Opinion statement
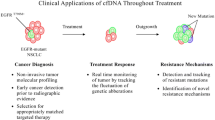
Isolation and analysis of circulating tumor DNA (ctDNA) have emerged as an effective and promising tool for genomic profiling in non-small cell lung cancer. Analysis of ctDNA can be particularly useful in situations where tissue biopsy is not safely obtainable due to poor physical condition or inaccessible tumor biopsy location. In addition to identifying oncogenic driver mutations which can be treated with targetable therapy in the treatment naïve advanced non-small cell lung cancer (NSCLC) setting, ctDNA is being utilized in novel ways including monitoring during an advanced NSCLC patient’s treatment course (real-time monitoring), determining mechanisms of resistance and, lastly, as a tool to identify minimal residual disease in early-stage NSCLC. Recent research demonstrates that ctDNA testing can provide a useful adjunct to tissue genotyping in NSCLC. Utilization of ctDNA into routine clinical practice for NSCLC should be strongly considered.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tumor molecular analysis has become an increasingly important component of lung cancer evaluation and management, particularly in the non-small cell lung cancer (NSCLC) setting [1]. Once primarily differentiated by histology (squamous versus non-squamous), NSCLC is now recognized as a diverse and complex group of malignancies with distinctive molecular signatures. Since the initial identification of activating mutations in the epidermal growth factor receptor (EGFR) among NSCLC patients who responded to EGFR tyrosine kinase inhibitors (EGFR TKI) [2, 3], extensive basic science and translational research has focused on identification of other oncogenic driver mutations and development of appropriate targeted therapies [1]. Routine testing of several oncogenic driver mutations with U.S. Food and Drug Administration (FDA)-approved targeted therapies is now recommended as standard of care for all patients diagnosed with advanced non-squamous NSCLC and in selected patients with advanced squamous NSCLC, which includes EGFR mutations, anaplastic lymphoma kinase (ALK) rearrangements, ROS1 rearrangements, and BRAF V600E mutations [4]. In addition, several other emerging biomarkers in NSCLC such as RET rearrangements, MET amplifications or mutations, and HER2 mutations have known targeted therapies which are not FDA-approved or have associated investigational agents which are being evaluated in clinical trials [1, 4]. Accurate testing of genomic alterations at the time of initial NSCLC diagnosis and at disease progression for those with select oncogenic driver mutations is a crucial aspect of therapeutic management in this patient population.
Traditionally, solid tumor tissue biopsy has been considered the gold standard for the detection of molecular alterations in many malignancies including NSCLC. More recently, isolation and analysis of circulating tumor DNA (ctDNA) from the plasma has emerged as an effective and promising tool for the performance of this molecular testing [5••]. The term “liquid biopsy” is commonly not only used to refer to molecular assays performed on blood rather than tumor tissue samples but can also include testing on other body fluids such as urine, pleural fluid, saliva, and cerebrospinal fluid. Liquid biopsy testing can measure circulating tumor cells, exosomes, platelet RNA, circulating tumor RNA (miRNA and long non-coding RNA), as well as ctDNA. The most robust research in the clinical setting has evaluated ctDNA technologies, and several innovative ctDNA platforms have been created to enhance the reliability and breadth of these molecular techniques. As ctDNA is thought to represent a relatively small proportion of cell-free DNA present within the blood (< 1%), technologies with high sensitivity are required in order to detect ctDNA. While not considered a replacement for tumor tissue testing, ctDNA analysis can act as a useful adjunct for NSCLC management in myriad clinical situations. Although genomic profiling in NSCLC is most applicable in the advanced disease setting, newer innovative approaches are being evaluated including utilization of plasma ctDNA as a tool to identify minimal residual disease in early-stage NSCLC and as a predictive biomarker for immunotherapy use.
Potential advantages of ctDNA analysis
Genomic profiling using ctDNA offers a significantly less invasive testing approach compared to tumor tissue biopsy. Among patients who are considered unfit to undergo an invasive tumor tissue biopsy due to medical comorbidities or inaccessible tumor location, ctDNA analysis provides an opportunity for biomarker analysis to be performed [5••]. Rates of successful tissue genotyping vary, with significantly lower rates among small tumor tissue samples such as fine needle aspiration. For lung cancers associated with certain oncogenic driver alterations such as EGFR mutations which can progress during initial targeted treatment and may be eligible for subsequent targeted therapies, repeat tissue biopsy is indicated to evaluate for tumor evolution and development of resistance mutations and therefore may present a diagnostic challenge. Re-biopsy is not always considered feasible or safe. For example, a recent study showed that among 69 patients with advanced EGFR mutation-positive NSCLC with disease progression while on EGFR TKI therapy, the feasibility of re-biopsy was dependent on the anatomical site of relapse, with progression of metastatic disease sites assessed as difficult to re-biopsy in a substantial portion of study participants (42% at time of RECIST-progressive disease (PD)) [6]. Use of ctDNA assays would allow for less invasive serial genomic assessments over the course of a patient’s treatment, sparing patients from having to undergo potentially difficult or unsuccessful procedures.
