Abstract
The wild type (Wt) and mutant form of yeast (sphingolipid compensation) genes, SLC1 and SLC1-1, have been shown to have lysophosphatidic acid acyltransferase (LPAT) activities (Nageic et al. in J Biol Chem 269:22156–22163, 1993). Expression of these LPAT genes was reported to increase oil content in transgenic Arabidopsis and Brassica napus. It is of interest to determine if the TAG content increase would also be seen in soybeans. Therefore, the wild type SLC1 was expressed in soybean somatic embryos under the control of seed specific phaseolin promoter. Some transgenic somatic embryos and in both T2 and T3 transgenic seeds showed higher oil contents. Compared to controls, the average increase in triglyceride values went up by 1.5% in transgenic somatic embryos. A maximum of 3.2% increase in seed oil content was observed in a T3 line. Expression of the yeast Wt LPAT gene did not alter the fatty acid composition of the seed oil.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Different plant species vary greatly in seed storage reserves with some grain seeds having >85% starch, while 50–70% oil in oil seeds, and legumes such as soybeans with 40% protein and 20% oil of the seed dry weight [2]. Soybeans are the largest oil seed crop in the world followed by Brassica, peanut, sunflower and cotton seeds [3, 4]. There is a significant amount of vegetable oil that is used for industrial purposes. For example, of the ~380 million tons of oilseeds produced in 2005, about 85% of the extracted oil is used for food and cooking applications, while nearly 15% is put to use in industry for, among others, lubricants, inks, coatings, plasticizers and biodiesel [5]. Since oil production has such tremendous importance, conventional and genetic engineering strategies are being applied to improve oil quantity and quality in oil seed crops [6].
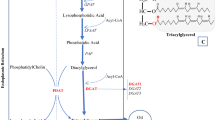
The Kennedy pathway is the major TAG pathway of plants [7, 8] with the three acyltransferases the glycerol-3-phosphate acyltransferase (GPAT, EC 2.3.1.15), the lyso-phosphatidic acid acyltransferase (LPAT, EC 2.3.1.51) and the diacylglycerol acyltransferase (DGAT, EC 2.3.1.20) operating in a sequential manner. The other enzymes that directly participate in the DAG to TAG conversion are DAG transacylase and phospholipid-DAGAT [9]. Over-expression of Kennedy pathway enzymes, GPAT [10]; yeast LPAT [11, 12]; DAGAT [13, 14] have reportedly increased oil content in plants and yeast. Maize high oil QTL (Qho6) encodes a mutated DAGAT1-2 allele [15]. Besides the Kennedy pathway, other pathway enzymes like glucose metabolism enzymes like glycerol-3-phosphate dehydrogenase (GPDH, EC 1.1.1.8) [16] and Dof-type transcription factors [17] were also implicated in the seed oil increase. Arabidopsis transcriptional factor FUSCA3 (FUS3) was implicated in the increased oil content in Arabidopsis seeds [18].
Saccharomyces cerevisiae sphingolipid suppressor gene, SLC1-1 and the wild type SLC1 gene proved to be acyltransferases with LPAT activity [1, 11]. Transforming plants with wild type (SLC1) and mutated (SLC1-1) forms of yeast genes resulted in increased triglyceride levels [11, 12, 19]. So far there are no reports on soybean seed oil enhancement except the high oil cultivar N88–480 developed by [20] through conventional breeding though several strategies were successfully reported for seed oil increase in plants [10, 14–16, 21]. Soybean seed expressing a diacylglycerol acyltransferase 2A from the soil fungus Umbelopsis (formerly Mortierella) ramanniana showed a 1.5% increase in oil levels [22].
An important method of soybean regeneration is somatic embryogenesis. Embryogenic tissues can be proliferated by subculture on a solid proliferation (MSD20) medium or a liquid suspension culture medium [23, 24]. Through somatic embryogenesis, genetic engineering of soybean has proved to be a powerful technique for improving seed compositions including the oil for enhanced edible and industrial purposes [25–27]. Furthermore, somatic embryo proliferation can result in a higher number of somatic embryos [24] thus probably increasing the rate of recovery. Another potential advantage of the somatic embryos system is that they are good targets in many cases of seed specific traits since they can be analyzed at the mature soybean somatic embryo stage prior to the zygotic embryonic stage, thus saving labor and time [28–33].
Here in this paper we report the generation of transgenic soybean somatic embryos (SS embryos) expressing the yeast SLC1 gene with increased total lipids. We also report the generation of fertile transgenic soybean plants with seeds showing increased oil content.
