30% of women, compared with 70–75% men [3, 74,75,76,77]. Women are older at diagnosis by 1–3 years compared with men, and are more likely to present with heart failure and left bundle branch block [74,75,76]. However, outcomes are better for women, as both all-cause and cardiovascular mortality, as well as rates of heart transplants, malignant ventricular arrhythmias (MVAs), and sudden cardiac death are reduced in women compared to men [74,75,76]. Right ventricular function is also better in women compared to men [78]. However, as DCM is a heterogenous disease caused by a multitude of genetic defects and other causes, these results must not necessarily hold true for all sub-forms of DCM. In DCM caused by TTN truncating variants, men have a significantly worse prognosis [79]. In lamin A/C (LMNA) cardiomyopathy, one of the most arrhythmogenic forms of DCM and thereby comparable to RBM20 cardiomyopathy, end-stage heart failure, rates of MVAs, and mortality are increased in men [80]. RBM20 cardiomyopathy also presents in a more severe fashion in male patients: In a patient collective of 80 RBM20 mutation carriers, men were both younger at diagnosis and presented with lower ejection fraction (“age, 29 ± 11 versus 48 ± 12 years; P < 0.01; ejection fraction, 29 ± 13% versus 38 ± 9%; P < 0.01”) [17•]. One-third of male patients required a heart transplant, and at young ages (33 ± 16 years old), while none of the female patients required a heart transplant [17•]. A total event-free survival of male patients was also significantly reduced compared with female patients [17•]. Interestingly, male RBM20 cardiomyopathy patients also fared worse when compared to a cohort of patients with DCM of unknown genetic cause, while female RBM20 cardiomyopathy patients had better event-free survival when compared to patients with DCM of unknown cause, suggesting that these sex and gender differences are not shared by all forms of DCM, but rather (at least to some extent) specific to RBM20 cardiomyopathy [17•]. The cause of these sex and gender differences is so far unknown. Behavior-related risk factors differ in prevalence between the genders; for example, smoking is more common in men, while obesity has a higher prevalence in women [81, 82]. However, sex is likely also an independent risk factor. In fact, many cardiac genes show sexual dimorphism in their expression, which could change their interactions with RBM20’s splice targets and thereby affect heart function [83]. There are also large differences in gene expression between pre- and post-menopausal women, which might result in a loss of cardioprotective effects that could relate to the late onset of RBM20 cardiomyopathy in females [83]. Animal models of RBM20 cardiomyopathy may be useful to better understand these sex-specific genetic differences. One promising line of research is the altered calcium handling in RBM20 cardiomyopathy patients. The different prevalence of arrhythmias seems to be a major part of the differing outcomes, and other arrhythmogenic forms of DCM show similar differences [17•, 80]. Interestingly, CaMKIIδ might play a major part in this: As previously described, CaMKIIδ performs an isoform switch towards the CaMKIIδA and CaMKIIδ9 isoforms upon loss of RBM20 [11•]. Transgenic overexpression of CaMKIIδA alone is sufficient to induce a phenotype similar to RBM20 cardiomyopathy, and strikingly this largely affects male mice, leading to ~ 50% lethality at 8 weeks of age, while females are unaffected [16]. CaMKIIδ has already been shown to be sex-specifically regulated in cardiac disease: In ischemia-reperfusion experiments, female hearts showed higher generation of activated phospho-CaMKIIδB(Thr287) and higher phosphorylation of its downstream target PLN, which was associated with arrhythmia suppression [84]. This might be due to differing expression of CaMK phosphatase (CaMKP), as CaMKP expression was increased in males and decreased in females after TAC surgery [85]. This cardioprotective effect of CaMKIIδB is unlikely to be the reason for gender differences in RBM20 cardiomyopathy, where CaMKIIδB is downregulated and the detrimental CaMKIIδA is much more common, but it shows that sex can affect the activity of specific CaMKIIδ isoforms [11•, 16]. It remains to be seen if CaMKIIδA and CaMKIIδ9 might also be differently activated. There is also evidence that calcium handling and excitation-contraction coupling in general are affected in a sex-dependent way. For example, female cardiomyocytes show lower calcium transients [86]. This may be caused by sex hormones, as testosterone can increase intracellular calcium and has been shown to be correlated with worse outcomes in other arrhythmogenic cardiac disease, while higher estrogen levels correlate with better outcomes [87, 88]. Likely, the observed gender differences are caused by a mixture of multiple genetic and non-genetic causes. Further research might not only advance treatment of RBM20 cardiomyopathy itself but also increase understanding of sex and gender differences in other forms of cardiac disease.
