Summary
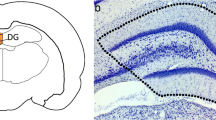
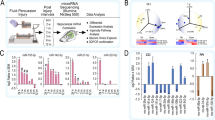
The changes of microRNA expression in rat hippocampus after traumatic brain injury (TBI) were explored. Adult SD rats received a single controlled cortical impact injury, and the ipsilateral hippocampus was harvested for the subsequent microarray assay at three time points after TBI: 1st day, 3rd day and 5th day, respectively. We characterized the microRNA expression profile in rat hippocampus using the microRNA microarray analysis, and further verified microarray results of miR-142-3p and miR-221 using quantitative real-time PCR. Totally 205 microRNAs were identified and up-/down-regulated more than 1.5 times. There were significant changes in 17 microRNAs at all three time points post-TBI. The quantitative real-time PCR results of miR-142-3p and miR-221 indicated good consistency with the results of the microarray method. MicroRNAs altered at different time points post-TBI. MiR-142-3p and miR-221 may be used as potentially biological markers for TBI assessment in forensic practice.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Feigin VL, Theadom A, Barker-Collo S, et al. Incidence of traumatic brain injury in New Zealand: a population-based study. Lancet Neurol, 2013,12(1):53–64
Jennekens N, de Casterlé BD, Dobbels F. A systematic review of care needs of people with traumatic brain injury (TBI) on a cognitive, emotional and behavioral level. J Clin Nurs, 2010,19(9–10):1198–1206
Fanselow MS, Dong HW. Are the dorsal and ventral hippocampus functionally distinct structures? Neuron, 2010, 65(1):7–19
Orrison WW, Hanson EH, Alamo T, et al. Traumatic brain injury: a review and high-field MRI findings in 100 unarmed combatants using a literature-based checklist approach. J Neurotrauma, 2009,26(5):689–701
Berezikov E, Chung WJ, Willis J, et al. Mammalian mirtron genes. Mol Cell, 2007,28(2):328–336
Filipowicz W, Bhattacharyya SN, Sonenberg N. Mechanisms of posttranscriptional regulation by microRNAs: are the answers in sight? Nat Rev Genet, 2008,9(2):102–114
Fineberg SK, Kosik KS, Davidson BL. MicroRNAs potentiate neural development. Neuron, 2009,64(3):303–309
Krichevsky AM. MicroRNA profiling: from dark matter to white matter, or identifying new players in neurobiology. Sci World J, 2007,7(S2):155–166
Bak M, Silahtaroglu A, Møller M, et al. MicroRNA expression in the adult mouse central nervous system. RNA, 2008,14(3):432–444
Farkas O, Povlishock JT. Cellular and subcellular change evoked by diffuse traumatic brain injury: a complex web of change extending far beyond focal damage. Prog Brain Res, 2007,161:43–59
Lim LP, Lau NC, Garrett-Engele P, et al. Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature, 2005,433(7027):769–773
Lewis BP, Burge CB, Bartel DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA target. Cell, 2005,120(1):15–20
Sempere LF, Freemantle S, Pitha-Rowe I, et al. Expression profiling of mammalian microRNAs uncovers a subset of brain-expressed microRNAs with possible roles in murine and human neuronal differentiation. Genome Biol, 2004,5(3):R13
Krichevsky AM, King KS, Donahue CP, et al. A microRNA array reveals extensive regulation of microRNAs during brain development. RNA, 2003,9(10):1274–1281
Smith B, Treadwell J, Zhang D, et al. Large-scale expression analysis reveals distinct microRNA profiles at different stages of human neurodevelopment. PLoS One, 2010, 5(6):e11109
Beezhold K, Liu J, Kan H, et al. miR-190-mediated downregulation of PHLPP contributes to arsenic-induced Akt activation and carcinogenesis. Toxicol Sci, 2011,123(2):411–420
Almog N, Briggs C, Beheshti A, et al. Transcriptional changes induced by the tumor dormancy-associated microRNA-190. Transcription, 2013,4(4):177–191
Huang B, Zhao J, Lei Z, et al. miR-142-3p restricts cAMP production in CD4+CD25-T cells and CD4+CD25+ TREG cells by targeting AC9 mRNA. EMBO Rep, 2009,10(2):180–185
Cai D, Qiu J, Cao Z, et al. Neuronal cyclic AMP controls the developmental loss in ability of axons to regenerate. J Neurosci, 2001,21(13):4731–4739
Wang F, Wang XS, Yang GH, et al. miR-29a and miR-142-3p downregulation and diagnostic implication in human acute myeloid leukemia. Mol Biol Rep, 2012,39(3):2713–2722
Kaduthanam S, Gade S, Meister M, et al. Serum miR-142-3p is associated with early relapse in operable lung adenocarcinoma patients. Lung Cancer, 2013,80(2):223–227
Lei P, Li Y, Chen X, et al. Microarray based analysis of microRNA expression in rat cerebral cortex after traumatic brain injury. Brain Res, 2009,1284:191–201
Liu X, Cheng Y, Zhang S, et al. A necessary role of miR-221 and miR-222 in vascular smooth muscle cell proliferation and neointimal hyperplasia. Circ Res, 2009,104(4):476–487
Galardi S, Mercatelli N, Giorda E, et al. miR-221 and miR-222 expression affects the proliferation potential of human prostate carcinoma cell lines by targeting p27kip1. J Biol Chem, 2007,282(32):23 716–23 724
Wong QW, Ching AK, Chan AW, et al. MiR-222 overexpression confers cell migratory advantages in hepatocellular carcinoma through enhancing AKT signaling. Clin Cancer Res, 2010,16(3):867–875
Urbich C, Kuehbacher A, Dimmeler S. Role of microRNAs in vascular diseases, inflammation, and angiogenesis. Cardiovasc Res, 2008,79(4):581–588
Chen Y, Banda M, Speyer CL, et al. Regulation of the expression and activity of the antiangiogenic homeobox gene GAX/MEOX2 by ZEB2 and microRNA-221. Mol Cell Biol, 2010,30(15):3902–3913
Castagnino P, Kothapalli D, Hawthorne EA, et al. miR-221/222 compensates for Skp2-mediated p27 degradation and is a primary target of cell cycle regulation by prostacyclin and cAMP. PLoS One, 2013,8(2):e56140
Le Sage C, Nagel R, Egan DA, et al. Regulation of the p27 (Kip1) tumor suppressor by miR-221 and miR-222 promotes cancer cell proliferation. EMBO J, 2007,26(15):3699–3708
Author information
Authors and Affiliations
Corresponding author
Additional information
The authors contributed equally to this work.
This project was supported by grants from the National Natural Science Foundation of China (No. 81102304), Doctoral Fund of Ministry of Education of China (No. 20090142120054), and Foundation of Huazhong University of Science and Technology (No. 20100573).
Rights and permissions
About this article
Cite this article
Sun, Ty., Chen, Xr., Liu, Zl. et al. Expression profiling of MicroRNAs in hippocampus of rats following traumatic brain injury. J. Huazhong Univ. Sci. Technol. [Med. Sci.] 34, 548–553 (2014). https://doi.org/10.1007/s11596-014-1313-1
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11596-014-1313-1