Abstract
The involvement of glutathione transferases (GSTs) in plant’s tolerance to abiotic stresses has been extensively studied; however, the metabolic changes occurring in the plants with altered GSTs expression have not been studied in detail. We have previously demonstrated that GmGSTU4 overexpression in tobacco plants conferred increased tolerance to herbicides, partly through its peroxidase activity. Here, we investigated GmGSTU4 transcriptional response to abiotic and chemical stimuli in soybean. Transgenic tobacco plants overexpressing GmGSTU4 were also evaluated regarding their phenotypic and metabolomics responses under salt stress. GmGSTU4 expression was highly induced after salt stress and atrazine treatment. Tobacco plants overexpressing GmGSTU4 were highly tolerant to 150 mM NaCl in vitro. Metabolomics comparison of plants growing under optimal conditions, indicating a shift of the transgenic plants metabolism towards the metabolic profiles observed under stress, increased concentration of precursors of glutathione biosynthesis and hexose concentration reduction. Under salt stress, transgenic plants maintained their cellular homeostasis in contrast to wild-type plants which exhibited deregulated energy metabolism. The metabolic response of the transgenic plants was characterized by higher concentration of protective metabolites such as proline and trehalose and greater induction of the oxidative pentose phosphate pathway. These results confirm GmGSTU4 contribution to salt stress tolerance, and outline a regulatory role that primes plants towards the up-regulation of protective and detoxification mechanisms under abiotic stress.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants’ responses to stress conditions constitute of both sensing the environmental adversity and the downstream activation of signaling cascades (Dietz 2008) which result in adaptive responses such as the up-regulation of the reactive oxygen species (ROS) detoxification network, to counteract oxidative stress, commonly generated under these conditions (Gill and Tuteja 2010; Suzuki et al. 2012).
Glutathione transferases (GSTs, EC 2.5.1.18) superfamily of enzymes forms a major part of plants detoxification network. They are commonly cytosolic, dimeric enzymes, and their major catalytic function is the conjugation of the tripeptide glutathione (GSH; γ-Glu–Cys–Gly) to various electrophilic (and frequently cytotoxic) substrates (Cummins et al. 2011; Edwards et al. 2000). Owing to this function, they were originally described as enzymes facilitating the detoxification of herbicides in crop plants through GSH conjugation (Dixon et al. 1998).
GSTs appear to have a significant role in plants’ adaptation under abiotic stress as many isoenzymes are found to be differentially expressed under these conditions (Chi et al. 2011; Sappl et al. 2009). Their protective function can be exerted by both their GSH conjugating activity and GSH-dependent peroxidase activity on cytotoxic alkenals and fatty acid hydroperoxides, respectively, generated under stress (Chronopoulou et al. 2011, 2012).
Multiple evidence concerning GSTs involvement in abiotic stress tolerance exists: first their induced expression after different stress stimuli (Csiszár et al. 2014; Lan et al. 2009), the association of different allelic forms of GST isoenzymes with stress tolerance (Kim et al. 2011) and the second comes by the overexpression of several stress responsive isoenzymes resulting in increased stress tolerance of the respective transgenic plants (Diao et al. 2011; George et al. 2010; Roxas et al. 2000).
Despite the numerous evidences for the involvement of GSTs in plants’ protection against abiotic stress, there is a limited knowledge on the underlying mechanisms. It is postulated to be mostly exerted through their GSH-dependent peroxidase activity (Cummins et al. 1999), however, pleiotropic effects have been observed in the activity of antioxidant enzymes (Roxas et al. 2000) and other components of the plants’ stress tolerance network (Diao et al. 2011). The cellular redox state is recently acknowledged as a central hub in the coordination of plants’ developmental and metabolic processes both in the presence or absence of stress conditions, through post-translational modification of proteins and transcription factors which alter their biological functions (Tanou et al. 2009; Yun et al. 2011). GSTs have been shown to modulate redox homeostasis by alterations in GSH content and redox state (Chen et al. 2012). It is suggested that through this function they can strongly affect plant development (Jiang et al. 2010) as well as tolerance to abiotic stresses (Chen et al. 2012). Given the complexity of plant stress responses, tools for high-throughput analysis such as metabolomics enable the dissection of stress responses and the identification of major modules that underlie increased tolerance (Obata and Fernie 2012). Metabolites are the end products of gene expression and protein activities and therefore are the penultimate regulatory components for growth and development. It has been shown recently that metabolites apart from their typical properties are also involved in major signaling processes (Liu et al. 2010; Pajerowska-Mukhtar et al. 2012) and therefore can have pleiotropic effects beyond their “regular” function. Moreover, metabolomics can uncover silent phenotypes, which can be informative in gene families which have members with redundant functions, such as the GSTs, as demonstrated by the observation of metabolic perturbations under stress in Arabidopsis plants with multiple silenced GSTs (Sappl et al. 2009). As GSTs increasingly appear to be involved in endogenous metabolic processes (Dixon et al. 2010), metabolomics could be the tool to uncover novel metabolic functions.
The isoenzyme GmGSTU4 (a tau class GST) from soybean has been well characterized in terms of substrate specificity, kinetic mechanism and 3D structure. GmGSTU4 which catalyzes a broad range of reactions exhibits high glutathione transferase and glutathione peroxidase activities (Axarli et al. 2009a, b, 2010). We have previously overexpressed the GmGSTU4 isoenzyme in tobacco plants (Benekos et al. 2010) examining their tolerance against herbicides. Herein, we further investigated GmGSTU4 transcriptional responses to various stress stimuli in wild-type (WT) soybean. Furthermore, we have evaluated under salt stress, the tolerance and the metabolite profile of transgenic tobacco plants overexpressing GmGSTU4. The metabolomics analyses revealed significant metabolic alterations in the transgenic tobacco plants, both under non-stress and salt stress conditions, towards stress anticipation and up-regulation of diverse protective mechanisms and homeostasis maintenance. These findings provide insights into the metabolic alterations underlying GSTs protective effects against salt stress and aid further biotechnological approaches in enhancing salt tolerance in crops by targeting the underlying metabolic pathways.
