Abstract
Paulownia witches’ broom (PaWB) caused by phytoplasma is a destructive disease of Paulownia in China and has been studied over the past 40 years. Recently, our research team found that methyl methanesulfonate (MMS) could help Paulownia plants recover from the symptoms of Paulownia witches’ broom in which the expression levels of genes related to photosynthesis, carbohydrate and nitrogen metabolism, plant-pathogen interaction, circadian rhythm changed. However, the molecular mechanisms underlying the interaction between PaWB and phytoplasma are still not fully understood. Here, high-throughput mRNA sequencing and de novo assembly were conducted in order to investigate gene expression profiles in three Paulownia tomentosa samples (healthy plantlets, diseased plantlets, and diseased plantlets treated with MMS. A total of 85,545 all-unigene were obtained, 2540 of which were significantly differentially expressed among the three samples. Some genes associated with folate and fatty acid synthesis, signal transduction pathways of plant hormones were identified, and their expression patterns were further validated using quantitative real-time PCR. To our knowledge, this is the first transcriptome-based study to demonstrate variations of gene expression related to morphological changes in metabolic pathways of phytoplasma-infected P. tomentosa plants. The information provided in this study will help understand the molecular pathogenesis of the Paulownia plant in response to phytoplasma infection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Phytoplasmas in the subgroup 16SrI-D of aster yellows are a group of microorganisms with no cell wall (Lee et al. 1993; Win et al. 2012). They range in size from 0.1 to 0.8 μm and have a 580–2200-kb circular DNA genome with a G + C content of less than 28 % (Hogenhout et al. 2008; Hoshi et al. 2007). Infected plants display a range of symptoms including witches’ broom, proliferation of axillary buds, and short internodes (Bayliss et al. 2005; Namba 2011).
Paulownia witches’ broom (PaWB) caused by a phytoplasma was first recognized in 1967 (Doi et al. 1967). Since then, the Paulownia-phytoplasma interaction has been of interest to researchers, but progress has been slow because phytoplasma are difficult to culture in vitro. The detection of a phytoplasma plasma-encoded protein and elongation factor EF-Tu (Lin et al. 2009; Wang et al. 2010), and information about the expression of host genes, such as those encoding plant hormones and proteins involved in cell wall biosynthesis and degradation, photosynthesis and carbohydrate metabolism, plant defense, plant-pathogen interaction, circadian rhythm, phytoplasma virulence, amino acid metabolism, and nitrogen metabolism have been reported so far (Cao et al. 2014a, b; Fan et al. 2014; Liu et al. 2013; Mou et al. 2013).
Paulownia tomentosa belongs to the family Paulowniaceae and is one of the fast growing trees native to China. Because it is a rich source of biologically active secondary metabolites, it is used in Chinese herbal medicine (Schneiderová and Šmejkal 2014). P. tomentosa has now been introduced into many other countries as a decorative ornamental tree (Erbar and Gülden 2011). To enrich the gene expression information associated with morphological changes in Paulownia plantlets after PaWB infection, here we investigated the expression patterns of genes that encode proteins involved in folate, fatty acid synthesis, and plant hormone signal transduction pathways. As reported previously, methyl methanesulfonate (MMS) at suitable concentrations could help Paulownia plants recover from the symptom of PaWB (Cao et al. 2014a). Hence, in this study, we performed a transcriptome analysis to determine the differences in gene expression in healthy P. tomentosa plantlets, diseased plantlets, and plantlets treated with 20 mg·L−1 MMS. The results will help in understand the mechanisms of Paulownia-phytoplasma interactions.
Materials and methods
Plant material treatment and RNA isolation
Healthy and diseased tissue culture plantlets of P. tomentosa were obtained from the Institute of Paulownia, Henan Agricultural University, Zhengzhou, China. The plantlets were cultured for 30 days on 1/2 MS medium (Murashige and Skoog 1962) before being clipped. The 1.5-cm-long terminal buds were clipped from the diseased plantlets and transplanted into 100-mL triangular flasks with 1/2 MS culture medium containing 0 and 20 mg·L−1 MMS. The same-sized terminal buds of the healthy plantlets were also transplanted into 1/2 MS medium without MMS, as the control. For each MMS concentration (0 and 20 mg·L−1) and for the control, three terminal buds were transplanted into one flask and a total of 20 flasks was used for each treatment. Each treatment was performed in triplicate.
The cultivation period for all these plantlets was 30 days for the MMS-treated plantlets, which was divided into two stages: first cultured at 20 °C in a darkroom for 5 days, and then transferred to a chamber at 25 ± 2 °C and 130 µmol·m−2 s−1 light intensity with a 16/8 h light/dark photoperiod for 25 days. The 1.5-cm long terminal buds were sheared from the healthy plantlets (PHP), diseased plantlets (DP), and the diseased plantlets treated with 20 mg·L−1 MMS (DP-20), and immediately frozen in liquid nitrogen and stored at −80 °C.
DNA isolation and phytoplasma detection
Total DNA was extracted from ten individual terminal buds of PHP, DP, and DP-20 using the cetyl trimethylammonium bromide (Beijing Chemical Co., Beijing, China) method as described by Zhang et al. (2009). The PaWB phytoplasma was detected by nested-PCR as described by (Lee et al. 1993). The PCR and agarose gel electrophoresis were performed according to the method of Fan et al. (2007).