Materials and Methods
Plant Material
Soybeans [Glycine max (L.) Merrill cv. ‘Jack’] were grown in a greenhouse at the University of Kentucky, Lexington under a 16 h photoperiod at 35 °C day and 25 °C night temperature. Pods with immature seeds were surface sterilized by immersing for 30 s in 70% 2-isopropyl alcohol followed by a 10 min immersion in 25% bleach solution (with 1.5% final hypochlorite concentration) with a few drops of Liquinox (detergent). The pods were then rinsed three times in sterile water for 5 min each time. Immature seeds 3–6 mm in length were removed from the pods. The end containing the embryonic axis was cut off and discarded. Then the two cotyledons were pushed out from the seed coat, separated and placed with abaxial side (round side) down on MSD40 (Murashige and Skoog salts, 40 mg/l 2,4-dichlorophenoxyacetic acid, B5 vitamins, sucrose (3% wt/vol) with 0.2% gelrite) medium [34]. Cultures were then incubated at 25 °C at a 23 h photoperiod (low light intensity, 5–10 µE). Globular staged somatic embryo clusters were harvested from the explant tissues 4–6 weeks after induction and then placed on MSD20 (MS salts, B5 vitamins, sucrose (3% wt/vol), 20 mg/l 2,4-D with 0.2% gelrite) solid medium for a period of 1 month for proliferation. Embryogenic tissues were then transferred from MSD20 to FNL (Finer and Nagasawa “lite”) “liquid medium” [24] for further proliferation. Suspension cultures were agitated at 100 rpm and maintained with a 2 week subculture period at 25 °C with a 23 h photoperiod. The T1 and T2 generation transgenic plants were grown in the green house under the same conditions as described above for the Jack control plants.
Vector Construction
The amplified SLC1 gene product was cloned into the PGEM T-vector. The product was sequenced using T7 and SP6 vector sequencing primers to confirm the sequence of the ligated product. A 0.9-kbp NcoI/HpaI T-Vector digested SLC1 fragment was cloned into a pPHI472 vector [29] digested with the same enzymes to put the SLC1 gene under the seed specific phaseolin promoter and phaseolin terminator. The whole cassette was digested with EcoRI/PstI and ligated to a pCAMBIA 1201 (Genbank accession number AF234293) plant transformation vector digested with the same restriction enzymes. The pCAMBIA 1201 binary vector has a chloramphenicol resistance gene for bacterial selection and a hygromycin resistance gene for plant selection. The GUS reporter gene was driven by the constitutive CaMV 35S promoter.
Microprojectile Bombardment
Green embryo clumps were slightly pressed with a spatula to partially separate them and were placed in the center of a moist filter paper in sterile petri plates (approximately 100–150 mg of somatic embryos per plate) and partially desiccated in a laminar flow hood for 15 min prior to bombardment. Transformation was carried out via particle bombardment with a gene gun (Dupont PDS1000; Bio-Rad Laboratories, Hercules, and CA) by gold/DNA microprojectile preparations as described by [34]. Briefly, for 9 shots, 25 µg of plasmid DNA was used to coat 7.5 mg of 0.6 µm gold particles. Cultures were bombarded at 10,687 kPa (1,550 psi) helium gas pressure under 91 kPa (27 in) of Hg vacuum, at a shooting distance of 11 cm from the rupture disk to the target tissue. Immediately after bombardment, embryogenic cultures were placed on D20 proliferation media without any selective agent for 7 days.
Selection and Regeneration of Transformants
Bombarded globular SS embryos that were cultured on D20 were transferred (approximately 100 clumps of 0.3–0.4 cm diameter per plate) to FN Lite medium Samoylov et al. [35] containing 25 mg/l hygromycin. After 3–4 weeks, visibly growing clumps were moved to fresh selection medium. After approximately 12 weeks, green looking hygromycin resistant SS embryo clumps were transferred from FN Lite into 500-ml Erlenmeyer flasks containing 100 ml of liquid MS0 (MS salts, B5 vitamins and 3% sucrose) medium as described by Samoylov et al. [24]. Prior to transfer into flasks, each embryogenic cluster was gently pressed with a spatula to partially separate the individual globular-stage embryos. At 4 weeks, the resulting cotyledon-stage embryos were analyzed for GUS staining (β-glucuronidase) and RNA isolation. Some of the embryos were desiccated as described in [24]. Ten to twelve matured embryos were placed in a 100 × 15 mm Petri dish and sealed with Nescofilm. To allow gradual desiccation of embryos over a period of 5–7 days, a small piece (approximately 1 cm3) of solid MS0 medium (MSO medium with 0.2% Gelrite) was placed in the middle of the plate away from the embryos. Desiccated embryos were germinated on 1/5th MS medium (1/5th concentration of MS salts, B5 vitamins, 0.4% Gelrite without sucrose).