Materials and methods
Expression analysis of GmGSTU4 gene in soybean
Soybean seedlings were grown in perlite in a growth chamber under controlled conditions (25 °C, 16 h light/8 h dark cycle) and were subsequently subjected to various stresses as a means to assess the expression of GmGSTU4 gene. Fourteen-day-old soybean plants were individually transferred into test tubes and exposed to the following xenobiotics: 10 mM atrazine, 10 mM 1-chloro-2,4-dinitrobenzene (CDNB), 10 mM H2O2. Osmotic stress was achieved by the addition of 100 mM NaCl. For temperature stress treatments, plants were incubated at 40 and 4 °C. Plants of the same age that were not exposed to stress were included as experimental controls. Following 4 h of incubation, leaf samples from individually treated and control plants were collected. Samples were immediately frozen in liquid nitrogen and maintained at −80 °C.
Total RNA isolation was performed on the collected leaf samples of soybean plants with the SV Total RNA Isolation Kit (Promega). To eliminate contamination of genomic DNA, RNA samples were treated with DNase I at 37 °C for 45 min. First-strand cDNA was reverse transcribed using SuperScript II and random hexanucleotides. GmGSTU4 transcripts were amplified using gene-specific primers. PCR reactions were performed on a Bio-Rad Mini Opticon cycler using Kapa SYBR Fast qPCR kit (Kapa Biosystems). Amplification cycles included a first denaturation cycle of 10 min at 95 °C, then 40 cycles composed of 15 s at 95 °C and 1 min at 60 °C. The expression G. max ubiquitin gene (GmUb) was used as a control for relative gene expression quantification. Relative expression was calculated with the \(2^{{ - \Delta \Delta C_{\text{t}} }}\) method (Livak and Schmittgen 2001).
Transgenic tobacco plants overexpressing GmGSTU4
Wild-type and T1 transgenic tobacco var. “Basmas” overexpressing GmGSTU4 plants were grown in vitro as described (Benekos et al. 2010). After preliminary experiments evaluating salt stress tolerance phenotypes of the transgenic lines reported in our previous study (Benekos et al. 2010) transgenic line L3 was used in all the subsequent experiments.
The expression of the GmGSTU4 gene under the control of the constitutive CaMV 35S promoter was examined in the transgenic tobacco plants. Shoot and root RNA isolation, cDNA synthesis and primers used were carried out as described in Benekos et al. (2010). Quantitative expression analysis of GmGSTU4 transgene was performed using Kapa SYBR Fast qPCR Kit (KapaBiosystems) in a Corbett Rotor Gene 6000 Real-Time PCR thermocycler. The expression of the tobacco β-actin gene was used as a control for relative gene expression quantification. Amplification of both genes was performed for 30 cycles of 5 s at 95 °C, 20 s at 61 °C, and 3 s at 72 °C. Relative expression was determined as described above.
Salt stress treatments
Three-week-old in vitro grown WT and transgenic tobacco plants were subjected to salinity stress after transplantation to MS medium supplemented with 150 and 300 mM NaCl. Stress tolerance was assessed after 30 days of growth measuring growth parameters such as shoot length and total root fresh weight.
Stress treatments in plants used for the metabolite analysis were slightly modified. Two-week-old plants were transplanted in MS medium containing 100 mM NaCl. Leaves used for the metabolite analyses were harvested 20 days after imposition of stress.
Metabolite extraction, derivatization and GC–MS analysis
Plant leaves were harvested 20 days after imposition of 100 mM NaCl in vitro and immediately frozen in liquid nitrogen and lyophilized. Approximately, 10 mg dry weight of tissue was ground in liquid nitrogen. Ground samples were extracted with 380 μL methanol and 20 μL ribitol in methanol (0.2 mg mL−1). Samples were incubated at 70 °C for 15 min with continuous shaking. The addition of 200 μL chloroform was followed by a further incubation at 37 °C for 5 min under continuous shaking. After addition of 400 μL ddH2O, samples were vortexed and then centrifuged at 18,000g for 5 min at room temperature. The aqueous phase containing the polar metabolite fraction was transferred into new eppendorf tubes and dried by nitrogen gas. For derivatization, dried samples were re-suspended in 25 µl methoxyamine-HCl (MOX) (20 mg mL−1 in pyridine), and incubated at 30 °C for 90 min with continuous gentle agitation. This was followed by addition of 50 µl of N-methyl-N-(trimethylsilyl)-trifluoroacetamide (MSTFA) and incubated at 37 °C for 30 min with continuous gentle agitation. Finally, 10 µl of n-alkane mix was added for determination of retention indexes (RIs). Gas chromatography coupled to mass spectrometry (GC–MS) measurements were performed in a HP6890 GC coupled to a HP 5973 MS.
Data analysis
For all treatments tested, five biological replications were performed. The chromatograms were evaluated automatically using the AMDIS software and metabolites were identified using the Golm metabolome database (Kopka et al. 2005; Schauer et al. 2005). Results were expressed as a response that corresponds to the ratio between the areas of the target metabolite divided by the area of the reference metabolite (ribitol, m/z 319) and reported relative to the dry weight.
Principal component analysis (PCA) was performed on the relative metabolite concentration data using the Unscrambler 9.5 software (CAMO Software Inc., NJ, USA). Statistically significant differences between genotypes and treatments for each metabolite were validated by the Student’s t test (p ≤ 0.05).