Utilization of ctDNA technology is also a more cost-effective means of performing molecular analysis, as a blood draw is significantly less expensive than an invasive tissue biopsy procedure. Patients may decline invasive tumor tissue biopsies, opting for non-invasive blood testing. Turnaround times for plasma genotyping are also shorter than tissue genotyping [7].
In addition, genetic heterogeneity within different areas of a given tumor as well as between primary and metastatic sites in NSCLC is well described [8,9,10], which may limit the accuracy of a single tumor tissue biopsy in detecting an actionable genomic alteration. Analysis of ctDNA can better address this issue of tumor heterogeneity by allowing for a more comprehensive assessment of a tumor’s molecular landscape.
Shortcomings and challenges of ctDNA analysis
Despite many advantages of ctDNA analysis, several possible issues with use of this technology exist. Molecular analysis using ctDNA cannot provide information on histology; therefore, formal tissue biopsy is required to obtain this information at the time of a lung cancer diagnosis. A positive plasma genotyping result relies on the fact that a tumor is actively shedding ctDNA into the systemic circulation; therefore, false negative results are possible in the setting of low tumor burden or low degree of tumor shedding [11]. Diagnostic accuracy of ctDNA is also influenced by the location of a patient’s metastatic disease. A pooled analysis in EGFR-mutated NSCLC demonstrated that the sensitivity of ctDNA EGFR mutation testing was significantly higher in patients with extrathoracic versus intrathoracic metastases [12].
As a result of improved sensitivity of this technology, testing of ctDNA can have the unintended consequence of identifying alterations which were not being evaluated for. For example, clonal hematopoiesis of indeterminate potential is a phenomenon in which somatic mutations result in a clonal expansion of cells detectable in the peripheral blood among those without a hematologic malignancy [13]; these alterations can be identified through ctDNA testing. This clonal hematopoiesis may be a possible recurrent source of discordance between plasma and tissue genotyping and may lead to false positive plasma genotyping results [14]. Among patients with EGFR-mutant NSCLC, most JAK2 mutations, some TP53 mutations and rare KRAS mutations identified in ctDNA were thought to be derived from clonal hematopoiesis rather than tumor [14]. It is also possible to identify incidental germline mutations among cancer patients who are undergoing ctDNA analysis for somatic mutation evaluation. Among 10,888 unselected patients with advanced malignancy (41% were lung cancers) who underwent ctDNA analysis, 1.4% were found to have suspected hereditary cancer mutations in 11 genes [15]. Identification of unintended germline mutations during ctDNA evaluation requires disclosure and referral for genetic counseling services.
In addition, the quality of ctDNA specimen handling and processing can affect accuracy of the results. In a multicenter study evaluating liquid biopsy workflow optimization in a clinical setting among EGFR-mutated NSCLC patients, longer transit time resulted in increased sample hemolysis and lower temperatures negatively impacted reached assay sensitivity [16].
Methods of ctDNA mutational analysis
Several laboratory techniques have been developed to evaluate for targetable alterations in NSCLC and can be utilized in ctDNA analysis. These methods can be broadly categorized into two groups, which include single gene testing and multiplex testing.
Molecular alterations can be evaluated individually with single gene testing, in which a specific alteration of interest is evaluated for and identified within a relatively small region of DNA. Single gene testing can be performed using polymerase chain reaction (PCR) technology such as quantitative PCR (qPCR) [17, 18]; droplet digital PCR (ddPCR) [19, 20]; and beads, emulsions, amplification, and magnetics (BEAMing) [21]. In contrast to qPCR, ddPCR, and BEAMing techniques also have the ability to provide direct quantification of mutant alleles, known as the variant allele fraction (VAF), which can play an important role in monitoring of a patient’s disease status over time and is particularly attractive in EGFR-mutated NSCLC [20, 21]. Currently, the cobas® EGFR mutation test v2 (Roche Molecular Diagnostics, Branchburg, NJ) is the only FDA-approved test for blood-based molecular analysis available [18], although this test’s sensitivity has already been surpassed by other platforms. Specifically, the sensitivities of this method for the detection of EGFR T790M mutation varied were 41% and 73% in the AURA1 trial [22] and 42% in the AURA17 trial [23], lower than newer advanced technologies.
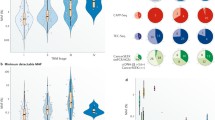
In contrast to single gene evaluation, next-generation sequencing (NGS) or massive parallel sequencing represents a more comprehensive approach to molecular analysis, in which larger regions of multiple genes are evaluated simultaneously using focused malignancy-specific gene panels (Fig. 1) [24]. PCR capture or “hotspot” testing is one NGS approach, which sequences predefined areas of genes and focuses on point mutations within several hotspot regions [5••, 24]. Hybrid capture NGS is an alternative approach in which more extensive gene sequencing is performed and can detect mutations along both introns and exons, insertions, deletions, amplifications, and rearrangements [5••, 24].
