Abstract
Purpose of Review
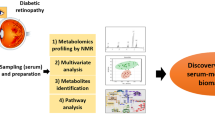
Metabolomics is the study of dysregulated metabolites in biological materials. We reviewed the use of the technique to elucidate the genetic and environmental factors that contribute to the development of diabetic retinopathy.
Recent Findings
With regard to metabolomic studies of diabetic retinopathy, the field remains in its infancy with few studies published to date and little replication of results. Vitreous and serum samples are the main tissues examined, and dysregulation in pathways such as the pentose phosphate pathway, arginine to proline pathway, polyol pathway, and ascorbic acidic pathways have been reported.
Summary
Few studies have examined the metabolomic underpinnings of diabetic retinopathy. Further research is required to replicate findings to date and determine longitudinal associations with disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The prevalence of diabetes is increasing worldwide, particularly in the Asia-Pacific region which has nearly 100 million persons with diabetes in China and 80 million in India alone [1]. Diabetic retinopathy remains one of the leading causes of blindness worldwide [1]. Much of the blindness from diabetic retinopathy is preventable with early detection and treatment.
Metabolomics is one of the newer “omics” fields and building on the principles of genomics, transcriptomics, and proteomics; it aims to identify and quantify the low molecular-weight metabolites in biological samples. In doing so, it moves further away from the genome and closer to providing a complete picture of the phenome. While the metabolomics techniques have been applied to a wide range of biological samples from food and beverages to tissue samples, its biological fluids such as blood plasma, serum, vitreous, tears, saliva, and urine are the most commonly studied. Tissue metabolites such as amino acids, oligonucleotides, carbohydrates, ketones, aldehydes, amines, and many forms of lipids are produced biologically as a result of cellular metabolism. Exogenous sources from diet, medications, gut microbe-host metabolism can also influence levels of these metabolites.
Metabolomic studies have uncovered blood metabolic signatures associated with complex diseases such as impaired glucose tolerance and diabetes. Recently, the technique has also been applied to study diabetic retinopathy. While genetic loci have been identified for diabetes susceptibility, diabetic retinopathy has proven to be more resistant to investigation with genome-wide association studies (GWAS) and results remain inconclusive [2]. Metabolomic studies into diabetic retinopathy are thus an attractive proposition. This review will cover the recent progress in metabolomic investigations of diabetic retinopathy.
Methods and Materials
Search Strategy
We conducted a Medline search of studies in participants with type 1 and type 2 diabetes published between 1980 and 2017. The search strategy involved the terms “diabetic retinopathy,” “diabetic macular edema,” “diabetic eye disease,” “metabolomics,” “metabonomics,” “metabolites,” “mass spectroscopy,” and “nuclear magnetic resonance”. Eighty-six abstracts were identified and reviewed, and from these, relevant articles were retrieved. Reference lists of review and original articles were examined to identify relevant publications. No studies published in a language other than English were found using this search strategy. Five original studies were identified which used metabolomic techniques to investigate diabetic retinopathy.
Overview of Metabolomics
The metabolome is analogous to the genome or proteome and refers to the full complement of metabolites in the tissue of interest. The Human Metabolome Database [3] currently lists 42,632 metabolite entries which are considerably less than the number of proteoforms, gene variants, and RNA transcripts identified to date. This reduced “search space” can make metabolomics an attractive field relative to other omic approaches. However, the diversity of chemical properties across different classes of metabolites and the prevalence of isomers mean that the field is not without its challenges. The data which is obtained is complementary to other “omics” studies and is often thought of as being closest to the phenotype. That is, it can often reveal the net change brought about by many competing upstream processes influencing protein activity, protein abundance, gene expression, and changes to the genome.