RNA isolation and construction of cDNA libraries
Total RNA was extracted from ten individual terminal buds of PHP, DP, and DP-20 using TRIzol reagent (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s protocol. The total RNA was treated with DNase I (RNase-free) to remove any contaminating DNA. The RNA quantity was assessed by the OD260/280 and OD260/230 ratios using a NanoDrop 2000 spectrophotometer (Thermo Scientific, Wilmington, DE). The integrity of the RNA was assessed by 1 % agarose gel electrophoresis. The high-quantity and high-integrity RNA was used to construct the cDNA libraries. Briefly, magnetic beads with oligo (dT) were used to isolate the poly (A)-containing mRNA from the total RNA. Then the mRNA was sheared into smaller fragments that were used as templates to synthesize the first-strand cDNA with SuperScript II reverse transcriptase (Life Technologies, Carlsbad, CA). Second-strand cDNA was synthesized using RNase H and DNA polymerase I. The resultant double-stranded cDNA was purified and washed with EB buffer for end reparation. Next, a single A (adenine) nucleotide was added to the 3′ ends and adapters were connected to the 5′ ends. The suitable fragments were used as templates for the PCR amplification. The PCR products were purified on agarose gel to obtain the final cDNA libraries, which were then quantified using an Agilent 2100 Bioanalyzer and ABI StepOnePlus Real-Time PCR System (ABI, New York, NY). The cDNA libraries were sequenced on an Illumina HiSeq™ 2000 platform (Illumina, San Diego, CA) by Beijing Genomics Institute (BGI) (Shenzhen, China) following the manufacturer’s standard cBot and sequencing protocols.
Unigene assembly and annotation
High-quality clean transcriptome sequencing reads were obtained after removing reads with adaptor sequences, more than 5 % unknown nucleotides, and low quality reads. The reads in the PHP, DP, and DP-20 libraries were assembled into all-unigene using the Trinity software (release-20121005) (Grabherr et al. 2011). The all-unigene sequences were aligned using BLASTX (E value < 1.0E-5) against the NCBI non-redundant (NR) protein database (release-20121005) (Altschul et al. 1997), the Swiss-Prot protein database (release-2012_08), the cluster of orthologous groups (COG) database (release-20090331), and the Kyoto Encyclopedia of Genes and Genomes (KEGG) Pathway database (Release 63.0). BLASTN (E value < 1.0E-5) was used to align the all-unigene sequences to the NCBI nucleotide (NT) database (release-20121005) (Xiang et al. 2010). All-unigene sequences that did not align to any of the above databases were processed using ESTScan (Iseli et al. 1999). To assign functions of the all-unigene that shared more than 70 % similarity with sequences in the NR database, the gene ontology (GO) (release 2012-10-01) functional annotations for the aligned NR sequences were retrieved using the WEGO software (Ye et al. 2006). The COG database was used to classify the assigned functional annotations of the all-unigene, and the KEGG Pathway database was used to determine the metabolic pathways that the all-unigene might be involved in.
Differential expression of genes among the three libraries
To assess the changes of gene expressions between DP vs. PHP and DP-20 vs. DP, the fragments per kilobase of transcript per million mapped reads (FPKM) method (Mortazavi et al. 2008) was used to calculate the expression levels as:
where FPKM is the expression of unigene, C is the number of fragments that aligned uniquely to unigene, N is the total number of fragments that aligned uniquely to all-unigene, and L is the number of bases in the coding sequence of unigene.
The differentially expressed genes (DEGs) between any two samples according to Audic and Claverie (1997).
To detect the DEGs, the p value threshold in multiple hypothesis testing and analyses was determined by manipulating the false discovery rate (FDR) value (Benjamini and Yekutieli 2001). Two standards (|log2 Ratio| ≥ 1 and FDR ≤ 0.001) were used to judge the significance of gene expression differences. The KEGG enrichment analysis was performed by applying the hypergeometric test to find significantly enriched KEGG pathways among the DEGs compared with the whole transcriptome background of P. tomentosa. The Q value (similar to p value) was calculated as:
where N is similar as the total number of genes with KEGG annotation; n is the number of DEGs in N; M is the total number of all genes that were annotated to a certain KEGG pathway; and m is the number of DEGs in M. After applying the Bonferroni correction to the calculated p value, we chose a corrected p value of ≤0.05 as the threshold, and KEGG pathways with q ≤ 0.05 were defined as significantly enriched KEGG pathways for the DEGs.
Quantitative real-time PCR (qRT-PCR) analysis
To validate the expression levels of the DEGs in the PHP, DP, and DP-20 libraries, 20 DEGs that were predicted to be involved in the Paulownia-phytoplasma interaction were chosen. First-strand cDNAs of all were synthesized using an iScript cDNA synthesis kit (Bio-Rad, Hercules, CA). The qRT-PCR primers were designed using the Beacon Designer 6.0 software (Premier Biosoft International, Palo Alto, CA). The cDNAs were then amplified on a Bio-Rad CFX96TM Real-Time System (Bio-Rad). The PCR cycles were as follows: 95 °C for 1 min, followed by 40 cycles of 95 °C for 10 s and 55 °C for 15 s. 18S rRNA served as the internal reference gene. The results were analyzed using the 2−ΔΔCt method (Livak and Schmittgen 2001). Each gene was analyzed in three replicates. All the primers used for the qRT-PCR are showed in Online Resource 1: Table S1. Statistical analyses were performed using SPASS 19.0 (SPASS, Inc., Chicago, IL).
Results
Detection of phytoplasma in P. tomentosa plantlets
The nest-PCR results showed that phytoplasma 16S rDNA fragments (1.2 kb) were detected in the DP and DP-20 libraries. Morphological changes were observed in DP-20 compared with DP (Fig. 1); that is, the color of the plantlets changed from light yellow to green, the sizes of the leaves changed from small turn to normal, the seta gradually appear again, and the short internodes became normal; however, very tiny axillary buds were found in the base of the DP-20 plantlets (Fig. 2). These observations indicated that the PaWB symptoms were attenuated with the 20 mg·L−1 MMS treatment.
