Abstract
Trichoderma stromaticum, T. rossicum and newly discovered species form a unique lineage in Trichoderma. Phylogenetic and phenotypic diversity in Trichoderma stromaticum are examined in the light of reported differences in ecological parameters and AFLP patterns. Multilocus phylogenetic analysis using 4 genes (tef1, rbp2, cal, chi18-5) did not reveal phylogenetic basis for the two reported divergent AFLP patterns or for ecological parameters; however, this analysis does indicate incomplete speciation with one supported clade derived from within T. stromaticum that corresponds to AFLP Group 2 of de Souza et al. (2006, Phytopathology 96:61–67). Trichoderma stromaticum is known only from tropical America and is typically found in association with Theobroma cacao infected with Moniliophthora perniciosa. It is reported here for the first time on pseudostromata of M. roreri in Peru. Strains of T. stromaticum also have been isolated as endophytes from stems of Theo. cacao. There are no recognized close relatives of T. stromaticum in tropical America. The closest relatives of T. stromaticum are collected in Africa and Thailand; somewhat more distantly related are T. rossicum and T. barbatum, both found in north temperate regions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Witches’ Broom Disease has caused considerable economic and social disruption in South America, but especially in the Brazilian state of Bahia (see reviews and references in de Souza et al. 2006, 2008; Loguercio et al. 2009a, b). The pathogen, Moniliophthora perniciosa, infects cacao trees when basidiospores germinate and penetrate all meristematic tissues, including pods and flower cushions, leading to hypertrophy and hyperplasia and loss of apical dominance (witches’ broom formation) and loss of fruit. The pathogen ultimately produces small mushrooms on the infected dead brooms and infected pods that continue the disease cycle (Purdy and Schmidt 1996; Hebbar et al. 2002; de Souza et al. 2006, 2008; Meinhardt et al. 2008). Trichoderma stromaticum colonizes dead brooms and other tissue infected with M. perniciosa and destroys the basidiocarps of the pathogen thereby reducing the inoculum (Bastos 1996; Hjorth et al. 2003). A biological control product ‘Trichovab®,’ which is based on a single strain of T. stromaticum, has been used on an experimental scale in Brazil since 1996 (de Souza et al. 2006). The strain used in this product (‘TVC’) was originally isolated in the Brazilian Amazonian region (Pará State) but its efficacy is subject to environmental conditions (Sanogo et al. 2002; Loguercio et al. 2009a, b) and its effectiveness is greatly enhanced when it is applied in combination with farm sanitation and copper-based fungicides (Medeiros et al. 2010).
de Souza et al. (2006) found that a large collection of strains of T. stromaticum from Brazil, Ecuador and Colombia could be divided between two amplified fragment length polymorphism (AFLP) groups that correlated with a number of biological differences (de Souza et al. 2006; 2008; Loguercio et al. 2009a, b) and the fact that a Hypocrea teleomorph, H. stromatica JL Bezerra et al. (Bezerra et al. 2003) is known for Group 2. These differences suggest that T. stromaticum as it is currently understood is composed of at least two cryptic species. In the present work, we apply multilocus phylogenetic and classical mycological analyses to a collection of strains of T. stromaticum to clarify the taxonomy of this species.
In the course of taxonomic work with Trichoderma/Hypocrea, we have collected or received from collaborators many strains, which are routinely identified through DNA sequence analysis. Several of these are closely related to T. stromaticum and are included in the present work.
Materials and methods
Cultures used
Although the primary emphasis of the present work is T. stromaticum, related strains and species were included when their close relationship to T. stromaticum was revealed during on-going taxonomic studies of Trichoderma. The strains used in this study are cited in Table 1. These include several cultures cited by de Souza et al. (2006) and perithecial collections. Cultures and perithecial specimens of Trichoderma stromaticum and other species were collected by the authors in Brazil (Bahia), Cameroon, Ecuador and Thailand. Cultures were also provided by colleagues in Austria, Canada, Côte d’Ivoire, Ecuador, Peru, Republic of South Africa, Russia, U.K., and USA. Single ascospores were isolated from perithecial collections on cornmeal agar (Difco or BD BBL, Franklin Lakes, NJ, USA) + 2% (w/v) dextrose (CMD) with the use of a micromanipulator. Representative cultures are deposited in Centraalbureau voor Schimmelcultures (CBS, Utrecht, and The Netherlands).
Morphological analyses
Observations of microscopic characters were made from cultures grown on CMD or SNA (low nutrient agar; Nirenberg 1976), less frequently from potato dextrose agar (PDA; Difco). Cultures to be used for micromorphological observations were incubated at 25°C under alternating cool white fluorescent light/darkness.
Material to be used for microscopic measurements was first immersed in 3% (aq.) potassium hydroxide (KOH), which was replaced with water or more KOH as the preparation dried. Observations were made with differential interference contrast, phase contrast or bright field microscopy. Helicon Focus® version 4.21.5 Pro (MP) (Helicon Soft, www.heliconfocus.com) was used to produce some composite images. To make sections of perithecial collections, small pieces of substratum with one or two stromata were rehydrated in 3% KOH. These were supported in Tissue-Tek® O.C.T. compound (Miles, Elkhart, IN, USA) on the stage of an IEC-CTF microtome cryostat; sections were made at a thickness of ca. 15 μm. Permanent preparations were made following Volkmann-Kohlmeyer and Kohlmeyer (1996).
Where possible, 30 units of each parameter were measured for each collection using Scion Image for Windows® (www.scioncorp.com). The continuous measurements are reported as extremes in brackets with the range calculated as mean plus and minus standard deviation. Computation of descriptive statistics, including 95% confidence intervals (ci), was performed using Systat 10 (Systat Software, San José, CA, USA).
Growth rates were determined on PDA and SNA at 15, 20, 25, 30 and 35°C in darkness (with intermittent light). Measurements were made at intervals of 24 up to 96 h. Colony characters were taken from colonies incubated on PDA and SNA at 25°C with alternating cool white fluorescent light and darkness (12 h:12 h) after 7−10 days; these conditions are referred to in descriptions as ‘under light.’ Color standards are from Kornerup and Wanscher (1978, K&W).
DNA extraction, PCR and sequencing
DNA from all strains included in this study was extracted using ArchivePure DNA cell/Tissue kit from 5 PRIME (Gaithersburg, MD, USA). The primers and their sequences used in this study are given in Table 2. The primers for ITS and α-actin (act) were described previously (Samuels and Ismaiel 2009). A portion of translation elongation factor 1-α (tef1) was amplified using the primers Ef728M and Ef-2. Ef7-28 M is a modified version of EF728F (Carbone and Kohn 1999) where nucleotide #4 is changed from C to Y = C/T. This modification was necessary to make the primer sequence match those for all Trichoderma species from all the sections of the genus. For rpb2, we designed primers RPB2-F2 and RPB2-R2 (Table 2). For calmodulin (cal), we used the primer CL1 (O’Donnell et al. 2000) and Cal737RM. The latter is a modified version of cal737R (Carbone and Kohn 1999) where we changed nucleotides 4, 9, and 10 from C, T, and G to Y(C/T), K (G/T), and K (G/T), respectively. For chitinase (chi18-5), we used the primers Chi18-5-1a and Chi18-5-2a (Kullnig-Gradinger et al. 2002).
The PCR mixture (20 μL) contained 10 μL of Taq 2X master mix (New England Biolabs, Ipswich, MA, USA), 1 μL of 25 mM MgCl2, 0.5 μL of 10 mM forward and reverse primers and 7.5 μL of distilled water. The reaction mixtures in 0.2-ml PCR tubes were placed in an MJ Research PCR PTC-200 thermo-cycler. The program used for all the genes was a touch down (Don et al. 1991) described previously in Samuels and Ismaiel (2009)
The PCR products were prepared for sequencing using an enzymatic purification system (Exosap-IT; USB Corporation, Cleveland, OH, USA). The purified PCR products were directly sequenced using BigDye Terminator v3.1 chemistry on an automated 3130xl Genetic Analyzer (Applied Biosystems, Foster City, CA, USA). Both strands of each amplicon were sequenced using the same primers used in the PCR reactions. For sequencing act and rpb2, two additional internal primers were used (Table 2). The sequences were assembled and edited to construct a consensus sequence using Sequencher 4.1 (Gene Codes, Madison, WI, USA). For each locus, sequences were preliminarily aligned using Clustal X version 1.8 (Thompson et al. 1997) under the default settings and saved in nexus format. The alignments were manually edited, if necessary, using MacClade 4.06 (Maddison and Maddison 2003).
Phylogenetic analyses
Data for each gene were analyzed separately and concatenated as a combined multilocus sequence (MLS). A reciprocal 70% bootstrap threshold (Mason-Gamer and Kellogg 1996; Reeb et al. 2004; de Queiroz 1993) was used to determine whether partitions could be combined into a single phylogeny. A conflict was assumed to be significant if two different relationships for the same taxa—one being monophyletic and the other non-monophyletic, both with BP ≥ 70%—were observed on each of the genealogies. If no conflict exists between the highly supported clades in individual gene genealogies, the genes sequenced likely share similar phylogenetic history and resolution, and combining the datasets can ultimately increase clade support. Trichoderma virens was selected as outgroup because of its close relationship of the species with T. stromaticum and T. rossicum (Bissett et al. 2003).
To infer the phylogenies of the selected taxa, four individual data sets (tef1, chi18-5, rpb2, cal) were used in Bayesian and maximum parsimony analyses. ITS and act were not included in the analyses because of their low polymorphism and thus few phylogenetically informative characters. Consequently, ITS and act did not provide any resolution for the Stromaticum Clade.
For the individual data sets, Bayesian inference analysis was performed as described in Samuels and Ismaiel (2009) in MrBayes version 3.1.2 (Huelsenbeck and Ronquist 2001). The substitution model for each locus was determined using jMODELTEST (Posada 2008). The models selected for tef1, rbp2, cal, chi18-5 were K80, TrN+G, TrN+I+G, and GTR+G, respectively. Two concurrent analyses of four chains (one cold and three heated) were both run for 1 million generations to ensure that the analyses were not trapped at local optima. Random starting trees were used and sampled every 100 generations. Among 10,000 trees produced, initial trees were discarded in the burn-in phase based on the plot of log-likelihood scores of the trees versus generation number as performed by TRACER v.1.5 (Rambaut and Drumond 2009) to ensure that the log-likelihood reached stable equilibrium. The exact numbers discarded in burn-in phase for each locus tree are given in Table 3. The remaining trees were used to construct a majority rule consensus tree with posterior probability of 0.95 or greater considered significant (Leache and Reeder 2002).
Maximum parsimony analysis (MP) was conducted using PAUP version 4.0b10 (Swofford 2002), using a heuristic search with the starting tree obtained via stepwise addition with random addition of 1,000 replicates, tree–bisection–reconnection (TBR) as the branch swapping algorithm, and Multrees off. All characters were unordered and of equal weight and gaps were treated as missing data. Stability of the clades (bootstrap) was assessed with 1,000 replicates, using the same MP settings. Bootstrap values greater than or equal to 70% were considered significant (Hillis and Bull 1993).
The combined dataset was also analyzed by Bayesian and maximum parsimony methods as described for the individual datasets with exceptions described below. The Bayesian analysis was performed with data partitioned into four, representing four loci. The substitution model for each locus obtained above was used with each partition. For the combined dataset, we used 5 × 106 generations in MrBayes. Convergence of log-likelihoods was also examined in TRACER v1.5.
To investigate the presence of recombination in the combined data set within the isolates in the Stromaticum clade, two tests were performed: firstly, the Partition Homogenetity Test (PHT; Farris et al. 1995) implemented in PAUP 4.0b10 (Swofford 2002). Ten thousand replicates were analyzed in a heuristic search with the addition of ten random sequences and one tree was saved per replicate. A P value of 0.01 (99% confidence) was used as a significance threshold (Cunningham 1997); and secondly, the Phi-test as implemented in SplitsTree software (Huson 1998).
