Abstract
Better management of allergic diseases needs a sharpened understanding of disease heterogeneity and mechanisms in relation to clinically significant outcomes. Phenotypes describing observable clinical and morphologic characteristics and unique responses to treatment have been developed; however, they do not relate to disease mechanisms. Recently, extended heterogeneous and disease-related metabolic, inflammatory, immunological, and remodeling pathways have been described, and reproducible patterns are defined as disease endotypes. An endotype might consist of several intricated mechanisms that cannot be clearly separated into “pure single molecular mechanism” thus being a “complex endotype.” The description of an endotype may rely on biomarkers, which can be the signature of a complex underlying pathway or a key molecule associated with or directly playing a role in a particular disease endotype. The Th2 type inflammation can be defined as a complex endotype in asthma and linked to mechanisms of disease development and response to treatment and to disease outcomes such as exacerbations and remodeling. The type 2 complex endotype in allergies and asthma includes innate lymphoid cells, T helper 2 cells, tissue eosinophilia, and IgE production. Currently, emerging endotype-driven strategies in asthma, particularly the development of biologicals that target a single molecular pathway, are being focused for solving individualized clinical problems on disease outcomes. Progress is also being made for endotyping rhinitis, chronic rhinosinusitis, and atopic dermatitis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Allergic diseases affect the lives of more than one billion people worldwide, and their prevalence is expected to reach up to 4 billion in 2050. Better understanding of disease heterogeneity and mechanisms in relation to clinically significant outcomes, such as disease progression or response to treatment, is essential to improve their outcome (Fig. 1). In recent years, induction of immune tolerance has become a prime target for prevention and treatment strategies for allergic diseases. Immune tolerance to allergens can be defined as establishment of a long-term clinical tolerance against allergens, which immunologically implies changes in memory type allergen-specific T and B cell responses as well as mast cell and basophil activation thresholds that no longer cause allergic symptoms [1–3]. T and B regulatory cells and production of allergen-specific IgE-blocking IgG4 isotype antibodies play an essential role in allergen tolerance. Selection of patients for allergen-specific immunotherapy is based today on clinical judgment, and unfortunately, there are no good predictors of response. An endotype-driven approach to select responders might improve results if properly validated, predictive biomarkers were available.
The Concept of Asthma Phenotypes, Endotypes, and Biomarkers
The heterogeneity of allergic diseases and asthma in relation to clinically significant outcomes, including response to treatment, has been established beyond any doubt. However, current guidelines ignore disease heterogeneity and causal pathways. These lead to unsuccessful clinical “bulk” trials or contradictory epidemiologic and genetic surveys, in which patient subgrouping and disease heterogeneity has not been taken into consideration.
At first, phenotypes describing clinical and morphologic characteristics as well as unique responses to treatment have been developed to address the complexities of the disease. Phenotypes may be clinically relevant in terms of presentation, triggers, and treatment response, but do not necessarily relate to or give insights into the underlying pathological mechanism [4]. For most of the allergic diseases, extended heterogeneous disease-related metabolic, inflammatory, immunological, and remodeling pathways have been described, and a reproducible underlying mechanism is defined as a disease endotype [5•, 6•]. Single molecular mechanism-linked endotypes can be defined such as periostin high, anti-IL-13 responsive asthma. In contrast, other endotypes involve concomitantly several pathways that might prevail one over the other (in a network model) and can be due to several modulators such as metabolic or different inflammatory pathways. Examples of such complex endotypes are Th2 inflammation or aspirin-intolerant asthma.
There are several benefits of endotyping in a clinical setting such as stringent consideration of entry criteria for epidemiological, genetic, or drug-related trials. In the future, the endotype-tailored management (personalized medicine, stratified medicine, individualized medicine) may identify patient subgroups at risk for severe disease or with a particular response to existing or new treatments [5•, 7••, 8]. In addition, developing mouse models reflecting human endotypes would help the understanding of pathophysiological mechanisms of allergic disease.