Detection of phytoplasma in P. tomentosa diseased plantlets with MMS treatment. The phytoplasma 1.2 kb fragment was detected in the diseased plantlets, and the same weakened fragment was also detected in the diseased plantlets with 20 mg·L−1MMS treatment, while no fragment was detected in the healthy plantlets. 1 diseased plantlets. 2 diseased plantlets with 20 mg·L−1MMS treatment. 3 healthy plantlets. 4 double distilled water. M: DNA Marker
Morphological changes of P. tomentosa diseased plantlets with MMS treatment. When the phytoplasma infected healthy plantlets, the plantlets showed the diseased morphology, but this morphology can attenuate with 20 mg·L−1MMS treatment. a healthy plantlets; b diseased plantlets; c diseased plantlets with 20 mg·L−1MMS treatment
Transcriptome sequencing and de novo assembly
The transcriptome sequencing produced 48,006,640 (PHP), 75,244,384 (DP), and 45,932,476 (DP-20) clean reads. The N percentages were 0.08, 0.11, and 0.04 % in PHP, DP, and DP-20, respectively, and the GC content was 46.59, 46.55, and 45.64 %, respectively (Online Resource 1: Table S2). The clean reads were assembled into 126,285 (PHP), 134,791 (DP), and 147,094 (DP-20) contigs with average length of 368, 367, and 351 nt, respectively. The N50s n each of the contigs were 750 bp (PHP), 78 5 bp (DP), and 667 bp (DP-20). The contigs were further assembled into 77,844 (PHP), 79,456 (DP), and 90,969 (DP-20) unigene with mean lengths of 780, 848, and 866 nt, and N50s of 1513, 1618, and 1664 bp, respectively. Finally, a total of 85,545 unique all-unigene with a mean length of 1127 nt and N50 of 1833 bp were obtained from among the unigene from the three samples (Online Resource 1: Table S2). The length distributions of these all-unigene are shown in Online Resource 2: Figure S1.
Functional annotations of the all-unigene
To predict the functions of the all-unigene, we matched 85,545 all-unigene sequences first to the NR database, and then to the NT, Swiss-Prot, COG, GO, and KEGG databases (E value < 1.0E-5). We found that 60,328 (70.5 %) all-unigene matched known sequences in at least one of these databases (Table 1). Among them, 58,423 (68.3 %) all-unigene matched NR sequences, 52,631 (61.5 %) matched NT sequences, and 38,000 (44.4 %) matched Swiss-Prot sequences. In addition, 24,158 (28.2 %) all-unigene were mapped to the COG database, 48,326 (56.5 %) were assigned GO annotations, and 35,367 (41.3 %) were assigned to KEGG pathways. The remaining approximately 30 % of all-unigene that did not map to any of the databases may represent new genes that have not yet been identified in other plant species.
The distributions of the E values, sequence similarity, and species similarity of the all-unigene sequences that mapped to NR sequences are shown in Fig. 3. To increase the robustness of the annotations, the 24,158 all-unigene that mapped to the COG database were classified into 25 COG subgroups based on the COG classification (Fig. 4). Among them, the R subgroup “general function prediction only” was the largest group (8568; 35.5 %), followed by the K (4445; 18.4 %), L (4226; 17.5 %), and T (3545; 14.7 %) subgroups of “transcription”, “replication”, “recombination and repair”, and “signal transduction mechanisms”, respectively. The Y subgroup “nuclear structure” (5; 0.02 %) was the smallest subgroup (Online Resource 1: Table S3).
Based on the GO annotations, 48,326 all-unigene were assigned GO terms under the three main GO categories: biological process, molecular function, and cellular component. Among the GO terms under biological process, “cellular process” (31,520; 65.2 %) was assigned to the largest number of all-unigene (Online Resource 1: Table S4); under cellular component, “cell” (39,081; 80.9 %) and “cell part” (39,081; 80.9 %) were assigned to the largest number of all-unigene; and under molecular function, “catalytic activity” (23,261; 48.1 %) was assigned to the largest number of all-unigene. Based on the KEGG pathway annotations, 35,367 (41.3 %) all-unigene were predicted to be involved in 128 biological process; “metabolic pathways” (7890; 22.3 %), was the largest group, and “betalain biosynthesis” (2, 0.01 %) was the smallest group (Online Resource 1: Table S5), (Fig. 5).
DEGs among the three P. tomentosa libraries
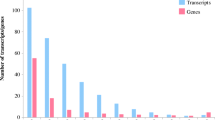
We compared the gene expression profiles in the DP versus PHP and DP-20 versus DP libraries to identify DEGs among the three samples (Online Resource 1: Table S6). A total of 2540 DEGs were detected; 2412 were down-regulated in DP versus PHP and up-regulated in DP-20 versus DP, and 128 were up-regulated in DP versus PHP and down-regulated in DP-20 versus DP (Fig. 6). The 2540 DEGs were assigned to 119 pathways in the KEGG database (Online Resource 1: Table S7). Among these DEGs, we identified several DEGs that may be related closely to PaWB disease, including genes encoding folylpolyglutamate synthase (FPGS) and bifunctional dihydrofolate reductase-thymidylate synthase (BDHFR-TS) that may be associated with the folate synthesis pathway. In addition, DEGs encoding acetyl-CoA carboxylase beta subunit (ACCase), beta-ketoacyl-ACP synthase II (KASII), 3-oxoacyl-[acyl-carrier-protein] reductase (KR), and acyl-ACP thioesterase (TE) that are related to the biosynthesis of fatty acid were altered after phytoplasma infection. DEGs associated with plant hormone signal transduction also were identified, namely genes encoding histidine kinase SHK278 (CRE1), histidine-containing phosphotransfer protein 5 (AHP), transcription repressor KAN1 (B-ARR), type-a response regulator (A-ARR), protein phosphatase (PP2C), serine/threonine-protein kinase SRK2B isoform 1 (SnRK2), ABA responsive element binding factor (ABF), interleukin-1 receptor -associated kinase 4 (BAK1), LRR receptor-like serine/threonine protein kinase (BRI1), serine/threonine protein kinase At4g35230 (BSK), serine/threonine protein kinase (BIN2), mid1- complementing activity 2 (CYCD3), and ubiquitin-protein ligase (TGA).