Results
Phylogenetic analyses
A total of 69 strains were included in the study (Table 1). Two strains of T. virens were used as outgroup for the phylogenetic analysis. For each strain, we sequenced six unlinked genes: five were protein-coding genes and one non-coding (ITS). Summaries of the alignments for the six loci characterized in this study are shown in Table 2. Tef1 had the highest number of informative characters with 23% followed by chi18-5, cal, rpb2, act and ITS with 16.6, 14.8, 8.6, 3.6 and 3.4%, respectively. Due to the paucity of phylogenetically informative characters in the ITS and act datasets, these two loci were not used for further analyses.
Partition homogeneity test detected significant heterogeneity within the combined four-gene data set at the 99% confidence level (P = 0.01). Nonetheless, we combined the multi-locus sequence (MLS) data for the phylogenetic analyses because combining all genes can improve the accuracy of the inferred phylogenies even in the presence of significant incongruence (Cunningham 1997; Darlu and Lecointre 2002). In addition, the 70% reciprocal bootstrap indicates no topological conflicts between gene genealogies and that the datasets can be combined.
The phylogenetic trees were obtained based on individual gene datasets by two methods (Bayesian inference and maximum parsimony). The trees obtained by the two methods had essentially identical topology; therefore, for each individual gene sequence, the tree obtained from the Bayesian inference is presented but the bootstrap support from the maximum parsimony analysis are depicted on respective trees. The Bayesian trees for the individual gene trees are shown in Fig 1a−d.
Majority-rule (50%) consensus tree resulting from Bayesian analysis of a rpb2, b chi18-5, c tef1, d cal, and e MLS dataset. Branches with thick black lines represent clades with posterior probability (PP) ≥0.95 and bootstrap maximum parsimony (BS) values ≥70%. Thick gray lines represent clades with either PP ≥ 0.95 or BS 70%. Trichoderma virens G.J.S. 01-287 was used as outgroup. Support for the branches refers to posterior probability of equal or greater than 0.95 and maximum parsimony bootstrap of 70% or greater (Leache and Reeder 2002; Hillis and Bull 1993). Tree statistics are presented in Table 3. Text in red indicates perithecial collections
The monophyly of T. stromaticum (Clade A, Fig. 1a−e) was supported in all the individual gene trees. Of the subclades A−C, seen in the MLS (Fig 1e), only A was supported in more than one of the individual gene trees (rpb2, chi18-5). The tree based on rpb2 (Fig 1a) was the only gene genealogy that recovered two lineages within the Stromaticum clade (subclades A and B) supported by both methods. The phylogenetic tree based on chi18-5 (Fig 1b) was the closest to the rpb2 tree. Subclade A of this tree was identical to that of rpb2 and was supported by the two methods. However, isolates of subclade B did not form a distinct cluster. In the tef1 tree (Fig 1c), most strains of T. stromaticum fell into a single, well-supported clade; however, seven strains that were collected in Peru and Ecuador (C in Fig 1c) formed a subclade that had no statistical support by either of the two methods. This subclade was also present in the cal phylogeny (Fig 1d), where it was supported by only one of the methods (Bayesian inference posterior probability 0.96, BS was 63%). Like tef1, the cal tree did not support division of T. stromaticum into subclades A and B, although a clade of four Brazilian strains found here was also found in MLS (Fig 1d,e).
The combined tree (MLS, Fig 1e) revealed a monophyletic T. stromaticum with three highly supported internal branches (A, C, D). Clade A corresponds to AFLP Group 2 (including the ex-type strain ‘TVC’ = G.J.S. 97-183) of de Souza et al. (2006). AFLP Group 1 of de Souza et al is equivalent to poorly supported Clade B. Clade C, which appeared with weak support in tef1, and was supported by Bayesian analysis in the cal tree, was highly supported in MLS; it includes only strains from Peru and Ecuador. Clade D received strong support only in cal and MLS; it was not seen in any other trees.
Clades E (Fig 1e, G.J.S. 01-171/G.J.S. 01-166) and F (G.J.S. 01-174, G.J.S. 01-176) received varying levels of support in all single gene trees. These clades have high support by both methods in the MLS tree. Clade F, which consists of two Cameroonian ascospore strains, G.J.S. 01-174 and G.J.S. 01-176, was maintained across all four individual gene trees with high support. Clades E and F had a sister relationship in three of four individual gene trees. Only the tree based on cal did not support the sister-relationship of these two species. Clade F had a sister relationship with single strain PPRI 3559 in the chi18-5 tree with high support. Clades E and F formed a larger clade in three out of four gene trees plus the MLS tree. South African strain PPRI 3559 clustered with these two clades in three of four single gene trees and in MLS. In the cal tree, clades E, F and PPRI 3559 were unresolved with respect to each other and basal to T. stromaticum. This large clade had a sister relationship with T. stromaticum with support in three of the four individual gene trees. Therefore these five African strains plus G.J.S. 01-225 from Thailand are considered to be the closest relatives of T. stromaticum.
Trichoderma rossicum and Clade G (DAOM 230008, G.J.S. 04-308) were consistently basal to the Stromaticum clade. These are species of north temperate regions. Trichoderma rossicum was described from Siberia and has been collected in Austria but was reported from Peru near Lake Titicaca at an elevation of ca. 3,800 m (Hoyos-Caravajal et al. 2009); Clade G strains were obtained from Siberia and Michigan, respectively. Clade G was maintained in all four single gene trees with high support by both methods. However, its sister-relationship with T. rossicum was supported only in two of the single gene trees.
Between the clades of T. stomaticum and T. rossicum three single strains, PPRI 3559, G.J.S. 01-238 and G.J.S. 01-312, did not fall into any clade. These three strains fall between T. stromaticum and T. rossicum clades in a majority of the gene trees and the MLS tree with high support (Fig 1a−e).
The ITS rRNA region could distinguish most of the species under the study except PPRI 3359, which cannot be distinguished from Clade E (Table 4). None of the various subclades of T. stromaticum seen in Fig. 1 a−e were diagnosed by ITS sequences. The differences in genetic distance among the strains of the Stromaticum/Rossicum clade range from 0.0035 (2 bp) –0.014 (7 bp) (Table 4). These differences are considerable given that some species of Trichoderma differ by only one base pair (Komon-Zelazowska et al. 2007; Degenkolb et al. 2008; Samuels et al. 2009).
Species delimitation
Multilocus phylogenetic analysis with four genes indicates that the Stromaticum/Rossicum clade is well supported and that it comprises six strongly supported internal lineages and four independently clustering single strains. We interpret these lineages and single strains as taxonomic species.
Trichoderma stromaticum is clearly distinguished from all other closely related fungi by its apparent host specialization on Moniliophthora species that parasitize cacao, by the stiff, erect hairs that arise from the conidial pustules and by the broad-celled hyphae that comprise the pustule. Moreover, it is conclusively known only from tropical America, although Zachow et al. (2009) reported it from soil in the Canary Islands based on ITS1 sequence and Druzhinina (personal communication) detected it on air filters in Austria based on ITS1 + 2 sequences. All other members of the clade except T. rossicum and T. barbatum, which are northern hemisphere species (or in the case of T. rossicum, high elevation Andean), are paleotropical.
De Souza et al (2006) and Loguercio et al. (2009b) distinguished two groups within T. stromaticum based on AFLP patterns; Group 1 corresponds to Clade B (Fig 1) here, and Group 2 corresponds to Clade A. They attributed biological and ecological differences to the respective groups. We have included in this study several of the strains studied by de Souza et al (Table 1). Members of Group 2 sporulated more profusely on rice and pieces of ‘dry broom’ (dead, proliferated branches of cacao that formed following infection of meristematic tissue by Moniliophthora perniciosa) than members of Group 1. Loguercio et al. (2009b) observed that generally members of Group 1 were able to sporulate better in the tree canopy than members of Group 2, and this increased sporulation was correlated with lower incidence of M. perniciosa. When representatives of the two groups were experimentally inoculated into cacao trees in the field, members of both groups were able to develop an endophytic relationship in sapwood of the trees, but Group 2 members were recovered up to 120 days after inoculation whereas Group 1 members were not reisolated (de Souza et al. 2008). Loguercio et al. (2009a) found that in the field Group 1 members more effectively reduced survival of M. perniciosa in hanging brooms than did members of Group 2. They concluded that, allowing for strain differences, overall members of Group 1 were more effective biological control agents than Group 2 members. We observed the following differences between the phylogenetic groups corresponding to AFLP Groups 1 and 2: conidial pustules of Group 1 are slightly larger (95% ci 0.5−0.6 vs 0.3−0.4 mm) and hairs arising from pustules of Group 1 are slightly longer (105−113 vs 80−85 μm). De Souza et al. (2006) reported longer conidia in Group 2 than in Group 1 but after measuring 700 and 600 conidia from the respective collections, we did not observe any difference in conidial length. To add to these differences, we have observed perithecia only for Clade A of T. stromaticum and a subset of subclade A, subclade C (Fig 1), only includes strains from Peru and Ecuador. Ascospores of subclade C are longer and narrower than in subclade A and members of subclade C were isolated from the pseudostroma of M. roreri, cause of Frosty Pod Rot of cacao (Evans et al. 2003a), in addition to M. perniciosa.
Can the two AFLP groups be recognized taxonomically? Although in de Souza et al. (2006) and Loguercio et al. (2009b), AFLP groups 1 (clade B here) and 2 (clade A here) are biologically distinct, and perithecia have not been found for one of those groups (Group 1, Clade A) multilocus phylogenetic data (Fig. 1e) support this clustering only weakly (Clade A is highly supported, but not clade B), and De Respinis et al. (2010) did not observe any differences between the groups in MALDI-TOF MS analysis of the proteome. Various studies have discussed comparisons and reliability of AFLP data versus phylogenetic (Meudt and Clarke 2007; Althoff et al. 2007). Many phylogeneticists are skeptical about the reliability of AFLPs for phylogenetic inferences because of homology issues (Althoff et al. 2007; Bussell et al. 2005). Because AFLP fragments are anonymous and sorted by size, there is a potential for homoplasy of fragments, producing different results from those of phylogenetic analyses. In contrast, other authors conclude that AFLP data may contain dependable phylogenetic signal and is best for examining within species relationships such as phylogeographic patterns or genotyping, or to study recently diverged species (Meudt and Clarke 2007; Althoff et al. 2007). Trichoderma stromaticum Clades A and B are possibly in the process of speciation. In support of this, we examined whether there was genetic exchange among the various clades of the species. When only the combined four-gene dataset for T. stromaticum was used, partition homogeneity test provided no evidence for heterogeneity of the data at the level of P = 0.1, and the PHI test, which is used for detection of recombination in the data, was also insignificant. These results suggest the absence of recombination and hence incipient isolation. Various authors hypothesize that even if isolation and speciation is effective immediately, the time required for evolutionary changes to appear in two distinct lineages might not be enough to allow for their recognition (Knowles and Carstens 2007; O'Meara 2010). As is the case of several species of Trichoderma (e.g., T. harzianum, T. hamatum, T. viridescens, T. asperelloides, etc.), species radiation seems to be an active and rapid process (Chaverri et al. 2003; Druzhinina et al. 2010), and T. stromaticum is just another example. Using the two criteria of Dettman et al. (2003), the phylogenetic data do not support division of T. stromaticum into two phylogenetically distinct lineages. According to the genealogical concordance criterion, the subclades must exist in the majority of single gene trees if they are to be considered distinct. Here, we observed the division of T. stromaticum into two clades in only one single gene tree (rpb2) and thus the phylogeny of the individual gene trees does not fulfill this criterion. According to the genealogical non-disconcordance criterion, the clades must be supported in at least one individual gene tree and not be contradicted in any other single locus tree at the same level of support. The two clades of the rpb2 tree (A, B) did not exist with support in any other trees. Moreover this result was contradicted in the cal tree, in which there are two additional supported subclades that do not match the clades of the rpb2 tree; therefore, the nondisconcordance criterion was not met either.