Yeast DNA Isolation and SLC1 Gene Amplification
A pellet from a 10-ml culture of Yeast strain InVsc1 (Invitrogen, Carlsbad, CA) was homogenized with mortar and pestle. Total DNA was isolated from the homogenized yeast following the procedure described by [36]. The SLC1 gene [1] was directly amplified from yeast genomic DNA since no introns sequences were found. With 100 ng of yeast genomic DNA as template, PCR was done to amplify the SLC1 gene using the primers 5′-CCATGGATGAGTGTGATAGGTAGGTTC-3′ and 5′-GTTAACAATGCATCTTTTTTACAGATGA-3′ for the sense and antisense strand, respectively. NcoI restriction site was added to the forward primer while the reverse primer was designed with a HpaI site. The PCR conditions were 94 °C for 2 min; 30 cycles at 94 °C, 30 s; 55 °C, 30 s; 72 °C, 1 min and a final extension at 72 °C for 8 min.
Plant DNA Isolation
Genomic DNA was isolated from soybean leaves and embryos as described by Reddy et al. [36]. Screening of transgenic soybean lines was done by PCR of the SLC1 gene using 200 ng of the genomic DNA in 50-µl reactions. The primers and the PCR conditions for SLC1 gene amplification were as described above.
Southern Blotting
Five micrograms of DNA was digested overnight with EcoRI, fractionated on a 0.8% agarose gel, and blotted onto a Zetaprobe membrane (Bio-Rad). Hybridization was done overnight at 42 °C in a hybridization solution containing 50% formamide, 0.12 M Na2HPO4, 0.25 M NaCl, and 1 mM EDTA with a SLC1 gene-specific fragment random prime labeled with [α32P]-dCTP (Prime-It II Random Primer Labeling Kit; Stratagene, La Jolla, CA) as a probe. The membrane was washed three times at room temperature in 0.1 × SSC (1 × SSC is 0.15 M NaCl plus 0.015 M sodium citrate) and 0.1% SDS and exposed in a phosphorImager cassette (Molecular Dynamics, Sunnyvale, CA). The intensity of hybridized DNA bands was estimated using the ImageQuant software program (Molecular Dynamics).
RT-PCR
Total RNA was extracted from the hygromycin resistant matured soybean somatic embryos using the Trizol reagent as advised by the manufacturer (Invitrogen Corporation, Carlsbad, USA). Using the isolated total RNA as a template, reverse transcription was performed for the synthesis of the first-strand cDNA using oligo dT as prescribed by manufacturers of the Kit (Sigma-Aldrich Corporation, St. Louis, MO, USA) in a 20-µl reaction at 48 °C for 45 min. A 2-µl aliquot of the RT reaction was used in a 50-µl PCR reaction with the following profile: 94 °C for 2 min; 30 cycles at 94 °C, 30 s; 55 °C, 30 s; 72 °C, 1 min and a final extension at 72 °C for 8 min. The primers used were the same as in the PCR reaction described for SLC1 gene amplification.
Lipid Analysis
Lipids were isolated as described in [37]. Total lipids were extracted independently from seed chips (~2 mg) cut from ten different matured seeds for each line. For fatty acid analysis of matured somatic embryos ten individual 5 week old (matured on 6% maltose containing MS medium plates) somatic embryos of a clone were used. The somatic embryos were desiccated before lyophilization. To allow gradual desiccation of embryos over a period of 5 days a small piece (approximately 1 cm3) of solid MS0 medium was placed in the middle of the plate away from the embryos. The desiccated embryos were freeze dried and weighed. TAG (triacylglycerol) contents were calculated from the addition of known amounts of tri -17:0 (1,2,3-Triheptadecanoylglycerol, Sigma-Aldrich Corporation, St. Louis, MO, USA) internal standard added to the somatic embryos and seed chips before lipids were extracted. The lipids were methylated with 0.5 ml of sodium methoxide (4.2%, wt/vol) with shaking at 800 rpm for 45 min. The fatty acid methyl esters were extracted with 1 ml of hexane twice. The hexane extracts were combined and then washed with 1 ml of 0.9% KCl. The fatty acid methyl esters in hexane were analyzed by gas chromatography (Hewlett Packard 5890 with a flame ionization detector) on FFAP column of 14 m × 0.25 mm, 0.33 µm film thickness. The temperature program started from 140 °C for 1 min, then increased to 235 °C at a rate of 10 °C/min and held at this temperature for 20 min.