Differentially expressed genes’ analysis of P. tomentosa. A differentially expressed genes in diseased plantlets versus healthy plantlets up-regulated and down-regulated in diseased plantlets with 20 mg·L−1MMS treatment versus diseased plantlets. B differentially expressed genes in diseased plantlets versus healthy plantlets down-regulated and up-regulated in diseased plantlets with 20 mg·L−1MMS treatment versus diseased plantlets
QRT-PCR analysis
To confirm the results of the transcriptome sequencing, 12 PaWB-related genes were selected randomly for qRT-PCR (Fig. 7). The results showed that the relative expressions of 11 of the 12 selected genes were significantly up-regulated in DP versus PHP and down-regulated in DP-20 versus DP, while one of the genes was significantly down-regulated in DP versus PHP and up-regulated in DP-20 versus DP. These expression patterns are similar to the expressions predicted from the transcriptome data analysis, indicating that the Illumina sequencing data were robust enough to be used to assess transcriptomic changes associated with the morphological changes that were observed in the PaWB plantlets.
qRT-PCR analysis of P. tomentosa selective DEGs. A relative expression of peroxidase 31 (Per31). B relative expression of metallothionein-like protein 1 (MetP1). C relative expression of carotenoid cleavage dioxygenase 4 (CarCD). D relative expression of polyphenol oxidase (PolO). E relative expression of multicopper oxidase. F relative expression of cryptochrome 2 (Cry2). G relative expression of dihydrofolate reductase (DihR). H relative expression of HSP90 (Hsp90). I relative expression of flavonoid 7-O-methyltransferase (FlaOM). J relative expression of caffeoyl-CoA O-methyltransferase (CCOMT). K relative expression of chalcone isomerase (ChaI). L relative expression of disease resistance protein 1-like protein b (DisRPb). DP-20 diseased plantlets with 20 mg·L−1 MMS treatment, PHP phytoplasma-free plantlets. The different letters within a gene repression level indicate significant difference, while the same letters within a gene repression level indicate no significant differences (p < 0.05)
Discussion
Phytoplasmas, microbes without cell walls, have to exclusively inhabit nutrient-rich plant phloem tissues in their hosts. At the same time, the hosts launch a series of measures against phytoplasma infection. It has been reported that phytoplasma invasion induced callose deposition in the sieve plates of sieve cells in host plants, which can block the spread of the phytoplasma through the vascular system of plants (Hren et al. 2009). However, callose deposition will reduce the transport of sugar, phytohormones, ions, and metabolites and affect the host’s metabolism (Sugio et al. 2011). Indeed, it has been reported that phytoplasma infection modulated the membrane system, modified the cell wall, altered the cell cycle and cell division, and altered membrane ionic permeability. These changes initiated the synthesis and release of secondary messengers, changed the expression patterns of related stress-induced genes, influenced carbohydrate metabolism, and changed cell wall biosynthesis and degradation. The expression of genes such as BAK1, MEKK1, and WRKY29 have been associated with plant-pathogen interactions and cytokinin (CK) and abscisic acid (ABA) biosynthesis, and genes such as TOC1 and LHY/CCA1 have been reported to influence circadian rhythm (Fan et al. 2014; Liu et al. 2013; Mou et al. 2013). Phytoplasma infection was reported to repress the expression of several genes in the photosynthesis process, and genes related to defense and energy metabolism were found to be differentially expressed (Liu et al. 2013; Mou et al. 2013), indicating that the expression of these gene was related closely to the smaller leaves and shorter internode symptoms that have been observed in PaWB-infected plants.
To fully understand the molecular mechanisms behind the morphological changes that have been observed in Paulownia tomentosa plantlets after phytoplasma infection, high-throughput mRNA sequencing and de novo assembly were conducted to found the relation of expression levels of some genes involved in folate, fatty acid synthesis and signal transduction of plant hormones with the morphological changes of the plantlets infected with phytoplasma.
DEGs related to folate synthesis
Folate is an essential B vitamin that functions as a cofactor for enzymes in one-carbon metabolism in plants and plays an important role in the biosynthesis of nucleic acids, amino acids, pantothenate, and photorespiration (Anukul et al. 2010; Basset et al. 2005). In plants and insects, it has been reported that the infecting phytoplasma harbor genes (folk, folP, folC, and folA) that encode enzymes involved in folate biosynthesis (Oshima et al. 2013), thereby allowing the phytoplasma to adapt to different environments by regulating gene expression in their hosts (Oshima et al. 2004).
In the present study, genes encoding FPGS and BDHFR-TS were significantly up-regulated in DP versus PHP and down-regulated in DP-20 versus DP. These two enzymes are involved in the folate synthesis pathway. FPGS catalyzes the conversion of 7,8-dihydropteroate to 7,8-dihydrofolate (DHF), which can be converted to folate and 5,6,7,8-tetrahydrofolate (THF) by BDHFR-TS. In this process, THF-L-glutamate and THF-polyglutamate are also produced. In the phytoplasma, tetrahydrofolate was found to be synthesized by a combination of the four phytoplasma folate biosynthesis-related enzymes and FPGS (Rébeillé et al. 1997); therefore, we speculated that the high expression of an FPGS-encoding gene in the PaWB-infected P. tomentosa plantlets would provide a good source of folate for phytoplasma survival. Nevertheless, successful phytoplasma might consume a lot of folate, which might result in a lack of folate in their host and trigger the disease symptoms. Additionally, folate deficiency in the host could decrease the ratio of S-adenosylmethionine to S-adenosylhomocysteine and affect DNA methylation reactions (Balaghi and Wagner 1993). BDHFR-TS, which is involved in the biosynthesis of tetrahydrofolic acid, is the carrier of 1-carbon transfer reactions that are essential for DNA methylation (Maejima et al. 2014). Hence, the higher expression of FPGS and BDHFR-TS may be related to the changes in DNA methylation in Paulownia plantlets after phytoplasma infection, which is consistent with our previous results (Cao et al. 2014a, b; Zhai et al. 2010).