While biological and AFLP data suggest the existence of more than one taxon in T. stromaticum, the lack of differences in the sequences of peptides in the proteome (De Respinis et al. 2010) and the multilocus phylogenetic analysis lead us to conclude that T. stromaticum is undergoing speciation, but we cannot recognize taxonomic partition of the species at this time.
All of the members of the Stromaticum/Rossicum clade produce hairs from their conidial pustules and there are essentially two distinct morphologies among these hairs. In T. stromaticum, the hairs are more or less awl-shaped, relatively short and often produce phialides from their tips. Trichoderma stromaticum is further distinguished by the anatomy of its pustules, which are formed of broad, short hyphae. Most of the remaining members of the clade produce more or less wooly pustules and the hairs are relatively long; they tend to be variously coiled or sinuous and most often are sterile, the exception being strains of Clade F (Fig 1). Each of the clades represented in Fig 1e, including the single strains G.J.S. 01-225, G.J.S. 01-312, G.J.S. 01-238 and PPRI 3359 represents a distinct species. These species are distinguished as follows.
The two strains comprising each of the Clades E and F were derived from Hypocrea specimens collected on decorticated wood in close proximity to each other in a single more or less undisturbed Cameroonian forest. The Hypocrea teleomorphs of these collections were indistinguishable but the clades differ in characters of the anamorphs and growth rates. Conidial pustules of the two members of Clade F (Fig 9a−c; see Trichoderma lanuginosum, below) are conspicuously wooly whereas pustules in Clade E produce more discrete sterile hairs (Fig 11b, c; see Trichoderma medusae, below). Conidia of Clade E are somewhat longer and narrower (95% ci 4.5−4.7 × 2.7−2.8 μm, L/W = 1.7−1.8) than conidia of Clade F (95% ci 3.8−4.0 × 2.5−2.7, L/W = 1.5−1.6). A verticillium-like synanamorph formed in SNA cultures of Clade F (Fig 9g−j; see Trichoderma lanuginosum, below) while Clade E lacks a synanamorph. Ascospores of Clade E are larger than those of Clade F. Finally, cultures of Clade E grow much faster on PDA and SNA at 30°C than cultures of Clade F. We recognize Clade E below as the new species T. medusae and Clade F as T. lanuginosum.
Members of Clade G cannot be separated from T. rossicum on the basis of their microscopic morphology, and actually strain DAOM 230008 was identified as T. rossicum in Bissett et al. (2003). However, the two available cultures of Clade G grow considerably faster than any of the six cultures of T. rossicum that we have studied, the difference especially noticeable on PDA and SNA at 30°C. Despite the similarity in their phenotypes, T. rossicum and T. barbatum are phylogenetically distinct and Clade G is described as a new species, T. barbatum. These two species are sympatric in central Europe and are known from temperate regions.
Four single strain lineages (PPRI 3359, G.J.S. 01-225, G.J.S. 01-238, G.J.S. 01-312) cluster independently in the Stromaticum/Rossicum Clade and are recognized here as distinct taxonomic species. The closest relationship of G.J.S. 01-225 is with T. medusae (Fig. 1e) but it differs from that species in its much faster growth rate on PDA and SNA; it is described as T. caesareum. PPRI 3559 is closely related to T. lanuginosum but differs from that species in its larger conidia and faster growth rate; PPRI 3559 is described as T. vermipilum. Two single strain lineages, G.J.S. 01-312 and G.J.S. 01-238 do not have any close relationships within the Stromaticum/Rossicum Clade but they are morphologically consistent with the other members of the clade in anamorph and teleomorph (G.J.S. 01-238) characters. Strain G.J.S. 01-312, isolated from soil in Côte d’Ivoire, is distinguished by its small conidia, the smallest conidia in the group, and is described as T. ivoriense. Despite the relative phylogenetic isolation of strain G.J.S. 01-238, an ascospore isolate from the same Thai forest as T. caesareum, no single character of its anamorph or teleomorph distinguishes it from other species in the Stromaticum/Rossicum Clade. However, its rate of growth on SNA at 30°C is most similar to Andean collections of T. stromaticum, T. barbatum, and T. ivoriense. It can be distinguished from T. stromaticum on the basis of its morphology; geography distinguishes it from the north temperate T. barbatum, and it has larger conidia than T. ivoriense. The strain G.J.S. 01-238 is named below as T. floccosum.
Biogeography
Trichoderma stromaticum is known conclusively only from Latin America. The greatest diversity of the species is found in the eastern Brazilian states of Bahia and Pará. The unresolved Clade A (Fig. 1), which is equivalent to AFLP Group 1 of de Souza et al. (2006), includes strains almost exclusively from the Bahia, exceptions being one strain from the state of Pará (G.J.S. 00-102) and one strain from Colombia (AM13 of de Souza et al. 2006 = G.J.S. 00-02). Clade B, Group 2 of de Souza et al. (2006), includes a mixture of strains from Bahia and Pará. Clade C, which is resolved in cal, tef1 and MLS (Fig 1c,d, and e, respectively) is known only from Amazonian Peru and Ecuador. This clade includes the only known strains that occur on both cacao pathogens Moniliophthora perniciosa and M. roreri. All others are found on M. perniciosa; T. stromaticum is isolated rarely as an endophyte from sapwood of cacao. The closest relatives of T. stromaticum are strains collected in, respectively, West (Côte d’Ivoire), Central (Cameroon), and South (Republic of South Africa) Africa, and Thailand. Somewhat more distantly related are T. rossicum and T. barbatum, both found in north temperate regions (Siberia, Austria, Andean Peru, USA: Michigan). Trichoderma lanuginosum and T. medusae were collected as Hypocrea specimens in close proximity in a more or less undisturbed Cameroonian rainforest (Reserve Faunal du Dja), and T. caesareum and T. floccosum were collected as Hypocrea specimens in an undisturbed forest in southern Thailand (Khao Yai National Park). Trichoderma rossicum and T. barbatum are sympatric in soil in Austria and are only known from soil collected in North temperate regions of Asia (Siberia), Europe (Austria) and the United States (Michigan, T. barbatum). Trichoderma rossicum has previously been known only from Siberia (Bissett et al. 2003) and Andean Peru (Hoyos-Carvajal et al. 2009); strain DAOM 230008 was originally identified as T. rossicum (Bissett et al. 2003).
Descriptions of the species
Trichoderma barbatum Samuels, sp. nov.Figs. 2a and 3
Colonies of Trichoderma species grown on PDA (above) and SNA (below) under light for 1 week at 25°C unless noted otherwise. a T. barbatum (G.J.S. 04-208, 10 days). b T. floccosum (G.J.S. 01-238). c T. ivoriense (G.J.S. 01-312). d T. lanuginosum (G.J.S. 01-176). e T. medusae (G.J.S. 01-171). f T. rossicum (DAOM 230011, 96 h). g T. vermipilum (PPRI 3559, 10 days). h−j T. stromaticum showing variation in groups. h Clade A (Fig 1e, G.J.S. 07-76). i Clade B (Fig 1e, G.J.S. 07-77). j Clade C (Fig 1e, G.J.S. 05-455)
Trichodermati rossici simile sed in agaris dictis PDA vel SNA temperatura 30°C magis celeriter crescens. Pustulae conidiales lanosae; conidia anguste ellipsoidea, (4.0−)4.2−5.0(−5.7) × (2.2−)2.5−3.5(−6.5) μm, LW = (1.4−)1.5−1.9(−2.0).
Holotype BPI 881029, designated here
Mycobank 519539
Telemorph
None known
Characteristics in culture
Optimum temperature for growth on PDA and SNA at 25−30°C. Colony radius after 96 h on PDA at 25−30°C 60−68 mm, on SNA 44−55 mm (n = 2 cultures). Not growing at 35°C. After 10 days at 25°C under light on PDA forming a continuous lawn of conidia, K&W 28D−F8 (Deep Green, Dark Green), no distinctive odor or diffusing pigment; on SNA pustules forming in abundance around the periphery of the colony. Pustules pulvinate to hemispherical, 0.5−1 mm diam, easily removed from the agar, gray green, with abundant protruding white hairs. Hairs extending beyond the surface of the pustule, sinuous tending to spiraled, septate, smooth, infrequently branched, base ca. 5 μm diam, tip subacute to acute, sterile. Fertile branches arising from near the base of the hairs, paired or solitary, typically a few cells long, but longer with distance from the tip, ca. 5 μm wide, producing phialides directly or producing unicellular 2º branches bearing phialides. Phialides flask-shaped, (3.7−)5.0−7.5(−10.0) μm long, (2.7−)3.2−4.2(−5.0) μm at the widest point, (1.7−)2.2−3.5(−4.2) μm at the base, L/W = (1.1−)1.3−2.1(−3.3), arising from a cell (2.7−)3.5−4.5(−5.0) μm diam, arising from 1º and 2º branches, solitary, paired or in dense botryose heads of several. Conidia (n = 152) oblong to narrowly ellipsoidal, (4.0−)4.2−5.5(−9.5) × (2.2−)2.5−3.5(−6.5) μm, L/W = (1.4−)1.5−1.9(−2.0) (95% ci = 4.8−5.0 × 2.9−3.1 μm, L/W = 1.6−1.7), smooth. Chlamydospores not observed on CMD or SNA within 10 days.
Etymology
‘Barbatum’ from Latin ‘barbatus’ meaning bearded, with reference to the long hairs arising from the pustules.
Habitat
Soil, roots of strawberry (Fragaria).
Known distribution
USA (Michigan), Russia (Siberia).
Holotype
USA, Michigan, Michigan State University, Horticulture Farm, isolated from roots of strawberry, date unknown, R. Olatinwo and A. Schilder T-10 (BPI 881029 G.J.S. 04-308; holotype designated herewith). Live ex-type culture CBS 125733.
Additional material examined
Russia, Siberia, Krasnoyarsk region, isolated from soil under apple, Oct 1997, G. Szakacs, TUB F698 (Culture DAOM 230008).
Comments
The culture DAOM 230008 (98-90) was considered to be T. rossicum by Bissett et al. (2003).
The ex-type strain (T-10) of T. barbatum was originally identified as T. stromaticum by Olatinwo et al. (2004) because of a similarity between the two species in their ITS sequences. These authors isolated the strain from healthy strawberry roots. It was found to reduce severity of root lesions caused by Rhizoctonia fragariae as compared to untreated control. When applied as a root dip to strawberry transplants at two sites in Michigan, the total fruit weight and number of berries the following season was significantly greater in treated plants than untreated plants or plants treated with PlantShield (T. harzianum) at one of the sites. Olatinwo et al. (2004) suggested that pre-planting treatment with T-10 might be beneficial to strawberries.
Trichoderma caesareum Samuels, sp. nov. Figs. 4 and 5
Trichoderma caesareum. a, b Pustules from SNA. The wooly nature of the pustule is evident in (b). c, d Hairs arising from the pustule. e Fertile branches arising from the base of a sterile hair. f Fertile branch with phialides. g Conidia. All from G.J.S. 01-225. Scale bars (a) 1 mm, (b) 0.5 mm, (c) 100 μm, (d, e) 20 μm, (f, g) 10 μm
Trichoderma caesareum, Hypocrea teleomorph. a, b Stromata. c Surface of the stroma. d Section through a stroma showing four perithecia in median longitudinal section. e Stroma surface showing perithecial ostiolar canal. f Stroma surface in section. g Cells of the interior of a stroma below perithecia. h Asci. i Ascospores. All from G.J.S. 01-225. Scale bars (a) 1 mm, (b) 0.5 mm, (c, e−h) 20 μm, (d) 200 μm, (i) 10 μm
Pustulae conidiales lanosae; conidia (4.0−)4.5−5.2(−6.0) × (2.0−)2.5−3.0, L/W = 1.5−2.1(−2.4). Trichodermati medusae Samuels simile sed in agaris dictis PDA vel SNA magis celeriter crescens.
Holotype BPI 863896, designated here
Mycobank 519540
Telemorph
Hypocrea sp.