To become clinically relevant, the endotype should be related to validated biomarkers that correspond to the underlying mechanism. The purpose of the biomarker is to identify disease endotype, predict onset and prognosis of a disease, measure exposure, monitor response to treatment, and forecast unfavorable evolution [6•]. The biomarker can be the signature of a complex underlying pathway or a key molecule of a particular disease endotype. To further complicate the picture, the predictive value of the same biomarker is highly variable across age groups, disease severity, and in relation to the outcome. The ideal biomarker should be pathway-specific, reproducible in the same individual and in an independent prediction cohort, and usable as a diagnostic test (easily measurable and affordable). New strategies for discovery and validation of biomarkers such as gene expression (microarrays) and omics provide combined signatures as per system medicine.
The concept of mechanism-tailored end points is surfacing in the field of allergic disease. In addition, the outcomes should be relevant both for the disease and for that particular patient. Unfortunately, no single intervention with a biological immune response modifier has been proven to improve disparate aspects of the disease, with significant differences within the same individual in terms of disease outcomes (the dissociated response). For example in asthma, one biological may well control exacerbations but has no effect on lung function, symptoms, or quality of life [1, 6•, 8].
The Type 2 Inflammatory Endotype
Physiologic Type 2 Inflammatory Responses
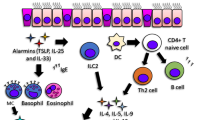
As part of the physiologic immune response, type 2 inflammatory responses are highly dynamic processes initiated by epithelial cell damage, resulting either in chronic injury or in healing of the tissue architecture. The type 2 milieu promotes wound healing processes, which are beneficial in parasitic infections or toxin exposure, but account for increasingly dysfunctional vital organs such as the lung in the case of asthma. This milieu is shaped by cytokines (IL-25, IL-31, IL-33, and thymic stromal lymphopoietin (TSLP)) released from epithelial cells, which stimulate Th2 cells, type 2 innate lymphoid cells (ILCs) and invariant natural killer T cells (NKT cells) to secrete Th2 cytokines and to activate dendritic cells, which results in further differentiation and clonal expansion of Th2 cells and further activation of ILC2 cells as well as tissue eosinophilia [9–14].
The type 2 inflammatory process requires communication between resident cells such as epithelial, endothelial, fibroblast, and muscle cells and the highly mobile cells of the innate and adaptive immunity. In the human airway mucosa, the TSLP receptor is constitutively and preferentially expressed by myeloid CD1c+ dendritic cells (DCs) [15].
The Type 2 Inflammatory Response in Allergic Diseases
The density of the DC subset expressing the TSLP receptor in the nasal mucosa increases significantly after allergen challenge. TSLP enhances the capacity of CD1c+ DCs to activate allergen-specific memory CD4+ T cells and induces CCR7 expression on CD1c+ DCs with their recruitment from the lymph nodes. However, in an already established Th2-mediated inflammatory reaction, the Th2 cytokines attenuate TSLP-mediated CCR7 induction with retention of CD1c+ DCs in the inflamed tissue to further exacerbate local inflammation by activating local antigen-specific memory Th2 cells [15]. Eosinophils also contribute to the initiating phase of the type 2-polarized pulmonary inflammation and to the recruitment of effector Th2 T cells while suppressing the Th1/Th17 pathways [16]. Epidermal γδ T cells might play an important role in the pathogenesis of Th2-dominant skin diseases because of their active production of IL-13 [17].
ILC2s are capable of producing high amounts of IL-5 and IL-13 in response to IL-25, IL-33, and TSLP, leading to eosinophilia and goblet cell hyperplasia at mucosal surfaces [18–20]. Moreover, ILC2s facilitate sensitization to local Th2-inducing allergen exposures and mediate Th2 cell differentiation via IL-33-dependent manner [21, 22].
In humans, the subset of ILC2 is expressed in peripheral blood, lung, skin, and inflamed tissues and release high amounts of important sources of IL-5 and IL-13 [23•, 24, 25]. IL-5- and IL-13-producing ILC2 have been found in the sputum of asthma patients [26], while overexpression of TSLP in the airway epithelium results in increased number of IL-13-producing ILC2s that correlated with severe asthma [27]. Taken together, these data indicate that ILC2s play a critical role for the induction of Th2-driven inflammation in asthma. In addition, ILC2s have been found in nasal polyps in patients with chronic rhinosinusitis [28–30].