Additionally oil and protein content of the zygotic seeds were analyzed using a near infrared (NIR) analyzer as described by [38]. NIR analyses were performed in triplicate compared to the known oil and protein content of control soybean cultivar Jack [39] and high oil seeds, N88–480 [20].
Results
PCR, GUS staining and Southern blot hybridization techniques were used to identify the transgenic nature of the embryos and plants obtained by hygromycin selection. Thirteen transgenic lines were PCR positive for the introduced SLC1 gene [S1 (supplementary SFig. 1)] when analyzed at the embryo stage. Only nine of those lines were further regenerated. The expected fragment size of 0.9 kbp was amplified in the transgenic samples but not in the negative control embryos. As a positive control, a SLC1 expression cassette containing pCAMBIA 1201 vector DNA was used. Genomic DNA from empty pCAMBIA 1201 vector transformed embryos and the untransformed Jack embryos were used as negative controls. A strong PCR band was observed in the positive controls. No SLC1 amplification was seen in the negative controls.
Southern blot hybridization was done for the first five independent transgenic lines (SFig. 1) for the presence of the SLC1 gene. Southern blot hybridization (Fig. 1) analysis of these transgenic plants indicated the integration of the SLC1 gene into the soybean genome. As expected, Southern blot hybridization bands from the transgenic plants were much larger than the size of the SLC1 fragment. Southern blot analysis also revealed that the introduced gene copy number varied from one to two. Four of the five transgenic plants analyzed have varying sized single insertions of the introduced SLC1 cassette. A single transgenic plant (Plant 1) had two copies of the introduced gene. All the transgenic plants regenerated grew normal and set flowers and pods (SFig. 2). Except plant 3 all other southern positive plants are turned out to be GUS positive (SFig. 3). GUS expression among different transgenics lines varied with the blue GUS stain was visible in the leaves and flowers of some lines while it was visible only in the flowers in others (SFig. 3). The reasons for such variation are not clear since the GUS gene was expressed under a constitutive CaMV 35S promoter. GUS staining was also visible in the cotyledons and in the axis of matured somatic embryos (SFig. 4).
Southern blot hybridization of yeast LPAT (SLC1) transgenic plants. Each lane represents a different transgenic line. Genomic DNA was digested with EcoRI. Since the yeast SLC1 gene has no internal EcoRI restriction sites total DNA was digested with EcoRI restriction enzyme. As a positive control the PCR amplified 0.9 kb SLC1fragment was used. Lane 1 has two hybridization bands while other transgenic lines have single copy insertions of the transgene. The bands representing the SLC1 transgenic lines are larger in size than the positive control. Molecular weights are given on the sides. Letters “P” and “C” represents positive and negative controls. “E” indicates an empty lane
Total lipid content was analyzed in matured SLC1 transgenic and vector control transformed soybean somatic embryos (Fig. 3). On average the total lipid content of the SLC1 transgenic embryos was found to be 1.5% higher than the transgenic vector controls and Jack control embryos. In addition to GUS analysis (SFig. 4) RT-PCR was done to determine the expression of the introduced SLC1 gene in matured somatic embryos (SFig. 5). Transgenic plant line 8 was established from the somatic embryos analyzed for total lipid content. As shown in Table 1 the seeds of transgenic line 8 also had a similar increase in total lipid content.
The total seed lipid content was analyzed from the seeds of greenhouse grown T1 and T2 plants. PCR was done to identify the presence of the SLC1 gene in the T2 generation transgenic plants. The TAG content of vector transformed Jack control seeds was ~20% on a dry weight basis (Table 1) while some of the T2 transgenic seeds had increased total lipid contents that varied from 0.6 to 2.5% (Table 1). The average total lipid content of T2 generation SLC1 transgenic line 9 were found to be lower than the vector-transformed Jack controls while transgenic line 1 had similar values (Table 1). TAG analysis of T3 seeds have shown that lines 1–11, 3–4, 3–9 and 6–20 have oil content increases of 1.6–3.2% compared to the vector control (Fig. 2). The T3 seeds of plants 1–11, 3–4, 3–9 and 6–20 represents the progeny of T1 transgenic lines 1, 3, and 6, respectively. The high oil cultivar N88–480 [20] showed TAG content of 22.5% by seed dry weight and the untransformed Jack plant 20.2%. The increase in lipid content of these SLC1 transgenics was accompanied by a decrease in protein content (Fig. 2).