Characteristics in culture
Optimum temperature for growth on PDA and SNA 25°C, colony radius after 96 h on PDA 65−70 mm, on SNA 35−50 mm. Typically not growing at 35°C. On SNA after 1 week at 25°C pustules forming in abundance throughout the colony. Pustules conspicuous, hemispherical, 1−2 mm diam, grayish green (K&W 28D5), with abundant protruding white hairs largely obscuring the mass of conidia. Hairs extending beyond the surface of the pustule, sinuous to spiraled or straight, septate, smooth or warted, infrequently branched, base ca. 5 μm wide, tip subacute. Fertile branches arising from near the base of the hairs, typically 1 or few cells in length, longer with distance from the tip and producing unicellular 2º branches, ca. 5 μm wide. Phialides doliform or flask-shaped, (3.7−)4.7−6.2(−7.2) μm long, (3.0−)3.5−4.0(−4.2) μm at the widest point, L/W = (1.2−)1.3−1.8(−2.1), (1.5−)2.0−3.2) μm at the base, arising directly from a cell (3.0−)3.2−4.5(−5.0) μm wide and terminating 1º and 2º fertile branches; all branches terminating in 1 to several densely clustered phialides. Conidia (n = 30) oblong, (4.0−)4.5−5.2(−6.0) × (2.0−)2.5−3.0 μm, L/W = 1.5−2.1(−2.4) (95% ci = 4.5−5.0 × 2.6−2.8 μm , L/W = 1.7−1.9), smooth, gray green. Chlamydospores not observed.
Characteristics of the teleomorph
Stromata scattered, gregarious, discoidal, ca. 1 mm diam, broadly attached, hyphae not visible, surface plane to convex, perithecial elevations appearing as low tubercles, perithecial openings appearing as darker dots against the surrounding tissue, yellowish brown to brown, not reacting to 3% KOH. Cells of the stroma surface in face view pseudoparenchymatous, elliptical in outline, 8−12(−15) μm diam, thin-walled. Perithecia circular in section, (n = 10) (225−)230−250(−265) μm high, 150−185(−200) μm wide, ostiolar canal 75−90 μm long. Perithecial papilla formed of small cells, clavate elements lacking. Surface region distinguished from the internal tissue of the stromata by pigmentation in the outermost 2−3 layers of cells; cells of the stroma surface in section pseudoparenchymatous, (5−)7−10(−12) μm diam, thin-walled. Tissue below the stroma surface, between perithecia pseudoparenchymatous, thin-walled, lacking hyphal elements. Tissue of the stroma below perithecia, perithecia textura epidermoidea, thin-walled, lacking long hyphal elements, (4−)7−14(−18) × (4−)5−8(−10) μm. Asci cylindrical, (77−)80−92(−103) × (4.0−)4.7−6.2(−7.5) μm (n = 30), apex with a conspicuous discharge ring, ascospores uniseriate. Part ascospores hyaline, conspicuously warted, dimorphic; distal part subglobose or conical, 4.0−5.0(−5.2) × (3.5−)3.7−4.2(−4.5) μm; proximal part oblong to wedge-shaped or ellipsoidal, (3.7−)4.2(−5.0(−5.5) × (2.7−)3.0−3.5(−4.0) μm.
Etymology
‘caesareum’ from Latin, in reference to the long hairs arising from the conidial pustule.
Habitat
Trichoderma caesareum is known only from cultures derived from ascospores of one Hypocrea specimen collected in primary forest; the Hypocrea develops on bark.
Known distribution
Thailand, known only from the original collection.
Holotype
Thailand, Prachinburi Province, Khao Yai National Park, along trail betweern Khlong E-Thao (14º28′N, 101º20′E, elev. 750 m) and 14º28′N, 101º12′E, elev. 800 m, on bark, 18 Aug 201, G.J.S. 9074, R. Nasit (BPI 863896, a dry culture ex-ascospore isolation). Live ex-type culture G.J.S. 01-225 = CBS 124369).
Trichoderma floccosum Samuels, sp. nov. Figs. 2b, 6 and 7
Trichoderma floccosum. a, b Pustules on SNA. The wooly nature of the pustule is evident in (b). c−f Sterile hairs arising from the pustules. Fertile branches arising from the base of the hairs can be seen in (c) and (f). f Sterile hair. g Fertile branch with phialides. h conidia. All from G.J.S. 01-238. Scale bars (a) 1 mm, (b) 0.5 mm, (c−f) 20 μm, (g, h) 10 μm
Trichoderma floccosum, Hypocrea teleomorph. a, b Stromata. c Surface of the stroma showing ostiolar openings. d Section through a stroma showing several perithecia in median, longitudinal section. e Cells of the stroma surface in section. f Stroma surface region showing the ostiolar opening of a perithecium. g Cells of the interior of a stroma below perithecia. i Asci. j Ascospores. All from G.J.S. 01-238. Scale bars (a, b) 1 mm, (c, e−g) 20 μm, (d) 200 μm, (i, j) 10 μm
Trichodermati medusae Samuels simile sed in agaris dictis PDA vel SNA temperatura 30°C magis celeriter crescens. Pustulae conidiales lanosae. Conidia ellipsoidea (3.5−)4.0−5.0(−5.5) × (2.5−)2.7−3.5(−3.7) μm, L/W = (1.3−)1.5−1.7(−1.9).
Holotype BPI 871616, designated here
Mycobank 519541
Telemorph
Hypocrea sp.
Characteristics in culture
Optimum temperature for growth on PDA 25°C, on SNA 25−30°C. Colony radius after 96 h on PDA at 25°C ca. 60 mm, on SNA at 25/30°C 40−45(−60) mm. Typically not growing at 35ºC. After 10 days at 25°C under light on PDA pustules forming in abundance around the margin of the colony; on SNA few large pustules forming at the margin of the colony. On SNA pustules conspicuous, pulvinate to nearly hemispherical, 1−2 mm diam, gray green with abundant protruding white hairs largely obscuring the mass of conidia. Hairs extending beyond the surface of the pustule, sinuous; septate; smooth, infrequently branched, base ca. 5 μm diam, tip subacute, sterile. Fertile branches arising from near the base of the hairs, typically 1 or few cells in length, longer with distance from the tip, ca. 5 μm wide, producing 2º branches. Phialides flask-shaped, (3.7−)4.7−6.2(−7.2) μm long, (3.2−)3.5−4.5(−4.7) μm at the widest point, L/W = (1.3−)1.5−1.7(−1.9), base (1.2−)2.2−3.5(−4.0), arising from a cell (3.2−)3.7−4.7(−5.0) μm wide; forming directly from and terminating 1º and 2º fertile branches; all branches terminating in 1 to several densely clustered phialides. Conidia (n = 30) ellipsoidal, (3.5−)4.0−5.0(−5.5) × (2.5−)2.7−3.5(−3.7) μm, L/W = (1.3−)1.5−1.7(−1.9) (95% ci = 4.4−4.7 × 2.9−3.1 μm, L/W 1.5−1.6), smooth, gray green. Chlamydospores abundant on CMD and SNA reverse, terminal and intercalary, subglobose, (6−)7−17(−30) μm diam.
Characteristics of the teleomorph
Stromata scattered, gregarious, discoidal, 1−1.5 mm diam, broadly attached, hyphae not visible, surface plane to convex, perithecial elevations appearing as low tubercles, perithecial openings appearing as darker dots against the surrounding tissue, yellowish brown to brown, not reacting to 3% KOH. Cells of the stroma surface in face view pseudoparenchymatous, elliptical in outline, (10−)13−21(−25) × (8−)11−17(−19) μm, thin walled. Perithecia circular to elliptic in section, (n = 12) (228−)240−285(−300) μm high, (143−)165−205(−212) μm high, ostiolar canal 70−100(−124) μm long. Perithecial papilla formed of small cells, clavate elements lacking. Surface region distinguished from the internal tissue of the stromata by pigmentation in the outermost 2−3 layers of cells; cells of the stroma surface in section pseudoparenchymatous, (6−)8−14(−16) × (5−)7−11(−12) μm, thin-walled. Tissue of the stroma below perithecia, perithecia textura epidermoidea, lacking long hyphal elements, thin-walled, (4−)9−17(−21) × (3−)5−9(−12) μm. Asci cylindrical, (n = 30), (89−)94−112(−124) × (4.5−)5.5−6.7(−7.5) μm, ascospores uniseriate, apex with a conspicuous discharge ring. Part ascospores spinose, hyaline, dimorphic; distal part subglobose or conical, (4.0−)4.5−5.2(−6.0) × (3.5−)3.7−4.5(−4.7) μm; proximal part oblong to wedge-shaped or ellipsoidal, (3.5−)4.0−5.2( −6.2) × (2.7−)3.0−3.7(−4.0) μm.
Etymology
‘floccosum’ from Latin, a lock or flock of wool on something such as clothing or fruit; in this case, on the pustule.
Habitat
Trichoderma floccosum is known only from cultures derived from ascospores of an unnamed Hypocrea species collected in primary forest; the Hypocrea develops on decorticated wood.
Known distribution
Thailand, known only from the original collection.
Holotype
Thailand, Nakhorn Nayok Province, Khao Yai National Park, Mo Sing To trail from shops at park headquarters, on decorticated wood, 4 Sep 2001, G.J.S. 9202, R. Nasit (BPI 871616, a dry culture ex-ascospore isolation). Live ex-type culture G.J.S. 01-238 = CBS 124372.
Trichoderma ivoriense Samuels, sp. nov. Figs. 2c and 8
Trichoderma ivoriense. a, b Pustules on CMD (a) and SNA (b). c−e Sterile hairs. Fertile branches arising from the base of the hairs seen in (d) and (e). Fertile branch arising from the base of a hair and phialides. g−i Synanamorph (CMD). j Conidia. k Chlamydospores. All from G.J.S. 01-312. Scale bars (a) 1 mm, (b, g) 0.5 mm, (c−e, h, I, k) 20 μm, (f , j) 10 μm
Pustulae conidiogenae lanosae, griseo-virides, pilei ex pustulis conidiogenis exorientes, steriles, sinuosi; conidia ellipsoidea, (3.0−)3.2−3.7(−4.0) × (2.0−)2.2−2.7 μm, L/W = (1.1−)1.2−1.6(−1.7).
Holotype BPI 881030, designated here
Mycobank 519542
Characteristics in culture
Optimum temperature for growth on PDA and SNA 25°C, on PDA colony radius ca. 60 mm after 96 h at 25°C, on SNA colony radius ca. 50 mm. Not growing at 35°C. On PDA and CMD after 10 days at 25°C under light colonies producing a continuous layer of white mycelium, no pigment diffusing through the agar, no distinctive odor; conidia forming throughout the colony on macronematous, mononematous conidiophores, conidia yellowish green (K&W 29D7). On SNA conidia forming in pustules dispersed in a 2−3 cm broad band around the margin of a 9-cm-diam Petri plate; mononematous conidiophores not observed. Pustules 0.5−1 mm diam, yellowish green (K&W 28D−E7), hemispherical, remaining discrete; long, sterile, sinuous to loosely coiled, white hairs arising in abundance from each pustule. Conidiophores in pustules comprising a long, sterile hair with short fertile branches arising from its base. Sterile hairs tapering from 5−7 μm at the base to an acute tip, and producing shorter sterile branches, often roughened, septate, tending to coil near the tip. Fertile branches arising at right angles from the base of the hairs, typically comprising a single basal cell with phialides clustered at its tip or producing 2º cells, each with terminal phialides; branches more distant from the tip 2 or 3 cells long. Phialides doliform, (3.7−)4.7−6.0(−7.5) μm long, (3.0−)3.2−4.0(−4.5) μm at the widest point, (1.7−)2.0−3.0(−5.0) μm at the base, L/W (1.1−)1.3−1.7(−2.1) (n = 30); arising from a cell (3.2−)3.5−4.5(−4.7) μm wide. Conidia (n = 30) ellipsoidal, (3.0−)3.2−3.5(−4.0) × (2.0−)2.2−2.5(−2.7) μm, L/W (1.1−)1.2−1.6(−1.7) (95% ci 3.3−3.5 × 2.5−2.7 μm, L/W 1.5−1.6). On CMD a gliocladium/verticillium-like synanamorph forming from erect hyphae, 40−170 μm long, 4−6 μm wide at base, phialides 5−15 μm long; conidia ellipsoidal, 2.5−4.0 × 2.0−3.2 μm, held in glistening green heads. Chlamydospores abundant on SNA reverse, terminal, obovoidal to fusiform, (4.2−)5.2−8.0(−10.0) × (4.0−)4.7−5.2(−9.2) μm.