In addition, ILC2 shares features with Th2 cells such as expressions of GATA3 transcription factor and CRTH2 [23•]. CRTH2 is also abundantly expressed on eosinophil and basophils [31–33]. It can directly stimulate ILC2s and can also potentiate IL-25/IL-33-mediated innate immune responses [34]. Nasal cat allergen challenge in allergic rhinitis induced an increased percentage of peripheral blood CRTH2 + ILC2 that express CD84 [35].
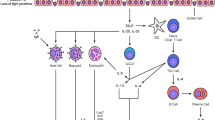
Effector Th2 cells release IL-4, IL-5, and IL-13. IL-4 and IL-13 are essential for class-switching to the ε immunoglobulin heavy chain in B cells and the production of allergen-specific IgE antibodies, while IL-5 is crucial for promoting and sustaining tissue eosinophilia. Other pathways involved in Th2 inflammation are the mast cell/IL-9 pathway and the PGD2 pathway [36–38, 39•]. Th1 or Th17 cells may add to the Th2-driven inflammation, with their role in apoptosis of the epithelium in asthma and atopic dermatitis, formation of granulomas (Th1), and in promoting neutrophilic inflammation (Th17) [40–44]. Further influences may be added by the remodeling phenotype, by associated microbiota or by activation of peculiar metabolic pathways such as the eicosanoid pathway in aspirin-exacerbated respiratory disease or the l-Arg/asymmetric dimethylarginine (ADMA) or the lectin pathway in obesity (Fig. 2) [45, 46, 47•]. It has been shown that human eosinophils express leptin receptor Ob-Rb and that leptin induces eosinophils to produce inflammatory cytokines [48, 49]. Data from the SARP cohort showed a higher median plasma ADMA level and lower median plasma l-arginine in subjects with late-onset asthma compared with early onset. A reduced l-arginine/ADMA was associated with less IgE [47•].
The complex network of Th2 endotype in allergic diseases involves the interaction between innate immune response and Th2 cells. Three major downstream effector pathways can be described: the IgE pathway, the IL-5/eotaxin pathway, and the IL-4/IL-13 pathway. Additional modulators of the Th2 endotype can be described such as Th17 or Th1 cells, the IL-9/mast cell axis, activation of the metabolic pathways, or the degree of airway remodeling
The type 2 inflammation is characterized by a high cellular plasticity that enables the cells to adapt to a specific inflammatory milieu. Innate cytokines such as IL-33 and TSLP modulate the mast cell phenotype [50], while type 2 cytokines promote a particular phenotype of smooth muscle cells in asthma and influence permissiveness of epithelium for allergens and of the endothelium for the recruitment of inflammatory cells to inflamed tissues and are involved in the production of mucus [51].
The Type 2 Complex Endotype in the Clinic
The type 2 inflammatory complex endotype has been related to response to treatment and to disease outcomes, such as exacerbations, and may have a role in remodeling in asthma [52•]. Several type 2 biomarkers have been described: sputum and blood eosinophils, fractional exhaled NO (FeNO), serum periostin, the Th2 gene signature (serpin B2, periostin, CLCA1 or CLC, CPA3, DNA-SE1L3) in bronchial and nasal epithelial cells, the sputum cells Th2 gene mean (IL-4, IL-5, IL-13), TSLP and mast cells in bronchial biopsies, and the salivary inflammatory profile (Table 1). Each biomarker reflects a compartment or a pathway involved in type 2 inflammation related to Th2 cells and ILC2. For non-specific interventions, such as inhaled corticosteroids, all of the above can be used to predict response. However, for more targeted interventions, such as anti-IL-5 or anti-IL-13, each of these Th2 or ILC2 biomarkers needs to be related to the specifically targeted pathway. For example, FeNO and sputum eosinophils on the one hand and blood eosinophils on the other hand reflect different endotypes of Th2 or ILC2-mediated inflammation. In a cross-sectional study, FeNO and blood eosinophil values offered independent information with respect to the prevalence of wheeze, asthma diagnosis, and asthma events [70•]. The authors suggest that blood eosinophilia is a marker of more severe systemic inflammation driven by a strong chemokine signal (such as IL-5) and more extensive eosinophilic airway inflammation involving the small airways and therefore inhaled corticosteroids (ICS) non-responsive. It may also highlight the risk of asthma exacerbations requiring oral corticosteroids. Further evidence was provided in the DREAM trial, which indicated that blood eosinophils were most closely related to a positive response to mepolizumab (anti-IL-5) compared to sputum eosinophils [70•, 71••]. In contrast, an increased FeNO value indicates predominance of the IL-4/IL-13-mediated pathway with eosinophilc inflammation localized in the bronchial mucosa and thus responding to ICS or to IL-4/IL-13 blockade. Increased FeNO value was a good predictor of a clinical response to lebrikizumab in the MILLY trial, besides serum periostin [72••]. Also in support, serum periostin was the best predictor of lung eosinophilia in severe asthma, when compared with blood eosinophils, again depicting the two types of systemic versus local inflammation [73•]. Longitudinal analysis of trends of increase or decrease in biomarkers, including fluctuation analysis, might offer additional information (Table 1). Combination of multiple biomarkers, including omics approaches and gene expression microarrays, may provide an added value.