Discussion
Some Brassica and Arabidopsis seeds that expressed the SLC1 and its mutant form, SLC1-1 genes with LPAT activity in yeast [1, 11, 19] showed increased oil levels. The present investigation examined the impact of the wild type SLC1 gene on total fatty acid levels in soybean somatic embryos and seeds. Some transgenic soybeans thus developed showed high oil in the T2 and T3 seeds. The initial aim behind transforming Brassica with yeast SLC1-1 gene was to increase erucic acid content though the transgenics also showed higher oil content [11]. As explained by Zou et al. [11] for Brassica and Arabidopsis, endogenous LPAT regulation might be one of the controlling steps in the TAG accumulation in soybeans also. Yeast LPAT with no significant homology to plant LPATs is likely not recognized by regulatory systems in plants thus creating a sink towards TAG accumulation in soybean seeds. Currently, there are at least 53 QTLs (quantitative trait loci) associated with oil content in different soybean cultivars, however, most of the QTLs are not confirmed [40]. As the accumulation of different storage components needs the coordination of several genes that encode the enzymes of the respective pathways [2] it will be interesting to know if any of the high oil QTLs encode for LPATs or for that matter any of the three Kennedy pathway acyltransferases. Besides yeast LPAT, safflower GPAT (glycerol-3-phosphate acyltransferase,) [10], Mortierella GPAT [41] and DGAT (Diacylglycerol acyltransferase) from Arabidopsis [21] have been reported to increase oil content in plants and yeast. A maize high oil QTL (Qho6) was recently analyzed [15] and known to encode DGAT.
Although larger seed sizes and ~40% increase in seed oil content have been reported for Brassica and Arabidopsis that expressed the yeast SLC1 gene [11, 19] no such dramatic results were noticed in the case of soybean seeds by us. The highest increases observed were similar in values reported for high oil line N88–480 [20]. Maybe the smaller increase in seed oil might be a reason for regular seed sizes in case of SLC1 transgenic soybean. To date the only soybean gene reported to have increased seed oil content when over- expressed are the two Dof-type transcription factor genes GmDof4 and GmDof11 [17]. When over-expressed in Arabidopsis these genes increased oil content. Over-expression of these genes activated the acetyl CoA carboxylase (ACC, EC 3.1.3.44) and long-chain-acyl CoA synthetase (ACS, EC 6.2.1.3). So one of the 58 reported QTLs of soybean oil enhancement might include these transcription factors also.
There is typically a negative correlation between protein and oil concentration in soybean seed [42].Wilcox [43] showed that increased protein amounts led to a decrease in oil content in soybeans. Interestingly the two soybean transcription factors GmDof4 and GmDof11 that elevated the activity of lipid biosynthesis enzymes also down regulated the seed storage protein gene CRA1 in Arabidopsis [17]. In the case of SLC1 transgenics also we have noticed that an increase in oil content was associated with reduced protein amounts (Fig. 2). Our results also indicate that the oil increase in soybean seeds expressing this yeast gene with LPAT activity was at the expense of protein content [44].
Somatic embryos mimic zygotic embryos in developmental and physiological aspects [45, 46]. Confirming the expected transgenic trait at the somatic embryo stage itself saves time and effort in soybean transgenic research. If a desired trait is observed in somatic embryos different clones can be screened for better producers of the end product and further taken to germination. Transgenic soybean somatic embryos were screened for fatty acid modification traits. Liu et al. [29] were the first to show palmitoleic acid in the soybean somatic embryos by over-expressing a mammalian ∆9 desaturase (SCD, EC 1.14.19.1). Cahoon et al. [32] produced vernolic acid and 12-epoxy-octadeca-9,15-dienoic acid; ∆5-eicosenoic acid and ∆5-hexadecenoic acid [31]; α-eleostearic acid and α-parinaric acids [30] in soybean somatic embryos by over-expressing the transgenes responsible for the synthesis of these compounds. As mentioned in the introduction several embryos can be generated and proliferated from a single clone of transgenic embryo. On an average when 10 embryo clones were analyzed the SLC1 expressing embryo line has 1.5% increase in oil content compared to the vector control and untransformed Jack embryos (Fig. 3). In the T2 seeds of the transgenic line 8 produced from these embryos the TAG content increase was found to be similar (Table 1). Besides the hygromycin resistance we have confirmed the transgenic status of the embryo line by RT-PCR by amplifying the introduced the SLC1 gene and GUS analysis. This is the report on the total fatty acid content of transgenic somatic embryos being correlated with an increase in oil content of transgenic soybean seeds.