Etymology
Named for the country of origin of the type, Côte d’Ivoire.
Habitat
Soil.
Known distribution
Côte d’Ivoire, known only from the type.
Holotype
Côte d’Ivoire, isolated from soil, Oct 2001, I. Kibbe 164 (BPI 881030). Live ex-type Culture: G.J.S. 01-312 = CBS 125734.
Comments
On the basis of morphology it would be very difficult to distinguish T. ivoriense from the only distantly related T. spirale Bissett. The only known strain of T. ivoriense is slower growing than T. spirale and produces far fewer chlamydospores. Cultures of T spirale may produce only mononematous conidiophores similar to those found in PDA and CMD cultures of T. ivoriense.
Trichoderma lanuginosum Samuels, sp. nov. Figs. 2d, 9 and 10
Trichoderma lanuginosum. a−c Pustules formed on SNA. d Sterile hairs arising from a pustule. e, f Hairs with fertile branches arising from the base. g−j Synanamorph on SNA. k Conidia. a, b, d−f, h, j, k from G.J.S. 01-174; c, g, i, k from G.J.S. 01-176. Scale bars (a−c) 0.5 mm, (d−f, h, i) 20 μm, (g) 150 μm, (k) 10 μm
Trichoderma lanuginosum, Hypocrea teleomorph. a−c Stromata. d Cells at the stroma surface; a perithecial opening visible in the upper left corner. e Section through a stroma showing a perithecium in median longitudinal section. f Surface of a stroma showing a perithecial opening. g Cell of the stroma interior below a perithecium. h−j Asci. A ring can be seen in the ascal apex in (i) and (j). k Discharged part ascospores. Images a, c−f, i−k from G.J.S. 01-174; g, h, k from G.J.S. 01-176. Scale bars (a) 1 mm; (b, c) 0.5 mm, (d, f−h) 20 μm, (e) 200 μm, (i−k) 10 μm
Trichodermati medusae Samuels simile sed conidia (3.0−)3.5−4.2(−4.7) × (2.0−)2.2−3.0(−3.5) μm, L/W = (1.2−)1.3−1.7(−2.0) et in agaro dicto SNA synanamorphosis verticillio simile formans.
Holotype BPI 863853, designated here
Mycobank 519543
Telemorph
Hypocrea sp.
Characteristics in culture
Optimum temperature for growth on PDA 20−25°C, colony radius after 96 h 28−42 mm; on SNA 25°C, colony radius after 96 h 30−35 mm. Typically not growing at 35°C. On PDA after 1 week at 25°C under light a continuous lawn of conidia forming in the aerial mycelium over the entire colony, margin sterile; pustules forming after two weeks; conidia grayish yellow green (K&W 28d6−7). Colony on SNA after 10 days under the same conditions producing mononematous conidiophores abundantly throughout the colony in the aerial mycelium; pustules just beginning to form. Mononematous conidiophores, (n = 38) 41−120(−187) μm long, (2.5−)3−5(−6.7) μm wide at base, verticillium-like, with 1−3 verticils of phialides and 2 branches. Phialides awl-shaped, (n = 60) (8−)12−21(−34) μm long, (2.0−)2.2−3.7(−4.5) μm wide at the base, arising singly from the condiophore or in verticils of 2−4. Conidia arising from mononematous conidiophores ellipsoidal, (n = 60) (3.7−)4.0−6.5(−8.2) × (2.5−)2.7−4.2(−5.2) μm, with a protuberant, flat basal abscission scar. Pustules hemispherical, 0.25−0.5 mm diam, with abundant protruding white hairs, hairs largely obscuring the mass of conidia. Hairs extending beyond the surface of the pustule, sinuous to straight, only a slight tendency to coiled or spiraled, sterile, septate, smooth, infrequently branched, base 3.5−4.5 μm wide, tip blunt. Fertile branches arising from near the base of the hairs, typically 1 or few cells in length, longer with distance from the tip and producing unicellular 2º branches, 3.5−4.5 μm wide. Phialides doliform to lageniform, (4.2−)5.2−7.2(−9.7) μm long, (3.0−)3.5−4.2(−4.5) μm at the widest point, L/W (1.0−)1,4−2.0(−2.7), arising from a cell (3.2−)3.5−4.5−5.0) μm wide; arising directly from, and terminating 1º and 2º fertile branches; all branches terminating in 1 to several densely clustered phialides. Conidia (n = 60) ellipsoidal, (3.0−)3.5−4.2(−4.7) × (2.0−)2.2−3.0(−3.5) μm, L/W (1.2−)1.3−1.7(−2.0) (95% ci 3.8−4.0 × 2.5−2.7 μm, L/W 1.5−1.6), smooth, yellow green. Chlamydospores not observed.
Characteristics of the teleomorph
Stromata scattered, gregarious, discoidal, ca. 1 mm diam, broadly attached, hyphae not visible, surface plane to convex, perithecial elevations appearing as low tubercles, perithecial openings appearing as darker dots against the surrounding tissue or concolorous, yellowish brown to brown, not reacting to 3% KOH. Cells of the stroma surface in face view pseudoparenchymatous, elliptical in outline, (5−)8−15(−18) μm diam, thin-walled. Perithecia circular to elliptical in section, (n = 15) (188−)200−225(−240) μm high, (100−)115−160(−180) μm wide, ostiolar canal 50−75 μm long. Perithecial papilla formed of small cells, clavate elements lacking. Surface region distinguished from the internal tissue of the stromata by pigmentation in the outermost 2−3 layers of cells; cells of the stroma surface in section pseudoparenchymatous, (4−)5−9(−11) × 4−8(−11) μm diam, thin-walled. Tissue of the stroma below perithecia, perithecia textura epidermoidea to t. angularis, lacking long hyphal elements, cells thin-walled, (6−)10−20(−30) × (6−)8−15(−20) μm. Asci cylindrical, (65−)70−85(−97) × (2.7−)3.7−5.2(−6.5) μm, ascospores uniseriate, apex with a conspicuous discharge ring. Part ascospores dimorphic, spinose, hyaline; distal part subglobose or conical, (n = 60) (3.2−)3.7−4.2(−4.7) × (2.7−)3.2−3.7(−4.0) μm; proximal part oblong to wedge-shaped or ellipsoidal, (3.0−)3.7−4.7(−5.0) × (2.2−)2.5−3.2(−3.5) μm.
Etymology
‘lanuginosum’ from Latin, in reference to the hairs that appear as wooly down on the surface of the pustule.
Habitat
Trichoderma lanuginosum is known only from cultures derived from ascospores of Hypocrea specimens; the Hypocrea teleomorph on rotting wood with bark.
Known distribution
Cameroon, known only from the type locality.
Holotype
Cameroon, Provinces du Sud et de l’Est, Dept. Haut-Nyong, Reserve Faunal du Dja, vic Somalomo, ca. 30 km E of Somalomo, Mintoum, 1.5 h walk S in forest, in primary forest, 03º18’N, 12º58’E, elev. 620 m, on bark, 13 Jul 2001, G.J.S. 9021, D. Begoude (BPI 863853, culture G.J.S. 01-176 = CBS 125718).
Additional specimen examined
Data as the holotype except collected on bark of rotten wood, G.J.S. 9019, live culture 01-174 = CBS 126100.
Trichoderma medusae Samuels sp. nov. Figs. 2e, 11 and 12
Trichoderma medusae. a−c Pustules from CMD. e Sterile hairs arising from a pustule. d−f Hairs with fertile branches arising from the base. g A fertile branch arising from the base of a hair. h Conidia. Images a, c, e, h from G.J.S. 01-166; b, c, f, g from G.J.S. 01-171; b provided by D Farr. Scale bars (a, b) 1 mm, (c−e) 20 μm, (f−h) 10 μm
Trichoderma medusae, Hypocrea teleomorph. a−c Stromata. d Cells of the stroma surface showing a perithecial opening. e Section through a stroma showing median longitudinal sections through two perithecia. f Stroma surface region showing a perithecial opening. g Cells of the interior of a stroma below a perithecium. h, i Asci. An apical ring can be seen in the apex of asci in i, j. Mature part ascospores. Images a, b, h−j from G.J.S. 01-166; c−g from G.J.S. 01-171. Scale bars (a, b) 1 mm, c 0.5 mm, (d, f, h) 20 μm, (e) 100 μm; (g, i, j) 10 μm
Trichodermati lanuginosi Samuels simile sed synanamorphosis verticillio simile abest et conidia (3.7−)4.2−5.0(−5.7) × (2.2−)2.5−3.0(−3.5) μm , L/W 1.5−1.9(−2.5).
Holotype BPI 863841, designated here.
Mycobank 519544
Telemorph
Hypocrea sp.
Characteristics in culture
Optimum temperature for growth on PDA 20−30º, on SNA 25−30°C; on PDA and SNA colony radius 25−35 mm after 96 h at 25°C. Typically not growing at 35°C. On PDA after 1 week at 25°C under 12 h cool white fluorescent light/12 h darkness colony tan, margin scalloped or lobbed, lacking diffusing pigment or distinctive odor; conidia forming in one or two nearly continuous rings around the original inoculum, grayish green (K&W 26c−d4); on SNA conidial pustules abundantly forming in broad concentric rings around the original inoculum. Pustules hemispherical, 0.5−1 mm diam, with protruding hairs. Hairs white, protruding beyond the surface of the pustule, sinuous to somewhat spiraled, especially toward the tip, smooth, septate, tapering slightly from 3.5−4.5 μm at the base, tip blunt, unbranched or infrequently branched, sterile (rarely producing a single phialide). Fertile branches arising near the base of the hairs, 1 to a few cells in length, longer with distance from the tip; phialides arising directly from the main branch or unicellular 2º branches arising from the 1º fertile branches. Phialides subglobose to lageniform, (3.7−)4.7−6.7(−8.2) μm, (2.5−)3.5−4.5(−5.0)μm at the widest point, L/W (0.8−)1.1−1.7(−2.4), (1.7−)2.2−3.5(−4.5) μm wide at base, arising from a cell (2.2−)3.2−4.5(−6.0) μm wide; from arising directly from, and terminating 1º and 2º fertile branches; all branches terminating in 1 to several densely clustered phialides. Conidia (n = 60) ellipsoidal to nearly oblong, (3.7−)4.2−5.0(−5.7) × (2.2−)2.5−3.0(−3.5) μm, L/W 1.5−1.9(−2.5) (95% ci 4.5−4.7 × 2.8−3.0 μm, L/W 1.−1.8), smooth, gray green. Chlamydospores not observed.
Characteristics of the teleomorph
Stromata scattered to gregarious, discoidal, ca. 1 mm diam, broadly attached, hyphae not visible, surface plane to slightly convex, perithecial elevations not evident or appearing as low tubercules, perithecial openings appearing as darker dots against the surrounding tissue, yellowish brown to brown, not reacting to 3% KOH. Cells of the stroma surface in face view pseudoparenchymatous, (6−)8−12(−16) μm diam, thin-walled. Perithecia circular to elliptical in section, (n = 6) (185−)210−275 μm high, (137−)140−180(−185) μm wide, ostiolar canal 75−90 μm long. Perithecial papilla formed of small cells, clavate elements lacking. Surface region distinguished from the internal tissue of the stromata by pigmentation in the outermost 2−3 layers of cells; cells of the stroma surface in section pseudoparenchymatous, (4−)5−9(−11) μm diam, thin-walled. Tissue of the stroma below the surface pseudoparenchymatous, lacking hyphal elements. Tissue of the stroma below perithecia textura epidermoidea, lacking long hyphal elements, thin-walled, (6−)10−18(−24) × (6−)8−15(−21) μm. Asci cylindrical, (69−)80−104(−118) × (4.0−)5.7−6.5(−8.0) μm (n = 60), ascospores uniseriate, apex with a conspicuous discharge ring. Part ascospores dimorphic, spinose, hyaline; distal part subglobose or conical, (n = 60) (3.7−)4.2−5.2(−6.2) × (3.5−)4.0−4.5(−5.0); proximal part oblong to wedge-shaped or ellipsoidal, (4.0−)5.0−6.0(−8.7) × (2.5−)3.2−4.0(−4.7) μm.