RBM20 as a Therapeutic Target
Heart failure with preserved ejection fraction (HFpEF) is a highly relevant disease with a prevalence in patients aged 60 years or older of about 5% [89]. However, compared to heart failure with reduced ejection fraction (HFrEF), few effective therapeutic options exist [89, 90]. HFpEF is accompanied by changes in both titin and the extra-cellular matrix (ECM), which together contribute to an increase in myocardial stiffness [91]. Titin is differently phosphorylated at multiple sites: The S11878(S26) PKC/CaMKII site is hyperphosphorylated, while the S4185(S469) PKA/PKG site is hypophosphorylated, which together leads to a reduction of titin compliance, although the N2BA/N2B ratio does not change [91]. Additionally, the total amount of collagen in the heart is highly increased, which also increases myocardial stiffness [91]. The discovery of RBM20 as a highly effective modulator of titin-based stiffness opens up the possibility of using RBM20 as a treatment option both in HFpEF and other cardiac diseases that are be accompanied by a decrease in TTN compliance and an increase in myocardial stiffness [57,58,59]. Reducing Rbm20 activity has already shown positive effects, especially on diastolic function, in a mouse model with a deletion of Rbm20’s RRM [63, 92]. However, in otherwise healthy mice, the negative effects of reduced Rbm20 function are more pronounced; for example, the Frank-Starling mechanism is impaired and the mouse cannot fully compensate for cardiac stress [63]. More interesting and clinically relevant are studies on models for cardiac disease. In a model of HFpEF using transverse aortic constriction (TAC) surgery and deoxycorticosterone acetate (DOCA) pellet implantation, deletion of the RRM largely rescued the phenotype [64]. Functionally, this translated into an increased running distance after TAC [64]. Mechanistically, passive tension of cardiomyocytes (which is mainly determined by titin) decreased, while collagen-based stiffness was unaltered [64]. A similar rescue effect could also be observed in another HFpEF model with altered titin with increased passive tension [92]. Remarkably, in both cases, it was sufficient to suppress Rbm20 activity after the pathological state had already been established. However, as these models of HFpEF were mainly investigated in demembranated cardiomyocytes, they may not accurately represent the full phenotype of HFpEF [113]. In particular, they lack the contribution of cross-bridge mechanics, which have recently been shown to account for a significant amount of diastolic stiffness [115]. Therefore, further experiments on loss of RBM20’s effect on intact cardiomyocytes seem advisable, as the reduction of Ca2+ sensitivity caused by loss of RBM20 (in conjunction with adequate amounts of PKA activation) may counteract the increase in Ca2+ sensitivity which normally accompanies slowing of cross-bridge detachment [65, 113]. Currently, there exists one published drug discovery study on RBM20 splicing regulation, in which the group of cardenolides (including digoxin and digitoxin) is shown to specifically and potently be able to inhibit RBM20-dependent titin splicing [93]. Application of cardenolides led to a 10-fold reduction in RBM20 protein levels, and also changed the expression levels of several RBM20 interacting factors [93]. Cardenolides are already clinically used for cardiac disease; however, they have strict therapeutic levels, and potentially severe side effects. In other models, the loss of Rbm20 also reduced hypertrophy after volume overload and improved diastolic function and cardiac dimensions in a model with increased titin stiffness [56, 94]. Together, these findings suggest RBM20 manipulation as a valid therapeutic option, but reducing Rbm20 function also has negative effects. In general, heterozygous modifications of Rbm20 led to the best effects, while homozygous Rbm20 modifications came with an increase in negative side effects without corresponding benefits, like an increase in ECM stiffness and a worsened response to systolic stress [63]. The regulatory pathways affecting RBM20 are still poorly understood, and inhibiting RBM20 function might impact multiple pathways, not all of them in a beneficial way. For example, a 50% reduction in RBM20 already leads to potentially pro-arrhythmic calcium handling anomalies [11•]. Nevertheless, a greater understanding of how RBM20 functions and is regulated could open new avenues of treatment. For example, modulating RBM20’s PTMs, such as phosphorylation of the RSRSP stretch, could serve as a therapeutic target, especially in patients with mutations in the mutational hotspot. Another therapeutic approach would be the use of anti-sense oligonucleotides (AONs) that can bind and block RBM20 splicing sites, thereby reducing RBM20-dependent splicing [95]. AONs have already shown promising results in other diseases caused by altered splicing like Duchenne’s muscular dystrophy and spinal muscular atrophy [96, 97]. Interestingly, AONs can also be used to attract RNA-binding proteins or copy their function, which could be a therapeutic option for RBM20 cardiomyopathy patients [95, 98]. The use of small molecules to repress RBM20 splicing activity is also possible, although these may not only affect RBM20 [95, 99]. Different splice isoforms could also be directly targeted, with detrimental isoforms (such as CaMKIIδA) being knocked down using AONs, and protective isoforms being increased through the use of adeno-associated viruses [95]. In summary, RBM20 remains an intriguing target to therapeutically change titin (and thereby cardiac) compliance; however, potential side effects of RBM20 inhibition, including the potential induction of arrhythmias, cannot be neglected.
Conclusion
Familial DCM is a heterogenous disease with different presentations depending on the genetic cause. However, treatment is currently not tailored to the specific genetic cause of DCM. RBM20 cardiomyopathy is an arrhythmogenic form of DCM, and early ICD implantation and anti-arrhythmic drug therapy may be fitting treatment options. A reduction of the cellular calcium overload, for example through the use of verapamil, may also lead to better outcomes for patients. In addition, male patients with RBM20 cardiomyopathy may need earlier and more intensive care, while female patients might be treated more conservatively. Understanding the cause of the gender-specific presentation of RBM20 cardiomyopathy will also allow better therapy for the heavily affected male patients. Eventually, therapies could focus either on a restoration of normal splicing through, for example, anti-sense oligonucleotides, or reintroduction of RBM20 itself through, for example, adeno-associated viruses. Much of the research on RBM20 so far has focused on its interactions with TTN, and while aberrant TTN splicing likely contributes to disease presentation, other avenues of research also should not be neglected.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance
Pinto YM, Elliott PM, Arbustini E, Adler Y, Anastasakis A, Bohm M, et al. Proposal for a revised definition of dilated cardiomyopathy, hypokinetic non-dilated cardiomyopathy, and its implications for clinical practice: a position statement of the ESC working group on myocardial and pericardial diseases. Eur Heart J. 2016;37(23):1850–8. https://doi.org/10.1093/eurheartj/ehv727.
Hershberger RE, Hedges DJ, Morales A. Dilated cardiomyopathy: the complexity of a diverse genetic architecture. Nat Rev Cardiol. 2013;10(9):531–47. https://doi.org/10.1038/nrcardio.2013.105.
Codd MB, Sugrue DD, Gersh BJ, Melton LJ 3rd. Epidemiology of idiopathic dilated and hypertrophic cardiomyopathy. A population-based study in Olmsted County, Minnesota, 1975-1984. Circulation. 1989;80(3):564–72. https://doi.org/10.1161/01.cir.80.3.564.