While PCR-based approaches typically have more rapid turnaround times, NGS technology can provide information on several different genetic abnormalities simultaneously, allowing for a more comprehensive picture of the genome from a single collected specimen. Important considerations when evaluating and comparing NGS platforms include type of assay (laboratory developed versus commercial), depth of coverage (number of bases, genes, exons, VAF), validation and quality controls, limit of detection, sensitivity/specificity, turnaround time, and costs.
Despite the wide application of this technology in clinical practice, there is no well-defined standard for ctDNA isolation. Blood collection for ctDNA isolation can involve use of either ethylenediaminetetraacetic acid (EDTA) tubes or specialized preservative tubes [5••]. Preservative tubes provide the ability to maintain the quality of small DNA fragments for several days prior to processing, while less expensive and readily available EDTA tubes require immediate processing within 1–2 h of sample collection. Plasma testing is preferred over serum, as sensitivity is higher with plasma-based approaches. Concordances rates between ctDNA and tissue genotyping vary depending on the method of ctDNA analysis and the specific panel used.
In order to formally assess the clinical implications of adding plasma-based ctDNA NGS to tissue-based NGS in the real-world setting, a prospective cohort study of 323 advanced NSCLC patients who underwent plasma ± tissue NGS testing was performed [25••]. Among included patients who underwent both plasma and tumor genotyping, tissue testing alone detected targetable alterations in 20.5% of patients, while the addition of plasma NGS testing increased targetable mutation detection to 35.8% of patients [25••]. Tissue–plasma concordance was 88.9% of patients at initial diagnosis and 70.2% for patients at the time of progression [25••].
Analysis of ctDNA in treatment-naïve NSCLC
Several studies have evaluated the use of baseline plasma ctDNA prior to first-line advanced NSCLC treatment in order to identify targetable alterations. Douillard et al. collected baseline tumor tissue and two plasma ctDNA samples (plasmas 1 and 2) from treatment-naïve EGFR-mutated NSCLC patients prior to gefitinib treatment [26]. Concordance between 652 matched tumor and plasma 1 samples was 94.3%; plasma 1 ctDNA testing yielded a sensitivity of 65.7% and a specificity of 99.8% [26]. There was high concordance in mutation status between 224 matched collected plasma ctDNA samples (96.9%) [26]. This study provided further support for EGFR mutation status assessment using ctDNA.
The BENEFIT trial was a prospective multicenter phase 2 study in which advanced lung adenocarcinoma patients with EGFR mutations detected in plasma ctDNA by ddPCR were treated with the first-generation EGFR TKI gefitinib [27]. Objective response was achieved in 72.1% of patients. Clearance of EGFR mutation in ctDNA occurred at week 8 of gefitinib treatment in 88% of patients with evaluable blood samples, with median overall survival (OS) found to be superior in patients who exhibited EGFR mutation ctDNA clearance compared to those with persistently detected ctDNA [27].
The FLAURA trial is a phase III randomized study evaluating the third-generation EGFR TKI osimertinib versus standard of care EGFR TKI (gefitinib or erlotinib) as first-line treatment for advanced EGFR-mutated NSCLC [28]. Baseline plasma ctDNA samples were collected and analyzed for EGFR mutations via cobas. Good concordance was noted between tissue and plasma genotyping (87% for EGFR exon 19 deletion, 88% for EGFR exon 21 L858R) [28]. Among plasma EGFR-positive patients, osimertinib resulted in decreased risk of progression or death (hazard ratio 0.44, 95% CI 0.34–0.57) [28]. This study supports plasma ctDNA use for determination of eligibility for first-line osimertinib treatment.
Analysis of ctDNA in NSCLC progressing on targeted therapy
Analysis of ctDNA has also proved useful in the setting of advanced EGFR-mutated disease which progressed on EGFR TKI therapy. As mentioned previously, the low sensitivity of cobas testing for the detection of EGFR T790M mutations motivated researchers to explore other detection technologies. One study evaluated serial plasma ctDNA samples with ddPCR to look for T790M mutations in advanced/recurrent EGFR-mutated NSCLC patients treated with EGFR TKI therapy [29]. Of 117 evaluable patients who developed acquired EGFR TKI resistance after initial EGFR TKI therapy, 47% had plasma ctDNA positive for T790M. Plasma T790M was detected a median 2.2 months prior to clinical PD among patients with acquired T790M-mediated TKI resistance and pre-PD plasma samples. In patients receiving second-line or later EGFR TKI treatment, plasma T790M detection was associated with decreased median OS (p = 0.0489) [29].
Ramalingam et al. explored mechanisms of acquired resistance using ctDNA among patients who progressed on first-line osimertinib therapy in the FLAURA trial [30]. Interestingly, the mechanisms of EGFR resistance noted in this population differed from those which typically develop in the setting of first- or second-generation EGFR TKI therapy. No acquired EGFR T790M mutations were identified. MET amplification was the most common acquired resistance mechanism detected, followed by EGFR C797S mutation [30].