For over a hundred years, major advances in biochemistry have led to the myriad of biochemical pathways within a living cell being mapped in unprecedented detail. Metabolomics leverages this knowledge base through advances in two analytical technologies: nuclear magnetic resonance (NMR) spectroscopy and mass spectrometry (MS). Each approach has distinct benefits, and together, they can provide a comprehensive view of the metabolome. NMR has very minimal sample preparation as all the molecules within a sample are interrogated simultaneously. It is also a non-destructive technique and with automation can be high throughput. The major drawbacks to NMR are data interpretation is limited by the quality of the libraries and the computer algorithms available and relative to MS, the depth of coverage is greatly reduced due to the poor sensitivity of NMR which is generally in the millimolar range. Contemporary high-resolution high-mass accuracy mass spectrometers are able to detect extremely low levels of a particular metabolite. With the use of tandems, MS (MS/MS or MS2) and MSn great specificity can be achieved and modern instrument offers a linear dynamic range over five orders of magnitude giving great quantitation. However, this all comes at a price, samples need to be fractionated by liquid chromatography (LC) or gas chromatography (GC), and generally, only one subclass of metabolites can be assayed at a time, leading to lower throughput. In addition, the samples are consumed during the process which can be an issue in the event of limited samples.
Metabolomics studies are can be classified as discovery (global profiling) or targeted (at a defined set of compounds). Discovery metabolomics is an unbiased approach that can uncover novel metabolites related to disease, analogous to other omics approaches. A typical discovery metabolomics experiment will map and determine relative quantitation for many thousands of peaks or features across the sample set. However, definitively identifying the compound which forms a particular peak is limited by the library or libraries available to search the data against and the availability of purified standards to confirm, identity, and provide absolute quantitation of features of interest. Some of these limitations can be remedied to an extent with large sample sizes [4•].
Targeted metabolomics, on the other hand, can take advantage of commercially available standards to develop MS assays and construct calibration curves to determine quantities more precisely [4•]. Due to the targeted nature of the approach, once established, they can be high throughput due to shorter data acquisitions times and relatively simple data analysis. Targeted analyses also reduce the need for statistical corrections for multiple comparisons due to the lower run-to-run variance. While targeted methods can be designed to screen a number of putative pathways that are believed to be implicated in a disease, it is dependent on specified hypotheses and may not detect unsuspected or novel pathways.
A limitation of all MS approaches is isobaric compounds. Typically, mass spectrometers separate charged compounds based on their mass/charge ratio. If two compounds have the same, or very similar mass/charge, it is not possible to tell them apart. By increasing the resolving power of the instrument, it is possible to separate two compounds of increasingly similar masses. For example, at a resolution of 100,000, it is possible to separate mass which differs by only 0.01 m/z. However, if the instrument does not have sufficient higher resolving power or if the compounds have the same chemical formula, the mass of the compound alone is not enough.
Most mass spectrometers have the ability to isolate, fragment, and then reanalyze the product ions in the second round of mass spectrometry to provide an extra layer of information (tandem MS or MS2). Depending on their structure, two compounds that share a chemical formula may have distinct structures and yield quiet distinct product ions. Hence, the particular pairs of precursor and product ion masses (or a transition) becomes diagnostic for each compound. This forms the basis of most targeted MS methods. With ion trap instruments, this approach can be extended with repeated rounds of fragmentation giving MS3, MS4, or more. While this can be useful in identifying unknown compounds, it can be laborious and has poor sensitivity.
One problem with the tandem MS is if multiple precursor ions exist within the isolation window of the mass spectrometer. For example, there are hundreds of lipids which have precursor ion mass within 0.6 m/z of 762.4. If one of these species is a putative biomarker, but some others are present in the sample but are not associated with the disease, they will be co-isolated with the lipid of interest leading to the issue of isobaric interference. A new approach to reduce this problem is differential ion mobility, which can be used to separate isobaric ions on the basis of their shape. When incorporated into a targeted lipidomics, work flow can eliminate this issue without the need for extensive sample fractionation.