Endotype-driven therapeutic strategies have been increasingly successful in asthma. Selection of patients in the DREAM trial based on their eosinophilic profile proved the efficacy of anti-IL-5 interventions. However, the issues related to the dissociated effect and drug efficacy at the target site remain unresolved. Unfortunately, so far very few attempts were made for the endotype-driven management of allergic diseases.
Applying the same model to allergic and non-allergic rhinitis could prove as successful in promoting personalized approaches, especially for the severe forms of the disease. The well-recognized link between rhinitis and asthma should be integrated and tackled within the framework provided by endotypes. Rhinitis endotypes can be defined in relation to the background inflammation or in terms of treatment responsiveness. The following endotypes can be proposed for allergic rhinitis, although their clinical relevance remains to be proven and associated biomarkers validated: IL-5-, IgE-, or IL-4/IL-13-driven Th2 type inflammation which can be steroid-responsive, anti-IgE responsive, anti-IL-5 responsive, and anti-IL-4/IL-13 responsive. For non-allergic rhinitis, the definition of endotypes (Th2 or non-Th2 type inflammation) should include the driving cause: superantigens, local IgE production, and autoantibodies.
A predominant Th2 type of inflammation characterized by increased levels in tissue homogenates of IgE, specific IgE to Staphylococcus aureus enterotoxin, ECP, and IL-5 was related to relapse after surgery for chronic rhinosinusitis (CRS) with nasal polyps, while a mixed profile with significant higher levels of IFN-γ and lower concentrations of IgE, ECP, and IL-5 was encountered for non-recurrent cases [74]. In the same line, a PRACTALL document described several endotypes for CRS characterized by differences in responsiveness to treatment, including topical intranasal corticosteroids and biological agents, such as anti-IL-5 and anti-IgE. The described CRS endotypes were based on different biomarkers linked to underlying mechanisms [75•].
Atopic dermatitis (AD) is a chronic inflammatory skin disease with complex genetic and immunological mechanisms. Several endotypes can be proposed according to the inflammatory background such as Th2/IL-22/periostin high or Th17/Th1 high or in relation to the expression of fillagrin, MATT, or vitamin D pathway gene mutations. For the Th2 type AD, serum periostin is related to disease severity [76, 77].
Targeting both the inflammatory immune-dysregulated pathways and the barrier defect in AD holds future promise. Several new targets such as toll-like receptors, type 2 ILCs, and tight junction proteins are emerging. Promising new therapeutic agents in the near future are sphinganine, cannabinoids, and highly targeted monoclonal antibodies.
Conclusion
The type 2 endotype was described for all major allergic diseases: asthma, rhinitis, chronic rhinosinusitis, and atopic dermatitis. This complex endotype is mainly mediated by ILC2 and Th2 cells as well as Th2 cytokine-producing NKT cells, whereas their individual contribution is not known. Major progress has been obtained in some clinics for endotype-guided treatment of asthma, but still, there are unsolved issues pending such as the dissociated effect on disease outcomes and the drug efficacy at target site.
The description of an endotype relies on biomarkers. Several biomarkers have been described for the Th2 endotype, such as exhaled NO, sputum or blood eosinophils, total serum IgE or specific IgE, serum periostin, sputum or epithelial cell (bronchi or nasal) gene signature, or the saliva inflammatory profile. Until now, none of these biomarkers matched the ideal profile: pathway-specific, reproducible, and affordable as diagnostic test in the clinic. The variance across age, asthma severity, and with the treatment target is still an unsolved issue. The use of a composite index of clinical biomarkers such as BMI, sputum or blood eosinophils, exhaled NO or derived from omics approaches, and gene expression microarrays seems to have a better predictive value.