Oxnard et al. performed plasma ctDNA genotyping with BEAMing prior to treatment with osimertinib in patients with a common EGFR-sensitizing mutation and acquired EGFR TKI resistance [31]. In patients with T790M-positive tumor testing, the sensitivity of plasma genotyping for T790M was 70%. Plasma T790M was detected in 31% of patients with T790M-negative tumors. Comparable overall response rates (ORR) or median progression-free survival (PFS) among patients with plasma T790M-positive and tumor T790M-positive results [31•]. The authors proposed a paradigm in which plasma and tumor genotyping have complementary roles for T790M testing in advanced EGFR-mutated NSCLC. Those who progress on EGFR TKI therapy can first undergo plasma genotyping, with initiation of osimertinib if plasma T790M mutation is detected; tissue biopsy would be performed only if plasma T790M testing returns negative [31•].
ALK rearrangements and resistance mutations can also be identified through ctDNA analysis. Longitudinal plasma ctDNA analysis was performed using hybrid capture NGS among 22 ALK-positive NSCLC patients who developed acquired resistance to ALK TKIs [32]. Plasma genotyping identified an ALK fusion in 86% of included patients and identified an ALK resistance mutation in 50% of patients. There was high concordance between plasma and matched tissue genotyping. Changes in plasma ALK mutation status and mutational allelic frequency were noted during longitudinal analysis [32]. This study supports further exploration of plasma genotyping in ALK-positive NSCLC. Furthermore, the identification of new ALK mutations such as G1202R, F1174, and L1196M as a mechanism of resistance plays an important role in treatment selection for ALK-positive PD [33]; these alterations can be identified through plasma genotyping.
Current recommendations
Several practice guidelines and professional societies have provided statements or recommendations regarding ctDNA integration into clinical practice. The most recent National Comprehensive Cancer Network (NCCN) practice guidelines state that ctDNA use can be considered if a patient is medically unfit for invasive tissue sampling or in the initial diagnostic setting after pathologic confirmation of NSCLC on tissue biopsy if there is insufficient tissue to perform molecular testing; a caveat is that follow-up tissue testing should be planned for all patients in which an oncogenic driver mutation is not identified [4]. The American Society of Clinical Oncology (ASCO) 2017 Clinical Cancer Advances Report stated that among cancer patients who are unable to undergo tumor tissue biopsy, liquid biopsy may be “safer, quicker and more convenient--and perhaps even more informative” [34], providing support for continued evaluation of this technology in the future. The Updated Molecular Testing Guideline for the selection of lung cancer patients for treatment with targeted TKIs published by the College of American Pathologists (CAP), the International Association for the Study of Lung Cancer (IASLC), and the Association for Molecular Pathology (AMP) advocated for the use of “cell-free DNA to “rule in” targetable mutations” in situations where tumor tissue is difficult to obtain or limited [35].
Additionally, IASLC released a statement paper on use of liquid biopsy for Advanced NSCLC in 2018 [5••]; a multidisciplinary panel of experts in thoracic oncology concluded that use of liquid biopsy has “significant potential to improve patient care, and immediate implementation in the clinic is justified in a number of therapeutic settings relevant to NSCLC” [5••]. In the advanced treatment naïve NSCLC setting, IASLC consensus recommendation is to perform molecular analysis on ctDNA (preferably using NGS) if tumor biopsy specimen is insufficient for molecular testing; if a targetable alteration is identified, one should treat with standard of care therapy for that alteration; if a targetable alteration is not identified on ctDNA analysis, tissue re-biopsy should be performed [5••]. In NSCLC patients with progressive or recurrent disease during TKI therapy, IASLC recommendation is to perform molecular analysis on ctDNA (preferably cobas/ddPCR for EGFR mutations, NGS for ALK and ROS1 alterations); if a targetable resistance mutation is identified, treatment with standard of care therapy targeting this mutation is indicated; and if a targetable resistance mutation is not identified, then tissue re-biopsy should be performed if feasible [5••].
Future directions
Detection of minimal residual disease
Recent research efforts evaluated ctDNA analysis as a tool for detection of minimal residual disease (MRD) among treated early-stage NSCLC patients. One group of investigators used multiplex exome sequencing of primary tumor tissue to develop an individualized panel of single nucleotide variants for each included patient and then monitored ctDNA from those patients longitudinally [36]. Among 24 patients in subgroup analysis, 13 of 14 patients (93%) with relapsed disease had detectable ctDNA before or at the time of disease relapse. The median interval between ctDNA detection and radiographic NSCLC relapse was 70 days [36]. Another study investigated the reliability of deep sequencing (CAPP-seq) ctDNA from 40 patients with stage I–III lung cancer after treatment with curative intent therapies [37]. There was ctDNA detected in the first posttreatment blood sample of 94% of evaluable patients experiencing disease recurrence. Detection of ctDNA in posttreatment blood samples preceded radiographic progression in 72% of patients and by a median of 5.2 months [37]. Evaluation of MRD using plasma ctDNA is an important area of future investigation, as detection of ctDNA after definitive treatment of early-stage disease can help predict for disease recurrence. One limitation of MRD evaluation is the capacity of current technology to identify ctDNA in early-stage NSCLC. The low VAF required to identify MRD is far below the detection of many current technologies [38].