Metabolomics and Diabetes
In the Framingham Heart Study, fasting blood levels of branched-chain and aromatic amino acids isoleucine, leucine, valine, tyrosine, and phenylalanine were elevated up to 12 years before the onset of clinical diabetes. Participants in the top quartile of a combination of three of these amino acids were at five-fold greater risk of developing diabetes, even after adjusting for age, fasting glucose, and other confounders [5•]. Higher levels of another metabolite, 2-aminoadipate, have also been found to predict incident diabetes [6]. Mendelian randomization techniques have been used to analyze these associations, and the results suggest that the branched-chain amino acid association with type 2 diabetes is causal [7]. Further work suggests that some of these branched-chain amino acids may originate from the microbiome [8].
Another substudy of the Framingham population found lipids with lower carbon number and fewer double bonds were associated with an increased risk of diabetes, whereas lipids of higher carbon number and more double bonds were associated with decreased risk [9]. The major lipid class implicated was triacylglycerols (TAGs). It has been hypothesized that a joint effect of both lipids, and amino acids may promote mitochondrial deficiency and disrupt insulin signaling particularly in skeletal muscle [10,11,12]. Further studies are needed to determine the relationship of these compounds to diabetes, and whether they play a causal role or are epiphenomenon associated with metabolic dysregulation in diabetes [4•].
Metabolomics and Diabetic Retinopathy
These studies postulate that a distinct metabolic signature of diabetic retinopathy exists and can be resolved from that of diabetes alone. Studies have fallen into two main groups depending on the tissue investigated (Table 1). Three studies have examined and identified metabolite markers using vitreous samples obtained following ocular surgery [15,16,17]. The control group in these studies are patients who have undergone ocular surgery for other reasons such as macular hole or retinal detachment. However, the invasive nature of vitreous sampling limits study replication and it would be difficult for vitreous biomarkers to be utilized widely for risk prediction. For these reasons, blood plasma and sera have remained the biological sample of choice in metabolomic studies. Two studies to date have examined this issue [13•, 14].
Plasma Markers and Diabetic Retinopathy
Chen et al. [13•] performed one of the first studies of gas chromatography-mass spectrometry (GC-MS) on plasma samples to study the metabolic signature of diabetic retinopathy. Forty samples from Singaporean Indians with type 2 diabetes and moderate non proliferative diabetic retinopathy (NPDR) were compared with 40 samples from patients with type 2 diabetes and no diabetic retinopathy, and the discovery phase reported significantly altered levels of 11 metabolites. These were decreased levels of 1,5-anhydroglucitol and increased levels of 1,5-gluconolactone, 2-deoxyribonic acid, 3,4-dihydroxybutyric acid, erythritol, gluconic acid, lactose/cellobiose, maltose/trehalose, mannose, ribose, and urea. In the validation set, 2-deoxyribonic acid, 3,4-dihydroxybutyric acid, erythritol, gluconic acid, and ribose were found to remain elevated, while maltose was decreased. Pathway mapping of these metabolites showed significant enrichment of the pentose phosphate pathway that is responsible for the generation of NADPH to combat oxidative stress. Increased concentrations of cytosine (p = 0.010), cytidine (p = 0.001), and thymidine (p = 0.001) were also found associated with diabetic retinopathy compared to those without. Of these compounds, cytidine had the highest area under the curve (AUC) of 0.849 ± 0.048, and at the optimal cutoff point of 0.076 mg/L; the sensitivity and specificity of cytidine as a biomarker for diabetic retinopathy were 73.7 and 91.9%, respectively, suggesting potential value as a biomarker for diabetic retinopathy.
This study had major strengths including an independent validation cohort, and adjustment for confounders such as diet and kidney disease, but had limited generalizability as only Singaporeans of South Indian ancestry were studied. Such work shows the importance of validating metabolomics studies in other populations and ethnic groups.