Pending identification of validated biomarkers reflecting the disease mechanism and predicting the natural course and response to treatment of the disease, an endotype-driven approach should be identified also for allergic rhinitis, chronic rhinosinusitis, and atopic dermatitis, especially for the severe forms of the disease. Selecting responders for allergen-specific immunotherapy might also benefit from an endotype-oriented strategy.
Abbreviations
- ADMA:
-
Asymmetric dimethylarginine
- CLC:
-
Charcot-Leyden crystal protein
- CPA3:
-
Carboxypeptidase A3
- DNA-SE1L3:
-
Deoxyribonuclease I-like 3
- DC:
-
Dendritic cells
- Eos:
-
Eosinophils
- FeNO:
-
Fractional exhaled NO
- ICS:
-
Inhaled corticosteroids
- ILC:
-
Innate lymphoid cells
- l-Arg:
-
l-Arginine
- NKT cells:
-
Natural killer T cells
- PG:
-
Prostaglandin
- SARP:
-
Severe Asthma Respiratory Program
- Th:
-
T helper cell
- TSLP:
-
Thymic stromal lymphopoietin
- VOCs:
-
Volatile organic compounds
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
Akdis CA. Therapies for allergic inflammation: refining strategies to induce tolerance. Nat Med. 2012;18:736–49.
Akdis M, Akdis C. Immune tolerance. In: Bochner BS, Adkinson Jr NF, Burks W, Busse WW, Holgate Jr ST, Lemanski RF, O’Hehir RE, editors. Middleton’s allergy. 8th ed. Cambridge: Elsevier; 2013.
Akdis M et al. Mechanisms of allergen-specific immunotherapy: multiple suppressor factors at work in immune tolerance to allergens. J Allergy Clin Immunol. 2014;133:621–31.
Pillai RA, Calhoun WJ. Introduction to asthma and phenotyping. Adv Exp Med Biol. 2014;795:5–15.
Lotvall J et al. Asthma endotypes: a new approach to classification of disease entities within the asthma syndrome. J Allergy Clin Immunol. 2011;127:355–60. The PRACTALL consensus was the first to propose the criteria to validate an asthma endotype.
Agache IO. From phenotypes to endotypes to asthma treatment. Curr Opin Allergy Clin Immunol. 2013;13:249–56. The review critically revises new approaches to classify asthma and the emerging endotype-driven strategies.
Agache I et al. Untangling asthma phenotypes and endotypes. Allergy. 2012;67:835–46. Comprehensive description of asthma phenotypes and corresponding endotypes grading the level of evidence for corresponding biomarkers.
Agache IO. Endotype driven treatment of asthma. Curr Treat Options Allergy. 2014;1:198–212.
Scanlon ST et al. Type 2 innate lymphoid cells: new players in asthma and allergy. Curr Opin Immunol. 2012;24:707–12.
Holgate ST. Innate and adaptive immune responses in asthma. Nat Med. 2012;18:673–83.
DeKruyff RH et al. Innate immunity in the lung regulates the development of asthma. Immunol Rev. 2014;260:235–48.
Akbari O et al. CD4+ invariant T-cell-receptor+ natural killer T cells in bronchial asthma. N Engl J Med. 2006;354:1117–29.
Gauvreau GM et al. Effects of an anti-TSLP antibody on allergen-induced asthmatic responses. N Engl J Med. 2014;370:2102–10.
Dua B et al. Myeloid dendritic cells type 2 in allergic asthma. Allergy. 2013;68:1322–6.
Melum GR, et al. A thymic stromal lymphopoietin-responsive dendritic cell subset mediates allergic responses in the upper airway mucosa. J Allergy Clin Immunol. 2014;134(3):613–21
Jacobsen EA, et al. Re-defining the unique roles for eosinophils in allergic respiratory inflammation. Clin Exp Allergy. 2014;44(9):1119–36
Kim K et al. Comparative analysis of human epidermal and peripheral blood gammadelta T cell cytokine profiles. Ann Dermatol. 2014;26:308–13.