Evaluation of predictive biomarkers for immunotherapy use
Plasma ctDNA analysis has also been utilized to evaluate predictive biomarkers for immunotherapy use. Retrospective evaluation of the relationship between blood-based tumor mutational burden (bTMB) and clinical outcomes with atezolizumab in advanced NSCLC was performed utilizing baseline plasma samples collected from the intention-to-treat populations of the POPLAR and OAK trials [39]. Among 583 patients with sufficient ctDNA for analysis in the OAK trial, the prevalence of high bTMB (defined as bTMB ≥ 16) was 27%. High bTMB patients had improved PFS benefit from atezolizumab versus docetaxel (HR 0.65, 95% CI 0.47–0.92, p = 0.013); the interaction between high bTMB score and treatment was statistically significant for PFS. High bTMB patients had a median OS of 13.5 months with atezolizumab versus 6.8 months with docetaxel [39]. The study also demonstrated that bTMB is not associated with high PD-L1 expression and independently predicts for PFS benefit [39].
The B-F1RST trial is a prospective study evaluating bTMB as a predictive biomarker for immunotherapy use in NSCLC. This trial sought to evaluate the clinical utility of bTMB as a predictive biomarker for the use of the PD-L1 inhibitor atezolizumab in first-line advanced NSCLC treatment [40]. Among 119 included patients with adequate ctDNA, the ORR was 28.6% in high bTMB patients (defined as bTMB ≥ 16) and 4.4% in low bTMB patients; median PFS was 4.6 months in high bTMB patients and 3.7 months in low bTMB patients, although not statistically significant [40]. These trials highlight the need for future studies to explore bTMB and other predictive biomarkers for immunotherapy use in NSCLC. One of the major issues of TMB (in tissue and plasma) is the lack of standardization for its detection and lack of consensus regarding an appropriate cutoff of mutations/Mb. For this reason, this biomarker is not yet appropriate for wide use in clinical practice. Several other biomarkers such as VAF determination, “ctDNA velocity” during immunotherapy treatment, or hypermutated ctDNA have been studied as potential predictive tools to evaluate the response to immune checkpoint inhibitors in cancer patients [41, 42].
Evaluation of ctDNA in non-blood body fluids in NSCLC
Although ctDNA is often performed on plasma, some research has evaluated ctDNA testing in other body fluids such as urine or cerebrospinal fluid (CSF). EGFR genotyping of matched urine, plasma, and tumor tissue specimens was performed in advanced EGFR-mutated NSCLC patients enrolled in the TIGER-X trial and treated with the EGFR TKI rociletinib [43]. Using tissue as a reference, EGFR T790M resistance mutation testing in urine samples of recommended volume of 90–100 ml had sensitivity of 93% and specificity of 96%, whereas plasma testing had sensitivity of 93% and specificity of 94% [43]. Another group of investigators postulated that dynamic changes in EGFR mutation levels within urine ctDNA could be indicative of tumor response in patients receiving second-line osimertinib therapy [44]. Urine ctDNA was analyzed daily, with an overall trend of spikes in ctDNA levels noted during the first week of treatment followed by subsequent overall decrease in ctDNA levels by day 7 [44]. Serial monitoring of urine ctDNA may be explored further as a means to evaluate patient responsiveness to treatment.
Analysis of ctDNA within the CSF has also been investigated. NGS was performed on matched plasma and CSF ctDNA of 40 NSCLC patients with suspected leptomeningeal disease [45]. ctDNA was detected in 93.8% of CSF and 66.7% of plasma. CSF ctDNA analysis identified several copy number variations which were not found in plasma, and the average maximum allelic fraction of CSF ctDNA was significantly higher than in plasma (56.7% versus 4.4%) [45]. This study demonstrated the potential utility of CSF in liquid biopsy in the future.
Conclusion
Liquid biopsy including plasma ctDNA analysis has emerged as a promising new tool in the diagnosis and monitoring of lung cancers, specifically in NSCLC. Although genomic profiling in NSCLC has been most extensively studied in the advanced NSCLC setting, newer innovative approaches are being evaluated including utilization of plasma ctDNA as a tool to identify minimal residual disease in early-stage NSCLC and as a predictive biomarker for immunotherapy use. Plasma ctDNA can provide an important adjunct to tissue-based genotyping in clinical practice, and further research efforts should investigate novel uses of this emerging technology.
References and Recommended Reading
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
Chan BA, Hughes BG. Targeted therapy for non-small cell lung cancer: current standards and the promise of the future. Transl Lung Cancer Res. 2015;4(1):36–54. https://doi.org/10.3978/j.issn.2218-6751.2014.05.01.
Lynch TJ, Bell DW, Sordella R, Gurubhagavatula S, Okimoto RA, Brannigan BW, et al. Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N Engl J Med. 2004;350(21):2129–39.
Paez JG, Janne PA, Lee JC, Tracy S, Greulich H, Gabriel S, et al. EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science. 2004;304(5676):1497–500.
National Comprehensive Cancer Network. Non-small cell lung cancer (Version 2.2019). https://www.nccn.org/professionals/physician_gls/pdf/nscl.pdf. Accessed 9 Jan 2019.