Elliott P, Andersson B, Arbustini E, Bilinska Z, Cecchi F, Charron P, et al. Classification of the cardiomyopathies: a position statement from the European Society Of Cardiology Working Group on Myocardial and Pericardial Diseases. Eur Heart J. 2008;29(2):270–6. https://doi.org/10.1093/eurheartj/ehm342.
Haas J, Frese KS, Peil B, Kloos W, Keller A, Nietsch R, et al. Atlas of the clinical genetics of human dilated cardiomyopathy. Eur Heart J. 2015;36(18):1123–35a. https://doi.org/10.1093/eurheartj/ehu301.
Refaat MM, Lubitz SA, Makino S, Islam Z, Frangiskakis JM, Mehdi H, et al. Genetic variation in the alternative splicing regulator RBM20 is associated with dilated cardiomyopathy. Heart Rhythm. 2012;9(3):390–6. https://doi.org/10.1016/j.hrthm.2011.10.016.
Brauch KM, Karst ML, Herron KJ, de Andrade M, Pellikka PA, Rodeheffer RJ, et al. Mutations in ribonucleic acid binding protein gene cause familial dilated cardiomyopathy. J Am Coll Cardiol. 2009;54(10):930–41. https://doi.org/10.1016/j.jacc.2009.05.038.
Kayvanpour E, Sedaghat-Hamedani F, Amr A, Lai A, Haas J, Holzer DB, et al. Genotype-phenotype associations in dilated cardiomyopathy: meta-analysis on more than 8000 individuals. Clin Res Cardiol. 2017;106(2):127–39. https://doi.org/10.1007/s00392-016-1033-6.
Guo W, Schafer S, Greaser ML, Radke MH, Liss M, Govindarajan T, et al. RBM20, a gene for hereditary cardiomyopathy, regulates titin splicing. Nat Med. 2012;18(5):766–73. https://doi.org/10.1038/nm.2693.
Filippello A, Lorenzi P, Bergamo E, Romanelli MG. Identification of nuclear retention domains in the RBM20 protein. FEBS Lett. 2013;587(18):2989–95. https://doi.org/10.1016/j.febslet.2013.07.018.
• van den Hoogenhof MMG, Beqqali A, Amin AS, van der Made I, Aufiero S, Khan MAF, et al. RBM20 mutations induce an arrhythmogenic dilated cardiomyopathy related to disturbed calcium handling. Circulation. 2018;138(13):1330–42. https://doi.org/10.1161/CIRCULATIONAHA.117.031947This paper showcases the arrhythmogenic nature of RBM20 cardiomyopathy, and shows the altered calcium handling of Rbm20-deficient cardiomyocytes.
Li D, Morales A, Gonzalez-Quintana J, Norton N, Siegfried JD, Hofmeyer M, et al. Identification of novel mutations in RBM20 in patients with dilated cardiomyopathy. Clin Transl Sci. 2010;3(3):90–7. https://doi.org/10.1111/j.1752-8062.2010.00198.x.
Wells QS, Becker JR, Su YR, Mosley JD, Weeke P, D'Aoust L, et al. Whole exome sequencing identifies a causal RBM20 mutation in a large pedigree with familial dilated cardiomyopathy. Circ Cardiovasc Genet. 2013;6(4):317–26. https://doi.org/10.1161/CIRCGENETICS.113.000011.
Herman DS, Lam L, Taylor MR, Wang L, Teekakirikul P, Christodoulou D, et al. Truncations of titin causing dilated cardiomyopathy. N Engl J Med. 2012;366(7):619–28. https://doi.org/10.1056/NEJMoa1110186.
Maier LS, Bers DM. Role of Ca2+/calmodulin-dependent protein kinase (CaMK) in excitation–contraction coupling in the heart. Cardiovasc Res. 2007;73(4):631–40. https://doi.org/10.1016/j.cardiores.2006.11.005.
Xu X, Yang D, Ding JH, Wang W, Chu PH, Dalton ND, et al. ASF/SF2-regulated CaMKIIdelta alternative splicing temporally reprograms excitation-contraction coupling in cardiac muscle. Cell. 2005;120(1):59–72. https://doi.org/10.1016/j.cell.2004.11.036.
• Hey TM, Rasmussen TB, Madsen T, Aagaard MM, Harbo M, Molgaard H, et al. Pathogenic RBM20-variants are associated with a severe disease expression in male patients with dilated cardiomyopathy. Circ Heart Fail. 2019;12(3):e005700. https://doi.org/10.1161/CIRCHEARTFAILURE.118.005700This clinical paper shows the gender-specific severity of RBM20 cardiomyopathy.
Watanabe T, Kimura A, Kuroyanagi H. Alternative splicing regulator RBM20 and cardiomyopathy. Front Mol Biosci. 2018;5:105. https://doi.org/10.3389/fmolb.2018.00105.
Zahr HC, Jaalouk DE. Exploring the crosstalk between LMNA and splicing machinery gene mutations in dilated cardiomyopathy. Front Genet. 2018;9:231. https://doi.org/10.3389/fgene.2018.00231.
Murayama R, Kimura-Asami M, Togo-Ohno M, Yamasaki-Kato Y, Naruse TK, Yamamoto T, et al. Phosphorylation of the RSRSP stretch is critical for splicing regulation by RNA-Binding Motif Protein 20 (RBM20) through nuclear localization. Sci Rep. 2018;8(1):8970. https://doi.org/10.1038/s41598-018-26624-w.
Maatz H, Jens M, Liss M, Schafer S, Heinig M, Kirchner M, et al. RNA-binding protein RBM20 represses splicing to orchestrate cardiac pre-mRNA processing. J Clin Invest. 2014;124(8):3419–30. https://doi.org/10.1172/JCI74523.
Dauksaite V, Gotthardt M. Molecular basis of titin exon exclusion by RBM20 and the novel titin splice regulator PTB4. Nucleic Acids Res. 2018;46(10):5227–38. https://doi.org/10.1093/nar/gky165.