Results
GmGSTU4 expression analysis under stress conditions in soybean
Given that the function of GSTs is linked to the detoxification of xenobiotics and the response to abiotic stresses (Chronopoulou et al. 2012, 2014), expression analyses for GmGSTU4 gene were carried out under normal as well as under stress conditions. Transcript levels were measured by means of real-time RT-qPCR analysis and normalized by the transcript levels of the reference GmUbiquitin gene. GmGSTU4 gene was found to be constitutively expressed at low level in soybean suggesting that the respective protein has a specific functional role under different conditions. To evaluate the expression abundance of GmGSTU4 in response to different chemical stimulants (atrazine, CDNB, H2O2) and stress conditions (osmotic stress, cold and heat shock), young soybean plants were exposed to various stress factors. Osmotic stress, caused by the addition of NaCl, induced an approximately 153-fold increase in transcript accumulation of GmGSTU4 (Fig. 1). Stress induced by the herbicide atrazine caused 17-fold increase in transcript accumulation. However, the abundance of transcripts was unaffected in response to different chemical stimulants (CDNB, H2O2) and heat shock.
Tolerance of tobacco plant overexpressing GmGSTU4 to salt stress
In a previous study, Benekos et al. (2010) had reported the development of three independent transgenic lines overexpressing GmGSTU4. Line L3 (hereafter mentioned as GST-OX) exhibited the greater increase in GST and GPOX activity and tolerance to herbicide treatments. While this line had no statistically significant difference with other lines tested on salt tolerance (Supp. Fig. 1a), it carried a single transgene insert, and thus it was selected and used in all the subsequent experiments presented here.
Transgene expression, determined quantitatively by real-time qPCR, was found to be 193-fold greater than actin in the shoots of the transgenic plants. Expression in the roots was in average twofold lower than in the shoots (yet, it was still 87-fold higher compared to actin), without being statistically significantly different compared to the shoot. We evaluated the tolerance of transgenic plants to salinity stress in vitro by transplanting three-week-old seedlings to MS medium supplemented with 150 and 300 mM NaCl, respectively. GST-OX plants, when grown in 150 mM NaCl, exhibited increased tolerance compared to WT plants (Fig. 2a), with growth traits measured, such as shoot length and total fresh weight, being almost equivalent with those of plants grown under normal conditions. Particularly, shoot length, total fresh weight and root fresh weight were 48.9, 22.9 and 52.9 %, respectively, greater in transgenic plants compared to WT plants (Fig. 2b). Doubling of the NaCl concentration to 300 mM resulted in severe growth reduction of both WT and GST-OX plants, with the latter exhibiting, although greatly inhibited, 60.5 % increased shoot length, compared to WT plants, while no significant differences were observed for total fresh weight and root fresh weight.
Tolerance of wild-type (WT) and GmGSTU4 overexpressing tobacco plants (GST-OX) to salt stress: a phenotype and b growth traits of WT and transgenic tobacco plants grown for 30 days in MS medium supplemented with 150 or 300 mM NaCl. Data are the mean ± SE (n = 4). Asterisks significant differences at p ≤ 0.05 between WT and transgenic tobacco plants at the same treatment
Metabolomics analyses
The increased tolerance of GST-OX plants to salinity stress in vitro led us to further investigate the response of WT and GST-OX plants through the study of their metabolite changes during stress. Plants were exposed to 100 mM NaCl. This concentration has caused significant differences in tolerance between control and GST-OX plants (Supp. Fig. 1a), while at the same time allows us to detect metabolite changes due to stress response and acclimatization, and not due to indirect effects of extreme stress conditions such as toxicity due to plasmolysis caused by high osmolarity (caused by the salt concentration) (Munns 2002) and cell death (Quan et al. 2008). Furthermore, plants were grown for a long period under stress (20 days), to monitor stable changes in metabolite concentration, in contrast to shock and rapid changes that occur under short period of stress, which allows us to study the acclimation and the achievement of cellular homeostasis under stress conditions (Sanchez et al. 2008; Shavrukov 2013; Skirycz et al. 2011).
Analysis of leaf extracts carried out by GC/MS identified a total of 76 polar metabolites (Supp. Table 1). On this basis, we examined differences in the metabolome between WT and GST-OX plants under control conditions and after salinity stress imposition.
Effect of GmGSTU4 overexpression on metabolome under non-stress conditions
GmGSTU4 overexpression had a significant effect on plant metabolism under stress-free conditions, as indicated by the 11 out of 76 metabolites (14.5 %) identified having significantly different concentration (Table 1). Metabolites significantly increased in GST-OX plants were acetyl-l-serine, a precursor of the cysteine and glutathione biosynthesis, and glycine (8.1- and 2.2-fold increase, respectively), the TCA cycle intermediate fumaric acid (6.3-fold), the osmoprotectant hydroquinone (5.3-fold) and the precursor of glycerolipid biosynthesis glycerol-3-phosphate (2.9-fold). The levels of hexoses, glucose and fructose were decreased significantly 2.9- and 2.8-fold, respectively, as well as the precursor of ascorbate biosynthesis mannose (8.3-fold). Two metabolites (palatinose, saccharic acid) were specifically identified in the transgenic plants under control conditions. When all the metabolite data were subjected to Principal Component Analysis (PCA), the effect of GmGSTU4 overexpression was revealed, indicating a shift of transgenic plants metabolic profile towards that observed under stress conditions. Thus, the overexpression of GmGSTU4 has functioned as a signal in the plant that it is under stress and responded accordingly by increasing certain metabolites to protect itself (Fig. 3a).