•• Rolfo C, Mack PC, Scagliotti GV, Baas P, Barlesi F, Bivona TG, et al. Liquid biopsy for advanced non-small cell lung cancer (NSCLC): a statement paper from the IALSC. J Thorac Oncol. 2018;13(9):1248–68. https://doi.org/10.1016/j.jtho.2018.05.030 Evidence-based recommendations regarding the use of liquid biopsy in NSCLC provided by a multidisciplinary panel of experts in thoracic oncology.
Uozu S, Imaizumi K, Yamaguchi T, Goto Y, Kawada K, Minezawa T, et al. Feasibility of tissue re-biopsy in non-small cell lung cancers resistant to previous epidermal growth factor receptor tyrosine kinase inhibitor therapies. BMC Pulm Med. 2017;17(1):175. https://doi.org/10.1186/s12890-017-0514-3.
Sacher AG, Paweletz C, Dahlberg SE, Alden RS, O’Connell A, Feeney N, et al. Prospective validation of rapid plasma genotyping for the detection of EGFR and KRAS mutations in advanced lung cancer. JAMA Oncol. 2016;2(8):1014–22. https://doi.org/10.1001/jamaoncol.2016.0173.
Zhang J, Fujimoto J, Zhang J, Wedge DC, Song X, Zhang J, et al. Intratumor heterogeneity in localized lung adenocarcinomas delineated by multiregion sequencing. Science. 2014;346(6206):256–9. https://doi.org/10.1126/science.1256930.
Kim EY, Cho EN, Park HS, Kim A, Hong JY, Lim S, et al. Genetic heterogeneity of actionable genes between primary and metastatic tumor in lung adenocarcinoma. BMC Cancer. 2016;16:27. https://doi.org/10.1186/s12885-016-2049-z.
Han HS, Eom DW, Kim JH, Kim KH, Shin HM, An JY, et al. EGFR mutation status in primary lung adenocarcinomas and corresponding metastatic lesions: discordance in pleural metastases. Clin Lung Cancer. 2011;12(6):380–6. https://doi.org/10.1016/j.cllc.2011.02.006.
Sacher AG, Komatsubara KM, Oxnard GR. Application of plasma genotyping technologies in non-small cell lung cancer: a practical review. J Thorac Oncol. 2017;12(9):1344–56. https://doi.org/10.1016/j.jtho.2017.05.022.
Passiglia F, Rizzo S, Rolfo C, Galvano A, Bronte E, Incorvaia L, et al. Metastatic site location influences the diagnostic accuracy of ctDNA EGFR-mutation testing in NSCLC patients: a pooled analysis. Curr Cancer Drug Targets. 2018;18(7):697–705. https://doi.org/10.2174/1568009618666180308125110.
Boettcher S, Ebert BL. Clonal hematopoiesis of indeterminate potential. J Clin Oncol. 2018:JCO2018793588;37:419–22. https://doi.org/10.1200/JCO.2018.79.3588.
Hu Y, Ulrich BC, Supplee J, Kuang Y, Lizotte PH, Feeney NB, et al. False-positive plasma genotyping due to clonal hematopoiesis. Clin Cancer Res. 2018;24(18):4437–43. https://doi.org/10.1158/1078-0432.CCR-18-0143.
Slavin TP, Banks KC, Chudova D, Oxnard GR, Odegaard JI, Nagy RJ, et al. Identification of incidental germline mutations in patients with advanced solid tumors who underwent cell-free circulating tumor DNA sequencing. J Clin Oncol. 2018:JCO1800328;36:3459–65. https://doi.org/10.1200/JCO.18.00328.
Sorber L, Zwaenepoel K, De Winne K, Van Casteren K, Augustus E, Jacobs J, et al. A multicenter study to assess EGFR mutational status in plasma: focus on an optimized workflow for liquid biopsy in a clinical setting. Cancers (Basel). 2018;10(9):E290. https://doi.org/10.3390/cancers10090290.
Wu YL, Lee V, Liam CK, Lu S, Park K, Srimuninnimit V, et al. Clinical utility of a blood-based EGFR mutation test in patients receiving first-line erlotinib therapy in the ENSURE, FASTACT-2, and ASPIRATION studies. Lung Cancer. 2018;126:1–8. https://doi.org/10.1016/j.lungcan.2018.10.004.
Malapelle U, Sirera R, Jantus-Lewintre E, Reclusa P, Calabuig-Farinas S, Blasco A, et al. Profile of the Roche cobas® EGFR mutation test v2 for non-small cell lung cancer. Expert Rev Mol Diagn. 2017;17(3):209–15. https://doi.org/10.1080/14737159.2017.1288568.
Oxnard GR, Paweletz CP, Kuang Y, Mach SL, O’Connell A, Messineo MM, et al. Noninvasive detection of response and resistance in EGFR-mutated lung cancer using quantitative next-generation genotyping of cell-free plasma DNA. Clin Cancer Res. 2014;20(6):1698–705. https://doi.org/10.1158/1078-0432.CCR-13-2482.