DEGs related to fatty acid synthesis
Fatty acids are the main building blocks for the phospholipid components of cell membranes and are determinants of intracellular communication (Prieschl and Baumruker 2000). Phytoplasma cannot synthesize fatty acids and, therefore, have to import them from their hosts (Bertaccini and Duduk 2010; Oshima et al. 2004). Phytoplasma infection was reported to modulate the plant cell membrane system by not only altering the membrane ionic (H+, Ca2+ and K+) permeability and changing the fatty acid composition, but also by releasing noxious reactive oxygen species (ROS), which is an important cell signal in phytoplasma-infected plants (Fan et al. 2014; Mou et al. 2013).
In our study, four genes involved in the fatty acid synthesis pathway were significantly up-regulated in DP versus PHP and down-regulated in DP-20 versus DP. In DP, the up-regulated DEGs associated with this pathway included genes encoding ACCase, KASII, KR and TE. In plants, ACCase is believed to be a major determinant of the overall rate of fatty acid synthesis. ACCase catalyzes the reaction between bicarbonate and acetyl-CoA to form malonyl-CoA, the precursor of malonyl-ACP, which is the carbon donor for the subsequent elongation reactions (Rismani-Yazdi et al. 2011), and also activated the expression of KASII and KR. KASII is believed to take part in the elongation of C16:0-ACP to C18:0-ACP, but KR catalyzes the reduction of this production of the elongation of the fatty acid (Slabas et al. 1992). Interestingly, this reduction reaction is reversible, the interaction of these two enzymes has been reported to further increase the synthesis of octanoyl-ACP, decanoyl-ACP, and hexa-deconoyl-ACP, the precursor substrates of octanoic acid, decanoic acid, and hexadecanoic acid, respectively. It has been reported that these substrates may be catalyzed by TE to break the fatty acid elongation cycle and release free fatty acids, such as octanoic acid, decanoic acid, and hexadecanoic acid, and induce the fatty acid degradative pathways (Zhang et al. 2011), which enhanced leakage of cytoplasmic solutes and damaged the plant membranes (McKersie et al. 1989). Moreover, because phytoplasma possess an enriched type III secretion system and ATP-binding cassette transporter system (Maejima et al. 2014; Oshima et al. 2004), the phytoplasma virulence factors may be transported easily through the damaged cell membranes, thereby enhancing the susceptibility of the plantlets to phytoplasma infection. At the same time, the gene encoding the cell membrane receptor, LRR receptor-like serine/threonine protein kinase (BRI1), was also up-regulated in DP. It has been reported that BRI1 can recognize a pathogen and trigger pathogen-associated molecular patterns that can bind the phytoplasma effector (EF-TU) and induce witches’ broom in infected plants (Schwessinger and Zipfel 2008; Wang et al. 2010). Hence, the formation of witches’ broom as a result of PaWB infection may be associated with changes in the expression levels of the genes that are involved in the release of free fatty acids.
DEGs related to plant hormone signal transduction
Plant hormones play important roles in regulating plant developmental processes and signaling networks in response to biotic and abiotic stresses (Bari and Jones 2009; Murmu et al. 2014). The disruption of hormonal balance has been related to abnormal morphology such as stunting and yellowing (Ćurković Perica 2008; Ehya et al. 2013; Leljak-Levanic et al. 2010). Hoshi et al. (2009) discovered that a phytoplasma virulence factor TENGU could induce proliferation and dwarfism by interfering with the expression of an auxin-related gene. Sugio et al. (2011) found phytoplasma infection reduced the production of jasmonic acid (JA), a hormone that plays fundamental roles in plant defense, which might decrease the hosts’ resistance and create an appropriate environment for phytoplasma survival. However, in a previous study, we identified genes involved in CK and ABA synthesis that were up-regulated in phytoplasma-infected Paulownia, whereas genes related to auxin were down-regulated. The expression of zeatin-related genes also was enhanced after phytoplasma infection (Fan et al. 2014; Liu et al. 2013; Mou et al. 2013). These observations suggested that changes in the plant hormones may play important roles in the formation of phytoplasma-mediated symptoms in the host.
In this study, we identified several genes in the multiple plant hormone signal transduction network that were markedly up-regulated in DP versus PHP and down-regulated in DP-20 versus DP. In DP, the up-regulated DEGs associated with this network included CRE1, AHP, B-ARR, A-ARR, PP2C, SnRK2, ABF, BAK1, BRI1, BSK, BIN2, CYCD3, and TGA.
Cytokinins are a class of plant hormones that influence cell proliferation, apical dominance, and leaf senescence (Ferreira and Kieber 2005). Alterations in the CK balance may help the pathogen to invade the stele region and vasculature (Moreau et al. 2014). Of the DEGs in the hormone signal transduction network, CRE1, AHP, B-ARR, and A-ARR may be involved in cytokinin signal transduction, and were differentially expressed in DP. CRE1 encodes a cytokinin receptor, a membrane histidine kinase that contains an extracellular sensing CHASE domain. CRE1 was reported to initiate the signal of phosphorylation, and regulate the osmoregulation, photosensitivity, and microbial pathogenesis (Bilwes et al. 1999; Inoue et al. 2001), thereby playing an important role in cell signal transduction. As reported in our previous studies (Cao et al. 2014a, b), CRE1 also may activate the expression of AHP. AHP encodes a histidine phosphotransfer protein that mediates the transfer of phosphoryl groups from the receptor kinases to the response regulators of nucleus, such as B-ARR and A-ARR (Ferreira and Kieber 2005). Over-expression of AHP can inhibit hypocotyl elongation of seedlings, showing the characteristic phenotype of cytokinin-hypersensitivity (Suzuki et al. 2002), and induce the expression of B-ARR and A-ARR in the cell nucleus. B-ARR usually implicated as DNA-binding transcription factors in the phosphorelay-mediated cytokinin signal transduction network (Ishida et al. 2008). A-ARR can negative regulation of CKs signaling, and have a role in regulating the homeostasis of the CK response (Gonzalez-Rizzo et al. 2006; Yokoyama et al. 2007), may be the production of large numbers of axillary shoots were implicated in the specific interaction between B-ARR and A-ARR.