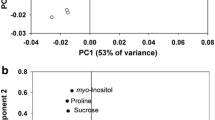
Principal component analysis (a) of leaf metabolites identified in WT and GmGSTU4 overexpressing tobacco plants grown for 20 days in MS medium and MS supplemented with 100 mM NaCl. Principal components (PC) 1 and 2 account for 82 % of the variance in the data. b Metabolite loadings plot for PC1 and PC2. Loading values for specific metabolites are indicated
Effect of salinity stress in vitro in the metabolome of WT tobacco plants
The imposition of salinity stress resulted in a major alteration of WT tobacco plants metabolism, affecting the concentration of 25 % of the metabolites identified, which were distributed in all the different functional categories suggesting strong effect of salt stress in the central metabolism (Table 1). Metabolite data were analyzed with PCA. The two highest ranking principal components accounted for 82 % of the total variance in the metabolite datasets for salinity stress. PCA revealed a clear separation of WT plants metabolome grown in 100 mM NaCl from those grown under non-stress conditions (Fig. 3a).
Differences between treatments were resolved by PC1, with the salt-treated plants being towards the negative side of PC1 and non-stressed plants on the positive side. Investigation of the loadings for PC1 revealed positive values for sucrose and fructose and negative for serine, meaning that differences can be attributed to the reduction in the concentration of these hexoses and the accumulation of serine in salt-stressed WT plants (Fig. 3b). Comparison of changes in individual metabolites concentration under salt stress using the t test (p ≤ 0.05) confirmed the results of the PCA. Specifically of the 19 metabolites whose concentration was significantly altered, nine increased and ten decreased (Table 1). Among the metabolites with increased concentration were the sugar alcohol mannitol (133.2-fold), the amino acid serine and its derivative acetyl-l-serine (2.8- and 6-fold, respectively) and the nitrogen metabolism intermediate allantoin (5.7-fold). On the other hand, a significant decrease was observed in the concentration of hexose sugars glucose and fructose and the heptose sedoheptulose (6.7-, 9.1- and 10.7-fold, respectively), the TCA cycle intermediate citric acid (2.8-fold) and the precursor of various amino acids aspartic acid (2.8-fold).
The metabolic response of GST-OX tobacco plants under salt stress and in comparison to the WT plants
GmGSTU4 overexpression had a profound effect on tobacco plants metabolome compared to WT plants, under salinity stress. A similar number of metabolites were significantly altered in the GST-OX plants in comparison with the WT (17 compared to 19). However, in transgenic plants, there was a significant reduction in the number of metabolites that were decreased under stress (3 compared to 10, respectively), indicating a protective effect of GmGSTU4 overexpression to the plants homeostatic mechanisms under salt stress (Table 1).
Comparison of changes in individual metabolites revealed that of the 14 metabolites increased in transgenic plants, six are unique (glycerol, nicotinamide, proline, sedoheptulose, trehalose, malonic acid), indicating distinct metabolic responses in the latter (Supp. Fig. 2a). Almost all of these metabolites have been previously shown to contribute in increased tolerance to salt stress, further justifying the phenotypes observed in GST-OX plants. On the other hand, all three metabolites (putrescine, valine and fructose) that decreased are common with the WT plants (Supp. Fig. 2b). These results point to a distinct response of GST-OX plants to salt stress, and an efficient up-regulation of the protective mechanisms.
Further quantitative comparison with PCA showed that GST-OX plants metabolic profile was clearly separated from that of WT plants under salt stress, with the former being on the positive side of PC2, while the latter on the negative (Fig. 3a). Metabolites that contributed to this separation were mainly glucose and proline whose concentration was significantly greater in GST-OX plants (2.7- and 1.8-fold, respectively). Other metabolites that were significantly increased compared to WT plants were the oxidative pentose phosphate pathway intermediates phopshogluconic acid (GST-OX specific) and sedoheptulose (17.1-fold increase), the osmoprotectants glycerol and hydroquinone (3.2- and 3.7-fold, respectively) and the hydrophilic amino acids glycine and hydroxyproline (both 2.2-fold). Notably, the compatible solute mannitol was 2.8-fold decreased compared to WT plants.
Discussion
The physiological roles of the plant GST superfamily of enzymes still remain to be fully understood, despite the functional characterization of a large number of GST genes from different plant species. Even though their function in herbicide detoxification through GSH conjugation is well characterized both enzymatically (Cummins et al. 2011; Skipsey et al. 2005), and in oxidative stress alleviation through GSH-dependent peroxidase activity (Kilili et al. 2004) their function in a cellular context and its homeostasis both under stress as well as non-stress conditions are not yet well defined (Dixon and Edwards 2010).
Salt stress resulted in a marked increase of the native GmGSTU4 gene expression in WT soybean plants, indicative of its involvement in salt stress adaptation. GmGSTU4 induction exhibited a marked specificity in response to different chemical and stress treatments, suggesting that its regulation is controlled by specific mechanisms (Wagner et al. 2002).
GmGSTU4 overexpressing transgenic tobacco plants displayed a remarkable increased tolerance to salt stress as their growth was not affected at 150 mM NaCl compared to non-stressed plants. These observations support the results from previous studies which have demonstrated increased growth capacity of plants overexpressing GST isoenzymes under salt stress (Diao et al. 2011; Roxas et al. 2000). GmGSTU4 exhibits increased GSH peroxidase activity, its overexpression in tobacco resulted in increased tolerance to photobleaching herbicides that are not detoxified by GSH conjugation (Benekos et al. 2010), and potentially provides transgenic plants with an efficient antioxidant mechanism to counteract oxidative stress generated under abiotic stresses like the increased salt conditions they were subjected to.
We have further dissected the increased stress tolerance of GST-OX plants using metabolomics. Metabolomics is a recent addition along the other -omics technologies, offering substantial coverage of a large number of metabolites belonging to different structural and functional groups (Hagel and Facchini 2008). As metabolites are the end products of gene expression, metabolomics are invaluable for the dissection of the observed phenotype (Schauer and Fernie 2006). Along with their functional properties as structural molecules or their involvement in biochemical reactions, metabolites have been recently acknowledged as signaling molecules with pleiotropic effects on major developmental and stress acclimation processes, both in the cellular and whole plant context (Wahl et al. 2013; Xiao et al. 2012).