Beqqali A, Bollen IA, Rasmussen TB, van den Hoogenhof MM, van Deutekom HW, Schafer S, et al. A mutation in the glutamate-rich region of RNA-binding motif protein 20 causes dilated cardiomyopathy through missplicing of titin and impaired Frank-Starling mechanism. Cardiovasc Res. 2016;112(1):452–63. https://doi.org/10.1093/cvr/cvw192.
Upadhyay SK, Mackereth CD. Structural basis of UCUU RNA motif recognition by splicing factor RBM20. Nucleic Acids Res. 2020. https://doi.org/10.1093/nar/gkaa168.
Li S, Guo W, Dewey CN, Greaser ML. Rbm20 regulates titin alternative splicing as a splicing repressor. Nucleic Acids Res. 2013;41(4):2659–72. https://doi.org/10.1093/nar/gks1362.
Tilgner H, Knowles DG, Johnson R, Davis CA, Chakrabortty S, Djebali S, et al. Deep sequencing of subcellular RNA fractions shows splicing to be predominantly co-transcriptional in the human genome but inefficient for lncRNAs. Genome Res. 2012;22(9):1616–25. https://doi.org/10.1101/gr.134445.111.
• Bertero A, Fields PA, Ramani V, Bonora G, Yardimci GG, Reinecke H, et al. Dynamics of genome reorganization during human cardiogenesis reveal an RBM20-dependent splicing factory. Nat Commun. 2019;10(1):1538. https://doi.org/10.1038/s41467-019-09483-5This paper describes the colocalization and interaction of RBM20 and its targets in the nucleus, and shows that the TTN pre-mRNA is necessary for a RBM20-splicing factory to form and for proper splicing of other targets.
Aufiero S, van den Hoogenhof MMG, Reckman YJ, Beqqali A, van der Made I, Kluin J, et al. Cardiac circRNAs arise mainly from constitutive exons rather than alternatively spliced exons. RNA. 2018;24(6):815–27. https://doi.org/10.1261/rna.064394.117.
Khan MA, Reckman YJ, Aufiero S, van den Hoogenhof MM, van der Made I, Beqqali A, et al. RBM20 regulates circular RNA production from the titin gene. Circ Res. 2016;119(9):996–1003. https://doi.org/10.1161/CIRCRESAHA.116.309568.
Guo W, Zhu C, Yin Z, Wang Q, Sun M, Cao H, et al. Splicing factor RBM20 regulates transcriptional network of titin associated and calcium handling genes in the heart. Int J Biol Sci. 2018;14(4):369–80. https://doi.org/10.7150/ijbs.24117.
Beraldi R, Li X, Martinez Fernandez A, Reyes S, Secreto F, Terzic A, et al. Rbm20-deficient cardiogenesis reveals early disruption of RNA processing and sarcomere remodeling establishing a developmental etiology for dilated cardiomyopathy. Hum Mol Genet. 2014;23(14):3779–91. https://doi.org/10.1093/hmg/ddu091.
Zhu C, Yin Z, Tan B, Guo W. Insulin regulates titin pre-mRNA splicing through the PI3K-Akt-mTOR kinase axis in a RBM20-dependent manner. Biochim Biophys Acta Mol basis Dis. 2017;1863(9):2363–71. https://doi.org/10.1016/j.bbadis.2017.06.023.
Zhu C, Yin Z, Ren J, McCormick RJ, Ford SP, Guo W. RBM20 is an essential factor for thyroid hormone-regulated titin isoform transition. J Mol Cell Biol. 2015;7(1):88–90. https://doi.org/10.1093/jmcb/mjv002.
Keshwani MM, Aubol BE, Fattet L, Ma CT, Qiu J, Jennings PA, et al. Conserved proline-directed phosphorylation regulates SR protein conformation and splicing function. Biochem J. 2015;466(2):311–22. https://doi.org/10.1042/bj20141373.
Lai MC, Lin RI, Tarn WY. Transportin-SR2 mediates nuclear import of phosphorylated SR proteins. Proc Natl Acad Sci U S A. 2001;98(18):10154–9. https://doi.org/10.1073/pnas.181354098.
Vega RB, Konhilas JP, Kelly DP, Leinwand LA. Molecular mechanisms underlying cardiac adaptation to exercise. Cell Metab. 2017;25(5):1012–26. https://doi.org/10.1016/j.cmet.2017.04.025.
McMullen JR, Amirahmadi F, Woodcock EA, Schinke-Braun M, Bouwman RD, Hewitt KA, et al. Protective effects of exercise and phosphoinositide 3-kinase(p110alpha) signaling in dilated and hypertrophic cardiomyopathy. Proc Natl Acad Sci U S A. 2007;104(2):612–7. https://doi.org/10.1073/pnas.0606663104.
Blaustein M, Pelisch F, Tanos T, Munoz MJ, Wengier D, Quadrana L, et al. Concerted regulation of nuclear and cytoplasmic activities of SR proteins by AKT. Nat Struct Mol Biol. 2005;12(12):1037–44. https://doi.org/10.1038/nsmb1020.
Nyirenda MJ, Clark DN, Finlayson AR, Read J, Elders A, Bain M, et al. Thyroid disease and increased cardiovascular risk. Thyroid. 2005;15(7):718–24. https://doi.org/10.1089/thy.2005.15.718.
Ascheim DD, Hryniewicz K. Thyroid hormone metabolism in patients with congestive heart failure: the low triiodothyronine state. Thyroid. 2002;12(6):511–5. https://doi.org/10.1089/105072502760143908.
Kim EK, Choi EJ. Pathological roles of MAPK signaling pathways in human diseases. Biochim Biophys Acta. 2010;1802(4):396–405. https://doi.org/10.1016/j.bbadis.2009.12.009.