Abscisic acid content has been positively related with the degree of disease, and high ABA content was found to promote the formation of witches’ broom (Fan et al. 2014; Hegele and Bangerth 1998; Liu et al. 2013; Mou et al. 2013). In this study, PP2C, SnRK2, and ABF were implicated in ABA signal transduction. These genes were up-regulated in the phytoplasma-infected Paulownia plantlets compared with their expression in DP-20 and PHP. PP2C was found to be an essential component of the ABA signaling pathway, and the over-expression of a PP2C isoform was reported to activate cell proliferation and expansion, especially in vascular tissues and guard cells (Sugimoto et al. 2014). PP2C also was shown to mediate the inactivation of SnRK2 via dephosphorylation (Umezawa et al. 2009). However, SnRK2 was not repressed by the over-expression of PP2C in the phytoplasma infected Paulownia plantlets, but it can directly phosphorylate ABF in response to activate ABA-responsive genes, which can induce stomatal pore closure (Kobayashi et al. 2005). Thus, the formation of witches’ broom may be related to the up-regulation of ABF associated with ABA signal transduction.
Brassinosteroid (BR) plays an important role in plant development. In this study, genes annotated as BAK1, BRI1, BSK, BIN2, and CYCD3 may be implicated in BR signal transduction. BRI1 and BAK1 are the main membrane receptor-like kinases in BR signaling pathway. BAK1, a member of the somatic embryogenesis receptor kinase family, has been described as a signal transducer that can recognize microbial pathogens and induce the expressions of defense-related genes (Uematsu et al. 2005). BRI1 usually acts as a cell recognition receptor that can trigger the recognition of pathogen-associated molecular patterns (Schwessinger and Zipfel 2008), which can bind the phytoplasma effector (EF-TU) and promote proliferation and dwarfism in the infected plants (Dodds and Rathjen 2010). Activation of BRI1 can also initiate the expressions of the BSK kinases, which is the substrates of BRI1 kinase, which can activate downstream BR signal transduction. However, not all of these genes are activated in such situations, for example, BSU1 was not activated by the up-regulation of BSK, while BIN2, one kinase that should be repressed by BSU1 showing up-regulation in the DP, and BIN2 has been reported to repress BR-responsive gene expression (Kim et al. 2011). Interestingly, BIN2 was found to indirectly target the host’s DNA by phosphorylation, and can induce the expression of CYCD3, which can induce cell division. Thus, the different expression of BAK1, BRI1 and CYCD3 might be related to the development of witches’ broom symptoms.
The present study showed that plant hormone changes may be one reason for the PaWB, combining the previous findings (Cao et al. 2014a, b; Fan et al. 2014; Liu et al. 2013; Mou et al. 2013). It can be speculated that the occurrence of PaWB is a complex process that may be regulated by multiple hormone signaling processes. Based on these results, this study suggested that the expression levels of CRE1, AHF, B-ARR, ABF, and CYCD3 may be closely related to PaWB. Further studies are required to elucidate the contributions of these genes to the morphological changes of Paulownia in response to phytoplasma.
Conclusions
The aim of the present study was to reveal the mechanisms behind phytoplasma pathogenicity based on a transcriptome sequencing analysis. A total of 2540 DEGs were obtained among the three P. tomentosa libraries. The functions of these DEGs were enriched in 119 KEGG pathways including folate and fatty acid biosynthesis and the plant hormone signal transduction pathway, which might play important roles in the occurrence of PaWB. By analysis of the functions of the DEGs in these pathways, we believed that folate deficiency, accumulation of free fatty acids, and disruption of plant hormones related to signal transduction might be closely related to the morphological changes of Paulownia plantlets in response to phytoplasma infection.
Author contribution statement
Conceived and designed the experiments: G.F. Performed the experiments: Z.Z. Analyzed the data: M.D. Contributed reagents/materials/analysis tools: X.C. Wrote the paper: G.F and X.C.
References
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucl Acids Res 25:3389–3402
Anukul N, Ramos R, Mehrshahi P, Castelazo A, Parker H et al (2010) Folate polyglutamylation is required for rice seed development. Rice 3:181–193
Audic S, Claverie J (1997) The significance of digital gene expression profiles. Genome Res 7(10):986–995
Balaghi M, Wagner C (1993) DNA methylation in folate deficiency: use of CpG methylase. Biochem Bioph Res Co 193:1184–1190
Bari R, Jones JD (2009) Role of plant hormones in plant defence responses. Plant Mol Biol 69:473–488
Basset GJC, Quinlivan EP, Gregory JF III, Hanson AD (2005) Folate synthesis and metabolism in plants and prospects for biofortification. Crop Sci 45:449–453
Bayliss KL, Saqib M, Dell B, Jones MG, Hardy GE St (2005) First record of ‘Candidatus Phytoplasma australiense’ in Paulownia trees. Australas Plant Path 34:123–124
Benjamini Y, Yekutieli D (2001) The control of the false discovery rate in multiple testing under dependency. Ann Stat 29:1165–1188
Bertaccini A, Duduk B (2010) Phytoplasma and phytoplasma diseases: a review of recent research. Phytopathol Mediterr 48:355–378
Bilwes AM, Alex LA, Crane BR, Simon MI (1999) Structure of CheA, a signal-transducing histidine kinase. Cell 96:131–141