Comparing the metabolome of WT and GmGSTU4 overexpressing plants under optimal growth conditions with PCA indicated a shift of the metabolism of the latter towards the metabolic fingerprints obtained under stress conditions. Many metabolites were differentially accumulated under control conditions, among them precursors of major cellular redox regulators such as O-acetyl-serine and glycine, precursors of glutathione (Noctor et al. 2012) and mannose, precursor of ascorbic acid (Wheeler et al. 1998). Possibly the increased abundance of GmGSTU4, which possesses a relatively high GSH-dependent peroxidase activity, perturbs the redox balance even under control conditions, which is translated to altered flux through GSH biosynthesis pathway (Mhamdi et al. 2010; Roxas et al. 2000). The decreased hexose content resembles the metabolic response under oxidative stress (Baxter et al. 2007) and the increased accumulation of osmoprotectants (hydroquinone) further points that transgenic plants metabolic homeostasis was altered towards a stress anticipation state, probably due to the differential response observed between WT and GST-OX plants under salt stress. Thus, apart from the per se protective effect of GmGSTU4 against oxidative stress that shielded transgenic plants against salt stress, GmGSTU4 overexpression under non-stress conditions resulted in a primed response that potentially aids for a rapid response against salt stress. This is evident by the metabolic response of WT plants in which salt stress resulted in the accumulation of O-acetyl-serine and hydroquinone, metabolites that were already in increased concentration in the transgenic plants under optimal conditions.
Interestingly, glycerol 3-phosphate levels, which have recently been implicated in the induction of systemic acquired resistance (Chanda et al. 2011), were threefold higher in the transgenic plants, and may be the link to the positive contribution of GSTs to disease resistance (Chassot et al. 2008; Dean et al. 2005; Wisser et al. 2011).
Salt stress resulted in the deregulation of major metabolic processes in WT plants, namely nitrogen assimilation, glycolysis and the TCA cycle that are crucial for energy homeostasis and resumption of plant growth under stress (Widodo et al. 2009). On the other hand, none of these perturbations were observed in the GST-OX plants. Moreover, the latter had uniquely upregulated metabolites with unambiguous relationship with stress tolerance such as proline, trehalose, and glycerol. Proline alone has been shown to have multiple functions under stress such as osmoprotective, scavenging of ROS, and maintaining the redox balance (Szabados and Savouré 2010). Interestingly, transgenic tobacco plants overexpressing a GST gene from the halophyte Limonium bicolor, also exhibited increased accumulation of proline under salt stress (Diao et al. 2011). Trehalose contribution to adaptation and tolerance to abiotic stresses, through protective and signaling functions, is also established (Paul et al. 2008). In addition, several metabolites with protective function such as hydroquinone and hydroxyproline (Warren et al. 2012) reached higher levels in GST-OX plants as well as metabolites involved in energy homeostasis such as glucose and nicotinic acid compared to the WT plants. Both metabolites are further implicated in signaling functions that regulate plant development (Warren et al. 2012). The accumulation of sedoheptulose and phosphogluconic acid implies increased flux through the oxidative pentose pathway that generates NADPH (Kruger and Von Schaewen 2003), a reductant source for major antioxidant pathways in plants. Mannitol was the only osmoprotectant with a decreased concentration in transgenic plants. However, there is evidence that its accumulation is mostly related to susceptibility to stress (Chen et al. 2007), therefore this finding adds up to the fact that transgenic plants were less affected by salt stress.
The compilation of these observations points to the conclusion that the increased stress tolerance of GST-OX was an outcome of the up-regulation of diverse metabolic components. Thus, the manipulation of plant stress tolerance by overexpression of antioxidant enzymes does not simply involve the effective scavenging of ROS generated under stress but rather complex mechanisms that have pleiotropic effects. In addition to this complexity, we have to consider that GSTs are also implicated actively in endogenous metabolic processes such as binding and transport of anthocyanin, flavonoids and porphyrins (Cummins et al. 2013; Dixon et al. 2010), phytoalexin biosynthesis (Su et al. 2011), as well as binding and glutathione conjugation of reactive electrophile species (Mueller et al. 2008). Almost all those above-mentioned metabolites appear to have protective and/or signaling functions under stress conditions (Böttcher and Pollmann 2009; Phung et al. 2011). As the crystallographic structural characterization of GmGSTU4 showed a ligand site (L-site) (Axarli et al. 2009a), we cannot exclude that this enzyme exerts an above-listed function, which can in part result in the metabolic alterations observed under optimal growth conditions and salt stress.
In conclusion, the functional characterization of GmGSTU4 by its overexpression in tobacco plants confirmed its protective role to salinity stress which was strongly suggested by its strong salt stress expression induction in soybean. Further metabolomics analysis of the transgenic tobacco plants revealed pleiotropic effects of GmGSTU4 overexpression with multiple pathways affected even under optimal growth conditions, demonstrating that increased tolerance to salt was in part a result of metabolic priming. Moreover, given the signaling functions of many metabolites that were in higher abundance in transgenic plants under salt stress, the protective function of GmGSTU4 was potentially further amplified by these pleiotropic metabolic responses (Fig. 4). Biotechnological manipulation and fine-tuning these underlying metabolic alterations may further increase the potential of GSTs in conferring abiotic stress tolerance.
The unbiased identification of GST isozymes by forward genetics screens for both abiotic (Kim et al. 2011) and biotic (Wisser et al. 2011) stress tolerance points their major universal functions under stress and justifies further detailed studies on their physiological roles for crop improvement.