Cai H, Zhu C, Chen Z, Maimaiti R, Sun M, McCormick RJ, et al. Angiotensin II influences pre-mRNA splicing regulation by enhancing RBM20 transcription through activation of the MAPK/ELK1 signaling pathway. Int J Mol Sci. 2019;20(20). https://doi.org/10.3390/ijms20205059.
Nordgren KKS, Hampton M, Wallace KB. Editor’s highlight: the altered DNA methylome of chronic doxorubicin exposure in Sprague Dawley rats. Toxicol Sci. 2017;159(2):470–9. https://doi.org/10.1093/toxsci/kfx150.
Liu J, Carnero-Montoro E, van Dongen J, Lent S, Nedeljkovic I, Ligthart S, et al. An integrative cross-omics analysis of DNA methylation sites of glucose and insulin homeostasis. Nat Commun. 2019;10(1):2581. https://doi.org/10.1038/s41467-019-10487-4.
Joseph PV, Jaime-Lara RB, Wang Y, Xiang L, Henderson WA. Comprehensive and systematic analysis of gene expression patterns associated with body mass index. Sci Rep. 2019;9(1):7447. https://doi.org/10.1038/s41598-019-43881-5.
Suckale J, Wendling O, Masjkur J, Jäger M, Münster C, Anastassiadis K, et al. PTBP1 is required for embryonic development before gastrulation. PLoS One. 2011;6(2):e16992. https://doi.org/10.1371/journal.pone.0016992.
Boutz PL, Stoilov P, Li Q, Lin CH, Chawla G, Ostrow K, et al. A post-transcriptional regulatory switch in polypyrimidine tract-binding proteins reprograms alternative splicing in developing neurons. Genes Dev. 2007;21(13):1636–52. https://doi.org/10.1101/gad.1558107.
Amir-Ahmady B, Boutz PL, Markovtsov V, Phillips ML, Black DL. Exon repression by polypyrimidine tract binding protein. Rna. 2005;11(5):699–716. https://doi.org/10.1261/rna.2250405.
Wang ET, Ward AJ, Cherone JM, Giudice J, Wang TT, Treacy DJ, et al. Antagonistic regulation of mRNA expression and splicing by CELF and MBNL proteins. Genome Res. 2015;25(6):858–71. https://doi.org/10.1101/gr.184390.114.
Lorenzi P, Sangalli A, Fochi S, Dal Molin A, Malerba G, Zipeto D, et al. RNA-binding proteins RBM20 and PTBP1 regulate the alternative splicing of FHOD3. Int J Biochem Cell Biol. 2019;106:74–83. https://doi.org/10.1016/j.biocel.2018.11.009.
LeWinter MM, Wu Y, Labeit S, Granzier H. Cardiac titin: structure, functions and role in disease. Clin Chim Acta. 2007;375(1–2):1–9. https://doi.org/10.1016/j.cca.2006.06.035.
Trinick J, Knight P, Whiting A. Purification and properties of native titin. J Mol Biol. 1984;180(2):331–56. https://doi.org/10.1016/s0022-2836(84)80007-8.
Trombitas K, Wu Y, Labeit D, Labeit S, Granzier H. Cardiac titin isoforms are coexpressed in the half-sarcomere and extend independently. Am J Physiol Heart Circ Physiol. 2001;281(4):H1793–9. https://doi.org/10.1152/ajpheart.2001.281.4.H1793.
Freiburg A, Trombitas K, Hell W, Cazorla O, Fougerousse F, Centner T, et al. Series of exon-skipping events in the elastic spring region of titin as the structural basis for myofibrillar elastic diversity. Circ Res. 2000;86(11):1114–21. https://doi.org/10.1161/01.res.86.11.1114.
Cazorla O, Freiburg A, Helmes M, Centner T, McNabb M, Wu Y, et al. Differential expression of cardiac titin isoforms and modulation of cellular stiffness. Circ Res. 2000;86(1):59–67. https://doi.org/10.1161/01.res.86.1.59.
Hutchinson KR, Saripalli C, Chung CS, Granzier H. Increased myocardial stiffness due to cardiac titin isoform switching in a mouse model of volume overload limits eccentric remodeling. J Mol Cell Cardiol. 2015;79:104–14. https://doi.org/10.1016/j.yjmcc.2014.10.020.
Makarenko I, Opitz CA, Leake MC, Neagoe C, Kulke M, Gwathmey JK, et al. Passive stiffness changes caused by upregulation of compliant titin isoforms in human dilated cardiomyopathy hearts. Circ Res. 2004;95(7):708–16. https://doi.org/10.1161/01.RES.0000143901.37063.2f.
van Heerebeek L, Borbely A, Niessen HW, Bronzwaer JG, van der Velden J, Stienen GJ, et al. Myocardial structure and function differ in systolic and diastolic heart failure. Circulation. 2006;113(16):1966–73. https://doi.org/10.1161/CIRCULATIONAHA.105.587519.
Neagoe C, Kulke M, del Monte F, Gwathmey JK, de Tombe PP, Hajjar RJ, et al. Titin isoform switch in ischemic human heart disease. Circulation. 2002;106(11):1333–41. https://doi.org/10.1161/01.cir.0000029803.93022.93.
Jansweijer JA, Nieuwhof K, Russo F, Hoorntje ET, Jongbloed JD, Lekanne Deprez RH, et al. Truncating titin mutations are associated with a mild and treatable form of dilated cardiomyopathy. Eur J Heart Fail. 2017;19(4):512–21. https://doi.org/10.1002/ejhf.673.
Parikh VN, Caleshu C, Reuter C, Lazzeroni LC, Ingles J, Garcia J, et al. Regional variation in RBM20 causes a highly penetrant arrhythmogenic cardiomyopathy. Circ Heart Fail. 2019;12(3):e005371. https://doi.org/10.1161/CIRCHEARTFAILURE.118.005371.