Kabata H et al. Thymic stromal lymphopoietin induces corticosteroid resistance in natural helper cells during airway inflammation. Nat Commun. 2013;4:2675.
Kim HY et al. Innate lymphoid cells responding to IL-33 mediate airway hyperreactivity independently of adaptive immunity. J Allergy Clin Immunol. 2012;129:216–27. e1-6.
Kondo Y et al. Administration of IL-33 induces airway hyperresponsiveness and goblet cell hyperplasia in the lungs in the absence of adaptive immune system. Int Immunol. 2008;20:791–800.
Gold MJ et al. Group 2 innate lymphoid cells facilitate sensitization to local, but not systemic, TH2-inducing allergen exposures. J Allergy Clin Immunol. 2014;133:1142–8.
Halim TY et al. Group 2 innate lymphoid cells are critical for the initiation of adaptive T helper 2 cell-mediated allergic lung inflammation. Immunity. 2014;40:425–35.
Mjosberg JM et al. Human IL-25- and IL-33-responsive type 2 innate lymphoid cells are defined by expression of CRTH2 and CD161. Nat Immunol. 2011;12:1055–62. This paper describe human type 2 innate lymphoid cells for the first time.
Kim BS et al. TSLP elicits IL-33-independent innate lymphoid cell responses to promote skin inflammation. Sci Transl Med. 2013;5:170ra16.
Teunissen MB, et al. Composition of innate lymphoid cell subsets in the human skin: enrichment of NCR ILC3 in lesional skin and blood of psoriasis patients. J Invest Dermatol. 2014;134(9):2351–60
Allakhverdi Z et al. CD34+ hemopoietic progenitor cells are potent effectors of allergic inflammation. J Allergy Clin Immunol. 2009;123:472–8.
Shikotra A et al. Increased expression of immunoreactive thymic stromal lymphopoietin in patients with severe asthma. J Allergy Clin Immunol. 2012;129:104–11. e1-9.
Shaw JL et al. IL-33-responsive innate lymphoid cells are an important source of IL-13 in chronic rhinosinusitis with nasal polyps. Am J Respir Crit Care Med. 2013;188:432–9.
Nagarkar DR et al. Thymic stromal lymphopoietin activity is increased in nasal polyps of patients with chronic rhinosinusitis. J Allergy Clin Immunol. 2013;132:593–600. e12.
Kamekura R et al. The role of IL-33 and its receptor ST2 in human nasal epithelium with allergic rhinitis. Clin Exp Allergy. 2012;42:218–28.
Gervais FG et al. Selective modulation of chemokinesis, degranulation, and apoptosis in eosinophils through the PGD2 receptors CRTH2 and DP. J Allergy Clin Immunol. 2001;108:982–8.
Pettipher R et al. Heightened response of eosinophilic asthmatic patients to the CRTH2 antagonist OC000459. Allergy. 2014;69:1223–32.
Hirai H et al. Prostaglandin D2 selectively induces chemotaxis in T helper type 2 cells, eosinophils, and basophils via seven-transmembrane receptor CRTH2. J Exp Med. 2001;193:255–61.
Xue L et al. Prostaglandin D2 activates group 2 innate lymphoid cells through chemoattractant receptor-homologous molecule expressed on TH2 cells. J Allergy Clin Immunol. 2014;133:1184–94.
Doherty TA et al. Allergen challenge in allergic rhinitis rapidly induces increased peripheral blood type 2 innate lymphoid cells that express CD84. J Allergy Clin Immunol. 2014;133:1203–5.
Dougherty RH et al. Accumulation of intraepithelial mast cells with a unique protease phenotype in T(H)2-high asthma. J Allergy Clin Immunol. 2010;125:1046–53. e8.
Oh CK et al. Biology of the interleukin-9 pathway and its therapeutic potential for the treatment of asthma. Inflamm Allergy Drug Targets. 2011;10:180–6.
Fajt ML et al. Prostaglandin D(2) pathway upregulation: relation to asthma severity, control, and TH2 inflammation. J Allergy Clin Immunol. 2013;131:1504–12.
Barnes N et al. A randomized, double-blind, placebo-controlled study of the CRTH2 antagonist OC000459 in moderate persistent asthma. Clin Exp Allergy. 2012;42:38–48. Prood-of-concept study testing the safety and efficacy of targeted PGD2 intervention in Th2 high asthma.