Etymology
‘medusae’ from Latin, in reference to the conidial pustules with ‘snake like’ hairs suggestive of Medusa, a gorgon of mythology that has snake-like hairs and turns beholders to stone.
Habitat
Trichoderma medusae known only from cultures derived from ascospores of Hypocrea specimens collected in primary forest; the Hypocrea teleomorph develops on wood with bark.
Known distribution
Cameroon, known only from the type locality.
Holotype
Cameroon, Provinces du Sud et de l’Est, Dept. Haut-Nyong, Reserve Faunal du Dja (a World Heritage site), 6 km E of Dja River, in forest a 2-h quick walk south from route to Bourneville, 03º17′N, 12º47′E, elev. 600 m, on small branches, 14 Jul 2001, G.J.S. 9006, D. Begoude, A Guinwith, Pascal Togo (BPI 863841, a dry culture ex-ascospore isolation). Live ex-type culture G.J.S. 01-171 = CBS 125719.
Additional collection
Same collecting data as the holotype, except on termite infested wood, G.J.S. 9001 (BPI 863836; live culture G.J.S. 01-166).
Trichoderma rossicum Bissett, Kubicek & Szakacs, Can J Bot 81:578. 2003. Figs. 2f and 13
Teleomorph not known.
Characteristics in culture
Optimum temperature for growth on PDA and SNA 25°C. On PDA colony radius after 96 h at 25ºC 56−65 mm, on SNA 36−40 mm; considerably slower at 30°C (in 96 h on PDA 12−25 mm, on SNA 12−18 mm, n = 6 strains studied); not growing at 35°C. After 10 days at 25°C under light on PDA a continuous lawn of conidiophores covers the Petri plate, conidia K&W 30 D−E 8 (Deep Green, Parrot Green), no distinctive odor or diffusing pigment noted; on SNA pustules forming in concentric rings, no synanamorph noted. Pustules 0.25−0.5 mm diam, hemispherical, discrete to confluent, easily removed from the agar, with hairs arising abundantly from each. Hairs sinuous, thin-walled, septate, smooth, infrequently branched, base 5−7 μm diam, tip subacute. Fertile branches arising from near the base of the hairs, paired or solitary, typically 1 or few cells in length, longer with distance from the tip, 4−5 μm wide, producing phialides directly, sometimes producing 2º branches typically comprising a single cell. Phialides flask-shaped to subglobose with a short contricted neck, (4.5−)5.0−7.0(−7.7) μm long, (2.7−)3.5−4.5(−4.7) μm at the widest point, (1.7−)2.2−3.2(−4.2) μm at the base, L/W = (1.2−)1.3−1.9(−2.2), arising from a cell (3.5−)4.0−5.2(−6.0) μm diam; arising directly from and terminating 1º and 2º fertile branches; all branches terminating in 1 to several densely clustered phialides. Conidia (n = 120) oblong, (4.0−)4.2−5.0(−5.7) × (2.2−)2.7−3.0(−3.2) μm, LW (1.4−)1.5−1.7(−2.0)(95% ci 4.5−4.7 × 2.8−3.0 μm, LW 1.6−1.7), smooth. Chlamydospores not observed (few chlamydospores observed in the protologue).
Habitat
Soil
Known distribution
Russia (Siberia), Austria, Peru (Puño State).
Holotype
Russia, Siberia, Krasnoyarsk region, soil in apple orchard, October 1997, G. Szakacs (DAOM 230011). Not examined, ex-type culture DAOM 230011 studied.
Additional material examined
See Table 1.
Comments
Bissett et al (2003) compared this species to the unrelated species T. longipile Bissett (Bissett 1991), which has somewhat broader conidia. It would be difficult to distinguish these species on the basis of their morphology alone.
Culture Berg PR26-12-6 (=G.J.S. 07-72) is cited in Grosch et al (2006, as T. viride) as being antagonistic to Rhizoctonia solani.
Trichoderma stromaticum Samuels & Pardo-Schultheiss, Mycol Res 104: 762. 2000. Figs. 2 h−j, 14 and 15
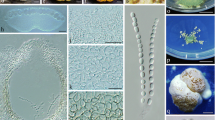
Trichoderma stromaticum. a Pustules in nature on a dead cacao tissue infected with Moniliophthora perniciosa (‘dry broom’). b−e Pustules on SNA (b, d), PDA (c) and CMD (d). f Tissue of a pustule showing broad hyphae terminating in phialides (on the left). g−1 Awl-like hairs arising from a pustule with fertile branches arising from near the base. The hair in (h) terminates in two phialides. j Phialides clustered on a fertile branch. k, l Synanamorph from SNA. Verticillium-like branching of the synanamorph seen in (l). m Conidia. n, o Chlamydospores. Images a from G.J.S. 97-181; b, G.J.S. 07-76; c, G.J.S. 01-92; d, G.J.S. 05-455; e, G.J.S. 04-327; f, G.J.S. 03-140; g, i, G.J.S. 00-132; h, G.J.S. 04-301; j, l, G.J.S. 04-190; k, G.J.S. 97-181; m, G.J.S. 03-134; n, o, G.J.S. 00-02. Scale bars (a, b) 1 mm, (c) 0.5 mm, (d, e) 0.5 mm, (f−i, l, n, o) 20 μm
Trichoderma stromaticum, Hypocrea stromatica teleomorph. a, b Stromata and conidial pustules on a dead cacao tissue infected with Moniliophthora perniciosa (‘dry broom’). c Cells of the stroma surface showing a perithecial opening. d Section through a stroma showing median longitudinal sections through several perithecia. e Section through a stroma showing details of the stroma surface and a perithecium in median longitudinal section. f Section through the surface region of a stroma showing the ostiolar canal of a perithecium. g Cells of the interior of a stroma below a perithecium. h−j Asci and part ascospores. A ring can be seen in the apex of asci in (i). Warted ascospores are seen in (j). Images a, c from G.J.S. 04-300, b from G.J.S. 04-305; c−j from G.J.S. 04-300
Teleomorph
Hypocrea stromatica Bezerra et al., Fitopatol Brasiliera 28:409. 2003.
Characteristics in culture
Optimum temperature for growth on PDA 25°C. Colony radius after 96 h at 25°C on PDA (54−)55−65(−68) mm, at 30°C 37−53(−61) mm; at 15°C 3−7(−16) mm, at 20°C 15−20(−31) mm (n = 9 strains). On SNA optimum temperature for growth 25−30°C. After 96 h at 25 and 30°C on SNA (25−)30−42(−52) mm, slightly slower at 30°C than at 25°C; 20°C 15−20(−30) mm, at 15°C 5−7(−15) mm. Colonies grown on PDA 1 week at 25 C under light producing pustules abundantly in 2−4 concentric rings, pustules in the center of the colony often with gold pigment. On SNA pustules forming abundantly in a broad marginal band, less typically pustules uniformly distributed throughout the colony. Individual pustules on PDA and SNA typically fully fertile within 2 weeks, easily dislodged from the agar, on SNA 0.2−0.5(−1.0) mm diam; conidia dark green but often yellow; hairs typically not visible on immature pustules, at maturity hairs arising from pustules, (45−)62−100(−135) μm long, (3.7−)4.7−6.7(−10.0) μm wide at the base, stiff, erect, septate, thin-walled, sterile or producing a single terminal phialide. Fertile branches arising at the base of hairs, typically one or a few cells in length, often densely clustered, producing unicellular lateral branches; phialides terminating all branches, densely clustered. Phialides ampulliform, (3.5−)5.3−7.7(−10.2) μm long, (1.0−)3.2−4.5(−5.5) μm at the widest point, (1.7−)2.5−3.7(−4.2) μm at the base, arising from a cell (3.0−)3.7−5.0(−7.0 μm wide. Conidia (n = 1,522) ellipsoidal, (3.0−)3.7−4.5(−7.2) × (1.7−)2.5−3.0(−3.7) μm, LW (1.0−)1.4−1.8(−2.3) (95% ci 4.3−4.4 × 2.80−2.82 μm, L/W 1.5−1.6), smooth. Chlamydospores (4.2−)6.5−9.7(−13.5) μm diam.
Characteristics of the teleomorph
Stromata dispersed among conidiomata, scattered or gregarious, discoidal, 1.0−1.5 mm diam, broadly attached, hyphae not visible, surface plain, perithecial elevations appearing as low tubercules, perithecial openings not visible, not reacting to 3% KOH. Cells of the stroma surface in face view pseudoparenchymatous, elliptical in outline, (4.5−)9−17(−20) μm diam, thin-walled. Perithecia circular in section, (185−)210−260(−300) μm high, (100−)130−190(−210) μm wide, ostiolar canal 60−90 μm long. Perithecial papilla formed of small cells, clavate elements lacking. Surface region distinguished from the internal tissue of the stromata by pigmentation in the outermost 2−3 layers of cells; cells of the stroma surface in section pseudoparenchymatous, (5−)8−13(−19) μm diam, thin-walled. Tissue of the stroma below perithecia pseudoparenchymatous, thin-walled, lacking long hyphal elements, (13−)15−27(−34) × (8−)10−16(−20) μm. Asci cylindrical, (64−)75−90(−100) × (3.2−)4.5−6.5(−9.0) μm, apex with a conspicuous discharge ring, ascospores uniseriate. Part ascospores hyaline, dimorphic; conspicuously warted, distal part conical to subglobose, (2.7−)3.7−5.2(−6.5) × (2.0−)3.5−4.5(−5.5) μm; proximal part wedge-shaped to ellipsoidal, (2.2−)3.7−5.0(−6.0) × (2.0−)3.0−4.02(−4.7) μm.
Habitat
Constantly associated with Moniliophthora perniciosa on Theobroma cacao, on dead cacao tissue infected with M. perniciosa (‘dry brooms) or fruit bodies of M. perniciosa. Collections from Peru were found on the pseudostroma of M. roreri. Isolated rarely as an endophyte from trunk of Theobroma cacao.
Known distribution
Brazil (Bahia, Pará), Colombia, Ecuador, Peru.
Holotype
Brazil, Para, Belem, from dead cocoa broom, C. N. Bastos, TVC, G.J.S. 97-183 (BPI 746496; live cultures: ATCC 204426, CBS 101875).
Comments
This species was fully described and illustrated in Samuels et al (2000).
Bezerra et al (2003) described Hypocrea stromatica Bezerra et al as the teleomorph of T. stromaticum. We have observed perithecia in Clade B and C (Fig 1e) but not in unresolved Clade A, which includes the ex-type culture (TVC = G.J.S. 97-183). We have not studied the ex-type culture of H. stromatica, which was collected in Pará State, Brazil, but following our phylogenetic analysis it is likely that it too will cluster in clade B.
Trichoderma vermipilum Samuels, sp. nov. Figs. 2g and 16
Trichodermati lanuginoso simile sed ob conidia (3.5−)4.0−5.2(−6.0) × (2.5−)2.7−3.0(−3.5) μm mensa, L/W (1.3−)1.5−1.9−2.0) differt, et in agaris dictis PDA vel SNA magis celeriter crescens.
Trichoderma vermipilum. a, b Pustules from SNA. Note the wooly nature of the pustule in (b). c Hairs arising from a pustule. d−h Hairs with fertile branches arising from near the base. Phialides are produced at the tip of hairs in (d, e). i Fertile branches arising from near the base of a hair. j Conidia. k chlamydospores. All from PPRI 3559. Scale bars (a) = 1 mm, (b) 0.5 mm, (c−h, k) 20 μm, (i, j ) 10 μm
Holotype BPI 881031, designated here.