Greaser ML, Warren CM, Esbona K, Guo W, Duan Y, Parrish AM, et al. Mutation that dramatically alters rat titin isoform expression and cardiomyocyte passive tension. J Mol Cell Cardiol. 2008;44(6):983–91. https://doi.org/10.1016/j.yjmcc.2008.02.272.
Methawasin M, Hutchinson KR, Lee EJ, Smith JE 3rd, Saripalli C, Hidalgo CG, et al. Experimentally increasing titin compliance in a novel mouse model attenuates the Frank-Starling mechanism but has a beneficial effect on diastole. Circulation. 2014;129(19):1924–36. https://doi.org/10.1161/CIRCULATIONAHA.113.005610.
Methawasin M, Strom JG, Slater RE, Fernandez V, Saripalli C, Granzier H. Experimentally increasing the compliance of titin through RNA binding motif-20 (RBM20) inhibition improves diastolic function in a mouse model of heart failure with preserved ejection fraction. Circulation. 2016;134(15):1085–99. https://doi.org/10.1161/CIRCULATIONAHA.116.023003.
Pulcastro HC, Awinda PO, Methawasin M, Granzier H, Dong W, Tanner BCW. Increased titin compliance reduced length-dependent contraction and slowed cross-bridge kinetics in skinned myocardial strips from Rbm (20ΔRRM) mice. Front Physiol. 2016;7:322. https://doi.org/10.3389/fphys.2016.00322.
Hamdani N, Krysiak J, Kreusser MM, Neef S, CGd R, Maier LS, et al. Crucial role for Ca2+/calmodulin-dependent protein kinase-II in regulating diastolic stress of normal and failing hearts via titin phosphorylation. Circ Res. 2013;112(4):664–74. https://doi.org/10.1161/CIRCRESAHA.111.300105.
Li KL, Methawasin M, Tanner BCW, Granzier HL, Solaro RJ, Dong WJ. Sarcomere length-dependent effects on Ca(2+)-troponin regulation in myocardium expressing compliant titin. J Gen Physiol. 2019;151(1):30–41. https://doi.org/10.1085/jgp.201812218.
Streckfuss-Bomeke K, Tiburcy M, Fomin A, Luo X, Li W, Fischer C, et al. Severe DCM phenotype of patient harboring RBM20 mutation S635A can be modeled by patient-specific induced pluripotent stem cell-derived cardiomyocytes. J Mol Cell Cardiol. 2017;113:9–21. https://doi.org/10.1016/j.yjmcc.2017.09.008.
Wyles SP, Li X, Hrstka SC, Reyes S, Oommen S, Beraldi R, et al. Modeling structural and functional deficiencies of RBM20 familial dilated cardiomyopathy using human induced pluripotent stem cells. Hum Mol Genet. 2016;25(2):254–65. https://doi.org/10.1093/hmg/ddv468.
Wyles SP, Hrstka SC, Reyes S, Terzic A, Olson TM, Nelson TJ. Pharmacological modulation of calcium homeostasis in familial dilated cardiomyopathy: an in vitro analysis from an RBM20 patient-derived iPSC model. Clin Transl Sci. 2016;9(3):158–67. https://doi.org/10.1111/cts.12393.
Schworer CM, Rothblum LI, Thekkumkara TJ, Singer HA. Identification of novel isoforms of the delta subunit of Ca2+/calmodulin-dependent protein kinase II. Differential expression in rat brain and aorta. J Biol Chem. 1993;268(19):14443–9.
• Zhang M, Gao H, Liu D, Zhong X, Shi X, Yu P, et al. CaMKII-delta9 promotes cardiomyopathy through disrupting UBE2T-dependent DNA repair. Nat Cell Biol. 2019;21(9):1152–63. https://doi.org/10.1038/s41556-019-0380-8This paper was the first in showing a detrimental effect of the CaMKIIδ9 isoform on the heart by a well-examined disruption of a DNA repair pathway.
Humphries KH, Izadnegahdar M, Sedlak T, Saw J, Johnston N, Schenck-Gustafsson K, et al. Sex differences in cardiovascular disease - impact on care and outcomes. Front Neuroendocrinol. 2017;46:46–70. https://doi.org/10.1016/j.yfrne.2017.04.001.
Cannata A, Fabris E, Merlo M, Artico J, Gentile P, Pio Loco C, et al. Sex differences in the long-term prognosis of dilated cardiomyopathy. Can J Cardiol. 2020;36(1):37–44. https://doi.org/10.1016/j.cjca.2019.05.031.
Halliday BP, Gulati A, Ali A, Newsome S, Lota A, Tayal U, et al. Sex- and age-based differences in the natural history and outcome of dilated cardiomyopathy. Eur J Heart Fail. 2018;20(10):1392–400. https://doi.org/10.1002/ejhf.1216.
Laura VS, Cristina L, Elena C, Giulia B, Massimo Z, Marco M, et al. Conflicting gender-related differences in the natural history of patients with idiopathic dilated cardiomyopathy. Epidemiol Biostat Public Health. 2017;14:e12527–1. https://doi.org/10.2427/12527.
Li X, Cai C, Luo R, Jiang R, Zeng J, Tang Y, et al. The usefulness of age and sex to predict all-cause mortality in patients with dilated cardiomyopathy: a single-center cohort study. Clin Interv Aging. 2015;10:1479–86. https://doi.org/10.2147/CIA.S88565.
Martinez-Selles M, Perez-David E, Yotti R, Jimenez-Borreguero J, Loughlin G, Gallego L, et al. Gender differences in right ventricular function in patients with non-ischaemic cardiomyopathy. Neth Hear J. 2015;23(12):578–84. https://doi.org/10.1007/s12471-015-0753-y.