In conclusion, expression of the yeast SLC1 gene that codes for a yeast lyso-phosphatidic acid acyltransferase led to an increase in soybean seed oil content. The increase in seed oil was accompanied by a reduction in seed protein content.
Analysis of soybean somatic embryos over-expressing the yeast SLC1gene led to an increase in total fatty acid content. Soybean somatic embryos can be used for analyzing quantitative changes in oil content before germinating and growing plants thus saving time and effort.
Abbreviations
- ACC:
-
Acetyl CoA carboxylase
- DGAT:
-
Diacylglycerol acyltransferase
- GPAT:
-
Glycerol-3-phosphate acyltransferase
- LPAT:
-
The lyso-phosphatidic acid acyltransferase
- ACS:
-
Long-chain-acyl CoA synthetase
- QTL:
-
Quantitative trait loci
- TAG:
-
Triacylglycerol
- Wt:
-
Wild type
References
Nageic M, Wells GB, Lester RL, Dickson RC (1993) A suppressor gene that enables Saccharomyces cerevisiae to grow without making sphingolipids encodes a protein that resembles and Escherichia coli fatty acyltransferase. J Biol Chem 269:22156–22163
Ruuska S, Girke T, Benning C, Ohlrogge J (2002) Contrapuntal networks of gene expression during Arabidopsis seed filling. Plant Cell 14:1191–1206
Scarth R, Tang J (2006) Modification of brassica oil using conventional and transgenic approaches. Crop Sci 46:1225–1236
FAO (2005) Agricultural Data, FAOSTAT. http://faostat.fao.org/faostat/collections?subset=agriculture
Dyer J, Mullen RT (2008) Engineering plant oils as high-value industrial feedstocks for biorefining: the need for underpinning cell biology research. Physiol Plant 132:11–22
Lung S, Weselake RJ (2006) Diacylglycerol acyltransferase: a key mediator of plant triacylglycerol synthesis. Lipids 41:1073–1088
Ohlrogge J, Browse J, Somerville CR (1991) The genetics of plant lipids. Biochim Biophys Acta 1082:1–26
Perry H, Bligny R, Gout E, Harwood JL (1999) Changes in Kennedy pathway intermediates associated with increased triacylglycerol synthesis in oil-seed rape. Phytochemistry 52:799–804
Saha S, Enugutti B, Rajakumari S, Rajasekharan R (2006) Cytosolic triacylglycerol biosynthetic pathway in oilseeds molecular cloning and expression of peanut cytosolic diacylglyerol acyltransferase. Plant Physiol 141:1533–1543
Jain R, Coffey M, Lai K, Kumar A, MacKenzie SL (2000) Enhancement of seed oil content by expression of glycerol-3-phosphate acyltransferase genes. Biochem Soc Trans 28:958–961
Zou J, Katavic V, Giblin EM, Barton DL, MacKenzie SL, Keller WA, Hu X, Taylor DC (1997) Modification of seed oil content and acyl composition in the Brassicaceae by expression of a yeast sn-2 acyltransferase gene. Plant Cell 9:909–923
Taylor DC, Katavic V, Zou J, MacKenzie SL, Keller WA, An J, Friesen W, Barton DL, Pedersen KK, Giblin EM, Ge Y, Dauk M, Sonntag C, Luciw T, Males D (2002) Field testing of transgenic rapeseed cv. Hero transformed with a yeast sn-2 acyltransferase results in increased oil content, erucic acid content and seed yield. Mol Breed 8:317–322
Jako C, Kumar A, Wei YD, Zou JT, Barton DL, Giblin EM, Covello PS, Taylor DC (2001) Seed-specific over-expression of an Arabidopsis cDNA encoding a diacylglycerol acyltransferase enhances seed oil content and seed weight. Plant Physiol 126:861–874
Bouvier-Navé P, Benveniste P, Oelkers P, Sturley SL, Schaller H (2000) Expression in yeast and tobacco of plant cDNAs encoding acyl CoA:diacylglycerol acyltransferase. Eur J Biochem 267:85–96
Zheng P, Allen WB, Roesler K, Williams ME, Zhang S, Li J, Glassman K, Ranch J, Nubel D, Solawetz W, Bhattramakki D, Llaca V, Deschamps S, Zhong GY, Tarczynski MC, Shen B (2008) A phenylalanine in DGAT is a key determinant of oil content and composition in maize. Nat Genet 40:269–270