Steinke JW et al. Prominent role of IFN-gamma in patients with aspirin-exacerbated respiratory disease. J Allergy Clin Immunol. 2013;132:856–65. e1-3.
Rebane A et al. Mechanisms of IFN-gamma-induced apoptosis of human skin keratinocytes in patients with atopic dermatitis. J Allergy Clin Immunol. 2012;129:1297–306.
Wenzel SE et al. Asthmatic granulomatosis: a novel disease with asthmatic and granulomatous features. Am J Respir Crit Care Med. 2012;186:501–7.
Linden A, et al. Interleukin-17 cytokine signalling in patients with asthma. Eur Respir J. 2014;44(5):1319–31
Agache I et al. Increased serum IL-17 is an independent risk factor for severe asthma. Respir Med. 2010;104:1131–7.
Szczeklik A et al. The broken balance in aspirin hypersensitivity. Eur J Pharmacol. 2006;533:145–55.
Desai D et al. Elevated sputum interleukin-5 and submucosal eosinophilia in obese individuals with severe asthma. Am J Respir Crit Care Med. 2013;188:657–63.
Holguin F et al. An association between l-arginine/asymmetric dimethyl arginine balance, obesity, and the age of asthma onset phenotype. Am J Respir Crit Care Med. 2013;187:153–9. The study advances a new hypothesis of metabolic disturbed pathway in obese asthma.
Conus S et al. Leptin is an eosinophil survival factor. J Allergy Clin Immunol. 2005;116:1228–34.
Wong CK et al. Leptin-mediated cytokine release and migration of eosinophils: implications for immunopathophysiology of allergic inflammation. Eur J Immunol. 2007;37:2337–48.
Han NR, et al. TSLP induces mast cell development and aggravates allergic reactions through the activation of MDM2 and STAT6. J Invest Dermatol. 2014;134(10):2521–30.
Soyka MB et al. Defective epithelial barrier in chronic rhinosinusitis: the regulation of tight junctions by IFN-gamma and IL-4. J Allergy Clin Immunol. 2012;130:1087–96. e10.
Woodruff PG et al. T-helper type 2-driven inflammation defines major subphenotypes of asthma. Am J Respir Crit Care Med. 2009;180:388–95. First in vivo study validating the concept of Th2 high and low asthma in relation to response to treatment and to lung remodelling.
Green RH, Brightling CE, McKenna S, et al. Asthma exacerbations and sputum eosinophil counts: a randomised controlled trial. Lancet 2002;360:1715–21.
Pijnenburg MW, De Jongste JC. Exhaled nitric oxide in childhood asthma: a review. Clin Exp Allergy. 2008;38:246–59.
Petsky HL, Cates CJ, Li A, et al. Tailored interventions based on exhaled nitric oxide versus clinical symptoms for asthma in children and adults. Cochrane Database Syst Rev. 2009;4, CD006340.
Kanemitsu Y, Matsumoto H, Mishima M, et al. Factors contributing to an accelerated decline in pulmonary function in asthma. Allergol Int. 2014;63:181–8.
Tran TN, Khatry DB, Ke X, et al. High blood eosinophil count is associated with more frequent asthma attacks in asthma patients. Ann Allergy Asthma Immunol. 2014;113(1):19–24.
ten Brinke A, Zwinderman AH, Sterk PJ et al. Factors associated with persistent airflow limitation in severe asthma. Am J Respir Crit Care Med. 2001;164:744–8.
Peters MC, Mekonnen ZK, Yuan S, et al. Measures of gene expression in sputum cells can identify TH2-high and TH2-low subtypes of asthma. J Allergy Clin Immunol. 2014;133:388–94.
Baines KJ, Simpson JL, Wood LG, et al. Sputum gene expression signature of 6 biomarkers discriminates asthma inflammatory phenotypes. J Allergy Clin Immunol 2014;133:997–1007.
Poole A, Urbanek C, Eng C, et al. Dissecting childhood asthma with nasal transcriptomics distinguishes subphenotypes of disease. J Allergy Clin Immunol. 2014;133(3):670–8
Little FF, Delgado DM, Wexler PJ, et al. Salivary inflammatory mediator profiling and correlation to clinical disease markers in asthma. PLoS One. 2014;9(1):e84449.