Franaszczyk M, Chmielewski P, Truszkowska G, Stawinski P, Michalak E, Rydzanicz M, et al. Titin truncating variants in dilated cardiomyopathy - prevalence and genotype-phenotype correlations. PLoS One. 2017;12(1):e0169007. https://doi.org/10.1371/journal.pone.0169007.
van Rijsingen IA, Nannenberg EA, Arbustini E, Elliott PM, Mogensen J, Hermans-van Ast JF, et al. Gender-specific differences in major cardiac events and mortality in lamin A/C mutation carriers. Eur J Heart Fail. 2013;15(4):376–84. https://doi.org/10.1093/eurjhf/hfs191.
Bilano V, Gilmour S, Moffiet T, d'Espaignet ET, Stevens GA, Commar A, et al. Global trends and projections for tobacco use, 1990-2025: an analysis of smoking indicators from the WHO Comprehensive Information Systems for Tobacco Control. Lancet. 2015;385(9972):966–76. https://doi.org/10.1016/S0140-6736(15)60264-1.
Garawi F, Devries K, Thorogood N, Uauy R. Global differences between women and men in the prevalence of obesity: is there an association with gender inequality? Eur J Clin Nutr. 2014;68(10):1101–6. https://doi.org/10.1038/ejcn.2014.86.
Fermin DR, Barac A, Lee S, Polster SP, Hannenhalli S, Bergemann TL, et al. Sex and age dimorphism of myocardial gene expression in nonischemic human heart failure. Circ Cardiovasc Genet. 2008;1(2):117–25. https://doi.org/10.1161/CIRCGENETICS.108.802652.
Bell JR, Raaijmakers AJ, Curl CL, Reichelt ME, Harding TW, Bei A, et al. Cardiac CaMKIIdelta splice variants exhibit target signaling specificity and confer sex-selective arrhythmogenic actions in the ischemic-reperfused heart. Int J Cardiol. 2015;181:288–96. https://doi.org/10.1016/j.ijcard.2014.11.159.
Previlon M, Pezet M, Vinet L, Mercadier JJ, Rouet-Benzineb P. Gender-specific potential inhibitory role of Ca2+/calmodulin dependent protein kinase phosphatase (CaMKP) in pressure-overloaded mouse heart. PLoS One. 2014;9(6):e90822. https://doi.org/10.1371/journal.pone.0090822.
Parks RJ, Howlett SE. Sex differences in mechanisms of cardiac excitation-contraction coupling. Pflugers Archiv-Eur J Physiol. 2013;465(5):747–63. https://doi.org/10.1007/s00424-013-1233-0.
Akdis D, Saguner AM, Shah K, Wei C, Medeiros-Domingo A, von Eckardstein A, et al. Sex hormones affect outcome in arrhythmogenic right ventricular cardiomyopathy/dysplasia: from a stem cell derived cardiomyocyte-based model to clinical biomarkers of disease outcome. Eur Heart J. 2017;38(19):1498–508. https://doi.org/10.1093/eurheartj/ehx011.
Vicencio JM, Ibarra C, Estrada M, Chiong M, Soto D, Parra V, et al. Testosterone induces an intracellular calcium increase by a nongenomic mechanism in cultured rat cardiac myocytes. Endocrinology. 2006;147(3):1386–95. https://doi.org/10.1210/en.2005-1139.
van Riet EE, Hoes AW, Wagenaar KP, Limburg A, Landman MA, Rutten FH. Epidemiology of heart failure: the prevalence of heart failure and ventricular dysfunction in older adults over time. A systematic review. Eur J Heart Fail. 2016;18(3):242–52. https://doi.org/10.1002/ejhf.483.
Ilieșiu AM, Hodorogea AS. Treatment of heart failure with preserved ejection fraction. Adv Exp Med Biol. 2018;1067:67–87. https://doi.org/10.1007/5584_2018_149.
Zile MR, Baicu CF, Ikonomidis JS, Stroud RE, Nietert PJ, Bradshaw AD, et al. Myocardial stiffness in patients with heart failure and a preserved ejection fraction: contributions of collagen and titin. Circulation. 2015;131(14):1247–59. https://doi.org/10.1161/CIRCULATIONAHA.114.013215.
Bull M, Methawasin M, Strom J, Nair P, Hutchinson K, Granzier H. Alternative splicing of titin restores diastolic function in an HFpEF-like genetic murine model (TtnDeltaIAjxn). Circ Res. 2016;119(6):764–72. https://doi.org/10.1161/CIRCRESAHA.116.308904.
Liss M, Radke MH, Eckhard J, Neuenschwander M, Dauksaite V, von Kries JP, et al. Drug discovery with an RBM20 dependent titin splice reporter identifies cardenolides as lead structures to improve cardiac filling. PLoS One. 2018;13(6):e0198492. https://doi.org/10.1371/journal.pone.0198492.
Hinze F, Dieterich C, Radke MH, Granzier H, Gotthardt M. Reducing RBM20 activity improves diastolic dysfunction and cardiac atrophy. J Mol Med (Berl). 2016;94(12):1349–58. https://doi.org/10.1007/s00109-016-1483-3.
van den Hoogenhof MM, Pinto YM, Creemers EE. RNA splicing: regulation and dysregulation in the heart. Circ Res. 2016;118(3):454–68. https://doi.org/10.1161/CIRCRESAHA.115.307872.
Hua Y, Sahashi K, Rigo F, Hung G, Horev G, Bennett CF, et al. Peripheral SMN restoration is essential for long-term rescue of a severe spinal muscular atrophy mouse model. Nature. 2011;478(7367):123–6. https://doi.org/10.1038/nature10485.
Goemans NM, Tulinius M, van den Akker JT, Burm BE, Ekhart PF, Heuvelmans N, et al. Systemic administration of PRO051 in Duchenne’s muscular dystrophy. N Engl J Med. 2011;364(16):1513–22. https://doi.org/10.1056/NEJMoa1011367.