Mycobank 519545
Telemorph
Not known
Characteristics in culture
Optimum temperature for growth on PDA 25°C, on SNA 30ºC; colony radius after 96 h on PDA at 25ºC ca. 55 mm, on SNA at 30°C ca. 60 mm. On PDA lacking diffusing pigment or distinctive odor. Typically not growing at 35°C. On PDA after 10 days at 25°C under 12 h cool white fluorescent light/12 h darkness conidia forming in a continuous mat of densely disposed, pulvinate pustules around the periphery of the Petri plate; conidia grayish green (K&W 30D5−8); on SNA producing numerous pustules scattered around the periphery of the Petri plate. Pustules pulvinate, 0.25−1.0 mm diam, with numerous protruding hairs. Hairs white, variable in length (50−175 μm), sinuous to somewhat spiraled, mostly producing a single terminal phialide, often producing a terminal verticil of phialides and sometimes producing a few solitary, lageniform phialides laterally, or sterile, tapering uniformly from 4−5 μm at the base. Fertile branches arising near the base of the hairs, each one or a few cells in length, longer fertile branches rebranching to produce unicellular secondary branches; phialides arising in dense clusters at the tips of all fertile branches and giving a botryose aspect. Phialides nearly subglobose to flask-shaped with a broad base, (4.7−)5.5−7.5(−9.5) μm long, (3.7−)4.0−5.0(−5.5) μm at the widest, L/W (1.0−)1.2−1/6(−2.1), (2.2−)2.7−3.5(−4.2) μm at the base, arising from a cell (2.5−)2.7−4.5(−5.5) μm wide. Conidia (n = 30) nearly oblong, (3.5−)4.0−5.2(−6.0) × (2.5−)2.7−3.2(−3.5) μm (95% ci 4.5−4.9 × 2.8−3.0 μm, L/W 1.6−1.7) smooth, gray green. Chlamydospores abundant on SNA within 1 week, intercalary in hyphal cells and conforming to the shape of the cell.
Etymology
‘vermipilum’ from Latin, reference to the worm-like hairs arising in abundance from conidial pustules.
Habitat
Known only from the original collection, mangrove soil.
Known distribution
Republic of South Africa
Holotype
Republic of South Africa, KwaZulu-Natal, Durban, isolated from soil under beachwood mangrove, 1986, R.Y. Auerlich s.n.(BPI 881031; ex-type culture PPRI 3359 = CBS 127103).
Comments
Using the sequence similarity search tool implemented in TrichoBLAST (www.ISTH.info) strain G.J.S. 01-312 (T. ivoriense, described here) could be identified as T. vermipilum. Strain PRI 3559 was reported as a Trichoderma sp. from South Africa by Kindermann et al. (1998) and Kullnig et al. (2000).
Discussion
Despite AFLP data, and phenotypic and biological evidence suggesting the existence of cryptic species in T. stromaticum, neither multilocus phylogenetic analysis nor MALDI-TOF MS peptide analysis (De Respinis et al. 2010) could support more than one taxon in this species. However, despite a lack of taxonomic resolution, the work of de Souza et al. (2006, 2008), Loguercio et al. (2009a, b), Medeiros et al. (2010) and Sanogo et al. (2002) suggests that strains of AFLP Group 1 (Clade B here) are better suited for biological control than are strains of Group 2 (Clade A here). Although T. stromaticum is very easily recognized through its morphology, the differences in phenotype between the two AFLP are not diagnostic. The only practical ways to determine to which group a newly collected strain of T. stromaticum belongs is through the AFLP procedure or through sequencing tef1, using the primers described here, followed by comparison with sequences that we have deposited in GenBank. In the present work, we have found strains of T. stromaticum isolated from the pseudostromata of the destructive cacao pathogen Moniliophthora roreri. This is the first report of T. stromaticum occurring on this pathogen, and these strains may provide a new source of control of the Frosty Pod Rot pathogen (Clade C, Fig 1e).
The literature abounds in reports of Trichoderma species that can be mycoparasites, including strains of species that have been isolated as endophytes, but of all these we only know two species that are host specific: T. aggressivum Samuels et al (Samuels et al. 2002), cause of green mold disease of Agaricus mushrooms and T. stromaticum Samuels et al. (2000), a parasite of Moniliophthora perniciosa, the cause of witches’ broom of cacao. There are more examples of more or less host specificity for perithecial formation, including Hypocrea latizonata Peck, which is only found on basidiomata of Cyathus species (Sundberg and Kost 1989), and members of Trichoderma sect. Hypocreanum Bissett, which occur only on members of the basidiomycete order Aphyllophorales (e.g. Hypocrea americanum Overton on Fomitopsis pinicola and Piptoporus betulinus (Overton et al. 2006b), H. sulphurea (Schw.) Sacc. on basidiomata or mycelium of Exidia nucleata (Overton et al. 2006a), or H. pulvinata Fuckel on various polypores (Overton et al. 2006a)). Trichoderma stromaticum is a parasite of Moniliophthora species and is utilized in a biological control program for Witches’ Broome Disease of cacao (‘Trichovab’; Samuels et al. 2000; Medeiros et al. 2010). It is host specific. It is always associated with cacao and, as was noted above, is almost always found growing on cacao tissue infected with the witches’ broom pathogen M. perniciosa; it is rarely isolated from sapwood of cacao or found on cacao pods infected with the Frosty Pod Rot pathogen of cacao, M. roreri. To our knowledge it has never been isolated from soil, which is the typical habitat of Trichoderma (Klein and Eveleigh 1998). Hoyos-Carvajal et al. (2009) did not list it among the many species of Trichoderma that they found in soil in Colombia and adjacent countries, and we did not find it among approximately 800 cultures of Trichoderma isolated from under wild cacao trees in Peru (Samuels, unpublished).
Has T. stromaticum coevolved with its host, Moniliophthora perniciosa on cacao? The center of origin, and the highest diversity of cacao is thought to be in the upper Amazon region of South America (Bartley 2005; Motomayor et al. 2008) and the highest diversity of M. perniciosa, at least in Brazil, is in the eastern, lower Amazon regions of the states of Amazonas and Pará (Rincones et al. 2006). Moniliophthora perniciosa is presumed to have coevolved with cacao in Amazonian America (Purdy and Schmidt 1996; Evans et al. 2003b). However in extensive exploration for wild cacao and its pathogens in several river systems in Amazonian Peru (Arevalo-Giardini, Meinhardt and Samuels, unpublished) T. stromaticum was not encountered despite the presence of M. perniciosa. Moniliophthora perniciosa is quite variable (Purdy and Schmidt 1996; de Arruda et al. 2005). In addition to pathogenic variations in cacao (‘C’ biotype) there are populations adapted to other plant species including the ‘S’ biotype, which is adapted to solanaceous hosts, and the ‘L’ biotype that is adapted to the liana Arrabidaea verrucosa (Bignoniaceae). The ‘H’ biotype was recently removed from M. perniciosa as the new species Crinipellis brasiliensis MCC de Arruda et al. (de Arruda et al. 2005); it forms basidiocarps on the fan brooms of Heteropterys acutifolia in Brazil and. according to Aime and Philips-Mora (2005). is most likely a species of Moniliophthora. Aime and Phillips-Mora (2005) suggested that additional parasitic basidiomycetes currently classified in Marasmiellus Murrill, might also be species of Moniliophthora.
The basidioma of M. perniciosa is a small marasmioid mushroom that forms on dry brooms of cacao and is sometimes found to be parasitized by T. stromaticum. Because the greatest diversity of the Stromaticum/Rossicum Clade is Paleotropical, possibly the origin of T. stromaticum, or an ancestor, is also Paleotropical, perhaps occurring on mushrooms of species in genera that are closely related to Moniliophthora, including Marasmius Fr., Marasmiellus, Crinipellis Pat. and Chaetocalathus Singer. Species of all of these genera occur in Africa (Singer 1976; Pegler 1977) and some species are Pantropical in distribution. Crinipellis stipitaria occurs on culms and roots of Gramineae, sometimes as a parasite, in Uganda but has a wide north temperate distribution as well (Pegler 1977). Dispersal of an agaric host could account for dispersal of T. stromaticum or a relative to America. The greatest diversity of T. stromaticum is found in the eastern Brazilian states of Bahia and Pará. This diversity may simply reflect intensity of sampling in a cacao-growing region, but eastern Brazil is also the closest land to West Africa and might be expected to be the first place that an invading organism from Africa would become established. This leaves open the possibility that some marasmioid species in the Atlantic coastal rain forest (Mata Atlantica) of Brazil may be a reservoir of T. stromaticum and that either there was a host switch to Moniliophthora species or that T. stromaticum infecting species in genera other than Moniliophthora has been overlooked by agaricologists. The range of possible hosts for T. stromaticum and its relatives is physically and taxonomically wide but unexplored.