Vigeolas H, Waldeck P, Zank T, Geigenberger P (2007) Increasing seed oil content in oil-seed rape (Brassica napus L.) by over-expression of a yeast glycerol-3-phosphate dehydrogenase under the control of a seed-specific promoter. Plant Biotechnol J 5:431–441
Wang H, Zhang B, Hao YJ, Huang J, Tian AG, Liao Y, Zhang JS, Chen SY (2007) The soybean Dof-type transcription factor genes, GmDof4 and GmDof11, enhance lipid content in the seeds of transgenic Arabidopsis plants. Plant J 52:716–729
Wang H, Guo J, Lambert KN, Li Y (2007) Developmental control of Arabidopsis seed oil biosynthesis. Planta 226:773–783
Zou J, Taylor DC, Katavic V, MacKenzie SL, Keller WA (2000) Modification of plant lipids and seed oils utilizing yeast SLC genes.US Patent no. 6051755
Burton JW, Wilson RF (1994) Registration of N88–480, a soybean germplasm line with a high concentration of oil in seeds. Crop Sci 34:313–314
Jako C, Kumar A, Wei Y, Zou J, Barton DL, Giblin EM, Covello PS, Taylor D (2001) Seed-specific over-expression of an Arabidopsis cDNA encoding a diacylglycerol acyltransferase enhances seed oil content and seed weight. Plant Physiol 126:861–874
Lardizabal K, Effertz R, Levering C, Mai J, Pedroso MC, Jury T, Aasen E, Gruys K, Bennett K (2008) Expression of Umbelopsis ramanniana DGAT2A in seed increases oil in soybean. Plant Physiol 148:89–96
Finer JJ, Nagasawa A (1988) Development of an embryogenic suspension culture of soybean (Glycine max Merrill.). Plant Cell Tissue Organ Cult 15:125–136
Samoylov VM, Tucker DM, Parrott WA (1998) A liquid medium-based protocol for rapid regeneration from embryogenic soybean cultures. Plant Cell Rep 18:49–54
Kinney AJ (1994) Genetic modification of the storage lipids of plants. Curr Opin Biotechnol 5:144–151
Kinney AJ (1997) Development of genetically engineered oilseeds from molecular biology to agronomics. In: Williams JP, Khan MU, Lem NW (eds) Physiology biochemistry and molecular biology of plant lipids. Kluwer, Dordrecht, pp 298–300
Kinney AJ (2001) Perspectives on the production of industrial oils in genetically engineered oilseeds. In: Kuo TM, Gardner HW (eds) Lipid biotechnology. Marcel Dekker Inc., New York
Perez-Grau L, Goldberg RB (1989) Soybean seed protein genes are regulated spatially during embryogenesis. Plant Cell 1:1095–1109
Liu W, Torisky RS, McAllister KP, Avdiushko S, Hildebrand D, Collins GB (1997) Somatic embryo cycling: evaluation of a novel transformation and assay system for seed-specific gene expression in soybean. Plant Cell Tissue Organ Cult 47:33–42
Cahoon EB, Carlson TJ, Ripp KG, Schweiger BJ, Cook GA, Hall SE, Kinney AJ (1999) Biosynthetic origin of conjugated double bonds: production of fatty acid components of high-value drying oils in transgenic soybean embryos. Proc Natl Acad Sci USA 96:12935–12940
Cahoon EB, Marillia EF, Stecca KL, Hall SE, Taylor DC, Kinney AJ (2000) Production of fatty acid components of meadowfoam oil in somatic soybean embryos. Plant Physiol 124:243–251
Cahoon EB, Ripp KG, Hall SE, McGonigle B (2002) Transgenic production of epoxy fatty acids by expression of a cytochrome P450 enzyme from Euphorbia lagascae seed. Plant Physiol 128:615–624
Cahoon E, Kinney AJ (2004) Dimorphecolic acid is synthesized by the coordinate activities of two divergent 12-oleic acid desaturases. J Biol Chem 279:12495–12502
Trick HN, Dinkins RD, Santarem ER, Di R, Samoylov VM, Meurer C, Walker D, Parrott WA, Finer JJ, Collins GB (1997) Recent advances in soybean transformation. Plant Tissue Cult Biotechnol 3:9–26
Samoylov VM, Tucker DM, Parrott WA (1998) Soybean [Glycine max (L.) Merrill] embryogenic cultures: the role of sucrose and total nitrogen content on proliferation. In Vitro Cell Dev Biol Plant 34:8–13