Hanania NA, Wenzel S, Rosén K, et al. Exploring the effects of omalizumab in allergic asthma: an analysis of biomarkers in the EXTRA study. Am J Respir Crit Care Med. 2013;187:804–11. First study to validate a composite biomarker measure for effectiveness of omalizumab in severe allergic asthma.
Ortega H, Li H, Suruki R, et al. Cluster analysis and characterization of response to mepolizumab: a step closer to personalized medicine for patients with severe asthma. Ann Am Thorac Soc. 2014 Jul 1. [Epub ahead of print].
Agache I, Ciobanu C. Persistent high FeNO phenotype in asthma. J Allergy Clin Immunol. 2011;127:AB4.
Agache I, Ciobanu C. Predictive value of lung function trend and FeNO for difficult asthma in children. J Investig Allergol Clin Immunol 2012;22:419–26.
Stern G, de Jongste J, van der Valk R, et al. Fluctuation phenotyping based on daily fraction of exhaled nitric oxide values in asthmatic children. J Allergy Clin Immunol 2011;128:293–300.
Newby C, Agbetile J, Hargadon B, Monteiro W, Green R, Pavord I et al. Lung function decline and variable airway inflammatory pattern: longitudinal analysis of severe asthma. J Allergy Clin Immunol. 2014;134(2):287–94.
van der Schee MP, Palmay R, Cowan JO, et al. Predicting steroid responsiveness in patients with asthma using exhaled breath profiling. Clin Exp Allergy. 2013;43:1217–25.
Malinovschi A et al. Exhaled nitric oxide levels and blood eosinophil counts independently associate with wheeze and asthma events in National Health and Nutrition Examination Survey subjects. J Allergy Clin Immunol. 2013;132:821–7. e1-5. The study shows that blood eosinophils and FeNO are independent predictors for asthma events and response to treatment supporting the 2 main sub-endotypes of Th2 inflammation.
Pavord ID et al. Mepolizumab for severe eosinophilic asthma (DREAM): a multicentre, double-blind, placebo-controlled trial. Lancet. 2012;380:651–9. DREAM is the first large randomized study validating the endotype-driven approach in severe asthma as a method to increase response to targeted treatment (anti IL-5). Patients were selected based on a composite measure of eosinophilic inflammation.
Corren J et al. Lebrikizumab treatment in adults with asthma. N Engl J Med. 2011;365:1088–98. MILLY is one of the first targeted treatments in asthma highlighting the value of a biomarker used as a pathway specific diagnostic test.
Jia G et al. Periostin is a systemic biomarker of eosinophilic airway inflammation in asthmatic patients. J Allergy Clin Immunol. 2012;130:647–54. e10. The BOBCAT data prove the value of serum periostin as a reliable and noninvasive biomarker of eosinophilic inflammation in asthma.
Van Zele T et al. Differences in initial immunoprofiles between recurrent and nonrecurrent chronic rhinosinusitis with nasal polyps. Am J Rhinol Allergy. 2014;28:192–8.
Akdis CA et al. Endotypes and phenotypes of chronic rhinosinusitis: a PRACTALL document of the European Academy of Allergy and Clinical Immunology and the American Academy of Allergy, Asthma & Immunology. J Allergy Clin Immunol. 2013;131:1479–90. The PRACTALL consensus evaluates the relation between the CRS endotypes and disease severity and response to treatment.
Mu Z, et al. Molecular biology of atopic dermatitis. Clin Rev Allergy Immunol. 2014.
Wollenberg A et al. Immunological and molecular targets of atopic dermatitis treatment. Br J Dermatol. 2014;170 Suppl 1:7–11.
Compliance with Ethics Guidelines
Conflict of Interest
Ioana Agache, Kazunari Sugita, Hideaki Morita, Mübeccel Akdis, and Cezmi A. Akdis declare that they have no conflicts of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is part of the Topical Collection on Asthma
Rights and permissions
About this article
Cite this article
Agache, I., Sugita, K., Morita, H. et al. The Complex Type 2 Endotype in Allergy and Asthma: From Laboratory to Bedside. Curr Allergy Asthma Rep 15, 29 (2015). https://doi.org/10.1007/s11882-015-0529-x
Published:
DOI: https://doi.org/10.1007/s11882-015-0529-x