Author contribution statement
Christos Kissoudis: performed the experiments, analyzed the data, wrote the paper. Chrissanthi Kalloniati: performed the experiments, analyzed the data. Emmanouil Flemetakis: performed the experiments, analyzed the data. Nikolaos E. Labrou: designed the study, supervised the study, wrote the paper. Panagiotis Madesis: performed the experiments, analyzed the data, wrote the paper. Athanasios Tsaftaris: supervised the study, wrote the paper. Irini Nianiou-Obeidat: designed the study, supervised the study, wrote the paper.
References
Axarli I, Dhavala P, Papageorgiou AC, Labrou NE (2009a) Crystallographic and functional characterization of the fluorodifen-inducible glutathione transferase from Glycine max reveals an active site topography suited for diphenylether herbicides and a novel L-site. J Mol Biol 385:984–1002
Axarli I, Dhavala P, Papageorgiou AC, Labrou NE (2009b) Crystal structure of Glycine max glutathione transferase in complex with glutathione: investigation of the mechanism operating by the tau class glutathione transferases. Biochem J 422(2):247–256
Axarli I, Georgiadou C, Dhavala P, Papageorgiou AC, Labrou NE (2010) Investigation of the role of conserved residues Ser13, Asn48 and Pro49 in the catalytic mechanism of the tau class glutathione transferase from Glycine max. Biochim Biophys Acta 1804(4):662–667
Baxter CJ, Redestig H, Schauer N, Repsilber D, Patil KR, Nielsen J, Selbig J, Liu J, Fernie AR, Sweetlove LJ (2007) The metabolic response of heterotrophic Arabidopsis cells to oxidative stress. Plant Physiol 143:312–325
Benekos K, Kissoudis C, Nianiou-Obeidat I, Labrou N, Madesis P, Kalamaki M, Makris A, Tsaftaris A (2010) Overexpression of a specific soybean GmGSTU4 isoenzyme improves diphenyl ether and chloroacetanilide herbicide tolerance of transgenic tobacco plants. J Biotechnol 150:195–201
Böttcher C, Pollmann S (2009) Plant oxylipins: plant responses to 12-oxo-phytodienoic acid are governed by its specific structural and functional properties. FEBS J 276:4693–4704
Chanda B, Xia Y, Mandal MK, Yu K, Sekine KT, Gao QM, Selote D, Hu Y, Stromberg A, Navarre D, Kachroo A, Kachroo P (2011) Glycerol-3-phosphate is a critical mobile inducer of systemic immunity in plants. Nat Genet 43:421–429
Chassot C, Buchala A, Schoonbeek HJ, Métraux JP, Lamotte O (2008) Wounding of Arabidopsis leaves causes a powerful but transient protection against Botrytis infection. Plant J 55:555–567
Chen Z, Cuin TA, Zhou M, Twomey A, Naidu BP, Shabala S (2007) Compatible solute accumulation and stress-mitigating effects in barley genotypes contrasting in their salt tolerance. J Exp Bot 58:4245–4255
Chen JH, Jiang HW, Hsieh EJ, Chen HY, Chien CT, Hsieh HL, Lin TP (2012) Drought and salt stress tolerance of an Arabidopsis glutathione S-transferase U17 knockout mutant are attributed to the combined effect of glutathione and abscisic acid. Plant Physiol 158:340–351
Chi Y, Cheng Y, Vanitha J, Kumar N, Ramamoorthy R, Ramachandran S, Jiang SY (2011) Expansion mechanisms and functional divergence of the glutathione S-transferase family in sorghum and other higher plants. DNA Res 18:1–16
Chronopoulou E, Axarli I, Nianiou-Obeidat I, Madesis P, Tsaftaris A, Labrou NE (2011) Structure and antioxidant catalytic function of plant glutathione transferases. Curr Chem Biol 5:64–74
Chronopoulou E, Madesis P, Asimakopoulou B, Platis D, Tsaftaris A, Labrou NE (2012) Catalytic and structural diversity of the fluazifop-inducible glutathione transferases from Phaseolus vulgaris. Planta 235:1253–1269
Chronopoulou E, Madesis P, Tsaftaris A, Labrou NE (2014) Cloning and characterization of a biotic-stress-inducible glutathione transferase from Phaseolus vulgaris. Appl Biochem Biotechnol 172:595–609
Csiszár J, Horváth E, Váry Z, Gallé T, Bela K, Brunner S, Tari I (2014) Glutathione transferase supergene family in tomato: salt stress-regulated expression of representative genes from distinct GST classes in plants primed with salicylic acid. Plant Physiol Biochem 78:15–26
Cummins I, Cole DJ, Edwards R (1999) A role for glutathione transferases functioning as glutathione peroxidases in resistance to multiple herbicides in black-grass. Plant J 18:285–292
Cummins I, Dixon DP, Freitag-Pohl S, Skipsey M, Edwards R (2011) Multiple roles for plant glutathione transferases in xenobiotic detoxification. Drug Metab Rev 43:266–280
Cummins I, Wortley DJ, Sabbadin F, He Z, Coxon CR, Straker HE, Sellars JD, Knight K, Edwards L, Hughes D, Kaundun SS, Hutchings SJ, Steel PG, Edwards R (2013) Key role for a glutathione transferase in multiple-herbicide resistance in grass weeds. Proc Natl Acad Sci USA 110:5812–5817
Dean JD, Goodwin PH, Hsiang T (2005) Induction of glutathione S-transferase genes of Nicotiana benthamiana following infection by Colletotrichum destructivum and C. orbiculare and involvement of one in resistance. J Exp Bot 56:1525–1533
Diao G, Wang Y, Wang C, Yang C (2011) cloning and functional characterization of a novel glutathione S-transferase gene from Limonium bicolor. Plant Mol Biol Report 29:77–87