Another study by Li et al. [14] applied systems biology-based approaches to study metabolomics of blood plasma in patients with diabetic retinopathy. This study was novel as patients were classified according to usual Western (International) classification systems of diabetic retinopathy, as well as to a Chinese Medicine classification. The authors reported that in 88 patients with type 2 diabetes, the Western classification was associated with ten metabolites (pyruvic acids, L-aspartic acid, β-hydroxybutyric acid, methymalonic acid, citric acid, glucose, stearic acid, trans-oleic acid, linoleic acid, arachidonic acid), while the Chinese classification was associated with four metabolites (pyruvic acids, L-aspartic acid, glycerol, and cholesterol). Pyruvic acid and L-aspartic acid were identified in both classification systems. What was unclear in this study was the control group and whether they were free from diabetic retinopathy. The authors acknowledged another limitation as lack of data on the presence of chronic kidney disease, which could lead to impaired renal excretion of aspartic acid and other non-essential amino acids and result in the elevated levels observed in cases.
The association of lower plasma levels of two omega-6 polyunsaturated fatty acids (i.e., the PUFAs arachidonic acid and linoleic acid) with proliferative diabetic retinopathy reported by Li. et al. [14] is of interest as it has been reported that lower levels of these two PUFAs may be associated with higher levels of circulating pro-inflammatory markers such as IL-1ra and IL-6 and lower anti-inflammatory marker TGFb [18]. Diabetic retinopathy is associated with pro-inflammatory cytokines and downregulation of omega-6 PUFAs may potentiate these effects and increase the risk of developing the condition.
Vitreous Markers and Diabetic Retinopathy
Paris et al. [15] have used global and targeted liquid chromatography-mass spectrometry (LC-MS) to generate and validate the metabolomic profile of vitreous samples from 20 patients with type 2 diabetes and proliferative diabetic retinopathy and 31 control patients with no diabetes. Results were compared to findings from the vitreous gel of oxygen-induced retinopathy mouse models. All patients underwent standard pars plana vitrectomy with a 25-gauge 3-port system and with a high-speed vitreous cutter (2500 cycle/min). Retinopathy was induced in C57BL/6 mice using standard protocols. Pathway enrichment analysis revealed that arginine metabolism and ammonia detoxification (urea cycle) were two of the most perturbed pathways in both species and were dysregulated to a similar magnitude. Elevated levels of methionine, allantoin, decanoylcarnitine, arginine, proline, citrulline, ornithine, and octanoylcarnitine were observed in patients with proliferative diabetic retinopathy. The authors speculated that these findings implicate compromised Mueller glial cell metabolism in disrupting neurovascular crosstalk within the retina, potentially promoting diabetic retinopathy progression [19, 20].
Arginine is metabolized through two different pathways in the retina: the arginase pathway that produces ornithine and urea, and the nitric oxide synthase (NOS) pathway, which generates citrulline and NO [21]. Overactivity of the enzyme arginase II in rodents causes a shortfall of arginine for the NOS pathway, resulting in reduced NO availability and uncoupling of the NOS pathway. This leads to consequent endothelial cell dysfunction and impaired vasodilation, which are characteristics of diabetic retinopathy. Lack of NO also results in increased generation of oxygen and nitrogen reactive species that accelerate diabetic retinopathy [22].
Ornithine and proline, both dysregulated in the vitreous samples examined by Paris et al. [15], can be generated from arginine via the arginase pathway, whereas citrulline is produced via the NOS pathway. Fatty acid oxidation was also perturbed in both human and mouse vitreous, in the form of dysregulated acylcarnitine. These results, however, have not been replicated in other studies of vitreous from patients with diabetic retinopathy [16].