Zhang R, Chen B, Tong X, Wang Y, Wang C, Jin J, et al. Diagnostic accuracy of droplet digital PCR for detection of EGFR T790M mutation in circulating tumor DNA. Cancer Manag Res. 2018;10:1209–18. https://doi.org/10.2147/CMAR.S161382.
Taniguchi K, Uchida J, Nishino K, Kumagai T, Okuyama T, Okami J, et al. Quantitative detection of EGFR mutations in circulating tumor DNA derived from lung adenocarcinomas. Clin Cancer Res. 2011;17(24):7808–15. https://doi.org/10.1158/1078-0432.CCR-11-1712.
Thress KS, Brant R, Carr TH, Dearden S, Jenkins S, Brown H, et al. EGFR mutation detection in ctDNA from NSCLC patient plasma: a cross-platform comparison of leading technologies to support the clinical development of AZD9291. Lung Cancer. 2015;90(3):509–15. https://doi.org/10.1016/j.lungcan.2015.10.004.
Zhou C, Wang M, Cheng Y, Chen Y, Ye X, Sun Y, et al. Detection of EGFR T790M in Asia-Pacific patients (pts) with EGFR mutation-positive advanced non-small cell lung cancer (NSCLC): circulating tumour (ct) DNA analysis across 3 platforms. Ann Oncol. 2017;28(suppl_5):v460–96.
Rozenblum AB, Ilouze M, Dudnik E, Dvir A, Soussan-Gutman L, Geva S, et al. Clinical impact of hybrid capture-based next-generation sequencing on changes in treatment decisions in lung cancer. J Thorac Oncol. 2017;12(2):258–68. https://doi.org/10.1016/j.jtho.2016.10.021.
Aggarwal C, Thompson JC, Black TA, Katz SI, Fan R, Yee SS, et al. Clinical implications of plasma-based genotyping with the delivery of personalized therapy in metastatic non-small cell lung cancer. JAMA Oncol. 2018. https://doi.org/10.1001/jamaoncol.2018.4305 Prospective cohort study of advanced NSCLC patients demonstrating that plasma-based NGS testing increased detection of targetable alterations and improved delivery of targeted therapy.
Douillard JY, Ostoros G, Cobo M, Ciuleanu T, Cole R, McWalter G, et al. Gefitinib treatment in EGFR mutated caucasian NSCLC: circulating-free tumor DNA as a surrogate for determination of EGFR status. J Thorac Oncol. 2014;9(9):1345–53. https://doi.org/10.1097/JTO.0000000000000263.
Wang Z, Cheng Y, An T, Gao H, Wang K, Zhou Q, et al. Detection of EGFR mutations in plasma circulating tumour DNA as a selection criterion for first-line gefitinib treatment in patients with advanced lung adenocarcinoma (BENEFIT): a phase 2, single-arm, multicentre clinical trial. Lancet Respir Med. 2018;6(9):681–90. https://doi.org/10.1016/S2213-2600(18)30264-9.
Gray J, Okamoto I, Sriuranpong V, Vansteenkiste J, Imamura F, Lee JS, et al. Osimertinib vs SoC EGFR-TKI as first-line treatment in patients with EGFRm advanced NSCLC (FLAURA): plasma ctDNA analysis. J Thorac Oncol. 2017;12(11):S1754–5. https://doi.org/10.1016/j.jtho.2017.09.348.
Zheng D, Ye X, Zhang MZ, Sun Y, Wang JY, Ni J, et al. Plasma EGFR T790M ctDNA status is associated with clinical outcome in advanced NSCLC patients with acquired EGFR-TKI resistance. Sci Rep. 2016;6:20913. https://doi.org/10.1038/srep20913.
Ramalingam SS, Cheng Y, Zhou C, Ohe Y, Imamura F, Cho BC, et al. Mechanisms of acquired resistance to first-line osimertinib: preliminary data from the phase III FLAURA study. Ann Oncol. 2018;29(suppl_9):ix173–8.
• Oxnard GR, Thress KS, Alden RS, Lawrance R, Paweletz CP, Cantarini M, et al. Association between plasma genotyping and outcomes of treatment with osimertinib (AZD9291) in advanced non-small-cell lung cancer. J Clin Oncol. 2016;34(28):3375–82. https://doi.org/10.1200/JCO.2016.66.7162 Retrospective study of EGFR-mutated NSCLC patients with acquired EGFR-TKI resistance, demonstrating comparable ORR and PFS among those with plasma T790M-positive and tumor T790M-positive results.
Dagogo-Jack I, Brannon AR, Ferris LA, Campbell CD, Lin JJ, Schultz KR, et al. Tracking the evolution of resistance to ALK tyrosine kinase inhibitors through longitudinal analysis of circulating tumor DNA. JCO Precis Oncol. 2018;2018:1–14. https://doi.org/10.1200/PO.17.00160.
Shaw AT, Martini JF, Besse B, Bauer TM, Lin CC, Soo RA, et al. Efficacy of lorlatinib in patients (pts) with advanced ALK-positive non-small cell lung cancer (NSCLC) and ALK kinase domain mutations. Cancer Res. 2018;78(13 Suppl):Abstract nr CT044.