Dietz KJ (2008) Redox signal integration: from stimulus to networks and genes. Physiol Plant 133:459–468
Dixon DP, Edwards R (2010) Glutathione transferases. Arabidopsis Book Am Soc Plant Biol 8:e0131. doi:10.1199/tab.0131
Dixon DP, Cummins L, Cole DJ, Edwards R (1998) Glutathione-mediated detoxification systems in plants. Curr Opin Plant Biol 1:258–266
Dixon DP, Skipsey M, Edwards R (2010) Roles for glutathione transferases in plant secondary metabolism. Phytochemistry 71:338–350
Edwards R, Dixon DP, Walbot V (2000) Plant glutathione S-transferases: enzymes with multiple functions in sickness and in health. Trends Plant Sci 5:193–198
George S, Venkataraman G, Parida A (2010) A chloroplast-localized and auxin-induced glutathione S-transferase from phreatophyte Prosopis juliflora confer drought tolerance on tobacco. J Plant Physiol 167:311–318
Gill SS, Tuteja N (2010) Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol Biochem 48:909–930
Hagel JM, Facchini PJ (2008) Plant metabolomics: analytical platforms and integration with functional genomics. Phytochem Rev 7:479–497
Jiang HW, Liu MJ, Chen IC, Huang CH, Chao LY, Hsieh HL (2010) A glutathione S-transferase regulated by light and hormones participates in the modulation of arabidopsis seedling development. Plant Physiol 154:1646–1658
Kilili KG, Atanassova N, Vardanyan A, Clatot N, Al-Sabarna K, Kanellopoulos PN, Makris AM, Kampranis SC (2004) Differential roles of Tau class glutathione S-transferases in oxidative stress. J Biol Chem 279:24540–24551
Kim SI, Andaya VC, Tai TH (2011) Cold sensitivity in rice (Oryza sativa L.) is strongly correlated with a naturally occurring I99V mutation in the multifunctional glutathione transferase isoenzyme GSTZ2. Biochem J 435:373–380
Kopka J, Schauer N, Krueger S, Birkemeyer C, Usadel B, Bergmüller E, Dörmann P, Weckwerth W, Gibon Y, Stitt M, Willmitzer L, Fernie AR, Steinhauser D (2005) GMD@CSB.DB: the Golm metabolome database. Bioinformatics 21:1635–1638
Kruger NJ, Von Schaewen A (2003) The oxidative pentose phosphate pathway: structure and organisation. Curr Opin Plant Biol 6:236–246
Lan T, Yang ZL, Yang X, Liu YJ, Wang XR, Zenga QY (2009) Extensive functional diversification of the populus glutathione S-transferase supergene family. Plant Cell 21:3749–3766
Liu G, Ji Y, Bhuiyan NH, Pilot G, Selvaraj G, Zou J, Wei Y (2010) Amino acid homeostasis modulates salicylic acid-associated redox status and defense responses in Arabidopsis. Plant Cell 22:3845–3863
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408
Mhamdi A, Hager J, Chaouch S, Queval G, Han Y, Taconnat L, Saindrenan P, Gouia H, Issakidis-Bourguet E, Renou JP, Noctor G (2010) Arabidopsis GLUTATHIONE REDUCTASE1 plays a crucial role in leaf responses to intracellular hydrogen peroxide and in ensuring appropriate gene expression through both salicylic acid and jasmonic acid signaling pathways. Plant Physiol 153:1144–1160
Mueller S, Hilbert B, Dueckershoff K, Roitsch T, Krischke M, Mueller MJ, Berger S (2008) General detoxification and stress responses are mediated by oxidized lipids through TGA transcription factors in arabidopsis. Plant Cell 20:768–785
Munns R (2002) Comparative physiology of salt and water stress. Plant Cell Environ 25:239–250
Noctor G, Mhamdi A, Chaouch S, Han YI, Neukermans J, Marquez-Garcia B, Queval G, Foyer CH (2012) Glutathione in plants: an integrated overview. Plant Cell Environ 35:454–484
Obata T, Fernie AR (2012) The use of metabolomics to dissect plant responses to abiotic stresses. Cell Mol Life Sci 69:3225–3243
Pajerowska-Mukhtar KM, Wang W, Tada Y, Oka N, Tucker CL, Fonseca JP, Dong X (2012) The HSF-like transcription factor TBF1 is a major molecular switch for plant growth-to-defense transition. Curr Biol 22:103–112
Paul MJ, Primavesi LF, Jhurreea D, Zhang Y (2008) Trehalose metabolism and signaling. Annu Rev Plant Biol 59:417–441
Phung TH, Jung H, Park JH, Kim JG, Back K, Jung S (2011) Porphyrin biosynthesis control under water stress: sustained porphyrin status correlates with drought tolerance in transgenic rice. Plant Physiol 157:1746–1764
Quan LJ, Zhang B, Shi WW, Li HY (2008) Hydrogen peroxide in plants: a versatile molecule of the reactive oxygen species network. J Integr Plant Biol 50:2–18
Roxas VP, Lodhi SA, Garrett DK, Mahan JR, Allen RD (2000) Stress tolerance in transgenic tobacco seedlings that overexpress glutathione S-transferase/glutathione peroxidase. Plant Cell Physiol 41:1229–1234
Sanchez DH, Siahpoosh MR, Roessner U, Udvardi M, Kopka J (2008) Plant metabolomics reveals conserved and divergent metabolic responses to salinity. Physiol Plant 132:209–219
Sappl PG, Carroll AJ, Clifton R, Lister R, Whelan J, Harvey Millar A, Singh KB (2009) The Arabidopsis glutathione transferase gene family displays complex stress regulation and co-silencing multiple genes results in altered metabolic sensitivity to oxidative stress. Plant J 58:53–68
Schauer N, Fernie AR (2006) Plant metabolomics: towards biological function and mechanism. Trends Plant Sci 11:508–516