References
Aime MC, Phillips-Mora W (2005) The causal agents of witches’ broom and frosty pod rot of cacao (chocolate, Theobroma cacao) form a new lineage of the Marasmiaceae. Mycologia 97:1012–1022
Althoff D, Gitzendanner M, Segraves K (2007) The utility of amplified fragment length polymorphisms in phylogenetics: A comparison of homology within and between genomes. Syst Biol 56:477–484
de Arruda MCC, Sepulveda Ch GF, Miller RNG, Ferreira MASV, Santiago DVR, Resende MLV, Dianese JC, Felipe MSS (2005) Crinipellis brasiliensis, a new species based on morphological and molecular data. Mycologia 97:1348–1361
Bartley BGD (2005) The genetic diversity of cacao and its utilization. CABI Publishing, Wallingford, UK
Bastos CN (1996) Potencial de Trichoderma viride no controle da vassoura-de-bruxa (Crinipellis perniciosa) do cacaueiro. Fitopatol Bras 21:509–512
Bezerra JL, Costa JC, Bastos CN, Faleiro FG (2003) Hypocrea stromatica sp. nov. teleomorfo de Trichoderma stromaticum. Fitopatol Bras 28:408–412
Bissett J (1991) A revision of the genus Trichoderma III Section Pachybasium. Can J Bot 69:2373–2417
Bissett J, Szakacs G, Nolan CA, Druzhinina I, Gradinger C, Kubicek CP (2003) New species of Trichoderma from Asia. Can J Bot 81:570–586
Bussell JD, Waycott M, Chappill JA (2005) Arbitrarily amplified DNA markers as characters for phylogenetic inference. Perspect Plant Ecol Evol Syst 7:3–26
Carbone I, Kohn LM (1999) A method for designing primer sets for speciation studies in filamentous Ascomycetes. Mycologia 91:553–556
Chaverri P, Castlebury LA, Overton BE, Samuels GJ (2003) Hypocrea/Trichoderma species with conidiophore elongations and green conidia. Mycologia 95:1100–1140
Cunningham CW (1997) Can three incongruence tests predict when data should be combined? Mol Biol Evol 14:733–740
Darlu P, Lecointre G (2002) When does the incongruence length difference test fail? Mol Biol Evol 14:432–437
Degenkolb T, Dieckmann R, Nielsen KF, Gräfenhan T, Theis C, Zafari D, Chaverri P, Ismaiel A, Brückner H, von Döhren H, Thrane U, Petrini O, Samuels GJ (2008) The Trichoderma brevicompactum clade: a separate lineage with new species, new peptaibiotics, and mycotoxins. Mycol Prog 7:177–209
Dettman J, Jacobson DJ, Taylor JW (2003) A multilocus genealogical approach to the phylogenetic species recognition in the model Eukaryote Neurospora. Evolution 57:2703–2720
de Queiroz A (1993) For consensus (sometimes). Syst Biol 42:368–372
De Respinis S, Voel G, Benagli C, Tonolla M, Petrini O, Samuels GJ (2010) MALDI-TOF MS of Trichoderma: a model system for the identification of microfungi. Mycol Prog 9:79–100
de Souza JT, Pomella AWV, Bowers JH, Pirovani CP, Loguercio LL, Hebbar PK (2006) Genetic and biological diversity of Trichoderma stromaticum, a mycoparasite of the witches’ broom pathogen. Phytopathology 96:61–67
de Souza JT, Bailey BA, Pomella AWV, Erbe EF, Murphy CA, Bae H, Hebbar PK (2008) Colonization of cacao seedlings by Trichoderma stromaticum, a mycoparasite of the witches’ broom pathogen, and its influence on plant growth and resistance. Biol Control 46:36–45
Don RH, Cox PT, Wainwright BJ, Baker K, Mattick JS (1991) Touchdown PCR to circumvent spurious priming during gene amplification. Nucl Acids Res 19:4008
Druzhinina IS, Kubicek CP, Komoń-Zelazowska M, Mulaw TB, Bissett J (2010) The Trichoderma harzianum demon: complex speciation history resulting in coexistence of hypothetical biological species, recent agamospecies and numerous relict lineages. BMC Evol Biol 10:94
Evans HC, Holmes KA, Reid AP (2003a) Phylogeny of the frosty pod rot pathogen of cocoa. Plant Pathol 52:476–485
Evans HC, Holmes KA, Thomas SE (2003b) Endophytes and mycoparasites associated with an indigenous forest tree, Theobroma gileri, in Ecuador and a preliminary assessment of their potential as biocontrol agents of cocoa diseases. Mycol Prog 2:149–160
Farris JS, Kallersjo M, Kluge AG, Bult C (1995) Testing significance of incongruence. Cladistics 10:315–319
Grosch R, Scherwinski K, Lottmann J, Berg G (2006) Fungal antagonists of the plant pathogen Rhizoctonia solani: selection, control efficacy and influence on the indigenous microbial community. Mycol Res 110:1464–1474
Hebbar P, Sanogo S, Pomella A, Soberanis W, Gomez I, Costa JC (2002) Biocontrol of cacao fungal diseases - example of disease management in a tropical tree crop. Bull OILB/SROP 25:359–361
Hillis DM, Bull JJ (1993) An empirical test of bootstrapping as a method for assessing confidence in phylogenetic analysis. Syst Biol 42:182–192
Hjorth S, Pomella AWV, Hockenhull J, Hebbar PK (2003) Biological control of Witches' Broom Disease, (Crinipellis perniciosa), with co-evolved fungus, Trichoderma stromaticum: testing different delivery regimes. Proceedings of the XIV International Cocoa Research Conference, Accra, Ghana, vol 2, pp 691–697
Hoyos-Carvajal L, Orduz S, Bissett J (2009) Genetic and metabolic biodiversity of Trichoderma from Colombia and adjacent neotropic regions. Fungal Genet Biol 46:615–631
Huelsenbeck JP, Ronquist F (2001) Mr Bayes: Bayesian inference of phylogeny. Bioinformatics 17:754–755
Huson DH (1998) SplitsTree: analyzing and visualizing evolutionary data. Bioinformatics 14:68–73
Kindermann J, El-Ayouti SGJ, Kubicek CP (1998) Phylogeny of the genus Trichoderma based on sequence analysis of the internal transcribed spacer region 1 of the rDNA cluster. Fungal Genet Biol 60:1–12
Klein D, Eveleigh DE (1998) Ecology of Trichoderma. In: Kubicek CP, Harman GE (eds) Trichoderma and Gliocladium Vol. 1 Basic biology, taxonomy and genetics. Taylor and Francis, London
Knowles LL, Carstens BC (2007) Delimiting species without monophyletic gene trees. Syst Biol 56:400–411
Komon-Zelazowska M, Bissett J, Zafari D, Hatvani L, Mancizinger L, Woo W, Lorito M, Kredicks L, Kubicek CP, Druzhinina IS (2007) Genetically closely related but phenotypically divergent Trichoderma species cause green mold disease in oyster mushroom farms worldwide. Appl Environ Microbiol 73:7415–7426
Kornerup A, Wanscher JH (1978) Methuen Handbook of Colour, 3rd edn. Methuen, London
Kullnig C, Szakacs G, Kubicek CP (2000) Molecular identification of Trichoderma species from Russia, Siberia and Himalaya. Mycol Res 104:1117–1125
Kullnig-Gradinger CM, Szakacs G, Kubicek CP (2002) Phylogeny and evolution of the genus Trichoderma: a multigene approach. Mycol Res 106:757–767
Leache AD, Reeder TW (2002) Molecular systematics of the eastern Fence Lizard (Sceloporus undulatus): a comparison of parsimony, likelihood, and Bayesian approaches. Syst Biol 51:44–68
Loguercio LL, de Carvalho AC, Niella GR, de Souza JT, Pomella AWV (2009a) Selection of Trichoderma stromaticum isolates for efficient biological control of witches' broom disease in cacao. Biol Control 51:130–139
Loguercio LL, Santos JS, Niella GR, Miranda RAC, de Souza JT, Collins RT, Pomella AWV (2009b) Canopy-microclimate effects on the antagonism between Trichoderma stromaticum and Moniliophthora perniciosa in shaded cacao. Plant Pathol 58:1104–1115
Maddison DR, Maddison WM (2003) MacClade 4 Analysis of phylogeny and character evolution (version 4.06). Sinauer, Sunderland, MA
Mason-Gamer RJ, Kellogg EA (1996) Testing for phylogenetic conflict among molecular data sets in the tribe Triticeae (Gramineae). Syst Biol 45:524–545
Medeiros FHV, Pomella AWV, de Souza JT, Niella GR, Valle R, Bateman RP, Fravel D, Vinyard B, Hebbar PK (2010) A novel, integrated method for management of witches’ broom disease in cacao in Bahia, Brazil. Crop Prot 29:704–711
Meinhardt LW, Rincones J, Bailey BA, Aime MC, Griffith GW, Zhang D, Pereira GAG (2008) Moniliophthora perniciosa, the causal agent of witches’ broom disease of cacao: what’s new from this old foe. Mol Plant Pathol 9:577–588
Meudt HM, Clarke AC (2007) Almost forgotten or latest practice? AFLP applications, analyses and advances. Trends Plant Sci 12:106–117
Motomayor JC, Lachenaud P, da Silva W, Mota J, Loor R, Kuhn D, Brown JS, Schnell RJ (2008) Geographic and genetic population differentiation of the Amazonian chocolate tree (Theobroma cacao L). PLoS ONE 3(10):e3311. doi:10.1371/journal.pone.0003311
Nirenberg HI (1976) Studies on the morphologic and biologic differentiation in Fusarium Section Liseola. Mitt Biol Bundesanst Land Forstwirtsch 169:1–117
O’Donnell K, Cigelnik E, Ninernburg HI, Aoki T (2000) A multigene phylogeny of the Gibberella fujikuroi species complex: detection of additional phylogeographically distinct species. Mycoscience 41:61–78
Olatinwo RO, Sabaratnam S, Schilder AMC (2004) Trichoderma stromaticum: A potential biological control agent for black root rot of Strawberries. Abstract. Phytopathology 94(6):s78
O'Meara BC (2010) New heuristic methods for joint species delimitation and species tree inference. Syst Biol 59:59–73
Overton BE, Stewart EL, Geiser DM (2006a) Taxonomy and phylogenetic relationships of nine species of Hypocrea with anamorphs assignable to Trichoderma section Hypocreanum. Stud Mycol 56:39–65
Overton BE, Stewart EL, Geiser DM, Jaklitsch WM (2006b) Systematics of Hypocrea citrina and related taxa. Stud Mycol 56:1–38
Pegler DN (1977) A preliminary agaric flora of East Africa. Kew Bull Add Ser VI:1–615
Posada O (2008) jMODELTEST: phylogenetic model averaging. Mol Biol Evol 25:1253–1256
Purdy LH, Schmidt RA (1996) Status of Cacao witches’ broom: biology, epidemiology, and management. Annu Rev Phytopathol 34:573–594
Rambaut A, Drumond AJ (2009) Tracer version 1.5, MCMC Trace analysis package available at http://tree.bio.ed.ac.uk/software/tracer/
Reeb V, Lutzoni F, Roux C (2004) Contribution of RPB2 to multilocus phylogenetic studies of the euascomycetes (Pezizomycotina, Fungi) with special emphasis on lichen-forming Acarosporaceae and evolution of polyspory. Mol Phylogenet Evol 32:1036–1060
Rincones J, Mazotti GD, Griffith GW, Pomela AW, Figueira A, Queiroz MV, Pereira JF, Azevedo RA, Pereira GAG, Meinhardt LW (2006) Genetic variability and chromosome-length polymorphisms of the witches’ broom pathogen Crinipellis perniciosa from various plant hosts in South America. Mycol Res 110:821–832
Samuels GJ, Ismaiel A (2009) Trichoderma evansii and T lieckfeldtiae: two new T hamatum-like species. Mycologia 101:142–156
Samuels GJ, Pardo-Schultheiss R, Hebbar KP, Lumsden RD, Bastos CN, Bezerra JL, Costa JC (2000) Trichoderma stromaticum sp. nov., a parasite of the cacao witches’ broom pathogen. Mycol Res 104:760–764
Samuels GJ, Dodd SL, Gams W, Castlebury LA, Petrini O (2002) Trichoderma species associated with the green mold epidemic of commercially grown Agaricus bisporus. Mycologia 94:146–170
Samuels GJ, Ismaiel A, Bon M-C, De Respinis S, Petrini O (2009) Trichoderma asperellum sensu lato consists of two cryptic species. Mycologia 102:944–966
Sanogo S, Pomella A, Hebbar PK, Bailey B, Costa JCB, Samuels GJ, Lumsden RD (2002) Production and germination of conidia of Trichoderma stromaticum, a mycoparasite of Crinipellis perniciosa on cacao. Phytopathology 92:103–1037
Singer R (1976) Marasmieae (Basidiomycetes − Tricholomataceae). Flora Neotrop Monogr 17:1–347
Sundberg WJ, Kost D (1989) Notes on Hypocrea latizonata. Mem NY Bot Gard 49:286–289
Swofford DL (2002) PAUP*: Phylogenetic analysis using parsimony (*and other methods). Version 4.06b10. Sinauer, Sunderland, MA
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The Clustal X windows interface; flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucl Acids Res 24:4876–4882
Volkmann-Kohlmdyer B, Kohlmeyer J (1996) How to prepare truly permanent microscope slides. Mycologist 10:107–108
Zachow C, Berg C, Müller H, Meincke R, Komon-Zelazowska DIS, Kubicek CP, Berg G (2009) Fungal diversity in the rhizosphere of endemic plant species of Tenerife (Canary Islands): relationship to vegetation zones and environmental factors. ISME J 3:79–92
Acknowledgments
The following individuals contributed cultures used in this research: Adriaana Jacobs and Michael Wingfield (Republic of South Africa); Ismaiel Kibbe (Côte d’Ivoire), John Bissett (Canada), Christian Kubicek and Irina Druzhinina (Austria), Carmen Suarez (Ecuador), Whillys Soberanis and Enrique Arevalo-Giardini (Peru), Alan Pomella (Brazil); Harry Evans, Keith Holmes and Sarah Thomas (UK); Rabio Olatinwo, Annemieke Schilder and Prakash Hebbar (USA). Didier Begoude (Cameroon) enabled collecting in the Reserve Faunal du Dja. Nigel Hywell-Jones and Rungtip Nasit facilitated collecting in the UNESCO World Heritage Site Khao Yai National Park (Thailand). Orlando Petrini corrected the Latin descriptions. Mention of trade names or commercial products in this publication is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the U.S. Department of Agriculture. USDA is an equal opportunity provider and employer.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Samuels, G.J., Ismaiel, A., de Souza, J. et al. Trichoderma stromaticum and its overseas relatives. Mycol Progress 11, 215–254 (2012). https://doi.org/10.1007/s11557-011-0743-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11557-011-0743-4
Keywords
- Hypocreales
- Theobroma cacao
- Pleomorphic fungi
- Species concepts
- Species complex
- Hypocrea
- Hypocreaceae
- Cacao
- Biological control
- Biogeography
- Systematics