Hydrogen NMR has also been used to investigate vitreous samples in diabetic retinopathy. Barba et al. [16] obtained vitreous samples from 22 patients with type 1 diabetes with proliferative diabetic retinopathy and from 22 nondiabetic patients who underwent macular hole surgery (controls). They reported higher lactate and lower galactitol and ascorbic acid levels in patients with proliferative diabetic retinopathy. As expected, glucose was significantly higher in samples from proliferative diabetic retinopathy patients than nondiabetic patients. They obtained a model using these and other metabolites that correctly classified 19 of 22 patients with proliferative diabetic retinopathy and 18 of 22 controls (sensitivity of 86% and a specificity of 81%). The authors highlighted several important limitations of their study. These included the presence of vitreous hemorrhage in the samples which introduced large amounts of blood metabolites which may be unrelated to diabetic retinopathy. The effect of this was minimized through the use of vitreous samples with low blood contamination. Patients with recent laser photocoagulation, which can alter retinal metabolism, were also excluded.
The high levels of lactate likely reflect increased tissue acidosis and anaerobic glycolysis, particularly as the retina is one of the most metabolically active tissues. After removing the lactate peak at 1.35 ppm, Barba et al. [16] found lower levels of galactitol which they attributed to polyol pathway activation, a metabolic pathway involved in the pathogenesis of diabetic retinopathy [23].
Aldose reductase (AR) is the first and rate-limiting enzyme in the polyol pathway, and both glucose and galactose are substrates for the enzyme, with galactose preferred under normoglycemic conditions. Under conditions of hyperglycemia, glucose displaces galactose which results in increased levels of sorbitol, the product of aldose reductase enzymatic activity on glucose. Galactitol, the reduced end product of galactose, is hence lowered in this situation [24]. The authors propose that they did not directly detect high levels of sorbitol in vitreous due to further metabolism of sorbitol to fructose, fructose-3-phosphate, and 3-deoxyglucosone, whereas galactitol is not further metabolized and thus accumulates [25]. Another possibility may be that sorbitol is impermeable to the cell membrane and accumulates intracellularly where it increases cellular osmolality and reduces vital cellular functions [23].
Ascorbic acid is a cofactor in the biosynthesis of collagen, catecholamines, neurohormones, and important antioxidant free radical scavenger. This role takes on particular significance in the retina where there is considerable light-induced free radical formation [26]. Vitamin C can only be acquired from dietary sources in humans and exists in two major forms. The charged form, ascorbic acid, enters cells through sodium-dependent facilitated transport. The uncharged form, dehydroascorbate, is similar structurally to glucose, and enters cells by way of glucose transporters (GLUTs) and is then converted back to ascorbic acid within the cell [27]. Retinal cells depend on GLUT-1 transport rather than sodium-dependent uptake of ascorbic acid, and this is competitively inhibited by glucose. Hence, in chronic hyperglycemia, ascorbic acid transport into retinal cells is impaired with consequent increased photo-oxidative damage [28].
Further, ascorbic acid is a cofactor for several hydroxylases important in neuropeptide synthesis including proline hydroxylase and dopamine hydroxylase [29]. The lack of ascorbic acid may hence play a role in the early neurodegeneration observed in diabetic retinopathy.
Finally, ascorbic acid may inhibit angiogenesis, a central event in diabetic retinopathy, as suggested by experiments with corneal neovascularization in rodent models [30]. This possibility is supported by a small study showing that ascorbic acid metabolism was impaired in persons with diabetes who developed diabetic retinopathy, compared to those who did not [31].
Young et al. [17] have also performed NMR spectroscopy on vitreous samples in a study of patients with ocular disease, a small number of whom (n = 2) had proliferative diabetic retinopathy while the majority had ocular inflammation. Although specific metabolites dysregulated in diabetic retinopathy were not reported in this study, the authors concluded that a distinct metabolomic profile existed for diabetic retinopathy which could discriminate between different ocular diseases. This provides further evidence of the feasibility that metabolomic analysis can identify distinct biomarkers for diabetic retinopathy.
Conclusion
Metabolomics is an attractive field with which to study the pathogenesis of the disease. With regard to diabetic retinopathy, the field remains in its infancy with few studies published to date and little replication of results. Vitreous and serum samples are the main tissues studied and dysregulation in pathways such as the pentose phosphate pathway, arginine to proline pathway, polyol pathway, and ascorbic acidic pathways have been reported. Further research is required to replicate these findings and determine longitudinal associations with disease.