Cartegni L, Krainer AR. Correction of disease-associated exon skipping by synthetic exon-specific activators. Nat Struct Biol. 2003;10(2):120–5. https://doi.org/10.1038/nsb887.
Pawellek A, McElroy S, Samatov T, Mitchell L, Woodland A, Ryder U, et al. Identification of small molecule inhibitors of pre-mRNA splicing. J Biol Chem. 2014;289(50):34683–98. https://doi.org/10.1074/jbc.M114.590976.
Kowalczyk MS, Hughes JR, Babbs C, Sanchez-Pulido L, Szumska D, Sharpe JA, et al. Nprl3 is required for normal development of the cardiovascular system. Mamm Genome. 2012;23(7–8):404–15. https://doi.org/10.1007/s00335-012-9398-y.
Ito J, Iijima M, Yoshimoto N, Niimi T, Kuroda S, Maturana AD. RBM20 and RBM24 cooperatively promote the expression of short enh splice variants. FEBS Lett. 2016;590(14):2262–74. https://doi.org/10.1002/1873-3468.12251.
Robyns T, Willems R, Van Cleemput J, Jhangiani S, Muzny D, Gibbs R, et al. Whole exome sequencing in a large pedigree with DCM identifies a novel mutation in RBM20. Acta Cardiol. 2019:1–6. https://doi.org/10.1080/00015385.2019.1674490.
Monaco I, Santacroce R, Casavecchia G, Correale M, Bottigliero D, Cordisco G, et al. Double de novo mutations in dilated cardiomyopathy with cardiac arrest. J Electrocardiol. 2019;53:40–3. https://doi.org/10.1016/j.jelectrocard.2018.12.015.
Pantou MP, Gourzi P, Gkouziouta A, Tsiapras D, Zygouri C, Constantoulakis P, et al. Phenotypic heterogeneity within members of a family carrying the same RBM20 mutation R634W. Cardiology. 2018;141(3):150–5. https://doi.org/10.1159/000494453.
Sedaghat-Hamedani F, Haas J, Zhu F, Geier C, Kayvanpour E, Liss M, et al. Clinical genetics and outcome of left ventricular non-compaction cardiomyopathy. Eur Heart J. 2017;38(46):3449–60. https://doi.org/10.1093/eurheartj/ehx545.
Long PA, Evans JM, Olson TM. Diagnostic yield of whole exome sequencing in pediatric dilated cardiomyopathy. J Cardiovasc Dev Dis. 2017;4(3). https://doi.org/10.3390/jcdd4030011.
Fedida J, Fressart V, Charron P, Surget E, Hery T, Richard P, et al. Contribution of exome sequencing for genetic diagnostic in arrhythmogenic right ventricular cardiomyopathy/dysplasia. PLoS One. 2017;12(8):e0181840. https://doi.org/10.1371/journal.pone.0181840.
Zhao Y, Feng Y, Zhang YM, Ding XX, Song YZ, Zhang AM, et al. Targeted next-generation sequencing of candidate genes reveals novel mutations in patients with dilated cardiomyopathy. Int J Mol Med. 2015;36(6):1479–86. https://doi.org/10.3892/ijmm.2015.2361.
Waldmuller S, Schroeder C, Sturm M, Scheffold T, Imbrich K, Junker S, et al. Targeted 46-gene and clinical exome sequencing for mutations causing cardiomyopathies. Mol Cell Probes. 2015;29(5):308–14. https://doi.org/10.1016/j.mcp.2015.05.004.
Rampersaud E, Siegfried JD, Norton N, Li D, Martin E, Hershberger RE. Rare variant mutations identified in pediatric patients with dilated cardiomyopathy. Prog Pediatr Cardiol. 2011;31(1):39–47. https://doi.org/10.1016/j.ppedcard.2010.11.008.
Millat G, Bouvagnet P, Chevalier P, Sebbag L, Dulac A, Dauphin C, et al. Clinical and mutational spectrum in a cohort of 105 unrelated patients with dilated cardiomyopathy. Eur J Med Genet. 2011;54(6):e570–5. https://doi.org/10.1016/j.ejmg.2011.07.005.
Hamdani N, Kooij V, van Dijk S, Merkus D, Paulus WJ, Remedios CD, et al. Sarcomeric dysfunction in heart failure. Cardiovasc Res. 2008;77(4):649–58. https://doi.org/10.1093/cvr/cvm079.
Sequeira V, Najafi A, Wijnker PJ, Dos Remedios CG, Michels M, Kuster DW, et al. ADP-stimulated contraction: a predictor of thin-filament activation in cardiac disease. Proc Natl Acad Sci U S A. 2015;112(50):E7003–12. https://doi.org/10.1073/pnas.1513843112.
Sequeira V, van der Velden J. The Frank-Starling Law: a jigsaw of titin proportions. Biophys Rev. 2017;9(3):259–67. https://doi.org/10.1007/s12551-017-0272-8.
King NM, Methawasin M, Nedrud J, Harrell N, Chung CS, Helmes M, et al. Mouse intact cardiac myocyte mechanics: cross-bridge and titin-based stress in unactivated cells. J Gen Physiol. 2011;137(1):81–91. https://doi.org/10.1085/jgp.201010499.
Funding
Open Access funding provided by Projekt DEAL. MvdH was supported by a DFG (German Research Foundation) Research Grant (HO 6446/1-1) and a DZHK (German Center for Cardiovascular Research) Postdoc Startup Grant (81X3500122). DL was supported by a Kaltenbach stipendium from the Deutsche Herzstiftung.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflicts of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Translational Research in Heart Failure
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lennermann, D., Backs, J. & van den Hoogenhof, M.M.G. New Insights in RBM20 Cardiomyopathy. Curr Heart Fail Rep 17, 234–246 (2020). https://doi.org/10.1007/s11897-020-00475-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11897-020-00475-x