Cao XB, Fan GQ, Deng MJ, Zhao ZL, Dong YP (2014a) Identification of genes related to Paulownia witches’ broom by AFLP and MSAP. Int J Mol Sci 15:14669–14683
Cao XB, Fan GQ, Zhao ZL, Deng MJ, Dong YP (2014b) Morphological changes of paulownia seedlings infected phytoplasmas reveal the genes associated with witches’ broom through AFLP and MSAP. Plos One 9:e1125331
Ćurković Perica M (2008) Auxin-treatment induces recovery of phytoplasma-infected periwinkle. J Appl Microbiol 105:1826–1834
Dodds PN, Rathjen JP (2010) Plant immunity: towards an integrated view of plant–pathogen interactions. Nat Rev Genet 11:539–548
Doi Y, Ternaka M, Yora K, Asuyama H (1967) Mycoplasma or PLT-group-like microorganisms found in the phloem elements of plants infected with mulberry dwarf, potato witches’ broom, aster yellows and Paulownia witches’ broom. Ann Phytopathol Soc Jpn 33:259–266
Ehya F, Monavarfeshani A, Fard EM, Farsad LK, Nekouei MK, Mardi M, Salekdeh GH (2013) Phytoplasma-responsive microRNAs modulate hormonal, nutritional, and stress signalling pathways in Mexican lime trees. Plos One 8:e66372
Erbar C, Gülden C (2011) Ontogeny of the flowers in Paulownia tomentosa—a contribution to the recognition of the resurrected monogeneric family Paulowniaceae Flora—morphology, distribution. Funct Ecol Plants 206:205–218
Fan GQ, Zhang S, Zhai XQ, Liu F, Dong ZQ (2007) Effects of antibiotics on the paulownia witches’ broom phytoplasmas and pathogenic protein related to witches’ broom symptom. Sci Silv Sin 43:138–142
Fan GQ, Dong YP, Deng MJ, Zhao ZL, Niu SY, Xu EK (2014) Plant-pathogen interaction, circadian rhythm, and hormone-related gene expression provide indicators of phytoplasma infection in Paulownia fortunei. Int J Mol Sci 15:23141–23162
Ferreira FJ, Kieber JJ (2005) Cytokinin signaling. Curr Opin Plant Biol 8:518–525
Gonzalez-Rizzo S, Crespi M, Frugier F (2006) The Medicago truncatula CRE1 cytokinin receptor regulates lateral root development and early symbiotic interaction with Sinorhizobium meliloti. Plant Cell 18:2680–2693
Grabherr MJ, Haas BJ, Yassour M et al (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29:644–652
Hegele M, Bangerth F (1998) Changes in IAA and ABA levels and IAA transport of proliferation-diseased apple trees. Acta Hortic 463:97–108
Hogenhout SA, Oshima K, Ammar ED, Kakizawa S, Kingdom HN, Namba S (2008) Phytoplasmas: bacteria that manipulate plants and insects. Mol Plant Pathol 9:403–423
Hoshi A, Ishii Y, Kakizawa S, Oshima K, Namba S (2007) Host-parasite interaction of phytoplasmas from a molecular biological perspective. B Insectol 60:105
Hoshi A, Oshima K, Kakizawa S et al (2009) A unique virulence factor for proliferation and dwarfism in plants identified from a phytopathogenic bacterium. Proc Natl Acad Sci USA 106:6416–6421
Hren M, Nikolić P, Rotter A et al (2009) ‘Bois noir’ phytoplasma induces significant reprogramming of the leaf transcriptome in the field grown grapevine. BMC Genom 10:460
Inoue T, Higuchi M, Hashimoto Y et al (2001) Identification of CRE1 as a cytokinin receptor from Arabidopsis. Nature 409:1060–1063
Iseli C, Jongeneel CV, Bucher P (1999) ESTScan: a program for detecting, evaluating, and reconstructing potential coding regions in EST sequences. Proc Int Conf Intel Syst Mol Biol 99:138–148
Ishida K, Yamashino T, Yokoyama A, Mizuno T (2008) Three type-B response regulators, ARR1, ARR10 and ARR12, play essential but redundant roles in cytokinin signal transduction throughout the life cycle of Arabidopsis thaliana. Plant Cell Physiol 49:47–57
Kim TW, Guan S, Burlingame Alma L, Wang ZY (2011) The CDG1 Kinase mediates brassinosteroid signal transduction from BRI1 receptor kinase to BSU1 phosphatase and GSK3-like kinase BIN2. Mol Cell 43:561–571
Kobayashi Y, Murata M, Minami H et al (2005) Abscisic acid-activated SNRK2 protein kinases function in the gene-regulation pathway of ABA signal transduction by phosphorylating ABA response element-binding factors. Plant J 44:939–949
Lee IM, Hammond RW, Davis RE, Gundersen DE (1993) Universal amplification and analysis of pathogen 16S rDNA for classification and identification of mycoplasmalike organisms. Phytopathology 83:834–842
Leljak-Levanic D, Jezic M, Cesar V et al (2010) Biochemical and epigenetic changes in phytoplasma -recovered periwinkle after indole-3-butyric acid treatment. J Appl Microbiol 109:2069–2078
Lin C, Zhou T, Li H, Fan Z, Li Y, Piao C, Tian G (2009) Molecular characterisation of two plasmids from paulownia witches’ broom phytoplasma and detection of a plasmid-encoded protein in infected plants. Eur J Plant Pathol 123:321–330
Liu RN, Dong YP, Fan GQ, Zhao ZL, Deng MJ, Cao XB, Niu SY (2013) Discovery of genes related to witches broom disease in Paulownia tomentosa × Paulownia fortunei by a de novo assembled transcriptome. Plos One 8:e80238
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408
Maejima K, Oshima K, Namba S (2014) Exploring the phytoplasmas, plant pathogenic bacteria. J Gen Plant Pathol 80:210–221
McKersie BD, Crowe JH, Crowe LM (1989) Free fatty acid effects on leakage, phase properties and fusion of fully hydrated model membranes. BBA Biomembranes 982:156–160
Moreau S, Fromentin J, Vailleau F et al (2014) The symbiotic transcription factor MtEFD and cytokinins are positively acting in the Medicago truncatula and Ralstonia solanacearum pathogenic interaction. New Phytol 201:1343–1357
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5:621–628
Mou HQ, Lu J, Zhu SF, Lin CL, Tian GZ, Xu X, Zhao WJ (2013) Transcriptomic analysis of Paulownia infected by Paulownia witches’ broom phytoplasma. Plos One 8:e77217