References
Papers of particular interest, published recently, have been highlighted as: •Of importance
Diabetes: facts and figures. 2017. (www.idf.org).
Kuo JZ, Wong TY, Rotter JI. Challenges in elucidating the genetics of diabetic retinopathy. JAMA Ophthalmol. 2014;132:96–107.
Wishart DS, Tzur D, Knox C, Eisner R, Guo AC, Young N, et al. HMDB: the human metabolome database. Nucleic Acids Res. 2007;35:D521–6.
• Filla LA, Edwards JL. Metabolomics in diabetic complications. Mol BioSyst. 2016;12:1090–105. This review provides a comprehensive survey of new results in metabolomic experiments and diabetic complications
• Wang TJ, Larson MG, Vasan RS, Cheng S, Rhee EP, McCabe E, et al. Metabolite profiles and the risk of developing diabetes. Nat Med. 2011;17:448–53. This study provides a good description of metabolomic techniques and relationship to diabetes
Wang TJ, Ngo D, Psychogios N, Dejam A, Larson MG, Vasan RS, et al. 2-Aminoadipic acid is a biomarker for diabetes risk. J Clin Invest. 2013;123:4309–17.
Lotta LA, Scott RA, Sharp SJ, Burgess S, Luan J, Tillin T, et al. Genetic predisposition to an impaired metabolism of the branched-chain amino acids and risk of type 2 diabetes: a mendelian randomisation analysis. PLoS Med. 2016;13:e1002179.
Pedersen HK, Gudmundsdottir V, Nielsen HB, Hyotylainen T, Nielsen T, Jensen BA, et al. Human gut microbes impact host serum metabolome and insulin sensitivity. Nature. 2016;535:376–81.
Rhee EP, Cheng S, Larson MG, Walford GA, Lewis GD, McCabe E, et al. Lipid profiling identifies a triacylglycerol signature of insulin resistance and improves diabetes prediction in humans. J Clin Invest. 2011;121:1402–11.
Newgard CB. Interplay between lipids and branched-chain amino acids in development of insulin resistance. Cell Metab. 2012;15:606–14.
Krebs M, Krssak M, Bernroider E, Anderwald C, Brehm A, Meyerspeer M, et al. Mechanism of amino acid-induced skeletal muscle insulin resistance in humans. Diabetes. 2002;51:599–605.
Anderson SG, Dunn WB, Banerjee M, Brown M, Broadhurst DI, Goodacre R, et al. Evidence that multiple defects in lipid regulation occur before hyperglycemia during the prodrome of type-2 diabetes. PLoS One. 2014;9:e103217.
• Chen L, Cheng CY, Choi H, Ikram MK, Sabanayagam C, Tan GS, et al. Plasma Metabonomic profiling of diabetic retinopathy. Diabetes. 2016;65:1099–108. One of the first studies using robust metabolomics techniques to study diabetic retinopathy
Li X, Luo X, Lu X, Duan J, Xu G. Metabolomics study of diabetic retinopathy using gas chromatography-mass spectrometry: a comparison of stages and subtypes diagnosed by Western and Chinese medicine. Mol BioSyst. 2011;7:2228–37.
Paris LP, Johnson CH, Aguilar E, Usui Y, Cho K, Hoang LT, et al. Global metabolomics reveals metabolic dysregulation in ischemic retinopathy. Metabolomics. 2016;12:15.
Barba I, Garcia-Ramirez M, Hernandez C, Alonso MA, Masmiquel L, Garcia-Dorado D, et al. Metabolic fingerprints of proliferative diabetic retinopathy: an 1H-NMR-based metabonomic approach using vitreous humor. Invest Ophthalmol Vis Sci. 2010;51:4416–21.