Reddy MSS, Dinkins RD, Redmond CT, Ghabrial SA, Collins GB (2001) Expression of bean pod mottle virus (BPMV) coat protein precursor results in resistance to (BPMV) in transgenic soybeans. Phytopathology 91:831–838
Bligh EG, Dyer WJ (1959) A rapid method of total lipid extraction and purification. Can J Biochem Physiol 37:911–917
McNeill S, Montross MD, Shearer SA (2005) Spatial variation of protein, oil, and starch in corn. Appl Eng Agric 21:619–625
Nguyen M, Nickell CD, Widholm JM (2001) Selection for high seed oil content in soybean families derived from plants regenerated from protoplasts and tissue cultures. Theor Appl Genet 102:1072–1075
Hyten D, Pantalone VR, Sams CE, Saxton AM, Landau-Ellis D, Stefaniak TR, Schmidt ME (2004) Quantitative trait loci for seed protein and oil concentration, and seed size in soybean. Theor Appl Genet 109:552–561
Macool DJ, Xue Z(2007) A mortierella alpina glycerol-3-phosphate o-acyltransferase for alteration of polyunsaturated fatty acids and oil content in oleaginous organisms. US Patent no. 7192762
Burton JW (1987) Quantitative genetics: results relevant to soybean breeding. In: Wilcox JR (ed) Soybeans: improvement, production and uses, 2nd edn. ASA, Madison, pp 211–247
Wilcox J (1998) Increasing seed protein in soybean with eight cycles of recurrent selection. Crop Sci 38:1536–1540
Wilcox J, Shibles RM (2001) Interrelationships among seed quality attributes in soybean. Crop Sci 41:11–14
Mathew M, Philip VJ (2003) Somatic embryogenesis versus zygotic embryogenesis in Ensete superbum. Plant Cell Tissue Organ Cult 72:267–275
Schmidt MA, Tucker DM, Cahoon EB, Parrott WA (2005) Towards normalization of soybean somatic embryo maturation. Plant Cell Rep 24:383–391
Jefferson RA (1987) Assaying chimeric genes in plants: the GUS gene fusion system. Plant Mol Biol Rep 5:387–405
Acknowledgments
The authors acknowledge the technical help provided by Wei Chen. The authors also acknowledge the financial support of the United Soybean Board and the support of the Kentucky Agricultural Experiment Station.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
11745_2009_3337_MOESM2_ESM.pdf
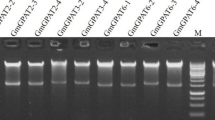
Fig. 1 SLC1 gene amplification by PCR using template DNA isolated from transgenic soybean embryos. The lane numbers indicate different transgenic lines. An untransformed Jack control is labeled − and the amplified SLC1 gene from the vector +. 1 kb plus molecular weight marker ladder is labeled as MW
11745_2009_3337_MOESM3_ESM.pdf
Fig. 2 SLC1 transgenic plant (left) with flowers and pods. All transgenic plants grew normally with no apparent differences from control Jack plants (right)
11745_2009_3337_MOESM4_ESM.pdf
Fig. 3 GUS staining in transgenic flowers and leaves of SLC1 transgenic plants 5 and 6. In case of transgenic line 6 only flowers showed GUS expression while flowers and leaves from plant 5 show GUS staining in both flowers and leaves
11745_2009_3337_MOESM5_ESM.pdf
Fig. 4 GUS expression in the matured soybean somatic embryos. GUS staining can be seen in matured SLC1 transgenic embryos. No GUS staining is visible in the untransformed control embryos
11745_2009_3337_MOESM6_ESM.pdf
Fig. 5 RT-PCR of SLC1 transgenic embryos shows the amplification of a 0.9 kbp SLC1 fragment. The vector control and untransformed Jack embryos show no amplification products. As a positive control the pCAMBIA-1201 vector with SLC1 cassette was used. MW molecular weight markers, Ctr controls
About this article
Cite this article
Rao, S.S., Hildebrand, D. Changes in Oil Content of Transgenic Soybeans Expressing the Yeast SLC1 Gene. Lipids 44, 945–951 (2009). https://doi.org/10.1007/s11745-009-3337-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11745-009-3337-z