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15:473–497
Murmu J, Wilton M, Allard G, Pandeya R, Desveaux D, Singh J, Subramaniam R (2014) Arabidopsis GOLDEN2-LIKE GLK transcription factors activate jasmonic acid JA-dependent disease susceptibility to the biotrophic pathogen Hyaloperonospora arabidopsidis, as well as JA-independent plant immunity against the necrotrophic pathogen Botrytis cinerea. Mol Plant Pathol 15:174–184
Namba S (2011) Phytoplasmas: a century of pioneering research. J Gen Plant Pathol 776:345–349
Oshima K, Kakizawa S, Nishigawa H et al (2004) Reductive evolution suggested from the complete genome sequence of a plant-pathogenic phytoplasma. Nat Genet 36:27–29
Oshima K, Maejima K, Namba S (2013) Genomic and evolutionary aspects of phytoplasmas. Front Microbiol 4:230
Prieschl EE, Baumruker T (2000) Sphingolipids: second messengers, mediators and raft constituents in signaling. Immunol Today 21:555–560
Rébeillé F, Macherel D, Mouillon JM, Garin J, Douce R (1997) Folate biosynthesis in higher plants: purification and molecular cloning of a bifunctional 6-hydroxymethyl-7,8-dihydropterin pyrophosphokinase/7,8-dihydropteroate synthase localized in mitochondria. EMBO J 16:947–957
Rismani-Yazdi H, Haznedaroglu BZ, Bibby K, Peccia J (2011) Transcriptome sequencing and annotation of the microalgae Dunaliella tertiolecta: pathway description and gene discovery for production of next-generation biofuels. BMC Genom 12:148
Schneiderová K, Šmejkal K (2014) Phytochemical profile of Paulownia tomentosa (Thunb). Steud. Phytochem Rev. doi:10.1007/s11101-014-9376-y
Schwessinger B, Zipfel C (2008) News from the frontline: recent insights into PAMP-triggered immunity in plants. Curr Opin Plant Biol 11:389–395
Slabas AR, Chase D, Nishida I et al (1992) Molecular cloning of higher-plant 3-oxoacyl-(acyl carrier protein) reductase. Sequence identities with the nodG-gene product of the nitrogen-fixing soil bacterium Rhizobium meliloti. Biochem J 283:321–326
Sugimoto H, Kondo S, Tanaka T et al (2014) Overexpression of a novel Arabidopsis PP2C isoform, AtPP2CF1, enhances plant biomass production by increasing inflorescence stem growth. J Exp Bot 65:5385–5400
Sugio A, MacLean AM, Kingdom HN, Grieve VM, Manimekalai R, Hogenhout SA (2011) Diverse targets of phytoplasma effectors: from plant development to defense against insects. Ann Rev Phytopathol 49:175–195
Suzuki T, Ishikawa K, Yamashino T, Mizuno T (2002) An Arabidopsis histidine-containing phosphotransfer HPt factor implicated in phosphorelay signal transduction: overexpression of AHP2 in plants results in hypersensitiveness to cytokinin. Plant Cell Physiol 43:123–129
Uematsu S, Sato S, Yamamoto M et al (2005) Interleukin-1 receptor-associated kinase-1 plays an essential role for Toll-like receptor TLR 7-and TLR9-mediated interferon-α induction. J Exp Med 201:915–923
Umezawa T, Sugiyama N, Mizoguchi M et al (2009) Type 2C protein phosphatases directly regulate abscisic acid-activated protein kinases in Arabidopsis. Proc Natl Acad Sci USA 106:17588–17593
Wang J, Marks BD, Kahindo C et al (2010) Genetic and serological analyses of elongation factor EF-Tu of paulownia witches’ broom phytoplasma (16SrI-D). Plant Pathol 59:972–981
Win N, Kyu K, Lee Y, Kim Y, Back C, Chung H, Jung H (2012) Reclassification of aster yellows group phytoplasmas in Korea. J Gen Plant Pathol 78:264–268
Xiang LX, He D, Dong WR, Zhang YW, Shao JZ (2010) Deep sequencing-based transcriptome profiling analysis of bacteria-challenged Lateolabrax japonicus reveals insight into the immune-relevant genes in marine fish. BMC Genom 11:472
Ye J, Fang L, Zheng H et al (2006) WEGO: a web tool for plotting GO annotations. Nucl Acids Res 34:W293–W297
Yokoyama A, Yamashino T, Amano YI, Tajima Y, Imamura A, Sakakibara H, Mizuno T (2007) Type-B ARR transcription factors, ARR10 and ARR12, are implicated in cytokinin-mediated regulation of protoxylem differentiation in roots of Arabidopsis thaliana. Plant Cell Physiol 48:84–96
Zhai XQ, Cao XB, Fan GQ (2010) Growth of Paulownia witches’ broom seedlings treated with methyl methane sulphonate and SSR analysis. Sci Silv Sin 46:176–181
Zhang YZ, Cao XB, Zhai XQ, Fan GQ (2009) DNA extraction of AFLP reaction system for the Paulownia plant. J Henan Agric Univ 43:610–614
Zhang X, Li M, Agrawal A, San KY (2011) Efficient free fatty acid production in Escherichia coli using plant acyl-ACP thioesterases. Metab Eng 13:713–722
Acknowledgments
This work was financially supported by the National Natural Science Foundation of China (Grant No. 30271082, 30571496), by the Outstanding Talents Project of Henan Province (Grant No. 122101110700), by the Transformation Project of the National Agricultural Scientific and Technological Achievement of China (Grant No. 2012GB2D000271), and by Science and Technology Innovation Team Project of Zhengzhou City (Grant No. 121PCXTD515).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by J.-H Liu.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary material 1
Figure S1 Length distribution of all-unigene in P. tomentosa (TIFF 305 kb)
Rights and permissions
About this article
Cite this article
Fan, G., Cao, X., Zhao, Z. et al. Transcriptome analysis of the genes related to the morphological changes of Paulownia tomentosa plantlets infected with phytoplasma. Acta Physiol Plant 37, 202 (2015). https://doi.org/10.1007/s11738-015-1948-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11738-015-1948-y