Young SP, Nessim M, Falciani F, Trevino V, Banerjee SP, Scott RA, et al. Metabolomic analysis of human vitreous humor differentiates ocular inflammatory disease. Mol Vis. 2009;15:1210–7.
Ferrucci L, Cherubini A, Bandinelli S, Bartali B, Corsi A, Lauretani F, et al. Relationship of plasma polyunsaturated fatty acids to circulating inflammatory markers. J Clin Endocrinol Metab. 2006;91:439–46.
Bringmann A, Pannicke T, Grosche J, Francke M, Wiedemann P, Skatchkov SN, et al. Muller cells in the healthy and diseased retina. Prog Retin Eye Res. 2006;25:397–424.
Metea MR, Newman EA. Glial cells dilate and constrict blood vessels: a mechanism of neurovascular coupling. J Neurosci. 2006;26:2862–70.
Narayanan SP, Xu Z, Putluri N, Sreekumar A, Lemtalsi T, Caldwell RW, et al. Arginase 2 deficiency reduces hyperoxia-mediated retinal neurodegeneration through the regulation of polyamine metabolism. Cell Death Dis. 2014;5:e1075.
Narayanan SP, Rojas M, Suwanpradid J, Toque HA, Caldwell RW, Caldwell RB. Arginase in retinopathy. Prog Retin Eye Res. 2013;36:260–80.
Lorenzi M. The polyol pathway as a mechanism for diabetic retinopathy: attractive, elusive, and resilient. Exp Diabetes Res. 2007;2007:61038.
Gabbay KH. The sorbitol pathway and the complications of diabetes. N Engl J Med. 1973;288:831–6.
Kador PF. The role of aldose reductase in the development of diabetic complications. Med Res Rev. 1988;8:325–52.
Tokuda K, Zorumski CF, Izumi Y. Effects of ascorbic acid on UV light-mediated photoreceptor damage in isolated rat retina. Exp Eye Res. 2007;84:537–43.
Hosoya K, Minamizono A, Katayama K, Terasaki T, Tomi M. Vitamin C transport in oxidized form across the rat blood-retinal barrier. Invest Ophthalmol Vis Sci. 2004;45:1232–9.
Minamizono A, Tomi M, Hosoya K. Inhibition of dehydroascorbic acid transport across the rat blood-retinal and -brain barriers in experimental diabetes. Biol Pharm Bull. 2006;29:2148–50.
Komeima K, Rogers BS, Lu L, Campochiaro PA. Antioxidants reduce cone cell death in a model of retinitis pigmentosa. Proc Natl Acad Sci U S A. 2006;103:11300–5.
Ashino H, Shimamura M, Nakajima H, Dombou M, Kawanaka S, Oikawa T, et al. Novel function of ascorbic acid as an angiostatic factor. Angiogenesis. 2003;6:259–69.
Sinclair AJ, Girling AJ, Gray L, Le GC, Lunec J, Barnett AH. Disturbed handling of ascorbic acid in diabetic patients with and without microangiopathy during high dose ascorbate supplementation. Diabetologia. 1991;34:171–5.
Acknowledgements
NHMRC Early Career Fellowship Grant APP1073530 to Gerald Liew
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Gerald Liew, Zhou Lei, Nichole Joachim, I-Van Ho, Tien Y. Wong, Paul Mitchell, Bamini Gopinath, and Ben Crossett declare that they have no conflict of interest.
Gavin Tan reports being on the Advisory board for Novartis, travel support from Bayer, research support from Santen, speaker for Abbott Medical, and speaker and travel support from Allergan.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
This article is part of the Topical Collection on Microvascular Complications—Retinopathy
Rights and permissions
About this article
Cite this article
Liew, G., Lei, Z., Tan, G. et al. Metabolomics of Diabetic Retinopathy. Curr Diab Rep 17, 102 (2017). https://doi.org/10.1007/s11892-017-0939-3
Published:
DOI: https://doi.org/10.1007/s11892-017-0939-3