Schauer N, Steinhauser D, Strelkov S, Schomburg D, Allison G, Moritz T, Lundgren K, Roessner-Tunali U, Forbes MG, Willmitzer L, Fernie AR, Kopka J (2005) GC-MS libraries for the rapid identification of metabolites in complex biological samples. FEBS Lett 579:1332–1337
Shavrukov Y (2013) Salt stress or salt shock: which genes are we studying? J Exp Bot 64:119–127
Skipsey M, Cummins I, Andrews CJ, Jepson I, Edwards R (2005) Manipulation of plant tolerance to herbicides through co-ordinated metabolic engineering of a detoxifying glutathione transferase and thiol cosubstrate. Plant Biotechnol J 3:409–420
Skirycz A, Vandenbroucke K, Clauw P, Maleux K, De Meyer B, Dhondt S, Pucci A, Gonzalez N, Hoeberichts F, Tognetti VB, Galbiati M, Tonelli C, Van Breusegem F, Vuylsteke M, Inzé D (2011) Survival and growth of Arabidopsis plants given limited water are not equal. Nat Biotechnol 29:212–214
Su T, Xu J, Li Y, Lei L, Zhao L, Yang H, Feng J, Liu G, Rena D (2011) Glutathione-indole-3-acetonitrile is required for camalexin biosynthesis in Arabidopsis thaliana. Plant Cell 23:364–380
Suzuki N, Koussevitzky S, Mittler R, Miller G (2012) ROS and redox signalling in the response of plants to abiotic stress. Plant Cell Environ 35:259–270
Szabados L, Savouré A (2010) Proline: a multifunctional amino acid. Trends Plant Sci 15:89–97
Tanou G, Job C, Rajjou L, Arc E, Belghazi M, Diamantidis G, Molassiotis A, Job D (2009) Proteomics reveals the overlapping roles of hydrogen peroxide and nitric oxide in the acclimation of citrus plants to salinity. Plant J 60:795–804
Wagner U, Edwards R, Dixon DP, Mauch F (2002) Probing the diversity of the Arabidopsis glutathione S-transferase gene family. Plant Mol Biol 49:515–532
Wahl V, Ponnu J, Schlereth A, Arrivault S, Langenecker T, Franke A, Feil R, Lunn JE, Stitt M, Schmid M (2013) Regulation of flowering by trehalose-6-phosphate signaling in Arabidopsis thaliana. Science 339:704–707
Warren CR, Aranda I, Cano FJ (2012) Metabolomics demonstrates divergent responses of two Eucalyptus species to water stress. Metabolomics 8:186–200
Wheeler GL, Jones MA, Smirnoff N (1998) The biosynthetic pathway of vitamin C in higher plants. Nature 393:365–369
Widodo Patterson JH, Newbigin E, Tester M, Bacic A, Roessner U (2009) Metabolic responses to salt stress of barley (Hordeum vulgare L.) cultivars, Sahara and Clipper, which differ in salinity tolerance. J Exp Bot 60:4089–4103
Wisser RJ, Kolkman JM, Patzoldt ME, Holland JB, Yu J, Krakowsky M, Nelson RJ, Balint-Kurti PJ (2011) Multivariate analysis of maize disease resistances suggests a pleiotropic genetic basis and implicates a GST gene. Proc Natl Acad Sci USA 108:7339–7344
Xiao Y, Savchenko T, Baidoo EEK, Chehab WE, Hayden DM, Tolstikov V, Corwin JA, Kliebenstein DJ, Keasling JD, Dehesh K (2012) Retrograde signaling by the plastidial metabolite MEcPP regulates expression of nuclear stress-response genes. Cell 149:1525–1535
Yun BW, Feechan A, Yin M, Saidi NBB, Le Bihan T, Yu M, Moore JW, Kang JG, Kwon E, Spoel SH, Pallas JA, Loake GJ (2011) S-nitrosylation of NADPH oxidase regulates cell death in plant immunity. Nature 478:264–268
Acknowledgments
Christos Kissoudis is supported by “Alexander S. Onassis” Public Benefit Foundation scholarship. The authors would also acknowledge the Ministry of Education, Lifelong Learning and Religious Affairs for the financial assistance provided. This work was performed within the action THALES: “Glutathione transferases: multifunctional molecular tools in red and green biotechnology”. This action falls under the Operational Programme ‘‘Education and Lifelong Learning’’ and is co-funded by the European Social Fund and National Resources.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by T. Moriguchi.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11738_2015_1852_MOESM1_ESM.tif
Supplementary Fig. 1. a) Fresh weight of WT and GmGSTU4 overexpressing tobacco lines L2 and L3 grown in MS medium supplemented with 150 or 300 mM NaCl, b) line L3 grown in MS medium supplemented with 100 mM NaCl. ± . Asterisks depict significant differences at P ≤ 0.05 between WT and transgenic tobacco plants at the same treatment. (TIFF 4499 kb)
11738_2015_1852_MOESM2_ESM.tif
Supplementary Fig. 2. Venn diagram representation of metabolites commonly or differentially increased (a) or decreased (b) in the leaves of WT and transgenic tobacco plants grown for 20 days in MS supplemented with 100 mM NaCl. (TIFF 180 kb)
11738_2015_1852_MOESM3_ESM.xls
Supplementary Table 1. List of total annotated metabolites identified in WT and GmGSTU4 overexpressing tobacco plants in control and salt stress conditions. Color highlighted relative response ratios indicate statistically significant differences (P ≤ 0.05). (XLS 35 kb)
Rights and permissions
About this article
Cite this article
Kissoudis, C., Kalloniati, C., Flemetakis, E. et al. Stress-inducible GmGSTU4 shapes transgenic tobacco plants metabolome towards increased salinity tolerance. Acta Physiol Plant 37, 102 (2015). https://doi.org/10.1007/s11738-015-1852-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11738-015-1852-5