Burstein HJ, Krilov L, Aragon-Ching JB, Baxter NN, Chiorean EG, Chow WA, et al. Clinical cancer advances 2017: annual report on progress against cancer from the American Society of Clinical Oncology. J Clin Oncol. 2017;35(12):1341–67. https://doi.org/10.1200/JCO.2016.71.5292.
Lindeman NI, Cagle PT, Aisner DL, Arcila ME, Beasley MB, Bernicker EH, et al. Updated molecular testing guideline for the selection of lung cancer patients for treatment with targeted tyrosine kinase inhibitors: guideline from the College of American Pathologists, the International Association for the Study of Lung Cancer, and the Association for Molecular Pathology. Arch Pathol Lab Med. 2018;142(3):321–46. https://doi.org/10.5858/arpa.2017-0388-CP.
Abbosh C, Birkbak NJ, Wilson GA, Jamal-Hanjani M, Constantin T, Salari R, et al. Phylogenetic ctDNA analysis depicts early-stage lung cancer evolution. Nature. 2017;545(7655):446–51. https://doi.org/10.1038/nature22364.
Chaudhuri AA, Chabon JJ, Lovejoy AF, Newman AM, Stehr H, Azad TD, et al. Early detection of molecular residual disease in localized lung cancer by circulating tumor DNA profiling. Cancer Discov. 2017;7(12):1394–403. https://doi.org/10.1158/2159-8290.CD-17-0716.
Abbosh C, Birkbak NJ, Swanton C. Early stage NSCLC – challenges to implementing ctDNA-based screening and MRD detection. Nat Rev Clin Oncol. 2018;15(9):577–86. https://doi.org/10.1038/s41571-018-0058-3.
Gandara DR, Paul SM, Kowanetz M, Schleifman E, Zou W, Li Y, et al. Blood-based tumor mutational burden as a predictor of clinical benefit in non-small-cell lung cancer patients treated with atezolizumab. Nat Med. 2018;24(9):1441–8. https://doi.org/10.1038/s41591-018-0134-3.
Kim ES, Velcheti V, Mekhail T, Leal TA, Dowell JE, Tsai ML, et al. Primary efficacy results from B-F1RST, a prospective phase II trial evaluating blood-based tumour mutational burden (bTMB) as a predictive biomarker for atezolizumab (atezo) in 1L non-small cell lung cancer (NSCLC). Ann Oncol. 2018;29(suppl_5):Abstr LBA55.
Raja R, Kuziora M, Brohawn PZ, Higgs BW, Gupta A, Dennis PA, et al. Early reduction in ctDNA predicts survival in patients with lung and bladder cancer treated with durvalumab. Clin Cancer Res. 2018;24(24):6212–22. https://doi.org/10.1158/1078-0432.CCR-18-0386.
Khagi Y, Goodman AM, Daniels GA, Patel SP, Sacco AG, Randall JM, et al. Hypermutated circulating tumor DNA: correlation with response to checkpoint inhibitor-based immunotherapy. Clin Cancer Res. 2017;23(19):5729–36. https://doi.org/10.1158/1078-0432.CCR-17-1439.
Reckamp KL, Melnikova VO, Karlovich C, Sequist LV, Camidge DR, Wakelee H, et al. A highly sensitive and quantitative test platform for detection of NSCLC EGFR mutations in urine and plasma. J Thorac Oncol. 2016;11(10):1690–700. https://doi.org/10.1016/j.jtho.2016.05.035.
Husain H, Melnikova VO, Kosco K, Woodward B, More S, Pingle SC, et al. Monitoring daily dynamics of early tumor response to targeted therapy by detecting circulating tumor DNA in urine. Clin Cancer Res. 2017;23(16):4716–23. https://doi.org/10.1158/1078-0432.CCR-17-0454.
Jiang BY, Yangsi LI, Chuai S, Zhang Z, Yang JJ, Zhong W, et al. NGS to reveal heterogeneity between cerebrospinal fluid and plasma ctDNA among non-small cell lung cancer patients with leptomeningeal carcinomatosis. J Clin Oncol. 2017;35(15):suppl:9022–9022. https://doi.org/10.1200/JCO.2017.35.15_suppl.9022.
Acknowledgements
The authors thank Joseph Pinto, PhD, for his cooperation in the creation of the figure.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Katherine A. Scilla declares that she has no conflict of interest.
Christian Rolfo has received speaker’s honoraria from Guardant Health, MSD, and Novartis; has received non-financial support from OncoDNA (exosomes research collaboration) and Mylan (scientific advisor); and is currently Vice President of the International Society of Liquid Biopsy (ISLB).
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Lung Cancer
Rights and permissions
About this article
Cite this article
Scilla, K.A., Rolfo, C. The Role of Circulating Tumor DNA in Lung Cancer: Mutational Analysis, Diagnosis, and Surveillance Now and into the Future. Curr. Treat. Options in Oncol. 20, 61 (2019). https://doi.org/10.1007/s11864-019-0653-2
Published:
DOI: https://doi.org/10.1007/s11864-019-0653-2